Abstract
Staphylococcus aureus is the most common clinical mastitis-associated pathogen in sheep which contributes to reduced welfare of affected animals and, therefore, compromises the quality and quantity of milk production. To prevent mastitis and its spread, it is essential to guarantee adequate breeding conditions and animal health, through the adoption of good farm management practices and the application of suitable biosecurity measures. Vaccination can play a strategic role in prevention, control, and eradication of diseases. The identification of secreted and cellular antigens of the predominant sheep-CC130/ST700/t1773 lineage would assist in the design of effective vaccine against mammary infections caused by S. aureus. In the current study, we carried out a 3D structural prediction analysis with the identification of the best B cell epitopes of the whole and secreted portion of S. aureus AtlA. Fragments of atlA, containing the main predicted epitopes, were amplified, cloned, and expressed in Escherichia coli for recombinant protein production. Two selected clones produced recombinant proteins (rAtl4 and rAtl8) showing strong reactivity with a hyperimmune serum against the native AtlA and with blood sera collected from sheep with clinical S. aureus mastitis. These may represent potential candidate protein-based vaccines able to elicit a protective immune response to be evaluated by vaccination and subsequent challenge of the vaccinated sheep.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Staphylococcus aureus is a major pathogen of dairy small ruminants, responsible for most of clinical intramammary infection (IMI) cases (Marogna et al. 2010; Dore et al. 2016). Along with the application of adequate flock management procedures including hygiene, biosafety measures, proper milking and equipment maintenance, regular monitoring of animals, treatment and or elimination of positive ewes, a strategic role in controlling S. aureus IMI can also be played by vaccination, as recommended by the guidelines for the prudent use of antimicrobials in veterinary medicine (2015/C 299/07). In Italy, two commercial vaccines are currently available for immunisation of small ruminants against S. aureus IMI. These are based on inactivated S. aureus strains expressing biofilm components alone or combined with alpha and beta haemolysins.
The Istituto Zooprofilattico della Sardegna (IZSSA) is authorized by Italian Health Ministry to produce an inactivated autogenous vaccine based on the S. aureus isolate involved in the outbreak, with the aim of limiting infection spread within the flock. However, vaccines based on epitopes are preferred to whole antigens for eliciting a specific immune response against viral and bacterial pathogens (Soria-Guerra et al. 2015). The production of an effective vaccine is subordinated to the knowledge of the natural S. aureus population circulating in the geographic area, since local strains may have evolved region-specific pathogenicity characteristics. In previous studies, we observed that the prevalent S. aureus lineage associated with ewe mastitis in Sardinia is CC130/ST700/t1773, a very ancestral lineage dating over five thousand years ago (Azara et al. 2017a). Our studies also demonstrated the absence of biofilm production traits in all the isolates analysed so far (Azara et al. 2017b). The absence of biofilm production genes and the presence of beta-haemolysin genes only in 44% of Sardinian isolates led us to search for alternative proteins. To this aim, we investigated the antibody response elicited in sheep with clinical mastitis by S. aureus (Longheu et al. 2020). In that study, we identified a total of 7 immunogens by immunoproteomics, including the extracellular protein autolysin A (AtlA).
AtlA is a well-known cell-surface associated multifunctional protein (Biswas et al. 2006) involved in cleaving the peptidoglycan (PG) layer and in cell separation during bacterial replication (Zoll et al. 2010). Specifically, AtlA includes two domains with hydrolytic activity, an amidase (AM) and a glucosaminidase (GM), that cleave the PG at different locations (Buttner et al. 2014). The active 62 kDa AM and 51 kDa GM proteins are generated by post-translational processing of the full-length precursor (Schlag et al. 2010). AM is the major contributor in staphylococcal pathogenesis as it binds various extracellular proteins such as fibronectin, heparin, and gelatin, thus facilitating colonization and infection (Porayath et al. 2018). Since cellular adhesion is the first step of bacterial invasion, the immunogenic AtlA protein represents a suitable candidate for sheep vaccination. In addition, the choice of this protein is strengthened by the fact that almost all (97.3%) biofilm-negative S. aureus isolates were PCR positive for atl gene (Azara et al. 2017b).
With these premises, we carried out protein structure and B-cell epitope prediction of the secreted portion of S. aureus AtlA and finally produced recombinant proteins including all the predicted epitopes of interest. These were then tested for reactivity with a hyperimmune serum against the native AtlA and then with blood sera collected from sheep with clinical S. aureus mastitis.
