Abstract
Sewage sludge was considered a critical reservoir for the propagation of antibiotic resistance genes (ARGs) and its treatment and disposal were important for the environment and human health. This study investigated the efficiency of the bioleaching technology for reducing eight typical antibiotic resistance genes in municipal sewage sludge, which was an emerging environmentally friendly technology for sludge dewatering. The prevalence of forty ARGs subtypes (including two mobile gene elements, MEGs) and one 16SrRNA gene were concerned with high throughput quantitative polymerase chain reaction during sludge bioleaching and a total of sixteen ARGs subtypes (including two transposase genes) were presented in the sludge samples. These genes were significantly decreasing after bioleaching, namely, one sulfonamide resistance gene (sul2), five tetracycline resistance genes (tetA-02, tetB-01, tetG-01, tetO-01, tetX), four macrolide-lincosamide-streptogramin B (MLSB) resistance genes (ermF, mphA-01, mphA-02, lnuB-01), two β-lactam resistance genes (blaOXA1/blaOXA30, blaPSE), two aminoglycoside resistance genes [aac(6´)-Ib, strB] and two MEGs (tnpA-01, tnpA-03). This result indicated that the bioleaching technology could significantly reduce the ARGs abundance of sewage sludge and the maximum removal efficiency was sixty-eight percent (mphA-02) and other ARGs subtypes such as tetA-02, tetB-01, mphA-01 and blaOXA1/blaOXA30 were decreased over fifty percent. The characteristics of Acidithiobacillus ferrooxidans contributed to the sludge bioleaching and may influence the diversity and composition of bacterial community and consequentially result in the change of ARGs abundance.
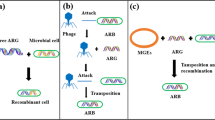
Graphical Abstract








Similar content being viewed by others
Availability of Data and Materials
Data and materials are available based on reasonable request.
References
Ajibola, A. S., & Zwiener, C. (2022). Occurrence and risk assessment of antibiotic residues in sewage sludge of two Nigerian hospital wastewater treatment plants. Water Air and Soil Pollution, 233(10). https://doi.org/10.1007/s11270-022-05875-4
Casiot, C., Morin, G., Juillot, F., Bruneel, O., Personne, J. C., Leblanc, M., et al. (2003). Bacterial immobilization and oxidation of arsenic in acid mine drainage (Carnoules creek, France). Water Research, 37(12), 2929–2936. https://doi.org/10.1016/S0043-1354(03)00080-0
Chen, Y., Wu, L., Boden, R., Hillebrand, A., Kumaresan, D., Moussard, H., et al. (2009). Life without light: Microbial diversity and evidence of sulfur- and ammonium-based chemolithotrophy in Movile Cave. Isme Journal, 3(9), 1093–1104. https://doi.org/10.1038/ismej.2009.57
Chen, Q., An, X., Li, H., Su, J., Ma, Y., & Zhu, Y. (2016). Long-term field application of sewage sludge increases the abundance of antibiotic resistance genes in soil. Environment International, 92–93, 1–10. https://doi.org/10.1016/j.envint.2016.03.026
Chu, L., & He, W. (2021). Toxic metals in soil due to the land application of sewage sludge in China: Spatiotemporal variations and influencing factors. Science of the Total Environment, 757. https://doi.org/10.1016/j.scitotenv.2020.143813
Cui, T. T., Zhang, S. Y., Ye, J. Y., Gao, L., Zhan, M. J., & Yu, R. (2022). Distribution, dissemination and fate of antibiotic resistance genes during sewage sludge processing-a review. Water Air and Soil Pollution, 233(4). https://doi.org/10.1007/s11270-022-05597-7
Forsberg, K. J., Patel, S., Gibson, M. K., Lauber, C. L., Knight, R., Fierer, N., et al. (2014). Bacterial phylogeny structures soil resistomes across habitats. Nature, 509(7502), 612. https://doi.org/10.1038/nature13377
Huang, J., Liang, J., Yang, X., Zhou, J., Liao, X., Li, S., et al. (2020). Ultrasonic coupled bioleaching pretreatment for enhancing sewage sludge dewatering: Simultaneously mitigating antibiotic resistant genes and changing microbial communities. Ecotoxicology and Environmental Safety, 193. https://doi.org/10.1016/j.ecoenv.2020.110349
