Abstract
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is handled in biosafety level 3 (BSL-3) facilities, whereas the antiviral screening of pseudotype virus is conducted in BSL-2 facilities. In this study, we developed a SARS-CoV-2 spike-pseudotyped virus based on a semi-replication-competent retroviral (s-RCR) vector system. The s-RCR vector system was divided into two packageable vectors, each with gag-pol and env genes. For env vector construction, SARS-CoV-2 SΔ19 env was inserted into the pCLXSN-IRES-EGFP retroviral vector to generate pCLXSN-SΔ19 env-EGFP. When pCLXSN-gag-pol and pCLXSN-SΔ19env-EGFP were co-transfected into HEK293 T cells to generate an s-RCR virus, titers of the s-RCR virus were consistently low in this transient transfection system (1 × 104 TU/mL). However, a three-fold higher amounts of MLV-based SARS-CoV-2 pseudotyped viruses (3 × 104 TU/mL) were released from stable producer cells, and the spike proteins induced syncytia formation in HEK293-hACE2 cells. Furthermore, s-RCR stocks collected from stable producer cells induced more substantial syncytia formation in the Vero E6-TMPRSS2 cell line than in the Vero E6 cell line. Therefore, a combination of the s-RCR vector and the two cell lines (HEK293-hACE2 or Vero E6-TMPRSS2) that induce syncytia formation can be useful for the rapid screening of novel fusion inhibitor drugs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pandemic of coronavirus disease 2019 (COVID-19) was caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). Coronaviruses are classified into four genera: alpha-CoV, beta-CoV, gamma-CoV, and delta-CoV-2. SARS-CoV-2, an enveloped, single-stranded, positive-sense RNA virus, was identified as a new member of the genus Betacoronavirus [1]. This virus binds to the cognate receptor–angiotensin-converting enzyme 2 (ACE2) using the spike glycoprotein (S), and the spike is proteolytically activated for entering the host cell by a type II transmembrane serine protease (TMPRSS2) [2, 3]. The ACE2 protein is localized on the apical plasma membrane of respiratory epithelial cells [4]. The colon carcinoma cell line (Caco-2), a lung carcinoma cell line (Calu-3), and a monkey kidney cell line (Vero E6) express ACE2 on the apical membrane domain. In SARS-CoV-2 infection, TMPRSS2 may play an important role. TMPRSS2 cleaves at single arginine or lysine residues (R/K↓) and TMPRSS2 inhibitor camostat mesylate can block the SARS-CoV-2 entry into cells [2]. Vero E6 cells constitutively expressing TMPRSS2 (Vero E6/TMPRSS2) are highly susceptible to SARS-CoV-2 infection [5]. HEK293T cells showed only modest viral replication because HEK293T cells did not express endogenous hACE2 [6]. However, HEK293T-hACE2 cells stably express human ACE2 receptor most widely used in SARS-CoV-2 infection.
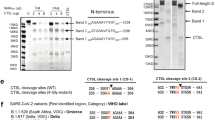
Cells infected with SARS-CoV-2 express the spike protein and form syncytia by fusing with ACE2 receptor-positive neighboring cells [7]. Expression of the spike protein alone, even in the absence of other proteins (M; membrane, E; envelope, and N; nucleocapsid), induces receptor-dependent syncytial formation [8, 9]. The spike protein, which contains two subunits, the S1 receptor-binding and the S2 fusion subunits, is cleaved by furin at the S1/S2 site and by TMPRSS2 at the S2’ site [10, 11]. Its high transmissibility results from a unique furin-like cleavage site (682-RRAR-685) [12]. Furin inhibitors block the SARS-CoV-2 spike-mediated syncytia formation [13]. Although “RRAR” was identified at the S1/S2 site by phylogenetic analysis of SARS-CoV-2, it is absent in SARS-CoV and other SARS-related coronaviruses [14]. In contrast, the S2’ cleavage site of SARS-CoV-2-S is similar to that of SARS-CoV-S.
