Abstract
Fecal contamination threatens human health and contributes to the eutrophication of water resources. In Oklahoma, approximately 75% of assessed stream miles in the state are listed as impaired for fecal indicator bacteria (FIB). We tested the performance of seven microbial source tracking (MST) markers in six Northeast Oklahoma streams. All samples were tested with human (HF183), bovine (COWM2, COWM3), porcine (Pig-2-Bac), avian (Av4143), Escherichia coli, and Enterococcus markers using digital PCR (dPCR), as well as culturable assays for E. coli (Colisure) and Enterococcus (Enterolert). Rural and agricultural land uses were characterized by bovine sources of bacterial contamination. Human fecal contamination was found to be prominent in developed landscapes with several indicators for chronic human fecal pollution in an urban stream. All the streams met the criterion for Enterococcus impairment in 2019 and 2020; however, we found no relationships between culturable Enterococcus and the MST markers except in the urban stream, which had chronic human fecal pollution issues. The urban stream met the criterion for E. coli impairment, and E. coli was significantly correlated with the dominant MST markers in both rural and urban streams. We find that the culturable Enterococcus assay is not specific enough to be used for FIB water quality standards. We support the continued use of culturable E. coli assays to monitor for fecal contamination, and we recommend following-up with MST to verify fecal sources so informed mitigative actions can be taken to improve stream water quality.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Approximately 23% of stream miles in the USA are listed as impaired by fecal pollution (USEPA, 2018), posing a risk to human health and contributing to the eutrophication of aquatic ecosystems. While the Clean Water Act has curtailed most point-source contributions to fecal contamination in the USA, failing waste-water infrastructure (Habteselassie et al., 2014; Paul et al., 2000), the application of livestock manure to landscapes (Cook et al., 2014; Laurensen & Houlbrooke, 2014), and direct livestock access to streams (O’Callaghan et al., 2018) continue to impair rural and urban streams nationwide.
Fecal pollution is commonly identified by the detection of fecal indicator bacteria (FIB) by culture-based methods that target Escherichia coli and Enterococcus spp. Although FIB such as E. coli and Enterococcus can be surrogates for human pathogens, the specificity of these methods for targeting fecal contamination and estimating human health risks have been brought into question with several studies finding conflicting results (Byappanahalli et al., 2010; Flood et al., 2011; Mika et al., 2014). Specifically, Enterococcus was originally intended for assessing point sources for FIB, and the application of it for non-point source detection has been scrutinized for not correlating with human illness (Bradshaw et al., 2016; Colford et al., 2007; Goh et al., 2019; McQuaig et al., 2012) or the detection of markers specific to human fecal pollution (Flood et al., 2011; Byappanahalli et al., 2012; Brooks et al., 2020). Using FIB assays alone can result in classifying water bodies as impaired for fecal pollution when they are not impaired (Derose et al., 2020).
Culture-based methods have been recommended as a preliminary method for identifying potential issues of fecal pollution, but more specific, quantitative genetic methods are necessary to truly assess the risks to human health and identify the sources of fecal pollution that should be targeted for remediation (Brooks et al., 2020; Frick et al., 2020). The practice of microbial source tracking (MST) aims to link the fecal pollution of water to the specific source animal(s), and it is a critical step for assessing human health risks for recreating in waters, identifying sources of organic nutrient pollution that contribute to eutrophication, and for implementing cost-effective best management practices to mitigate fecal pollution (Carry et al., 2021; Gonzalez et al., 2020; Nshimyimana et al., 2018; Shanks et al., 2008). One of the challenges with MST is that the markers are regionally specific (Gawler et al., 2007; Schiaffino et al., 2020); therefore, the markers must be validated using fecal sources from the geographic region under study. Once validated, those markers can be used by local, state, and federal government agencies, tribal nations, and academic institutions for future studies or monitoring programs.
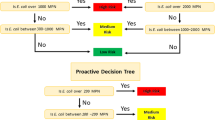
Approximately 74.5% of assessed stream miles in the state of Oklahoma are listed for not supporting their Primary Body Contact beneficial use designation by the Oklahoma Department of Environmental Quality (2018 accessed 2019). For a water body to support beneficial primary body contact uses in the State of Oklahoma, the current water quality standards require that a geometric mean of at least 5 samples within a 30-day period during the recreational season (May 1–September 30) must be under 126 MPN/100 mL for E. coli and under 33 MPN/100 mL for Enterococcus (Oklahoma Water Resources Board Title 785, Chapter 45, accessed 2022). The daily standard use criterion for E. coli and Enterococcus in recreational waters are 235 MPN/100 mL and 61 MPN/100 mL respectively (OWRB Title 785, Chapter 45, accessed 2022). Of assessed stream miles, 71.9% are impaired for Enterococcus and 25.7% are impaired for E. coli (ODEQ, 2018 accessed 2019). The prevalence of Oklahoma streams impaired for Enterococcus compared to E. coli suggests that Enterococcus may be too broad to be used as a fecal indicator in recreational streams, particularly those in rural landscapes.
The Ozark streams of Northeast Oklahoma provide a suite of ecosystem services, which are characterized by their gravel bottoms and clear, turquoise water, and they are culturally, economically, and recreationally significant to the region. In fact, some streams are classified by the state of Oklahoma as “wild and scenic” rivers, garnering them enhanced protections from point-source discharges (Meo, 2007; Siyoum, 2013; Chapagain et al., 2020); however, no such protections exist from non-point sources. Several of these streams are listed as impaired for Enterococcus or E. coli with no discernible source of chronic fecal contamination (ODEQ, 2018 accessed 2019), and thus, no strategy for effective mitigation. Molecular source tracking markers for fecal indicator bacteria have not been validated or utilized in this region or the surrounding states, which exhibit similar geographical features, land cover types, and land cover uses. Therefore, the objectives of this study are to (1) test the effectiveness of culturable assays on detecting fecal contamination in Ozark streams in Oklahoma, (2) validate MST markers for northeast Oklahoma, and (3) use the validated MST markers to identify sources of fecal contamination associated with bacteria-impaired Ozark streams. We hypothesized that the culturable Enterococcus assay is over-estimating FIBs in these streams. We also hypothesized that the urban stream, Town Branch, would be dominated by human markers, and the rural streams (Chewey, Piney, Tyner, Honey, and Sycamore) would be dominated by non-human markers.