Huber, B., Herzog, B., Drewes, J.E., Koch, K., & Mueller, E. (2016). Characterization of sulfur oxidizing bacteria related to biogenic sulfuric acid corrosion in sludge digesters. Bmc Microbiology, 16. https://doi.org/10.1186/s12866-016-0767-7
Inagaki, F., Takai, K., Nealson, K. H., & Horikoshi, K. (2004). Sulfurovum lithotrophicum gen. nov., sp nov., a novel sulfur-oxidizing chemolithoautotroph within the epsilon-Proteobacteria isolated from Okinawa Trough hydrothermal sediments. International Journal of Systematic and Evolutionary Microbiology, 54, 1477–1482. https://doi.org/10.1099/ijs.0.03042-0
Jiang, Y., Gao, F., Zhang, N., Li, J., Xu, M., & Jiang, Y. (2023). Dehydration Performance of Municipal Sludge and Its Dewatering Conditioning Methods: A Review. Industrial & Engineering Chemistry Research, 62(29), 11337–11357. https://doi.org/10.1021/acs.iecr.3c01553
Ju, F., Li, B., Ma, L., Wang, Y., Huang, D., & Zhang, T. (2016). Antibiotic resistance genes and human bacterial pathogens: Co-occurrence, removal, and enrichment in municipal sewage sludge digesters. Water Research, 91, 1–10. https://doi.org/10.1016/j.watres.2015.11.071
Li, Q., Tian, Y., Fu, X., Yin, H., Zhou, Z., Liang, Y., et al. (2011). The community dynamics of major bioleaching microorganisms during chalcopyrite leaching under the effect of organics. Current Microbiology, 63(2), 164–172. https://doi.org/10.1007/s00284-011-9960-y
Liu, F., Zhou, L., Zhou, J., Song, X., & Wang, D. (2012). Improvement of sludge dewaterability and removal of sludge-borne metals by bioleaching at optimum pH. Journal of Hazardous Materials, 221, 170–177. https://doi.org/10.1016/j.jhazmat.2012.04.028
Liu, Y., Wang, J., Hou, H., Chen, G., Liu, H., Liu, X., et al. (2020). Effect of introduction of exogenous strain Acidithiobacillus thiooxidans A01 on structure and function of adsorbed and planktonic microbial consortia during bioleaching of low-grade copper sulfide. Frontiers in Microbiology, 10. https://doi.org/10.3389/fmicb.2019.03034
Liu, C., Li, B., Wu, B., Lin, H., Jiang, L., & Qiu, Y. (2022). How heavy metal stress promotes dissemination of antibiotic resistance genes in the activated sludge process. Journal of Hazardous Materials, 437. https://doi.org/10.1016/j.jhazmat.2022.129279
Lu, Y., Meng, X., Wang, J., Dieketseng, M. Y., Xiao, Y., Yan, S., et al. (2022). Bioleaching rather than chemical conditioning using Fe[III ]/ CaO or polyacrylamide mitigates antibiotic resistance in sludge composting via pre-removing antibiotic resistance genes and limiting horizontal gene transfer. Waste Management, 137, 89–99. https://doi.org/10.1016/j.wasman.2021.10.029
Nascimento, L. P. D., Goncalves, J., & Duarte, I. C. (2022). Acidithiobacillus sp . applied to sewage sludge bioleaching : perspectives for process optimization through the establishment of optimal operational parameters. 3 Biotech, 12(11). https://doi.org/10.1007/s13205-022-03354-5
Nguyen, V. K., Ha, M. G., Shi, S., Seo, M., Jang, J., Jo, S., et al. (2018). Electrochemical effect on bioleaching of arsenic and manganese from tungsten mine wastes using Acidithiobacillus spp. Journal of Environmental Management, 223, 852–859. https://doi.org/10.1016/j.jenvman.2018.06.040
Ouyang, W., Huang, F., Zhao, Y., Li, H., & Su, J. (2015). Increased levels of antibiotic resistance in urban stream of Jiulongjiang River, China. Applied Microbiology and Biotechnology, 99(13), 5697–5707. https://doi.org/10.1007/s00253-015-6416-5
Pathak, A., Dastidar, M. G., & Sreekrishnan, T. R. (2009). Bioleaching of heavy metals from sewage sludge: A review. Journal of Environmental Management, 90(8), 2343–2353. https://doi.org/10.1016/j.jenvman.2008.11.005
Qin, S., Guo, L., Xie, Y., Zhang, D., & Ma, F. (2015). Improvement of municipal sludge dewaterability by Acidithiobacillus ferrooxidans. Journal of Harbin Institute of Technology, 47(08), 101–105.