Envelope proteins from nonrelated viruses can be “pseudotyped” into safer nonreplicative viral particles. These proteins facilitate the entry of pseudotype virus into cells [15,16,17,18]. The production of pseudotyped viral particles is a powerful tool for studying viral tropism and immunogenicity. Although SARS-CoV-2 requires handling in biosafety level 3 (BSL-3) facilities, the pseudotype virus allows antiviral screening to be conducted in BSL-2 facilities [19,20,21]. Recently, a SARS-CoV-2 pseudotyped virus was developed based on human immunodeficiency virus (HIV), murine leukemia virus (MLV), and vesicular stomatitis virus (VSV) [22]. Although the titers of the VSV-based SARS-CoV-2 pseudotyped virus are about 100-fold higher than those of HIV-based pseudotyped lentiviral particles, the former is not as practical as HIV-1-based SARS-CoV-2 pseudotyped viruses [21, 23, 24]. An HIV-1-based SARS-CoV-2 pseudotyped virus can assess the neutralization efficiency and entry inhibition. In the spike protein, 19 amino acids at the C-terminal were cleaved to overcome the low yield of the pseudotyped virus [25]. Previous studies have shown that C-terminal truncation of the spike protein and the D614G mutation enhances pseudoviral titers [26,27,28].
Murine leukemia virus (MLV) is a small RNA virus and contains three genes (gag, pol, and env). Replication-defective retroviral (RDR) vectors have insufficient gene transfer efficacy(103–106 transduction units/mL) and the inability to transfer genes into nondividing cells. To improve the low-level transduction efficiency of these vectors, MLV-based replication-competent retroviral (RCR) vectors have been developed [29]. RCR vectors have demonstrated improved efficacy of gene delivery, however, they still present the risk of accidental spread. To decrease this risk, semi-replication-competent retroviral (s-RCR) vectors were developed. They propagate transgenes as efficiently as RCR vectors and have an insert capacity of up to 7.3 kb of transgene. In a semi-replication-competent retroviral(s-RCR) system, the gag-pol and env genes were split into two vectors [30,31,32]. Our novel chimeric replication-competent Mo-MLV-10A1-EGFP vector was used as backbone plasmid to construct s-RCR [29].
In this study, incorporation of SARS-CoV-2 S protein into MLV particles was studied to construct an s-RCR vector that induces syncytial formation. The enhanced green fluorescent protein (EGFP) reporter gene was expressed in a gag-pol vector or an env vector to facilitate the detection of syncytial formation after transduction of target cells. In addition, an s-RCR vector released from stable producer cells was developed, as MLV-based SARS-CoV-2 pseudotyped viruses do not exhibit high titers through transient transfection [23].
When we compared the transient transfection system with stable producer cells to obtain high-titer s-RCR vectors that induce syncytia formation, titers of s-RCR virus—obtained from stable producer cells—were higher than those of the virus obtained from transient transfection. This study also shows that the s-RCR virus progressively replicates and induces syncytia formation in HEK293-hACE2 and Vero E6-TMPRSS2 cell cultures.
Materials and methods
Cell lines
The human embryonic kidney cell line HEK293 (ATCC, CRL-1573), HEK293 T cells expressing the SV40 T-antigen (ATCC, CRL-3216), and the African green monkey kidney cell line Vero E6 (ATCC, CRL-1586) were maintained in DMEM with 10% fetal bovine serum, 100 U/mL penicillin, and 100 μg/mL streptomycin.
Creation of HEK293-hACE2 and Vero E6-TMPRSS2 cells
From Hyeryun Choe [4] (Addgene plasmid #1786) and Roger Reeves [33] (Addgene plasmid # 53,887), pcDNA3.1-hACE2 and pCSDestTMPRSS2 were obtained, respectively. To create HEK293-hACE2 cells, pCLXSN-hACE2 was generated by cloning hACE2 from pcDNA3.1-hACE2 into the EcoRI and BamHI sites of pCLXSN retroviral vector. Next, VSV-G-pseudotyped retrovirus packaging human ACE2 was generated by co-transfecting 3.5 × 105 HEK293 T cells in 6-well plates with pCLXSN-hACE2 and packaging plasmids (2 µg of pVpack-GP and 2 µg of pCMV-VSV-G). The resulting retrovirus was used to infect HEK293 cells in the presence of 8 µg/ml polybrene. Transduced cells were selected in the growth medium containing G418 (800 µg/mL) for a period of 14 days.