2 Materials and Methods
2.1 Sites and Sampling
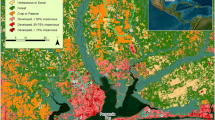
Over a 2-year period from 2019 to 2020 during the Oklahoma recreational season (May 1–September 30), we sampled five rural Ozark streams that were listed by the state of Oklahoma as impaired for Enterococcus and one urban stream with a history of fecal contamination that is listed as impaired for E. coli (ODEQ, 2018 accessed 2019) (Fig. 1; Table 1). We sampled each of the streams five times within a 30-day period, following the OWRB standards (Oklahoma Water Resources Board, 2022). Streams were sampled in the morning to limit potential marker decay from UV exposure even though the influence of UV exposure on FIB markers has not been resolved (Green et al., 2011; Sokolova et al., 2012). We used 120-mL IDEXX sample bottles with sodium thiosulfate to collect water for the Colisure and Enterolert IDEXX assays and 500-mL sterile polypropylene bottles to collect water for microbial source tracking. The water samples were immediately put on ice until arrival at the laboratory. One field replicate sample was taken during each sampling trip to ensure samples were representative for that site at the time of sampling. For the 2019 and 2020 recreational seasons, we sampled Chewey, Piney, and Town Branch in two separate clusters for within-year replication. We were unable to collect two clusters of samples from Honey, Sycamore, and Tyner in 2019 due to excessive flooding in Honey and Sycamore in the early summer, and the absence of water in Tyner during late summer. We used data from the 2016 national land cover database layer from the Multi-Resolution Land Characteristics Consortium (MRLC) to characterize the HUC 12 watersheds for each stream.
2.2 Evaluation of Markers
We used two general FIB MST markers (E. coli and genera-specific Enterococcus spp.) and five animal-specific MST markers: human (HF183), bovine (COWM2 and COWM3), porcine (Pig2Bac), and poultry (Av4143) (Table 2). We validated the markers with dPCR by testing them on DNA extracted from fecal samples from known sources (i.e., human, bovine, chicken, goose, and swine). Fresh animal fecal samples (n = 26) were collected in sterile 50-mL tubes and were processed immediately upon returning to the lab, or they were stored at − 80 °C until they were filtered, and the DNA were extracted. Cow (n = 8), pig (n = 8), chicken (n = 6), and dog (n = 6) samples were collected from local farms in Northeastern Oklahoma. Goose (n = 2) fecal samples were collected from a local swimming beach (Table 3). Human fecal samples (n = 30) were collected from the Grand River Dam Authority’s Grand River Energy Center wastewater treatment pond (Table 3). Upon bringing the fecal samples to the lab, 1 g of feces was mixed with pure DI water to form a 1 g/L fecal sample solution. These samples were then treated like regular stream samples and filtered through a 47-mm diameter polycarbonate filter with a 0.4-μm pore. The filters were then placed into capped tubes with glass beads and stored in a − 80 °C freezer until DNA extraction. A total of 56 fecal samples and 3 plasmids were tested for our MST validation study. For each marker, we calculated sensitivity, specificity, positive predictive value, negative predictive value, and accuracy (see supplemental materials for equations). The primers and probes were manufactured by Eurofins Genomics (Table 1) and rehydrated using TE buffer to master stock concentrations of 1000 μM. Thirty micromolar working stock primers and probes were made daily from this master stock.
2.3 Sample Processing
We processed the stream samples within 6 h of collection for EPA Methods 1600 (Santo-Domingo et al., 2004) and ASTMD6503-99 for Enterococcus (IDEXX Enterolert) and EPA method 9223 B for E. coli (IDEXX Colisure). For MST, we filtered 100 mL of sample water through a 47-mm diameter polycarbonate filter with a 0.4-μm pore, following Cao et al., (2016a). The filters were then placed into capped tubes with glass beads and stored in a − 80 °C freezer until extraction. We extracted DNA from the samples using GeneRite DNA EZ Extraction kits (GeneRite, North Brunswick, NJ, USA) following The California Microbial Source Identification Manual (Griffith et al., 2013).
2.4 MST Assay Efficiency
To demonstrate MST assay efficiency, standard curves were created using the observed and expected values from dilution series performed with both plasmids and contextually relevant fecal samples. Plasmids created by Integrated DNA Technologies were used for the E. coli and Enterococcus MST markers’ dilution series.
2.5 Detection and Quantification
All samples were checked by a fluorometric assay for total DNA concentrations as to not exceed 13 ng/μL. Samples that were found to be in exceedance of 13 ng/μL were diluted with PCR grade DI water at a ratio of 1:10 (Cao et al., 2018; Karlin-Neumann & Bizouarn 2018). The samples from the wastewater treatment pond were diluted at a 1:10 ratio as necessary, since they often yielded higher microbial loads relative to the stream sites. We used a QX100 Droplet Digital PCR System (Bio-Rad, Pleasanton, CA, USA) for DNA detection and quantification. We ran all the samples in duplicate. Our master mix consisted of DNA-free water, supermix (Bio-Rad ddPCR supermix for probes no dUTP), 0.9 μm of each primer, and 0.25 μM of each probe. We followed the plate setup procedure of Cao et al., (2016a). Each 24-μL reaction setup contained 18 μL of master mix and 6 μL of sample/template DNA. The droplet generator used 20 μL of this setup to create droplets, yielding a final total volume of 40 μL in the reaction wells. After the droplets were generated, the plate was sealed at 180 °C for 10 s, and it was either immediately placed into the thermal cycler for analysis or stored at 4 °C for analysis within 3 days. We followed Cao et al., (2016a) for the thermal cycling protocol, which consisted of 10 min at 95 °C; 40 cycles of 30 s at 94 °C and 1 min at 60 °C; and a 10-min hold at 98 °C. See Table 2 for annealing temperatures.
