Abstract
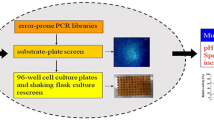
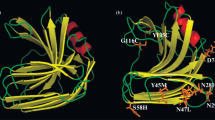
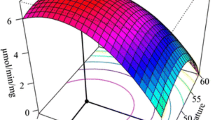
The thermal and alkaline pH stability of Streptomyces lividans xylanase B was improved greatly by random mutagenesis using DNA shuffling. Positive clones with improved thermal stability in an alkaline buffer were screened on a solid agar plate containing RBB-xylan (blue). Three rounds of directed evolution resulted in the best mutant enzyme 3SlxB6 with a significantly improved stability. The recombinant enzyme exhibited significant thermostability at 70°C for 360 min, while the wild-type lost 50% of its activity after only 3 min. In addition, mutant enzyme 3SlxB6 shows increased stability to treatment with pH 9.0 alkaline buffer. The K m value of 3SlxB6 was estimated to be similar to that of wild-type enzyme; however k cat was slightly decreased, leading to a slightly reduced value of k cat/K m, compared with wild-type enzyme. DNA sequence analysis revealed that eight amino acid residues were changed in 3SlxB6 and substitutions included V3A, T6S, S23A, Q24P, M31L, S33P, G65A, and N93S. The stabilizing effects of each amino acid residue were investigated by incorporating mutations individually into wild-type enzyme. Our results suggest that DNA shuffling is an effective approach for simultaneous improvement of thermal and alkaline pH stability of Streptomyces lividans xylanase B even without structural information.



Similar content being viewed by others
Abbreviations
- SlxB:
-
Streptomyces lividans xylanase B
- GH:
-
Glycoside hydrolase
- LB:
-
Luria-Bertani
- RBB-xylan:
-
4-O-methyl-d-glucurono-d-xylan-Remazol Brilliant Blue R
References
Arase A, Yomo T, Urabe I, Hata Y, Katsube Y, Okada H (1993) Stabilization of xylanase by random mutagenesis. FEBS Lett 316:123–127. doi:10.1016/0014-5793(93)81199-A
Biely P, Kluepfel D, Morosoli R, Shareck F (1993) Mode of action of three endo-beta-1, 4-xylanases of Streptomyces lividans. Biochim Biophys Acta 1162:246–254
Coughlan MP, Hazlewood GP (1993) β-1, 4-D-Xylan-degrading enzyme system: biochemistry, molecular biology and applications. Biotechnol Appl Biochem 17:259–289
Dupont C, Roberge M, Shareck F, Morosoli R, Kluepfel D (1998) Substrate-binding domains of glycanases from Streptomyces lividans: characterization of a new family of xylan-binding domains. Biochem J 330:41–45
Eriksson KEL (1990) Biotechnology in the pulp and paper industry. Wood Sci Technol 24:79–101. doi:10.1007/BF00225309
Gerstein M, Lesk AM, Chothia C (1994) Structural mechanisms for domain movements in proteins. Biochemistry 33:6739–6749. doi:10.1021/bi00188a001
Gonzalez-Blasco G, Sanz-Aparicio J, Gonzalez B, Hermoso JA, Polaina J (2000) Directed evolution of beta-glucosidase A from Paenibacillus polymyxa to thermal resistance. J Biol Chem 275:13708–13712. doi:10.1074/jbc.275.18.13708
Gruber K, Klintschar G, Hayn M, Schlacher A, Steiner W, Kratky C (1998) Thermophilic xylanase from Thermomyces lanuginosus: high-resolution X-ray structure and modeling studies. Biochemistry 37:13475–13485. doi:10.1021/bi980864l
Hakulinen N, Turunen O, Janis J, Leisola M, Rouvinen J (2003) Three-dimensional structures of thermophilic beta-1, 4-xylanases from Chaetomium thermophilum and Nonomuraea flexuosa. Comparison of twelve xylanases in relation to their thermal stability. Eur J Biochem 270:1399–1412. doi:10.1046/j.1432-1033.2003.03496.x
Havukainen R, Torronen A, Laitinen T, Rouvinen J (1996) Covalent binding of three epoxyalkyl xylosides to the active site of endo-1, 4-xylanase II from Trichoderma reesei. Biochemistry 35:9617–9624. doi:10.1021/bi953052n
Henrissat B, Bairoch A (1993) New families in the classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem J 293:781–788
Hori H, Elbein AD (1985) The biosynthesis of plant cell wall polysaccharides. In: Higuchi T (ed) Biosynthesis and biodegradation of wood components. Academic Press, Inc, Orlando, pp 109–135