The TMPRSS2 was cloned into the EcoRI and NotI sites of the retroviral vector pLPCX-IRES-EGFP (pLPCX-TMPRSS2). A VSV-G-pseudotyped retrovirus packaging human TMPRSS2 was constructed in the same way as described above. Vero E6 cells expressing TMPRSS2 were generated by transducing 4 × 105 Vero E6 cells in 6-well plates with VSV-G-pseudotyped retroviral vector pLPCX-TMPRSS2. Transduced cells were selected in the growth medium containing puromycin (3 µg/mL) for a period of 10 days.
Construction of C-terminal 19 aa-truncated SARS-CoV-2 S protein
The full-length SARS-CoV-2 S protein was codon-optimized and synthesized using GenScript (Piscataway, NJ, USA; https://www.molecularcloud.org/plasmid/pUC57-2019-nCoV-SHuman/MC-0101081.html). The SARS-CoV-2 S gene was subcloned into the EcoRI and NotI sites of the pcDNA3.1 vector (pcDNA3.1-S env). Hemagglutinin (HA)-tagged and C-terminal-truncated SARS-CoV-2 S expression constructs were generated by amplifying a fragment of approximately 760 bp using the forward primer 5’-GTG CTG GGC CAG TCT AAG AGA-3’ and the reverse primer 5’-GGA TCC TTA AGC GTA ATC TGG AAC ATC GTA TGG GTA ACA GCA GGA GCC ACA GCT ACA-3’ (BamHI restriction sites are underlined). Products of polymerase chain reaction (PCR) of C-terminal truncated SARS-CoV-2 S protein constructs were ligated into the pGEM-T Easy Vector System (Promega, Madison, WI, USA). Using the restriction enzymes DraIII and BamHI, pGEM-T-SARS-CoV-2 SΔ19 was digested, and the fragments were ligated into pcDNA3.1-S env to generate pcDNA3.1-SΔ19 env that encodes for the C-terminal 19 aa-truncated SARS-CoV-2 S and HA tag.
Construction of s-RCR vectors
A previously developed s-RCR vector system was used for the development of high-titer retroviral vectors [30]. To generate the s-RCR vector, pCLXSN-S env was constructed by cloning the EcoRI/BamHI fragment of SARS-CoV-2 into the retroviral vector pCLXSN. Construction of the pCLXSN-gag-pol-IRES-EGFP plasmid has been described previously [29, 30]. To investigate the complementation of gag-pol and env vectors, 3 × 105 Vero E6 cells in 6-well plates were transiently co-transfected with 3 µg of pCLXSN-gag-pol-IRES-EGFP and 3 µg of pCLXSN-S env using the CalPhos Mammalian Transfection Kit (TaKaRa Bio, Shiga, Japan). The replication of the s-RCR virus was measured by assessing the spread of syncytia and increase in EGFP-positive cells. To obtain s-RCR viruses from a stable producer cell line, 7 × 105 HEK293 T cells in 6-well plates were transiently co-transfected with 3 µg of pCLXSN-gag-pol, 3 µg of pCMV-VSV-G, and 3 µg of pCLXSN-S env-EGFP or 3 µg of pCLXSN-SΔ19 env-EGFP. Next, 6 × 105 HEK293 T cells in 6-well plates were infected with 1 mL of the s-RCR virus obtained from transfected HEK293 T cells. After two days, HEK293 cells were selected in G418 for 14 days (800 μg/mL). To produce s-RCR virus in transient transfection, 7 × 105 HEK293 T cells in 6-well plates were co-transfected with 3 µg of envelope expression plasmids (pCLXSN-S env or pCLXSN-VSV-G) and 3 µg of pCLXSN-gag-pol-IRES-EGFP. To investigate the release of virions from stable producer cells, viral lysates and cell lysates were detected with an anti-HA antibody (Sigma-Aldrich, MO, USA), a mouse monoclonal antibody anti-MLV p30 (1:1,000; Abcam, Cambridge, UK) and a rabbit polyclonal antibody anti-β-actin (1:10,000; AbClon, Seoul, Korea). To determine the viral titers, EGFP-positive cells were analyzed using a FACSCalibur flow cytometer (Becton, Dickinson and Company, NJ, USA). Vector titers were calculated using the following equation (N x P)/(V x D). N = cell number in each well used for infection; P = percentage of GFP-positive cells; V = viral volume used for infection; and D = dilution factor.