2.6 Data Analysis and Reporting
In dPCR, quantification is achieved by partitioning the sample into thousands of nanoliter to picoliter reactions in oil droplets and applying Poisson statistics to achieve a final concentration (Cao et al., 2016a). To be included in later analysis, samples needed to have at least 10,000 of these droplets per reaction well or at least a combined minimum of 20,000 droplets between two duplicate wells. The positive threshold is set at approximately one standard deviation, or around 500–700 units above the baseline formed by the NTC samples. Samples were interpreted as positive if there were 2 or more positive droplets (Steele et al., 2018). Enterococcus and HF183 were run in duplex; however, the other MST markers were run in simplex due to different annealing temperatures.
2.7 Statistical Analyses
We performed a forward-selection redundancy analysis (RDA) in Canoco 5.12, using HUC 12 land cover data to identify relationships between land cover types and FIB sources and abundances. Only land cover types that constituted more than 5% of the total land cover were included in the analysis. Furthermore, for the RDA, we only included land-cover types that significantly contributed (p ≤ 0.05) to the explained variation.
We performed regressions in SigmaPlot 12.5 to further assess the relationships between the MST markers and culturable FIB in rural versus urban stream sites by targeting markers that showed strong gradients in the RDA. To avoid redundancy between the two bovine markers, we only used COWM3, which had a higher detection rate than COWM2. We used Cook’s distance to identify outliers or influential observations. We found that the June 9, 2020 sampling event at Town Branch Creek yielded anomalous data for the Colisure and Enterolert assays that were having disproportional effects on the regressions. The Colisure sample from June 9, 2020 far exceeded what we had detected in any of the stream sites with a Cook’s distance of 20. The Enterolert sample had a Cook’s distance of 2. We also documented elevated dissolved organic matter, turbidity, nutrients, and chlorophyll concentrations during that June 9, 2020 sampling event, indicating there was an influx of organic matter into the stream (see Supplementary Materials). We are unsure of the source of the organic matter since there was no rainfall preceding the sampling event, and we had low detections for our MST markers.
3 Results
The daily standard for Enterococcus (61 MPN/100 mL) was exceeded in 61.8% of the stream samples, and the daily standard for E. coli (235 MPN/100 mL) was exceeded in 12.7% of the stream samples (Fig. 2). Honey Creek was the only stream that never exceeded the daily E. coli standard. All the streams exceeded the daily Enterococcus standard at least five times during the 2019–2020 recreational seasons. All the streams exceeded the Oklahoma Enterococcus geometric mean standard (33 MPN/100 mL) for recreational waters at least once during the 2019–2020 recreational seasons (Fig. 2). Town Branch Creek was the only stream that exceeded the OWRB E. coli geometric mean standard (126 MPN/100 mL), and it did so in both 2019 and 2020 (Fig. 2).
All the MST markers demonstrated high sensitivities to the regional fecal samples (Table 3). COWM2 and COWM3 were the highest performing MST markers exhibiting 100% sensitivity and 100% specificity, showing no cross-reactivity with non-bovid fecal samples. The Pig2Bac and AV4143 markers had 100% sensitivities, but they were cross-reactive with non-target fecal samples, yielding specificities of 98% and 94.1% respectively (Table 3). The HF183 marker exhibited the lowest sensitivity (96.8%) and specificity (53.6%) of all the markers tested, reacting to all the bovine fecal samples, four porcine samples, and one goose fecal sample (Table 3).
The human marker, HF183, was detected in all the streams at least once over the two summers, and it was detected in 95.8% of urban stream samples and 17.7% of rural stream samples (Fig. 3). The bovid markers, COWM3 and COWM2, were only detected in rural streams of which they were detected in 38% and 21.2% of stream samples respectively (Fig. 3). The avian marker, AV4143, was detected in 4.4% of the stream samples, which consisted of the urban stream (Town Branch) and three rural streams (Tyner, Sycamore, and Piney) (Fig. 3). We did not detect the porcine marker, Pig2Bac, in any of the streams.
Percent cover of developed land explained most of the variation in FIB concentrations (22.7%) followed by forest (8.6%), agriculture (5.1%), and grasslands (2.5%) (Fig. 4; RDA biplot, eigenvalue = 0.24, total explained variation = 36.97%). The HF183 and E. coli markers were more prevalent in watersheds with high urban development. The two bovid markers (COWM2 and COWM3) were more abundant in agriculture-dominated watersheds. The Enterococcus spp. and AV4143 markers were not associated with any of the land cover types (Fig. 4). Culturable E. coli was more abundant in more developed landscapes, whereas culturable Enterococcus was equally prevalent in developed and agricultural landscapes (Fig. 4).
Redundancy analysis of watershed land cover and microbial sources; eigenvalue = 0.24, total explained variation = 36.97%. Forward-selection was used to display land cover types that explained significant variation: developed (22.7% explained variation), agriculture (5.1% explained variation), forest (8.6% explained variation), and grassland (2.5% explained variation)
The E. coli and Enterococcus spp. MST markers correlated strongly with their respective culturable assays in the urban stream (regression: E. coli R2 = 0.76, p < 0.001; Enterococcus R2 = 0.7, p < 0.001; Fig. 5); however, those relationships were weaker or non-existent for the rural streams (regression: E. coli R2 = 0.37, p < 0.001; Enterococcus R2 < 0.01, p = 0.41; Fig. 5). The HF183 marker was significantly correlated with both culturable E. coli and Enterococcus in the urban stream (Regression: E. coli R2 = 0.54, p < 0.001; Enterococcus R2 = 0.47, p = 0.001; Fig. 5), but there were no significant relationships between the HF183 marker and culturable E. coli or Enterococcus in the rural streams (regressions: p > 0.05; Fig. 5). The COWM3 marker was never detected in the urban stream (Fig. 5). There was a significant relationship between culturable E. coli and the COWM3 marker in rural streams with low predictability (Regression: R2 = 0.14, p < 0.001). There was not a significant relationship between COWM3 and culturable Enterococcus in rural streams (regression: R2 < 0.001, p = 0.87; Fig. 5). For stream samples that exceeded the daily standard for E. coli, HF183 was detected in 92.9% of the samples, COWM3 was detected in 21.4% of the samples, and AV4143 was detected in 14.3% of the samples. For stream samples that exceeded the daily standard for Enterococcus, HF183 was detected in 44.1% of the samples, COWM3 was detected in 38.2% of the samples, and AV4143 was detected in 7.4% of the samples.