Materials and methods
S. aureus culture and antigen preparation
Twenty field sheep S. aureus isolates were randomly selected from our collection to verify the presence of AtlA in both pellet and culture supernatants. Isolates were grown in 5 ml of Brain Heart Infusion broth (BHI, Oxoid LTD, Basingstoke, UK) at 37 °C for 18 h with shaking. Overnight culture was pelleted by centrifugation at 1945 X g for 10 min. The pellet was resuspended in phosphate buffered saline (PBS) at the final concentration of one-tenth of the original volume while the supernatant was concentrated using Amicon® Ultra spin columns (Sigma-Aldrich, St. Louis, MO, USA) at 3800 X g for 20 min. The presence of secreted AtlA protein in the pellet and supernatant was evaluated by immunoblotting using a hyperimmune anti-secreted AtlA serum produced in lamb, according to the procedure described by Longheu et al. (2020).
Determination of S. aureus AtlA structure
The complete sequence of AtlA (1257 amino acids) was retrieved via the NCBI protein database (https://www.ncbi.nlm.nih.gov) as FASTA format. Protein structure prediction of AtlA (i.e. 1257 aa and 577 aa portions) based on the crystal structure of the homologous protein bifunctional autolysin from the S. aureus strain Mu50/ATCC 700,699 (PDB entry 6fxo) was carried out in July 2020 using two online search tools: I-TASSER (Yang et al. 2015) and In-Fold 5 (McGuffin et al. 2019). All the residues differing between the two protein sequences were then removed from each preliminary search model using CHAINSAW (Schwarzenbacher et al. 2004). Amended protein models were automatically created using BUCCANEER (Cowtan 2006). Additional manual rebuilding of the model was undertaken when required using Coot (Emsley and Cowtan 2004). Crystallographic refinement was finally carried out using REFMAC5 (Murshodov et al. 2011).
B-cell epitope prediction analysis
The following B-cell epitope prediction tools were used: (a) ElliPro (based on a protein antigen’s 3D structure - Protrusion Index method) (Ponomarenko et al. 2008); (b) BcePred (based on exposed surface - Janin method) (Saha and Raghava 2006); (c) BepiPred 2.0 (based on Jespensen method) (Jespersen et al. 2017); (d) IEDB (based on antigenity score-Kolaskar and Tongaonkar method) (Kolaskar and Tongaonkar 1990). The overall accuracy of computational results was deemed based on the level of general agreement observed among the different tools applied in parallel. The epitopes more frequently identified by most of the tools were finally selected (Table 1).
DNA extraction, primers design and cloning of selected AtlA fragments
Genomic DNA was extracted from the field sheep S. aureus isolate 34,074, according to the protocol described by Onni et al. (2012). The aminoacid sequences and the oligonucleotide primers used for recombinant protein production are indicated in Table 2. The oligonucleotide sets were designed for the directional and ORF cloning of the gene fragments by inserting engineered restriction sites in forward and reverse primers. Both PCR products and the pQE-30 expression vector (Qiagen, Chatsworth, CA, USA) were digested with BamHI and KpnI restriction enzymes. The double-digested pQE-30 plasmid was dephosphorylated with calf intestinal alkaline phosphatase (CIP, Sigma-Aldrich) for 60 min at 37 °C and purified using the Micropure-EZ enzyme removers (Sigma-Aldrich). After determining the ratio of each DNA insert to pQE vector, the ligation was performed by the T4 DNA ligase enzyme (Thermo Scientific, Vilnius, LT) at 25 °C for 1 h. Each construct was used to transform Escherichia (E.) coli DH5α cells containing pREP-4 repressor plasmid (lacIq). Cloning into pQE-30 makes it possible to produce recombinant proteins linked to a polyhistidine (His) stretch that binds strongly to nichel-chelated columns (Qiagen). pQE-30 derivatives were selected in LB agar plates supplemented under with 100 µg/ml ampicillin and 50 µg/ml kanamycin. Plasmid DNA was extracted with the Qiagen Plasmid mini kit (Qiagen).
Expression and purification of recombinant proteins
E. coli transformants containing the expected inserts were grown in LB medium until log phase and then induced with 1 mM isopropyl-thio-β-d-galactoside (IPTG) at 37 °C for 1, 3 and 18 h. After centrifugation at 4000 X g for 15 min, pellets were resuspended in PBS and analysed by SDS-PAGE and immunoblotting using anti-His serum (Qiagen), pooled serum from naturally S. aureus infected sheep (Longheu et al. 2020) and anti-AtlA hyperimmune serum, as described below. The His-tagged proteins were extracted from selected recombinant clones using the buffer B (8 M urea, 0.1 M NaH2PO4, 0.01 M Tris-HCl, pH 8.0) at room temperature for 1 h under continuous shaking agitation and then purified with the Ni-NTA Spin kit (Qiagen), according to the manufacturer’s instructions.