Iffland A, Gendreizig S, Tafelmeyer P, Johnsson K (2001) Changing the substrate specificity of cytochrome c peroxidase using directed evolution. Biochem Biophys Res Commun 286:126–132. doi:10.1006/bbrc.2001.5366
Kaneko S, Kuno A, Muramatsu M, Iwamatsu S, Kusakabe I, Hayashi K (2000) Purification and characterization of a family G/11 beta-xylanase from Streptomyces olivaceoviridis E-86. Biosci Biotechnol Biochem 64:447–451. doi:10.1271/bbb.64.447
Kluepfel D, Vats-Mehta S, Aumont F, Shareck F, Morosoli R (1990) Purification and characterization of a new xylanase (xylanase B) produced by Streptomyces lividans 66. Biochem J 267:45–50
Lawson SL, Wakarchuk WW, Withers SG (1996) Effects of both shortening and lengthening the active site nucleophile of Bacillus circulans xylanase on catalytic activity. Biochemistry 35:10110–10118. doi:10.1021/bi960586v
Leskinen S, Mantyla A, Fagerstrom R, Vehmaanpera J, Lantto R, Paloheimo M et al (2005) Thermostable xylanases, Xyn10A and Xyn11A, from the actinomycete Nonomuraea flexuosa: isolation of the genes and characterization of recombinant Xyn11A polypeptides produced in Trichoderma reesei. Appl Microbiol Biotechnol 67:495–505. doi:10.1007/s00253-004-1797-x
Lever M (1972) A new reaction for colorimetric determination of carbohydrates. Anal Biochem 47:273–279. doi:10.1016/0003-2697(72)90301-6
Meyer A, Schmid A, Held M, Westphal AH, Rothlisberger M, Kohler HP et al (2002) Changing the substrate reactivity of 2-hydroxybiphenyl 3-monooxygenase from Pseudomonas azelaica HBP1 by directed evolution. J Biol Chem 277:5575–5582. doi:10.1074/jbc.M110018200
Moore JC, Arnold FH (1996) Directed evolution of a p-nitrobenzyl esterase for aqueous-organic solvents. Nat Biotechnol 14:458–467. doi:10.1038/nbt0496-458
Muilu J, Torronen A, Perakyla M, Rouvinen J (1998) Functional conformational changes of endo-1, 4-xylanase II from Trichoderma reesei: a molecular dynamics study. Proteins 31:434–444. doi:10.1002/(SICI)1097-0134(19980601)31:4<434::AID-PROT9>3.0.CO;2-H
Schimmel PR, Flory PJ (1968) Conformational energies and configurational statistics of copolypeptides containing l-proline. J Mol Biol 34:105–120. doi:10.1016/0022-2836(68)90237-4
Shareck F, Roy C, Yaguchi M, Morosoli R, Kluepfel D (1991) Sequences of three genes specifying xylanases in Streptomyces lividans. Gene 107:75–82. doi:10.1016/0378-1119(91)90299-Q
Song JK, Rhee JS (2000) Simultaneous enhancement of thermostability and catalytic activity of phospholipase A(1) by evolutionary molecular engineering. Appl Environ Microbiol 66:890–894. doi:10.1128/AEM.66.3.890-894.2000
Stemmer WP (1994) Rapid evolution of a protein in vitro by DNA shuffling. Nature 370:389–391. doi:10.1038/370389a0
Suenaga H, Mitsuoka M, Ura Y, Watanabe T, Furukawa K (2001) Directed evolution of biphenyl dioxygenase: emergence of enhanced degradation capacity for benzene, toluene, and alkylbenzenes. J Bacteriol 183:5441–5444. doi:10.1128/JB.183.18.5441-5444.2001
Torronen A, Rouvinen J (1997) Structural and functional properties of low molecular weight endo-1, 4-beta-xylanases. J Biotechnol 57:137–149. doi:10.1016/S0168-1656(97)00095-3
Wakarchuk WW, Sung WL, Campbell RL, Cunningham A, Watson DC, Yaguchi M (1994) Thermostabilization of the Bacillus circulans xylanase by the introduction of disulfide bonds. Protein Eng 7:1379–1386. doi:10.1093/protein/7.11.1379
You L, Arnold FH (1996) Directed evolution of subtilisin E in Bacillus subtilis to enhance total activity in aqueous dimethylformamide. Protein Eng 9:77–83. doi:10.1093/protein/9.1.77
Acknowledgements
This work was sponsored by National Natural Science Foundation of China (30770837) to Qin Wang. We thank Ms. Ying Li for the assistance with the DNA sequence analysis and mutant library construction.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Xia, T., Wang, Q. Directed evolution of Streptomyces lividans xylanase B toward enhanced thermal and alkaline pH stability. World J Microbiol Biotechnol 25, 93–100 (2009). https://doi.org/10.1007/s11274-008-9867-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-008-9867-3