4 Discussion
4.1 MST Validation
Due to the wide variation in the specificities and sensitivities of MST markers across studies, it is critical that MST markers are validated for the geographical region of interest (Gawler et al., 2007). While most of the MST markers we used had high specificity and sensitivity, cross-reactivity with non-target fecal sources still occurred with the human (HF183), avian (AV4143), and porcine (Pig2Bac) markers.
The human fecal marker, HF183, has been widely documented to cross-react with non-human feces; however, its high sensitivity to human feces makes it a valuable marker for detecting human fecal contamination (Ahmed et al., 2016; Harwood et al., 2014; Schiaffino et al., 2020). In our validation study, the HF183 marker had an overall accuracy rating of 76.3%, with sensitivity and specificity at 96.8% and 53.6%, respectively (Table 3). The HF183 marker reacted to bovid, porcine, and goose feces (Table 3); however, the bovid markers were the only regularly detected non-human marker in the stream samples. Although HF183 cross-reacted with all our bovid fecal samples, bovid feces yielded higher concentrations of the COWM2 and COWM3 markers than the HF183 marker (Supplemental), and HF183 typically produces low copies when it cross reacts with non-target samples (Harwood et al., 2014). In rural sites where both human and bovid markers were detected, HF183 never exceeded 400 copies/100 mL (Supplemental). While we cannot rule out human fecal contamination in the presence of HF183, we posit that low HF183 concentrations that coincide with high bovid marker concentrations may be indicative of HF183 cross-reacting with bovid feces, particularly in areas without obvious sources of human fecal pollution.
The non-human markers exhibited higher specificities than the human marker, and they did not cross-react with human feces. Both bovid markers COWM2 and COWM3 performed identically in the validation study with 100% sensitivity, specificity, and accuracy. COWM2 produced more copies in the target fecal samples than COWM3; however, COWM3 was detected more often and at higher concentrations in field samples than COWM2, so we recommend using COWM3 for MST projects in the Ozark Plateau region. The porcine and avian markers both cross-reacted with the dog feces; however, the dog feces yielded substantially fewer copies relative to the target fecal samples (Supplemental); thus, cross-reactivity with dog feces in stream samples is unlikely.
4.2 Effectiveness of Culturable FIB Assays
All of the sites exceeded Oklahoma’s geometric mean criterion for being listed as impaired for Enterococcus (33MPN/100 mL); however, we found weak relationships between culturable Enterococcus and the MST markers (Fig. 5). We found a significant positive relationship between the HF183 marker and the culturable Enterococcus assay in the urban stream, but the predictability of that model was low compared to the direct comparisons between the culturable Enterococcus and the Enterococcus spp. marker (Fig. 5). Culturable Enterococcus and the Enterococcus spp. marker were only correlated at the Town Branch site, where human fecal contamination was detected at levels harmful to human health (HF183: > 4200 copies/100 mL (Boehm et al., 2015); > 3200 copies/100 mL (Ahmed et al., 2018)). The lack of correlation between the culturable Enterococcus and the Enterococcus spp. marker in the rural streams indicates that both assays are not always measuring the same thing. Many samples with high Enterococcus gene copies yielded few colonies in the Enterolert assay, and samples with high colony counts from the Enterolert assay produced low Enterococcus gene copies (Fig. 5). We have two possible explanations for this lack of a relationship that warrant further investigation: (1) the Enterolert assay is culturing bacteria other than Enterococcus causing false positives (Ferguson et al., 2013) and (2) the MST marker for Enterococcus spp. is interacting with genetic material from dead or dormant Enterococcus cells causing an over-estimation of biologically active Enterococcus (Byappanahalli et al., 2012). Flood et al. (2011) conducted a similar study, looking at the relationship between human markers and culturable Enterococcus in coastal streams, and while they often found high concentrations of culturable Enterococcus and human markers in the streams, they were not correlated with each other.
The urban stream, Town Branch Creek, was listed as impaired for E. coli prior to this study (ODEQ, 2018), and we found that it still exceeded the geometric mean criterion (126MPN/100 m L) for both 2019 and 2020. Unlike culturable Enterococcus, culturable E. coli was significantly positively correlated with its respective genetic marker and the dominant MST markers in the rural and urban streams (Fig. 5). However, the model fit for culturable E. coli with the markers was highest in the urban stream compared to the rural streams (Fig. 5), indicating that it is more responsive to human fecal contamination. This is corroborated by three isolated instances of daily exceedances of E. coli (> 235 MPN/100 mL) in the rural streams in 2019 in which HF183 was the dominant marker (Supplemental).
We posit that culturable Enterococcus lacks the specificity required to be used to assess standards for FIB. This is evident by the high percentage of bacteria-impaired streams in Oklahoma that are impaired for Enterococcus (96%) relative to E. coli (34%), and the frequency at which the sites in our study exceeded the standards for Enterococcus in the absence of notable fecal contamination. We found that culturable E. coli is conservative enough to continue to be used as a reliable indicator of fecal contamination, since the only site that exceeded the geometric mean standard was the Town Branch site, which we verified was the only site with chronically harmful levels of human feces. Culturable E. coli also maintained a significant relationship in rural streams with the E. coli genetic marker and other MST markers. It is important to note that the MST markers and the culturable FIB assays are acting as surrogates for a wide breadth of harmful pathogens; therefore, although multiple surrogates can be measured at high concentrations in the same water sample, their individual abundances are not directly related to each other, which is why finding relationships between these methods often fails to produce predictive models.