SDS-PAGE and immunoblotting (IB)
Whole-cell antigens from selected recombinant E. coli and each His-tagged protein were electrophoresed on 12% (w/v) polyacrylamide gels. The apparent molecular mass of recombinant proteins was determined using markers (kaleidoscope pre-stained standards, Bio-Rad, Hercules, CA, USA). Electrophoresed proteins were transferred to nitrocellulose membranes in a Trans-Blot-semidry-apparatus (Bio-Rad), as described by the manufacturer. Blots were incubated for 1 h at 37 °C with anti-His serum diluted 1:500, sheep sera diluted 1:100 in PBS-2% skim milk and anti-AtlA serum diluted 1:500. After several washings with phosphate-buffered saline with 2% skim milk, blots were incubated for 1 h at 37 °C with alkaline phosphatase-conjugated anti-mouse IgG (Sigma-Aldrich) diluted 1:30.000, or alkaline phosphatase-conjugated anti-sheep antibody (Sigma-Aldrich) diluted 1:10.000. After three more washes, blots were developed with bromochloroindolyl phosphate/nitroblue tetrazolium (BCIP/NBT, Promega, Madison, WI) in alkaline phosphatase buffer (100 mM NaCl, 5 mM MgCl2, 100 mM Tris, pH 9.5).
Results
Detection of secreted AtlA protein in sheep S. aureus isolates
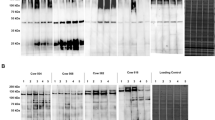
The anti-secreted AtlA serum was tested against the concentrated supernatant and cellular proteins obtained from twenty sheep S. aureus field isolates. As shown in Fig. 1, the hyperimmune serum detected the secreted-AtlA in all field isolates, both in the cellular pellet and in the secreted protein fractions.
Western Immunoblotting reactivity of anti-AtlA antibodies on the secreted (panel A) and cellular proteins (panel B) of field S. aureus isolates. Immune reactivity was observed on isolates 2089 (lane 1), 1495 (lane 2), 2620 (lane 3), 9625 (lane 4), 4902 (lane 5), 4992 (lane 6), 4460 (lane 7), 3817 (lane 8), and 34,074 (lane 9). Lane M, kaleidoscope protein standards (Bio-Rad).
Protein modelling and epitope prediction analysis of AtlA
The complete sequence of Atl (1257 amino acids) available in the NCBI protein database (https://www.ncbi.nlm.nih.gov) and used for this study is reported in Fig. 2, with the previously sequenced 577 aminoacid secreted portion (Longheu et al. 2020) indicated in red. Several algorithms based on different physico-chemical properties including hydrophilicity, flexibility, accessibility, polarity, and exposed surface have increasingly become available over the last decade for predicting the continuous and discontinuous B-cell epitopes (Soria-Guerra et al. 2015). Accordingly, we first used available tools to predict AtlA protein models for both 1257 aa and 577 aa portions based on the crystal structure of the homologous protein bifunctional autolysin from the S. aureus strain Mu50/ATCC 700,699 (PDB entry 6fxo). Linear B-cell prediction analysis was then carried out using 4 individual tools such as Ellipro, BcePred, BepiPred2 and IEDB applied in parallel (see material and methods). Predicted epitopes are indicated in grey in Fig. 2 and detailed in Table 1.
Amino acid sequence of full-length (in black) and secreted portion (in red) of AtlA (NCBI protein, Accession number: WP_001074519.1). Predicted epitopes are highlighted in grey. Forward and reverse primers were designed into the bold green and purple areas, respectively. Primer sequences are detailed in Table 2
General features of S. aureus AtlA and position of epitopes
The S. aureus AtlA protein (Accession number: WP_001074519.1) consists of 1257 a.a. with a molecular weight of 137.44 kDa. The secreted protein (577 a.a.), containing the N-acetylmuramoyl-L-alanine-amidase (314 a.a.) fragment, has a molecular mass of about 64 kDa. The three-dimensional (3D) structure of whole and secreted protein is shown in Fig. 3, while the spatial position of selected top scoring epitopes is shown in Fig. 4.