Infectivity assay
Receptor-dependent syncytia formation was induced via SARS-CoV-2 S expression. To examine syncytial formation induced by s-RCR viruses, 6 × 105 HEK293-hACE2 cells in 12-well plates and 6 × 105 Vero E6-TMPRSS2 cells in 12-well plates were infected with 0.5 mL of this virus obtained from stable producer cells. Both EGFP-positive and syncytia-forming cells were analyzed using fluorescence microscopy. Virus titers (syncytial-forming units [sfu]/mL) were determined by infecting Vero E6-TMPRSS2 target cells with a tenfold serial dilution of the viral supernatant obtained from stable producer cells.
Results
Construction of s-RCR vectors
The s-RCR vector system was constructed wherein two replication-defective vectors could generate infectious viruses from a cell infected by both vectors (Fig. 1). The two complementary replication-defective vectors are based on the Moloney-MLV (Mo-MLV) vector. To create the env vector (pCLXSN- S env-EGFP or pCLXSN-SΔ19 env-EGFP), the spike protein gene was inserted into the retroviral vector pCLXSN, with IRES-EFGP positioned directly downstream of the gene. To generate the gag-pol vector, pCLXSN-gag-pol was constructed by deleting the EGFP gene from the previously developed pCLXSN-gag-pol-IRES-EGFP recombinant vector [29, 34]. The 19 amino acids at the C-terminal, which codes for the ER/Golgi retention signal, are absent in SΔ19 env.
Structure of s-RCR vectors. Vectors derived from Moloney-Murine Leukemia virus (Mo-MLV). The plasmid pCLXSN-gag-pol contains the entire Mo-MLV gag-pol coding sequence and a neomycin resistance gene. An IRES-EGFP construct was inserted downstream of the gag-pol sequence of pCLXSN-gag-pol to construct pCLXSN-gag-pol-IRES-EGFP. The plasmid pCLXSN-S env contains the codon-optimized spike protein gene. C-terminal HA-tagged SΔ19 was inserted to pCLXSN-IRES-EGFP to produce pCLXSN-SΔ19 env-EGFP
Production of s-RCR viruses
To confirm the propagation of complementary defective vectors, the gag-pol vector (pCLXSN-gag-pol-IRES-EGFP) and env vector (pCLXSN-S env) were co-transfected into Vero E6 cells. As shown in Fig. 2, a progressive increase in EGFP-positive cells and syncytia formation was detected, whereas syncytia formation was not observed on Vero E6 cells transfected by pCLXSN-gag-pol-IRES-EGFP alone. This result suggests that s-RCR can spread through transfected Vero E6 cells, and that this viral propagation is accompanied by progressive syncytial formation.