4.3 Implications for Watershed Management
Fecal contamination can originate from a variety of sources with differing impacts to human health and downstream water quality. Identifying the source of contamination is important for assessing that risk and implementing mitigation strategies to reduce further contamination from nutrients and other pollutants that can affect biological integrity. The human health risks from being exposed to feces depend on the source animal, the mode of contamination, the source animals’ health, and the “freshness” of the fecal material (Soller et al., 2010; Laurensen & Houlbrooke, 2014; O’Callaghan et al., 2018). Human feces are the most harmful to human health (Soller et al., 2010); therefore, the high concentrations of FIB and the human marker at the urban stream, Town Branch Creek, were a human health concern, particularly since this stream runs through several public parks and is commonly used for recreation. During this study, we found that human sewage was discharging from a damaged sewage line into the stream, and city officials remediated the issue. Gonzalez et al., (2020) provides several case studies in which they used MST to identify damaged sewage infrastructure and implement repairs to prevent sewage pollution, emphasizing the power of this tool for water managers and public health officials. In a few instances, we found the human marker in the absence of the bovid marker with elevated culturable E. coli numbers in the rural streams, indicating a source of human fecal contamination. Malfunctioning on-site septic systems have been found to be a significant contributor to human fecal pollution in rural areas near streams. For example, Verhougstraete et al., (2015) found human fecal contamination in several rural streams in Michigan, and that human fecal contamination increased with the total number of septic systems in the watershed that were within 60 m of a stream. Derose et al., (2020) found that most of their sampling sites that were associated with rural residential areas exceeded their state and federal standards for FIB, and they suspected malfunctioning on-site septic systems to be the source. Septic systems are prevalent in rural Oklahoma, so the detection of human markers in areas without municipal sewage systems may be indicative of on-site septic system failures.
Soller et al., (2010) found that bovid feces pose a similar risk to humans as human feces, since several bovine-borne pathogens that are infectious to humans like E. coli, Giardia, Cryptosporidium, and Salmonella are transmitted through fecal matter; whereas feces from poultry and pigs are less likely to pose a risk for human health. Aside from Townbranch Creek, cattle were the most prevalent source of fecal contamination in the stream sites. It is unknown how the gene copies of the bovid markers compare to those of the human marker regarding risks to human health. Overall, the number of gene copies for the bovid markers in the rural streams was lower than the number of HF183 gene copies in the urban stream. All of the rural streams had stretches of stream bank that were accessible to cattle for drinking water, so the cattle fecal sources were probably a combination of runoff from pasturelands and direct defecation in the stream. The microbial load from cattle feces deposited on dry land decreases rapidly within 10 days (Laurensen & Houlbrooke, 2014). Cattle defecate more frequently in the proximity of water, and microbes from feces deposited directly in streams are more likely to survive and be pathogenic (Bremner et al., 2016; O’Callaghan et al., 2018). Some pathogenic protists like Giardia and Cryptosporidium rely on aquatic environments to infect new hosts such that entire watersheds can become infected with these protists (McAllister et al., 2005), increasing the risk to human and animal health. Ruminants that are infected with water-borne pathogens often have diarrhea, require more feed, and weigh less than those not infected (Willms et al., 2002; Lardner & Willms, 2005; Aloisio et al., 2006), barring a direct cost to ranchers. Excluding cattle from streams would not only reduce fecal contamination, but also improve cattle health and yields (Budu-Amoako et al., 2012; McAllister et al., 2005). Therefore, we recommend using conservation easements that exclude cattle from direct stream access to reduce the prominence of cattle FIB in the rural Ozark streams.
We did not detect the porcine marker at any of the stream sites. Hog farms are not common in the region; however, there are wild hog populations. We did not use wild hog feces to validate the Pig2Bac marker, so it is possible that the marker is ineffective at detecting wild hog FIB, and this is something that should be explored further. Poultry farms are prevalent in the area, and the chicken litter from those farms is commonly applied to pastures by local ranchers (Kemper et al., 2006), so we were surprised that we did not detect more avian feces in the stream samples. However, avian sources of FIB may require substantially more fecal material to produce equivalent concentrations of bacteria as larger animals like humans and large domestic mammals (Wright et al., 2009). Poultry litter is often applied to pastures in the spring to prepare the soil for new plant growth, so there may not be enough live material to contribute to FIB in the streams during the summer recreation season.
5 Conclusion
Here, we used MST to investigate the effectiveness of culturable FIB as indicators for fecal contamination and to identify the major sources of fecal pollution in recreational Ozark streams in Northeastern Oklahoma, USA. We successfully validated the HF183, AV4143, COWM2, COWM3, and Pig2Back MST markers for Northeastern Oklahoma, and we found that the most prevalent sources of fecal pollution in Ozark streams were cattle and humans. Our hypothesis that culturable Enterococcus is too general to be used as an assessment tool for FIB was supported by our results. While we found that all six of the sites exceeded the Enterococcus geometric mean standard (33MPN/100 mL) at least once during the 2019–2020 study period, culturable Enterococcus counts were only correlated with MST markers in the urban stream, Town Branch Creek, where there was known high human fecal contamination (Figs. 3 and 5). Conversely, culturable E. coli counts only exceeded the geometric mean criterion (Colisure 126MPN/100 mL) at Town Branch Creek, and they were correlated with the dominant MST markers in both urban and rural streams, indicating that culturable E. coli assays are viable surrogates for both human and bovid fecal pollution. In conclusion, we support the continued use of culturable E. coli assays for assessing fecal contamination, but it is critical to follow-up with the MST analysis to identify the source of contamination so informed mitigative steps can be taken.
Data Availability
All data generate or analyzed during this study are included in this published article and its supplementary information file.