Ribbon diagram representation of Atl protein complete sequence 1257 aa depicted using PyMOL version 2.5.1 (Schrödinger Inc, USA). (A) Protein structure is coloured using a rainbow colour scheme starting from blue for the N-terminal portion then gradually switching to red as approaching the C-terminal extremity of the molecule. (B) Spatial position of secreted AtlA protein (577 aa) in the crystal structure of Atl protein complete sequence (1257aa). Sequenced portions of the secreted AtlA protein (577 aa) are represented in red. Remaining portions of Atl protein complete sequence (1257 aa) are coloured in grey
Ribbon diagram (A, C, E) and orthographic molecular surface representation (B, D, F) of Atl protein complete sequence (1257 aa) and spatial position of selected top scoring epitopes. A and B show the right side of the protein; C and D represent a view from the back obtained by rotating the protein anti-clockwise by 90 degrees; E and F show a view from the top. Not relevant epitopes were identified in the front of the protein (i.e. not included in this section). Epitope areas represented in the three sides of the protein include: Atl4-20 (dark blue); Atl34-46 (light blue); Atl181-225 (red); Atl399-409 (magenta); Atl427-436 (yellow) and Atl469-520 (pale yellow)
Recombinant protein production
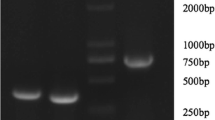
Figure 5 shows the five 240, 261, 492, 753 and 1050-bp fragments of AtlA gene amplified from genomic S. aureus isolate 34,074 DNA and cloned into pQE-30 plasmid inserted in E. coli DH5α.
Agarose electrophoresis of PCR products from the Atl gene of S aureus isolate 34,074. Amplicon sizes were listed in the Table 2. Lane 1, amplicon obtained with primers 1 + 4 (240 bp); lane 2, amplicon obtained with primers 1 + 5 (753 bp); lane 3, amplicon obtained with primers 1 + 6 (1050 bp); lane 4, amplicon obtained with primers 2 + 5 (261 bp); lane 5, amplicon obtained with primers 3 + 6 (492 bp), lane 6, undigested pQE; lane 7, pQE digested with BamHI and KpnI enzymes; M, Marker VI (Roche); M1, Marker II (Roche)
All recombinant E. coli clones derivated from the 5-PCR products were analyzed by SDS-PAGE and IB. Only two clones, containing the 492-bp (clone 4) and 753-bp (clone 8) inserts respectively, produced recombinant proteins that reacted with anti-His serum, pooled sheep sera and anti-AtlA serum (Fig. 6).
Panels A and B, Coomassie blue-stained SDS-PAGE of the two E. coli clones containing the 492-bp (clone 4) and the 753-bp (clone 8) inserts, respectively. Lane 1, E. coli culture without IPTG induction; lane 2, E. coli culture after 1 h of IPTG induction; lane 3, after 3 h of induction; lane 4, after overnight of IPTG induction. M, kaleidoscope protein standards (Bio-Rad). Panels A1, A2 and A3, IB analysis showing reactivities of anti-His serum (A1), pooled sera from naturally infected sheep (A2) and, anti-AtlA serum (A3) with whole-cell E.coli (clone 4) w/o and with IPTG induction. Panels B1, B2 and B3, IB analysis showing reactivities of anti-His serum (B1), pooled sera from naturally infected sheep (B2) and, anti-AtlA serum (B3) with whole-cell E.coli (clone 8) w/o and with IPTG induction. Lane C, positive controls: recombinant protein with a poly-His-tag (panels A1- B1), proteins from S. aureus supernatant (panels A2- B2 and A3-B3). M, kaleidoscope protein standards (Bio-Rad).
The two His-tagged proteins, named rAtl-4 and rAtl-8, presented a molecular mass of approximately 23 and 35 kDa, respectively; more larger than the predicted masses of 18 and 27 kDa. Both recombinant proteins, extracted under denaturing conditions and purified by metal-affinity chromatography, confirm the strong reactivity with the three tested sera (Fig. 7).
Purification and antigenicity characterization of recombinant AtlA-clone 8 (rAtl-8) and recombinant AtlA-clone 4 (rAtl-4). Panel C, Coomassie blu-stained SDS-PAGE gel with purified rAtl-8 (lane 1) and rAlt-4 (lane 2) from E. coli DH5α. Panel C1, Western blotting (WB) results with anti-6His; Panel C2, WB of rAtl-8 and rAlt-4 reactivity with the pooled sera from naturally infected sheep and anti-AtlA serum (panel C3). Lane C, positive controls: recombinant protein with a poly-His-tag (panel C1), proteins from S. aureus supernatant (panels C2 and C3). M, kaleidoscope protein standards (Bio-Rad).