Rivas-Castillo, A. M., Gomez-Ramirez, M., Lucas-Gomez, I. M., Carrillo-Vega, Y., & Rojas-Avelizapa, N. G. (2022). A new technique to evaluate Acidithiobacillus thiooxidans growth during a bioleaching process based on DNA quantification. Journal of Microbiological Methods, 198. https://doi.org/10.1016/j.mimet.2022.106494
Sanapareddy, N., Hamp, T. J., Gonzalez, L. C., Hilger, H. A., Fodor, A. A., & Clinton, S. M. (2009). Molecular diversity of a North Carolina wastewater treatment plant as revealed by pyrosequencing. Applied and Environmental Microbiology, 75(6), 1688–1696. https://doi.org/10.1128/AEM.01210-08
Wakeman, K. D., Honkavirta, P., & Puhakka, J. A. (2011). Bioleaching of flotation by-products of talc production permits the separation of nickel and cobalt from iron and arsenic. Process Biochemistry, 46(8), 1589–1598. https://doi.org/10.1016/j.procbio.2011.04.016
Wang, F., Qiao, M., Su, J., Chen, Z., Zhou, X., & Zhu, Y. (2014). High throughput profiling of antibiotic resistance genes in urban park soils with reclaimed water irrigation. Environmental Science & Technology, 48(16), 9079–9085. https://doi.org/10.1021/es502615e
Wang, J. L., Mao, D. Q., Mu, Q. H., & Luo, Y. (2015). Fate and proliferation of typical antibiotic resistance genes in five full-scale pharmaceutical wastewater treatment plants. Science of the Total Environment, 526, 366–373. https://doi.org/10.1016/j.scitotenv.2015.05.046
Wang, S., Ma, C., Zhu, Y., Yang, Y., Du, G., & Li, J. (2019). Deep dewatering process of sludge by chemical conditioning and its potential influence on wastewater treatment plants. Environmental Science and Pollution Research, 26(33), 33838–33846. https://doi.org/10.1007/s11356-018-2351-1
Xu, R., Yang, Z., Zheng, Y., Wang, Q., Bai, Y., Liu, J., et al. (2019). Metagenomic analysis reveals the effects of long-term antibiotic pressure on sludge anaerobic digestion and antimicrobial resistance risk. Bioresource Technology, 282, 179–188. https://doi.org/10.1016/j.biortech.2019.02.120
Yan, M., Prabowo, B., He, L., Fang, Z., Xu, Z., & Hu, Y. (2017). Effect of inorganic coagulant addition under hydrothermal treatment on the dewatering performance of excess sludge with various dewatering conditions. Journal of Material Cycles and Waste Management, 19(3), 1279–1287. https://doi.org/10.1007/s10163-016-0522-z
Yan, Y., Qin, L., Gao, J., Nan, R., & Gao, J. (2020). Protein extraction and sludge dewatering performance of ultrasound-assisted enzymatic hydrolysis of excess sludge. Environmental Science and Pollution Research, 27(15), 18317–18328. https://doi.org/10.1007/s11356-020-08208-2
Yang, Y., Li, B., Zou, S., Fang, H. H. P., & Zhang, T. (2014). Fate of antibiotic resistance genes in sewage treatment plant revealed by metagenomic approach. Water Research, 62, 97–106. https://doi.org/10.1016/j.watres.2014.05.019
Yang, T., Jiang, L., Bi, X., Cheng, L., Zheng, X., Wang, X., et al. (2022). Submicron aerosols share potential pathogens and antibiotic resistomes with wastewater or sludge. Science of the Total Environment, 821. https://doi.org/10.1016/j.scitotenv.2022.153521
Zeng, J., Gou, M., Tang, Y., Li, G., Sun, Z., & Kida, K. (2016). Effective bioleaching of chromium in tannery sludge with an enriched sulfur-oxidizing bacterial community. Bioresource Technology, 218, 859–866. https://doi.org/10.1016/j.biortech.2016.07.051
Zhai, W., Yang, F., Mao, D., & Luo, Y. (2016). Fate and removal of various antibiotic resistance genes in typical pharmaceutical wastewater treatment systems. Environmental Science and Pollution Research, 23(12), 12030–12038. https://doi.org/10.1007/s11356-016-6350-9
Zhang, T., Zhang, X., & Ye, L. (2011). Plasmid metagenome reveals high levels of antibiotic resistance genes and mobile genetic elements in activated sludge. PLoS ONE, 6(10), e26041. https://doi.org/10.1371/journal.pone.0026041
Zhang, T., Shao, M., & Ye, L. (2012). 454 Pyrosequencing reveals bacterial diversity of activated sludge from 14 sewage treatment plants. Isme Journal, 6(6), 1137–1147. https://doi.org/10.1038/ismej.2011.188
Zhang, M., Liu, X., Li, Y., Wang, G., Wang, Z., & Wen, J. (2017). Microbial community and metabolic pathway succession driven by changed nutrient inputs in tailings: Effects of different nutrients on tailing remediation. Scientific Reports, 7. https://doi.org/10.1038/s41598-017-00580-3
Zhou, G., Gu, Y., Yuan, H., Gong, Y., & Wu, Y. (2020). Selecting sustainable technologies for disposal of municipal sewage sludge using a multi-criterion decision-making method: A case study from China. Resources Conservation and Recycling, 161. https://doi.org/10.1016/j.resconrec.2020.104881
Acknowledgements
This work was supported by the Chengde major scientific and technological achievements transformation special project pre-decoking to strengthen three wastes recycling technology industrialisation for the flavour synthesis industry (202306B003).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Ethics Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent for Publication
All authors consent to publish the manuscript upon acceptance by the journal.
Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhao, L., Gao, X., Liu, X. et al. Reduction of Typical Antibiotic Resistance Genes and Mobile Gene Elements in Sewage Sludge During Sludge Bioleaching with Acidithiobacillus ferrooxidans. Water Air Soil Pollut 235, 263 (2024). https://doi.org/10.1007/s11270-024-07081-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11270-024-07081-w