References
Ahmed, W., Sidhu, J. P. S., Smith, K., Beale, D. J., Gyawali, P., & Toze, S. (2016). Distributions of fecal markers in wastewater from different climatic zones for human fecal pollution tracking in Australian surface waters. Applied and Environmental Microbiology, 82, 1316–1323.
Ahmed, W., Hamilton, K. A., Lobos, A., Hughes, B., Staley, C., Sadowsky, M. J., & Harwood, V. J. (2018). Quantitative microbial risk assessment of microbial source tracking markers in recreational water contaminated with fresh untreated and secondary treated sewage. Environment International, 117, 243–249.
Aloisio, F., Filippini, G., Antenucci, P., Lepri, E., Pezzotti, G., Cacciò, S. M., & Pozio, E. (2006). Severe weight loss in lambs infected with Giardia duodenalis assemblage B. Veterinary Parasitology, 142, 154–158.
Boehm, A. B., Soller, J. A., & Shanks, O. C. (2015). Human-associated fecal quantitative polymerase chain reaction measurements and simulated risk of gastrointestinal illness in recreational waters contaminated with raw sewage. Environmental Letters, 2, 270–275.
Bradshaw, J. K., Snyder, B. J., Oladeinde, A., Spidle, D., Berrang, M. E., Meinersmann, R. J., Oakley, B., Sidle, R. C., Sullivan, K., & Molina, M. (2016). Characterizing relationships among fecal indicator bacteria, microbial source tracking markers, and associated waterborne pathogen occurrence in stream water and sediments in a mixed land use watershed. Water Research, 101, 498–509.
Bremner, K., Gordon, R. J., Powers, J., Rooney, N., & Madani, A. (2016). Partial or fully restricted cattle watering access: Water quality considerations. Applied Engineering in Agriculture, 32(6), 811–821.
Brooks, Y. M., Spirito, C. M., Bae, J. S., Hong, A., Mosier, E. M., Sausele, D. J., Fernandez-Baca, C. P., Epstein, J. L., Shapley, D. J., Goodman, L. B., Anderson, R. R., Glaser, A. L., & Richardson, R. E. (2020). Fecal indicator bacteria, fecal source tracking markers, and pathogens detected in two Hudson River tributaries. Water Research. https://doi.org/10.1016/j.watres.2019.115342
Budu-Amoako, E., Greenwood, S. J., Dixon, B. R., Barkema, H. W., & McClure, J. T. (2012). Giardia and Cryptosporidium on dairy farms and the role these farms may play in contaminating water sources in Prince Edward Island, Canada. Journal of Veterinary International Medicine, 26, 668–673.
Byappanahalli, M. N., Whitman, R. L., Shively, D. A., & Nevers, M. B. (2010). Linking non-culturable (qPCR) and culturable enterococci densities with hydrometeorological conditions. Science of the Total Environment, 408, 3096–3101.
Byappanahalli, M. N., Nevers, M. B., Korajkic, A., Staley, Z. R., & Harwood, V. J. (2012). Enterococci in the environment. Microbiology and Molecular Biology Reviews, 76, 685–706.
Cao, Y., Griffith, J. F., & Weisberg, S. B. (2016a). The next generation PCR-based quantification method for ambient waters: digital PCR. Methods in Molecular Biology, https://doi.org/10.10007/978-1-4939-3774-5_7
Cao, Y., Raith, M. R., & Griffith, J. F. (2016b). A duplex digital PCR assay for simultaneous quantification of the Enterococcus spp. and the human fecal-associated HF183 marker in waters. Journal of Visualized Experiments, 109, e53611. https://doi.org/10.3791/53611
Cao, Y., Raith, M. R., & Griffith, J. F. (2018). Testing of general and human-associated fecal contamination in waters. Methods in Molecular Biology, 1768, 127–140.
Carry, R., Ballesté, E., Blanch, A. R., Lucena, F., Pons, P., López, J. M., Rull, M., Solà, J., Micola, N., Fraile, J., Garrido, T., Munné, A., Soler, A., & Otero, N. (2021). Combining multi-isotopic and molecular source tracking methods to identify nitrate pollution sources in surface and groundwater. Water Research. https://doi.org/10.1016/j.watres.2020.116537
Chapagain, B., Long, J., Taylor, A., & Joshi, O. (2020). Variation in black bass angler characteristics by stream size and accessibility in Oklahoma’s Ozark Highland streams. North American Journal of Fisheries Management. https://doi.org/10.1002/nafm.10565
Chern, E. C., Siefring, S., Paar, J., Doolittle, M., & Haugland, R. A. (2011). Comparison of quantitative PCR assays for Escherichia coli targeting ribosomal RNA and single copy genes. Letters of Applied Microbiology, 52, 298–306.
Colford, J. M., Wade, T. J., Schiff, K. C., Wright, C. C., Griffith, J. F., Sandhu, S. K., Burns, S., Sobsey, M., Lovelace, G., & Weisberg, S. B. (2007). Water quality indicators and the risk of illness at beaches with nonpoint sources of fecal contamination. Epidemiology, 18, 27–35.
Cook, K. L., Netthisinghe, A. M. P., & Gilfillen, R. A. (2014). Detection of pathogens, indicators, and antibiotic resistance genes after land application of poultry litter. Journal of Environmental Quality. https://doi.org/10.2134/jeq2013.10.0432
Derose, K. L., Roche, L. M., Lile, D. F., Eastburn, D. J., & Tate, K. W. (2020). Microbial water quality conditions associated with livestock grazing, recreation, and rural residences in mixed-use landscapes. Sustainability. https://doi.org/10.3390/su12125207
Ferguson, D. M., Griffith, J. F., McGee, C. D., Weisberg, S. B., & Hagedorn, C. (2013). Comparison of Enterococcus species diversity in marine water and wastewater using Enterolert and EPA Method 1600. Journal of Environmental and Public Health. https://doi.org/10.1155/2013/848049
Flood, C., Ufnar, J., Wang, S., Johnson, J., Carr, M., & Ellender, R. (2011). Lack of correlation between enterococcal counts and the presence of human specific fecal markers in Mississippi creek and coastal waters. Water Research, 45, 872–878.