Discussion
The Italian region of Sardinia produces the 68.92% and 57.30% of all Italian sheep and goat milk, respectively. Furthermore, 10% of the sheep milk collected in Europe is Sardinian. The regional production of sheep and goat cheese is estimated at 60,000 tons, of which about 30,000 are Protected Designation of Origin (DOP) cheese (https://www.sardegnaagricoltura.it/index.php?xsl=443&s=413001&v=2&c=6039&vd=1). Mastitis is one of the main causes of the decrease in the efficiency of milk production with consequent repercussions on the dairy industry sector. To prevent mastitis and its spread, and consequently reduce the use of antibiotics, it is essential to guarantee adequate breeding conditions and animal health, placing the emphasis on the adoption of good farm management practices and on the application of adequate biosecurity measures. In the control of mastitis, a strategic role can be played also by vaccination. Vaccines are essential for elicing the immune response and protecting against disease (Vasileiou et al. 2022).
Unfortunately, there is a lack of knowledge regarding the major immunogenic antigens associated with each lineage and which antigens might provide protection against heterologous isolates. In our recent study, we identified the major dominant antigens associated with the CC130/ST700/t1773 S. aureus ancestral lineage by an immunoproteomic approach (Longheu et al. 2020). Here, among the secreted and cellular antigens that had functional annotation, we selected the immunogenic AtlA protein, present in almost all the analyzed isolates (Longheu et al. 2020), as a suitable candidate for the development of recombinant proteins to be used for vaccination against S. aureus mastitis. Protein-based vaccines have been found to have all the necessary components to initiate T cell-dependent activation of B cells, a process characterized by a more robust immune response, affinity maturation, immunological memory and, simultaneously maintain a good safety profile (Vartak and Sucheck 2016). The bioinformatics approach can perform an appropriate in silico selection of epitopes for protein-based vaccine (Saylor et al. 2020). In the present study, computation analysis, one of the most important bioinformatics branches, was used to analyze both the whole and the AM secreted portion of the S. aureus AtlA, providing information on the main predicted epitopes and enabling the production of recombinant proteins including these epitopes. Six top-scoring epitope areas within the AM portion were predicted with the highest likelihood of proper epitope presentation, orientation, and exposure. These epitopes were cloned and expressed in E. coli, which is one of the earliest and most widespread hosts for the production of heterologous proteins (Terpe 2006). The advantages of this system include ease of cultivation, rapid growth and expression, high product yields and productivity, and low-cost production. It is used for massive production of many commercialized proteins, in particular non-glycosylated proteins (Cid and Bolivar 2021). Post-translational modifications play an important role in protein folding, processing, and stability, as well as biological activity and even the immunogenity/immunoreactivity of the protein (Walsh and Jefferis 2006). To overcome these disadvantages, we only selected the clones that produce recombinant proteins capable of binding the antibodies from naturally infected sheep, anti-AtlA and, anti-His sera.
Of the 5 constructed DNA fragments, only clones 4 and 8, however containing all the 6 epitopes, produced strongly immunoreactive proteins (rAtl4 and rAtl8). Using blood sera collected from sheep with clinical S. aureus mastitis (Longheu et al. 2020), we demonstrated that these epitopes are effectively recognized by naturally infected sheep.