Generation of s-RCR viruses encoding spike proteins. Vero E6 cells were transfected with 3 µg pCLXSN-S env and 3 µg pCLXSN-gag-pol-IRES-EGFP in 6-well plates to produce s-RCR viruses. Transfection of pCLXSN-gag-pol-IRES-EGFP alone in Vero E6 cells used as negative control. Syncytia formation was detected using fluorescence microscopy at 3, 6, 7, and 8 days post transfection
s-RCR viruses are efficiently released from stable producer cells
Roy et al. showed that stable HEK293GP cells (HEK293 cells that express MLV gag-pol) expressing SΔ19 are more useful than cells transiently transfected for generating SARS-CoV-2 pseudotyped viruses [35]. Stable HEK293 cells expressing MLV gag-pol and S or SΔ19 were generated via transduction. The presence of SARS-CoV-2 SΔ19 pseudotyped viruses released in the supernatant of stable HEK293 cells was analyzed using western blotting. The presence of spike protein at the stable HEK293 cells and the MLV viral particles—produced from stable producers—was detected using an antibody against HA (Fig. 3). These results show that SΔ19 was efficiently incorporated into MLV viral particles. It has been reported that SARS-CoV-2 spike-pseudotyped viruses are inefficiently released by transient transfection [21,22,23,24, 35]. In agreement with the previous results, titers of 5 × 105 transducing units (TU)/mL and 1 × 104 transducing units (TU)/mL were obtained for VSV-G and SARS-CoV-2 spike-pseudotyped viruses, respectively (Fig. 4a). A titer of SARS-CoV-2 spike-pseudotyped viruses was 50-fold lower than VSV-G-pseudotyped positive control vector. As shown in Fig. 4b, the SARS-CoV-2 SΔ19 env pseudovirus titer in the culture supernatant of stable producer cells was 3 × 104 transducing units (TU)/mL—detected by fluorescence-activated cell sorting (FACS), whereas the viral titer of SARS-CoV-2 S was 2 × 104 transducing units (TU)/mL. Both GFP-positive cells and syncytia formation were detected.
Western blot analysis of SARS-CoV-2 SΔ19 in viral lysates and cell lysates. To verify whether SARS-CoV-2 SΔ19 was incorporated into s-RCR viruses, concentrated supernatants obtained from stable producer cells were used as viral lysates. The presence of spike protein at the stable producer cells was also analyzed by western blot. Viral lysates: lane 1, bands corresponding to S and S2. lane M, protein marker. Cell lysates: lane 1, negative control, lane 2, bands corresponding to S and S2
Comparison of the yields of pseudotyped virus generated from transient transfection or stable producer cells. A Vero E6 cells were infected with s-RCR viruses obtained in transient transfection. B HEK293-hACE cells were infected with s-RCR viruses obtained from stable producer cells. Two days later, syncytia formation was detected using light and fluorescence microscope. To determine the viral titers, EGFP-positive cells were analyzed using a FACSCalibur™ flow cytometer (Becton, Dickinson and Company, NJ, USA). Magnification, 100 ×
s-RCR viruses induced syncytia formation in Vero E6-TMPRSS2
To investigate the effect of TMPRSS2 on syncytia formation, VeroE6-TMPRSS2 cells were infected with s-RCR viruses. As shown in Fig. 5, the syncytia formed in VeroE6-TMPRSS2 cells were larger than those in Vero E6 cells. The SARS-CoV-2 SΔ19 env pseudovirus titer was 2 × 103 sfu/mL. In addition, syncytia formation increased over time as the s-RCR virus spread throughout the culture. These results suggest that TMPRSS2 enhances syncytial formation.
Impact of TMPRSS2 on syncytia formation. Vero cells and Vero-TMPRSS2 cells were infected with s-RCR viruses obtained from stable producer cells at an MOI = 0.1. Syncytia formation was compared using light microscopy at 5, 7, and 9 days post infection. Virus titers were measured as syncytial-forming units (sfu/mL). Magnification, 100 ×
Discussion
Retroviral vectors pseudotyped with the SARS-CoV-2 spike protein are a powerful tool for screening SARS-CoV-2 entry inhibitors and neutralization assays [19,20,21, 24]. However, Mo-MLV cannot efficiently pseudotype the full-length SARS-CoV-2 spike protein in transient transfection systems [35]. We also obtained very low titers of the Mo-MLV-based SARS-CoV-2 pseudotyped virus using a transient transfection system (Fig. 4a). Recent studies reported that MLV pseudotyped with the C-terminal 19 aa-truncated SARS-CoV-2 S proteins obtained from stable producer cells infect HEK293-hACE2 cells more efficiently than with wild-type SARS-CoV-2 S protein [35]. In this study, we developed an s-RCR vector system that induces syncytia formation for large-scale production of s-RCR viruses in stable producer cells.