Frick, C., Vierheilig, J., Nadiotis-Tsaka, T., Ixenmaier, S., Linke, R., Reischer, G. H., Komma, J., Kirschner, A. K. T., Mach, R. L., Savio, D., Seidl, D., Blaschke, A. P., Sommer, R., Derx, J., & Farnleitner, A. H. (2020). Elucidating fecal pollution patterns in alluvial water resources by linking standard fecal indicator bacteria to river connectivity and genetic microbial source tracking. Water Research. https://doi.org/10.1016/j.watres.2020.116132
Gawler, A. H., Beecher, J. E., Brandão, J., Caroll, N. M., Falcão, L., & Meijer, W. G. (2007). Validation of host-specific Bacteriodales 16S rRNA genes as markers to determine the origin of faecal pollution in Atlantic Rim countries of the European Union. Water Research, 41, 3780–3784.
Goh, S. G., Saeidi, N., Gu, X., Vergara, G. G. R., Liang, L., Fang, H., Kitajima, M., Kushmaro, A., & Gin, K. Y. (2019). Occurrence of microbial indicators, pathogenic bacteria and viruses in tropical surface waters subject to contrasting land use. Water Research, 150, 200–215.
Green, H. C., Shanks, O. C., Sivaganesan, M., Haugland, R. A., & Field, K. G. (2011). Differential decay of human fecal Bacteroides in marine and freshwater. Environmental Microbiology, 13, 3235–3249.
Griffith, J.F., Layton, B.A., Boehm, A.B., Holden, P.A., Jay, J.A., Hagedorn, C., McGee, C.D., & Weisberg, S.B. (2013). The California Microbial Source Identification Manual: a tiered approach to identifying fecal pollution sources to beaches (Technical Report 804). Southern California Coastal Water Research Project.
Gonzalez, D., Keeling, D., Thompson, H., Larson, A., Denby, J., Curtis, K., Yetka, K., Rondini, M., Yeargan, E., Egerton, T., Barker, D., & Gonzalez, R. (2020). Journal of Microbiological Methods. https://doi.org/10.1016/j.mimet.2020.106068
Habteselassie, M. Y., Kirs, M., Conn, K. E., Blackwood, A. D., Kelly, G., & Noble, R. T. (2014). Tracking microbial transport through four onsite wastewater treatment systems to receiving waters in eastern North Carolina. Journal of Applied Microbiology. https://doi.org/10.1111/j.1365-2672.2011.05105.x
Harwood, V. J., Staley, C., Badgley, B. D., Borges, K., & Korajkic, A. (2014). Microbial source tracking markers for detection of fecal contamination in environmental waters: Relationships between pathogens and human health outcomes. Microbiology Reviews, 38, 1–40.
Karlin-Neumann, G., & Bizouarn, F. (Eds.). (2018). Digital PCR: methods and protocols. Springer. https://doi.org/10.1007/978-1-4939-7778-9
Kemper, N. P., Popp, J. S., Goodwin, H. L., Miller, W. P., & Doeksen, G. A. (2006). The economic power of poultry in the Ozarks. Journal of Applied Poultry Research, 15, 502–510.
Kildare, B. J., Leutenegger, C. M., McSwain, B. S., Bambic, D. G., Rajal, V. B., & Wuertz, S. (2007). 16S rRNA-based assays for quantitative detection of universal, human-, cow-, and dog-specific fecal Bacteroidales: A Bayesian approach. Water Research. https://doi.org/10.1016/jwastres.2007.06.037
Lardner, H. A., & Willms, W. D. (2005). The effect of water quality on cattle performance on pasture. Australian Journal of Agricultural Research, 56, 97–104.
Laurensen, S., & Houlbrooke, D. J. (2014). Nutrient and microbial loss in relation to timing of rainfall following surface application of dairy farm manure slurries to pasture. Soil Research, 52, 513–520.
McAllister, T. A., Olson, M. E., Fletch, A., Wetzstein, M., & Entz, T. (2005). Prevalence of Giardia and Cryptosporidium in beef cows in southern Ontario and in beef calves in southern British Columbia. The Canadian Veterinary Journal, 46, 46–55.
McQuaig, S., Griffith, J., & Harwood, V. J. (2012). Association of fecal indicator bacteria with human viruses and microbial source tracking markers at coastal beaches impacted by nonpoint source pollution. Applied and Environmental Microbiology, 78, 6423–6432.
Meo, M. (2007). The Illinois River Project and Oklahoma’s quest for environmental quality. Journal of Contemporary Water Research and Education, 136, 56–67.
Mieszkin, S., Furet, J. P., Corthier, G., & Gourmelon, M. (2009). Estimation of pig fecal contamination in a river catchment by real-time PCR using two pig-specific Bacteroidales 16S rRNA genetic markers. Applied and Environmental Microbiology, 75, 3045–3054.
Mika, K. B., Ginsburg, D. W., Lee, C. M., Thulsiraj, V., & Jay, J. A. (2014). Fecal indicator bacteria levels do not correspond with incidence of human-associated HF183 Bacteroides 16S rRNA genetic marker in two urban southern California watersheds. Water, Air, and Soil Pollution,. https://doi.org/10.1007/s11270-014-1960-7
Nshimyimana, J. P., Martin, S. L., Flood, M., Verhougstraete, M. P., Hyndman, D. W., & Rose, J. B. (2018). Regional variations of bovine and porcine fecal pollution as a function of landscape, nutrient, and hydrological factors. Journal of Environmental Quality, 47, 1024–1032.
O’Callaghan, P., Kelly-Quinn, M., Jennings, E., Antunes, P., O’Sullivan, M., Fenton, O., & hUuallacháin, D. Ó. (2018). The environmental impact of cattle access to watercourses: a review. Journal of Environmental Quality, 48, 340–351.