Atl mediates adherence of S. aureus and exerts peptidoglycal hydrolase activity with associated amidase and glucosaminidase domains. Amidase and glucosaminidase, however, can also bind to host matrices, including fibronectin, thrombospondin 1, vitronectin, and Hsc70, as well as heparin and gelatine (Porayath et al. 2018). Therefore, antibodies against this protein may play an important role in reducing S. aureus adhesion, especially when considering that according to our studies biofilm production traits are lacking in all the Sardinian isolates analysed so far (Azara et al. 2017b). Interestingly, it was recently demonstrated that Atl regulates the virulence of S. aureus by controlling the sorting of pore-forming leukocidins (Zheng et al. 2022). These toxins mediate leukocyte killing and play a major role in S. aureus pathogenesis (Zheng et al. 2021). Previous studies by our group demonstrated that ovine S. aureus produce and release significant amounts of lukF-PV/lukM in the extracellular milieu (Longheu et al. 2020), and that the abundance of leukocidin production might be related to the severity of clinical mastitis caused by S. aureus in dairy cows (Addis et al. 2022). Therefore, by eliciting the production of neutralizing antibodies, vaccination against Atl might also interfere with this regulatory mechanism, enabling the reduction of S. aureus virulence and of clinical mastitis severity. This acquires a particular relevance when considering that most clinical mastitis cases in small ruminants are caused by S. aureus (Marogna et al. 2010; Machado 2018). There are mainly three S. aureus lineages associated with ovine mastitis: CC133, CC130 e CC522. The first was predominant in Denmark (Eriksson et al. 2013), Netherlands (Hoekstra et al. 2019), Switzerland (Merz et al. 2016); the second in Italy (Azara et al. 2017a), Algeria (Azzi et al. 2020) whereas the third in Spain (Porrero et al. 2012). In dairy cows, vaccination against S aureus mastitis has been associated with reduced clinical severity and duration of clinical disease post-challenge (Middleton et al. 2006). Furthermore, reduced transmission within the herd was observed upon vaccination (Schukken et al. 2014). Accordingly, although vaccination strategies may not be completely successful in preventing S. aureus infection according to studies carried out in dairy cows (Rainard et al. 2022), these may reduce the incidence and severity of S. aureus IMI. When considering that most clinical mastitis cases in sheep are due to S. aureus, vaccination may indeed represent a more successful strategy for reducing the impact of mastitis in this dairy animal, in combination with the application of good mastitis prevention protocols, controlling the level of infection in the flock, and monitoring the type of circulating strains.
Conclusion
Developing a vaccine that can prevent S. aureus mastitis in small ruminants is a major challenge. In the current study, we have assessed in silico prediction metodologies for identifying the best B cell epitopes within the AtlA protein. In light of the findings obtained with the AtlA protein, further work is being undertaken to extend the computational analysis and the production of recombinant proteins to the other antigens identified in non-biofilm-producing S. aureus (Longheu et al. 2020). Then, the ability of candidate protein-based vaccine to elicit a protective immune response could be evaluated by vaccination and subsequent challenge of the vaccinated sheep.
Data availability
All data generated or analyzed during this study are includes within the manuscript.
References
Addis MF, Pisanu S, Monistero V, Gazzola A, Penati M, Filipe J, Di Mauro S, Cremonesi P, Castiglioni B, Moroni P, Pagnozzi D, Tola S, Piccinini R (2022) Comparative secretome analysis of Staphylococcus aureus strains with different within-herd intramammary infection prevalence. Virulence 13:170–190. https://doi.org/10.1080/21505594.2021
Azara E, Piras MG, Parisi A, Tola S (2017a) Antimicrobial susceptibility and genotyping of Staphylococcus aureus isolates collected between 1986 and 2015 from ovine mastitis. Vet Microbiol 205:53–56. https://doi.org/10.1016/j.vetmic.2017.05.006
Azara E, Longheu C, Sanna G, Tola S (2017b) Biofilm formation and virulence factor analysis of Staphylococcus aureus isolates collected from ovine mastitis. J Appl Microbiol 123:372–379. https://doi.org/10.1111/jam.13502