To investigate the generation of s-RCR virus through the transcomplementation of two defective retroviral vectors, the gag-pol vector carrying an EGFP reporter gene and an env vector were co-transfected into Vero E6 cells. As shown in Fig. 2, a progressive increase in the number of EGFP-positive cells and syncytium formation was observed, suggesting a successful generation of s-RCR viruses. Syncytia showing green fluorescence suggest co-expression of SARS-CoV-2 S and EGFP.
To generate stable producer cells, pCLXSN-gag-pol and pCLXSN-S env-EGFP or pCLXSN- SΔ19 env-EGFP were used to transduce HEK293 cells. A clone that was both EGFP-positive and resistant to G418 was selected. The presence of SΔ19 env in the supernatant of stable producer cells was confirmed via western blotting. Western blots of s-RCR viruses showed that most spike proteins were cleaved in the S1/S2 conformation (Fig. 3). In agreement with the results of previous studies [35], MLV pseudotyped with the C-terminal 19 aa-truncated SARS-CoV-2 S infected HEK293-hACE2 cells more efficiently than with SARS-CoV-2 S (Fig. 4). Interestingly, MLV pseudotyped with full-length spike protein also induced syncytium formation. These data suggest that two replication-deficient packageable vectors complement each other to create a semi-replication-competent virus.
Cells infected with SARS-CoV-2 viruses express the spike protein in the plasma membrane and fuse with hACE2-positive neighboring cells. In addition, the expression of SARS-CoV-2 spike protein alone (in the absence of other viral proteins) is known to induce receptor-dependent syncytia formation [8]. TMPRSS2 is involved in the fusion between spike proteins and receptors, increasing syncytia formation [7]. To examine the impact of TMPRSS2 on syncytia formation, Vero E6-TMPRSS2 cells were infected with s-RCR viruses. As shown in Fig. 5, larger syncytia were formed in Vero E6-TMPRSS2 cells than in Vero cells. These results suggest that SΔ19 env was better incorporated into s-RCR than S env and that TMPRSS2 was involved in SΔ19 env-receptor fusion. Vero E6 cells are known to highly express African green monkey ACE2 but expression level of TMPRSS2 is quite low in this clone. TMPRSS2-overexpressing Vero E6 cells are widely used to replicate and isolate SARS-CoV-2 [5]. To determine whether expression levels of hACE2 or TMPRSS2 contribute to spike-mediated syncytia formation, the construction of the 293 T-hACE2-TMPRSS2 stable cell line is required for further studies. ImageJ software can be used to measure the size and number of syncytial formations during drug screening [36].
In conclusion, an s-RCR vector that induces syncytium formation was constructed. Syncytia formation was readily detected upon infection of HEK293-hACE2 and Vero E6-TMPRSS2 cells with s-RCR viruses obtained from stable producer cells. Thus, the screening of novel fusion inhibitor drugs is possible using an s-RCR vector system and the two cell lines used in this study.
References
Lu RJ, Zhao X, Li J, Niu PH, Yang B, Wu HL et al (2020) Genomic characterisation and epidemiology of 2019 novel coronavirus: Implications for virus origins and receptor binding. Lancet 395:565–574
Hoffmann M, Kleine-Weber H, Schroeder S, Krüger N, Herrler T, Erichsen S, Schiergens TS, Herrler G, Wu NH, Nitsche A, Müller MA, Drosten C, Pöhlmann S (2020) SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181:271–280
Shang J, Ye G, Shi K, Wan Y, Luo C, Aihara H, Geng Q, Auerbach A, Li F (2020) Structural basis of receptor recognition by SARS-CoV-2. Nature 581:221–224