Odagiri, M., Schriewer, A., Hanley, K., Wuertz, S., Misra, P. R., Panigrahi, P., & Jenkins, M. W. (2015). Validation of Bacteroidales quantitative PCR assays targeting human and animal fecal contamination in the public and domestic domains in India. Science of the Total Environment, 502, 462–470.
Ohad, S., Ben-Dor, S., Prilusky, J., Kravitz, V., Dassa, B., Chalifa-Caspi, V., Kashi, Y., & Rorman, E. (2016). The development of a novel qPCR assay-set for identifying fecal contamination originating from domestic fowls and waterfowl in Israel. Frontiers of Microbiology. https://doi.org/10.3389/fmicb.2016.00145
Oklahoma Department of Environmental Quality. (2018). Integrated Report – 303(D) & 305(B). https://www.deq.ok.gov/water-quality-division/watershed-planning/integrated-report/ (accessed 2019)
Oklahoma Water Resources Board. (2022). Title 785, Chapter 45: Oklahoma’s Water Quality Standards. https://casetext.com/regulation/oklahoma-administrative-code/title-785-oklahoma-water-resources-board/chapter-45-oklahomas-water-quality-standards (accessed 2022)
Paul, J. H., Mclaughlin, M. R., Griffin, D. W., Lipp, E. K., Stokes, R., & Rose, J. B. (2000). Rapid movement of wastewater from on-site disposal systems into surface waters in the lower Florida Keys. Estuaries, 23, 662–668.
Ryu, H., Elk, M., Khan, I. U., Harwood, V. J., Molina, M., Edge, T. A., & Domingo, J. S. (2014). Comparison of two poultry litter qPCR assays targeting the 16S rRNA gene of Brevibacterium sp. Water Research, 48, 613–621.
Santo-Domingo, J., Hansel, J., Molina, M., Oshiro, R., Shanks, O. C., Stelma, G. N., Edge, T., Griffith, J., Hardwood, V., Jenkins, M., Layton, A., Nakatsu, C., Sadoswky, M., Stewart, J., Stoeckel, D., Wiggins, B., & Wilbur, J. (2004). Microbial source tracking guide document. (EPA 600-R-05-064). United States Environmental Protection Agency. https://cfpub.epa.gov/si/si_public_record_Report.cfm?Lab=NRMRL&dirEntryID=133523
Schiaffino, F., Pisanic, N., Colston, J. M., Rengifo, D., Paredes Olortegui, M., Shapiama, V., PeñataroYori, P., Heaney, C. D., Davis, M. F., & Kosek, M. N. (2020). Validation of microbial source tracking markers for the attribution of fecal contamination in indoor-household environments of the Peruvian Amazon. Science of the Total Environment. https://doi.org/10.1016/j.scitotenv.2020.140531
Shanks, O. C., Atikovic, E., Blackwood, A. D., Lu, J., Noble, R. T., Domingo, J. S., Seifring, S., Sivaganesan, M., & Haugland, R. A. (2008). Quantitative PCR for detection and enumeration of genetic markers of bovine fecal pollution. Applied Environmental Microbiology, 74, 745–752.
Siyoum, A. M. (2013). Essays on economic valuation of recreation and ecosystem services in the Illinois River Basin in Northeastern Oklahoma. Masters Thesis. Oklahoma State University, Stillwater, Oklahoma.
Sokolova, E., Astrom, J., Pettersson, T. J., Bergstedt, O., & Hermansson, M. (2012). Decay of Bacteroidales genetic markers in relation to traditional fecal indicators for water quality modeling of drinking water sources. Environmental Science and Technology, 46, 892–900.
Soller, J. A., Schoen, M. E., Bartrand, T., Ravenscroft, J. E., & Ashbolt, N. J. (2010). Estimated human health risks from exposure to recreational waters impacted by human and non-human sources of faecal contamination. Water Research, 44, 4674–4691.
Steele, J. A., Blackwood, A. D., Griffith, J. F., Noble, R. T., & Schiff, K. C. (2018). Quantifications of pathogens and markers of fecal contamination during storm events along popular surfing beaches in San Diego, California. Water Research, 1, 137–149.
USEPA. (2018). 2017 Five-year review of the 2012 recreational water quality criteria. (EPA 823-R-18-001). Environmental Protection Agency. Office of Water. Office of Science and Technology. https://www.epa.gov/wqc/five-year-reviews-epas-rwqc
Verhougstraete, M.P., Martin, S.L., Kendall, A.D., Hyndman, D.W., & Rose, J.B. (2015). Linking fecal bacteria in rivers to landscape, geochemical, and hydrologic factors and sources at the basin scale. Proceedings of the Natural Academy of Sciences, 112:10419-10424
Willms, W. D., Kenzie, O. R., McAllister, T. A., Colwell, D., Veira, D., Wilmshurst, J. F., Entz, T., & Olson, M. E. (2002). Effects of water quality on cattle performance. Journal of Range Management, 55, 452–460.
Wright, M. E., Solo-Gabriele, H. M., Elmir, S., & Fleming, L. E. (2009). Microbial load from animal feces at a recreational beach. Marine Pollution Bulletin, 58, 1649–1656.
Acknowledgements
We would like to thank Matt Conrad, Kenny Kerns, Travis Hinshaw, Melvin Pritchett, Jared Griffith, Aaron Roper, Vicki Bradley, Trisha Snyder, Brittany Kincaid, Whitney Collins, and Joy Taylor for donating various livestock fecal samples. Keener Brumble, Sydney McSlarrow, and Hailey Seago helped collect stream samples during the 2019 recreational season.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Browning, D.A., Mausbach, W.E., Stookey, C. et al. Validating Microbial Source Tracking Markers and Assessing the Efficacy of Culturable E. coli and Enterococcus Assays in Ozark Streams, USA. Water Air Soil Pollut 234, 348 (2023). https://doi.org/10.1007/s11270-023-06355-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11270-023-06355-z