Azzi O, Lai F, Tennah S, Menoueri MN, Achek R, Azara E, Tola S (2020) Spa-typing and antimicrobial susceptibility of Staphylococcus aureus isolated from clinical sheep mastitis in Médéa province, Algeria. Small Rumin Res 192:10168. https://doi.org/10.1016/j.smallrumres.2020.106168
Biswas R, Voggu L, Simon UK, Hentschel P, Thumm G, Gotz F (2006) Activity of the major staphylococcal autolysin atl. FEMS Microbiol Lett 259:260–268. https://doi.org/10.1111/j.1574-6968.2006.00281.x
Buttner FM, Zoll S, Nega M, Gotz F, Stehle T (2014) Structure-function analysis of Staphylococcus aureus amidase reveals the determinants of peptidoglycan recognition and cleavage. J Biol Chem 289(16):11083–11094. https://doi.org/10.1074/jbc.M114.557306
Cid R, Bolivar J (2021) Platforms for production of protein-based vaccines: from classical to next-generation strategies. Biomolecules 11:1072. https://doi.org/10.3390/biom11081072
Cowtan K (2006) The buccaneer software for automated model building. 1. Tracing protein chains. Acta Crystallogr D Biol Crystallogr 62:1002–1011. https://doi.org/10.1107/S0907444906022116
Dore S, Liciardi M, Amatiste S, Bergagna S, Bolzoni G, Caligiuri V et al (2016) Survey on small ruminant bacterial mastitis in Italy, 2013–2014. Small Rumin Res 141:91–93. https://doi.org/10.1016/j.smallrumres.2016.07.010
Emsley P, Cowtan K (2004) Coot: model-building tools for molecular graphics. Acta Crystallogr D Biol Crystallogr 60:2126–2132. https://doi.org/10.1107/S0907444904019158
Eriksson J, Espinosa-Gongora C, Stamphøj I, Larsen AR, Guardabassi L (2013) Carriage frequency, diversity and methicillin resistance of Staphylococcus aureus in danish small ruminants. Vet Microbiol 163:110–115. https://doi.org/10.1016/j.vetmic.2012.12.006
Hoekstra J, Rutten VPMG, van den Hout M, Spaninks MP, Benedictus L, Koop G (2019) Differences between Staphylococcus aureus lineages isolated from ovine and caprine mastitis but not between isolates from clinical or subclinical mastitis. J Dairy Sci 102:5430–5437. https://doi.org/10.3168/jds.2018-16196
Jespersen MC, Peters B, Nielsen M, Marcatili P (2017) BepiPred-2.0: improving sequence-based B-cell epitope prediction using conformational epitopes. Nucleic Acids Res 45:24–29. https://doi.org/10.1093/nar/gkx346
Kolaskar AS, Tongaonkar PC (1990) A semi-empirical method for prediction of antigenic determinants on protein antigens. FEBS Lett 276:172–174. https://doi.org/10.1016/0014-5793(90)80535-q
Longheu CM, Azara E, Marogna G, Addis MF, Tola S (2020) Identification of secreted and cellular antigens of Staphylococcus aureus causing dairy sheep mastitis and their potential for vaccine development. Vet Immun Immunopathol 230:110149. https://doi.org/10.1016/j.vetimm.2020.110149
Machado GP (2018) Mastitis in small ruminants. Anim Husb Dairy Vet Sci 2:1–9. https://doi.org/10.15761/AHDVS.1000144
Marogna G, Rolesu S, Lollai S, Tola S, Leori SG (2010) Clinical findings in sheep farms affected by recurrent bacterial mastitis. Small Rumin Res 88:119–125. https://doi.org/10.1016/j.smallrumres.2009.12.019
McGuffin LJ, Adiyaman R, Maghrabi AHA, Shuid AN, Brackenridge DA, Nealon JO et al (2019) IntFOLD: an integrated web resource for high performance protein structure and function prediction. Nucleic Acids Res 47:408–413. https://doi.org/10.1093/nar/gkz322
Merz A, Stephan R, Johler S (2016) Staphylococcus aureus isolates from goat and sheep milk seem to be closely related and differ from isolates detected from bovine milk. Front Microbiol 7:1–7. https://doi.org/10.3389/fmicb.2016.00319
Middleton JR, Ma J, Rinehart CL, Taylor VN, Luby CD, Steevens BJ (2006) Efficacy of different Lysigin™ formulations in the prevention of Staphylococcus aureus intramammary infection in dairy heifers. J Dairy Res 73:10–19. https://doi.org/10.1017/S0022029905001354
Murshodov GN, Skubak P, Lebedev AA, Pannu NS, Steiner RA, Nicholls RA et al (2011) REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr D Biol Crystallogr 67:355–367. https://doi.org/10.1107/S0907444911001314
Onni T, Vidili A, Bandino E, Marogna G, Schianchi G, Tola S (2012) Identification of coagulase-negative staphylococci isolated from caprine milk samples by PCR-RFLP of groEL gene. Small Rumin Res 104:185–190. https://doi.org/10.1016/j.smallrumres.2011.10.004
Ponomarenko JV, Bui H, Li W, Fusseder N, Bourne PE, Sette A, Peters B (2008) ElliPro: a new structure-based tool for the prediction of antibody epitopes. BMC Bioinformatics 9:514. https://doi.org/10.1186/1471-2105-9-514
Porayath C, Suresh MK, Biswas R, Nair BG (2018) Autolysin mediated adherence of Staphylococcus aureus with fibronectin, gelatin and heparin. Int J Biol Macromol 110:179–184. https://doi.org/10.1016/j.ijbiomac.2018.01.047