Li W, Moore MJ, Vasilieva N, Sui J, Wong SK, Berne MA, Somasundaran M, Sullivan JL, Luzuriaga K, Greenough TC, Choe H, Farzan M (2003) Angiotensin-converting enzyme 2 is a functional receptor for the SARS coronavirus. Nature 426:450–454
Matsuyama S, Nao N, Shirato K, Kawase M, Saito S, Takayama I et al (2020) Enhanced isolation of SARS-CoV-2 by TMPRSS2-expressing cells. Proc Natl Acad Sci USA 117:7001–7003
Harcourt J, Tamin A, Lu X, Kamili S, Sakthivel SK, Murray J et al (2020) Isolation and characterization of SARS-CoV-2 from the first US COVID-19 patient. bioRxiv. 10:914
Buchrieser J, Dufloo J, Hubert M, Monel B, Planas D, Rajah MM, Planchais C, Porrot F, Guivel-Benhassine F, Van der Werf S, Casartelli N, Mouquet H, Bruel T, Schwartz O (2020) Syncytia formation by SARS-CoV-2-infected cells. EMBO J 39:e106267
Cheng YW, Chao TL, Li CL, Wang SH, Kao HC, Tsai YM, Wang HY, Hsieh CL, Lin YY, Chen PJ, Chang SY, Yeh SH (2021) D614G Substitution of SARS-CoV-2 spike protein increases syncytium formation and virus titer via enhanced furin-mediated spike cleavage. MBio 12:e0058721
Heald-Sargent T, Gallagher T (2012) Ready, set, fuse! the coronavirus spike protein and acquisition of fusion competence. Viruses 4:557–580
Nguyen HT, Zhang S, Wang Q, Anang S, Wang J, Ding H, Kappes JC, Sodroski J (2020) Spike glycoprotein and host cell determinants of SARS-CoV-2 entry and cytopathic effects. J Virol 95:e02304-e2320
Walls AC, Park YJ, Tortorici MA, Wall A, McGuire AT, Veesler D (2020) Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell 181:281–292
Hoffmanna M, Kleine-Weber H, Pöhlmann S (2020) A multibasic cleavage site in the spike protein of SARS-CoV-2 is essential for infection of human lung cells. Mol Cell 78:779–784
Cheng YW, Chao TL, Li CL, Chiu MF, Kao HC, Wang SH et al (2020) Furin inhibitors block SARS-CoV-2 spike protein cleavage to suppress virus production and cytopathic effects. Cell Rep 33:108254
Coutard B, Valle C, de Lamballerie X, Canard B, Seidah NG, Decroly E (2020) The spike glycoprotein of the new coronavirus 2019-nCoV contains a furin-like cleavage site absent in CoV of the same clade. Antiviral Res 176:104742
Carnell GW, Ferrara F, Grehan K, Thompson CP, Temperton NJ (2015) Pseudotype-based neutralization assays for influenza: a systematic analysis. Front Immunol 6:1–17
Millet JK, Whittaker GR (2016) Murine Leukemia Virus (MLV)-based coronavirus spike-pseudotyped particle production and infection. Bio Protoc 6:e2035
Millet JK, Tang T, Nathan L, Jaimes JA, Hsu HL, Daniel S, Whittaker GR (2019) Production of pseudotyped particles to study highly pathogenic coronaviruses in a biosafety level 2 setting. J Vis Exp. https://doi.org/10.3791/59010
Moore MJ, Dorfman T, Li W, Wong SK, Li Y, Kuhn JH, Coderre J, Vasilieva N, Han Z, Greenough TC, Farzan M, Choe H (2004) Retroviruses pseudotyped with the severe acute respiratory syndrome coronavirus spike protein efficiently infect cells expressing angiotensin-converting enzyme 2. J Virol 78:10628–10635
Hu J, Gao Q, He C, Huang A, Tang N, Wang K (2020) Development of cell-based pseudovirus entry assay to identify potential viral entry inhibitors and neutralizing antibodies against SARS-CoV-2. Genes Dis 7:551–557
Schmidt F, Weisblum Y, Muecksch F, Hoffmann HH, Michailidis E, Lorenzi JCC et al (2020) Measuring SARS-CoV-2 neutralizing antibody activity using pseudotyped and chimeric viruses. J Exp Med 217:e20201181
Yang R, Huang B, Ruhan A, Li W, Wang W, Deng Y, Tan W (2020) Development and effectiveness of pseudotyped SARS-CoV-2 system as determined by neutralizing efficiency and entry inhibition test in vitro. Biosaf Health 2:226–231
Lei C, Qian K, Li T, Zhang S, Fu W, Ding M, Hu S (2020) Neutralization of SARS-CoV-2 spike pseudotyped virus by recombinant ACE2-Ig. Nat Commun 11:2070
Johnson MC, Lyddon TD, Suarez R, Salcedo B, LePique M, Graham M, Ricana C, Robinson C, Ritter DG (2020) Optimized pseudotyping conditions for the SARS-COV-2 spike glycoprotein. J Virol 94:e01062-e1120
Nie J, Li Q, Wu J, Zhao C, Hao H, Liu H, Zhang L, Nie L, Qin H, Wang M, Lu Q, Li X, Sun Q, Liu J, Fan C, Huang W, Xu M, Wang Y (2020) Establishment and validation of a pseudovirus neutralization assay for SARS-CoV-2. Emerg Microbes Infect 9:680–686
Giroglou T, Cinatl J Jr, Rabenau H, Drosten C, Schwalbe H, Doerr HW, von Laer D (2004) Retroviral vectors pseudotyped with severe acute respiratory syndrome coronavirus S protein. J Virol 78:9007–9015
Fu X, Tao L, Zhang X (2021) Comprehensive and systemic optimization for improving the yield of SARS-CoV-2 spike pseudotyped virus. Mol Ther Methods Clin Dev 20:350–356
Havranek KE, Jimenez AR, Acciani MD, Lay Mendoza MF, Reyes Ballista JM, Diaz DA, Brindley MA (2020) SARS-CoV-2 spike alterations enhance pseudo particle titers and replication-competent VSV-SARS-CoV-2 virus. Viruses 12:1465
Yurkovetskiy L, Wang X, Pascal KE, Tomkins-Tinch C, Nyalile TP, Wang Y et al (2020) Structural and functional analysis of the D614G SARS-CoV-2 spike protein variant. Cell 183:739–751
Lee ES, Jin SY, Kang BK, Jung YT (2019) Construction of replication-competent oncolytic retroviral vectors expressing R peptide-truncated 10A1 envelope glycoprotein. J Virol Methods 268:32–36
Kang BK, Jung YT (2020) Semi-replication-competent retroviral vectors expressing Gibbon Ape Leukemia Virus fusogenic membrane glycoprotein (GALV FMG) gene for cancer gene therapy. J Bacteriol Virol 50:273–281
Qiao J, Moreno J, Sanchez-Perez L, Kottke T, Thompson J, Caruso M, Diaz RM, Vile R (2006) VSV-G pseudotyped, MuLV-based, semi-replication-competent retrovirus for cancer treatment. Gene Ther 13:1457–1470
Trajcevski S, Solly SK, Frisén C, Trenado A, Cosset FL, Klatzmann D (2005) Characterization of a semi-replicative gene delivery system allowing propagation of complementary defective retroviral vectors. J Gene Med 7:276–287
Edie S, Zaghloul NA, Leitch CC, Klinedinst DK, Lebron J, Thole JF, McCallion AS, Katsanis N, Reeves RH (2018) Survey of human chromosome 21 gene expression effects on early development in danio rerio. G3 (Bethesda) 8:2215–2223
Jin SY, Jung YT (2020) Construction of replication-competent retroviral vector expressing VSV-G envelope glycoprotein for cancer gene therapy. Arch Virol 165:1089–1097
Roy S, Ghani K, de Campos-Lima PO, Caruso M (2021) A stable platform for the production of virus-like particles pseudotyped with the severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) spike protein. Virus Res 295:198305
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9(7):671–675
Acknowledgements
This work was supported by the National Research Foundation of Korea(NRF) grant funded by the Korea government(MSIT) (R2021001571).
Author information
Authors and Affiliations
Contributions
YTJ designed research and finalized the manuscript; SYL and DWK performed research and analyzed data.
Corresponding author
Ethics declarations
Conflict of interest
The author of this study has no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Edited by Simon D. Scott.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Lee, S.Y., Kim, D.W. & Jung, Y.T. Construction of SARS-CoV-2 spike-pseudotyped retroviral vector inducing syncytia formation. Virus Genes 58, 172–179 (2022). https://doi.org/10.1007/s11262-022-01890-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-022-01890-z