Porrero MC, Hasman H, Vela AI, Fernandez-Garayza JF, Domınguez L, Aarestrup FM (2012) Clonal diversity of Staphylococcus aureus originating from the small ruminants goats and sheep. Vet Microbiol 156:157–161. https://doi.org/10.1016/j.vetmic.2011.10.015
Rainard P, Gilbert FB, Martins RP, Germon P, Foucras G (2022) Progress towards the elusive mastitis vaccines. Vaccines 10:296. https://doi.org/10.3390/vaccines10020296
Saha S, Raghava GP (2006) BcePred: prediction of continuous B-cell epitopes in an antigen using recurrent neural network. Proteins 1:40–48. https://doi.org/10.1002/prot.21078
Saylor K, Gillam F, Lohneis T, Zhang C (2020) Designs of antigens structure and composition for improved protein-based vaccine efficacy. Front Immunol 11:283. https://doi.org/10.3389/fimmu.2020.00283
Schlag M, Biswas R, Krismer B, Kohler T, Zoll S, Yu W et al (2010) Role of staphylococcal wall teichoic acid in targeting the major autolysin atl. Mol Microbiol 75:864–873. https://doi.org/10.1111/j.1365-2958.2009.07007.x
Schukken YH, Bronzo V, Locatelli C, Pollera C, Rota N, Casula A, Testa F, Scaccabarozzi L, March R, Zalduendo D, Guix R, Moroni P (2014) Efficacy of vaccination on Staphylococcus aureus and coagulase-negative staphylococci intramammary infection dynamics in 2 dairy herds. J Dairy Sci 97(8):5250–5264. https://doi.org/10.3168/jds.2014-8008
Schwarzenbacher R, Godzik A, Grzechnik SK, Jaroszewski L (2004) The importance of alignment accuracy for molecular replacement. Acta Crystallogr D Biol Crystallogr 60:1229–1236. https://doi.org/10.1107/S0907444904010145
Soria-Guerra RE, Nieto-Gomez R, Govea-Alonso DO, Rosales-Mendoza S (2015) An overview of bioinformatics tools for epitope prediction: implications on vaccine development. J Biom Informatics 53:405–114. https://doi.org/10.1016/j.jbi.2014.11.003
Terpe K (2006) Overview of bacterial expression systems for heterologous protein production: from molecular and biochemical fundamentals to commercial systems. Appl Microbiol Biotechnol 72:211–223. https://doi.org/10.1007/s00253-006-0465-8
Vartak A, Sucheck SJ (2016) Recent advances in subunit vaccine carriers. Vaccines 4:E12. https://doi.org/10.3390/vaccines4020012
Vasileiou NGC, Lianou DT, Michael CK, Fthenakis GC, Mavrogianni VS (2022) Vaccination against bacterial mastitis in sheep. Vaccines 10:2088. https://doi.org/10.3390/vaccines10122088
Walsh G, Jefferis R (2006) Post-translational modifications in the context of therapeutic proteins. Nat Biotechnol 24:1241–1252. https://doi.org/10.1038/nbt1252
Yang J, Yan R, Roy A, Xu D, Poisson J, Zhang Y (2015) The I-TASSER suite: protein structure and function prediction. Nat Methods 12:7–8. https://doi.org/10.1038/nmeth.3213
Zheng X, Marsman G, Lacey KA, Chapman JR, Goosmann C, Ueberheide BM, Torres VJ (2021) The cell envelope of Staphylococcus aureus selectively controls the sorting of virulence factors. Nat Commun 12:6193. https://doi.org/10.1038/s41467-021-26517-z
Zheng X, Ma SX, St. John A, Torres VJ (2022) The major autolysin atl regulates the virulence of Staphylococcus aureus by controlling the sorting of LukAB. Infect Immun 90:e00056–e00022. https://doi.org/10.1128/iai.00056-22
Zoll S, Patzold B, Schlag M, Gotz F, Kalbacher H, Stehle T (2010) Structural basis of cell wall cleavage by a staphylococcal autolysin. PLoS Pathog 6:e1000807. https://doi.org/10.1371/journal.ppat.1000807
Funding
This work was supported by a grant from the Italian Health Ministry (Ricerca Corrente IZS SA 05/19).
Author information
Authors and Affiliations
Contributions
E.A. and C.M.L.: carried out production of recombinant proteins; performed the experiments; analyzed the data. A.C.F.: undertook protein structure and B-cell epitope prediction analyses. M.F.A.: interpreted the data; drafted and revised the manuscript. S.: conceived the study, analyzed, and interpreted the data; drafted the manuscript; supervised the project. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Ethics approval
No ethical approval was required.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Azara, E., Foddai, A.C., Longheu, C.M. et al. Production of recombinant proteins including the B-cell epitopes of autolysin A of Staphylococcus aureus isolated from clinical sheep mastitis and their potential for vaccine development. Vet Res Commun 47, 1665–1674 (2023). https://doi.org/10.1007/s11259-023-10121-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11259-023-10121-1