Abstract
The aim of this study was to determine the prevalence of the htrA, htrB and ppk1 genes -all of which are related to environmental persistence- in C. jejuni and C. coli isolates obtained from abattoir samples at the arrival of broilers (initial stage) and in meat products after processing (final stage). A total of 119 DNA extracts (55 C. jejuni and 64 C. coli) were included in the study. Identification of genes was performed by conventional PCR (one for each gene). The overall prevalence was 40.3%, 93.3% and 68.9% for the htrA, htrB and ppk1 genes, respectively. Statistically significant differences were found (p < 0.05) between prevalence of C. jejuni and C. coli for all three genes. In C. coli the prevalence was significantly higher for the htrA (p = 0.007) and htrB (p = 0.015) genes, while ppk1 gene prevalence was significantly higher in C. jejuni (p < 0.001). In addition, statistically significant increase in the frequency of htrA (p = 0.007) and htrB (p = 0.013) genes in the final product compared to broilers on arrival at the abattoir was observed in C. jejuni, but not in C. coli. These results suggest that htrA and htrB genes are involved in environmental persistence of Campylobacter jejuni.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In 2018, 246,571 cases of human campylobacteriosis were confirmed in the European Union. It is the most reported zoonosis since 2005 (EFSA and OCDE 2019). The species most frequently implicated in campylobacteriosis outbreaks are Campylobacter jejuni and Campylobacter coli (around 90% and 10% of the cases, respectively) (Wagenaar et al. 2013; Mossong et al. 2016). There are many domestic and wild reservoirs of Campylobacter species, but poultry are considered the main ones (Wagenaar et al. 2013; Josefsen et al. 2015). Meat contamination by Campylobacter spp. usually increases in broiler abattoirs along the processing (Ghafir et al. 2007; Torralbo et al. 2015). The main reason is the cross contamination that happens more frequently during scalding, evisceration and quartering stages (Torralbo et al. 2015; Seliwiorstow et al. 2016).
Bacteria belonging to Campylobacter genus are markedly labile in the external environment. Several conditions, such as the presence of oxygen, variations in pH or temperature, dryness and chemical products, can produce cell damage (Silva et al. 2011; Hofreuter 2014). However, a higher resistance of C. coli when compared with C. jejuni during processing at abattoir has been reported (Ma et al. 2014; Oh et al. 2015; Vinueza-Burgos et al. 2017).
The presence and the expression of some genes, such as htrB, htrA and ppk1, have been associated with a greater resistance to adverse environmental conditions. Previous studies about Campylobacter jejuni attributed different functions to htrA gene, such as greater tolerance to oxygen, resistance to thermal shock or invasion and colonization functions (Hanning et al. 2010; Baek et al. 2011; Boehm et al. 2015). Regarding htrB gene, it has an important role in promoting Campylobacter resistance against changes in pH, thermal stress, aerobic media and osmotic stress (Phongsisay et al. 2007; Bronowski et al. 2014). Finally, the ppk1 gene participates in the expression of poly-P, an inorganic polyphosphate responsible for the responses of the bacterium to conditions of lack of nutrients and osmotic stress, as well as the formation of biofilms (Candon et al. 2007; Drozd et al. 2014).
The objective of this study was to confirm the next hypothesis: if htrA, htrB and ppk1 genes are involved in environmental persistence of C. jejuni and C. coli, isolates having these genes will be selected under adverse conditions, such as those present during the processing at abattoir. Consequently, prevalence of isolates with these genes after processing would be significantly higher compared with prevalence before processing.
Material and methods
DNA isolates from a previous study (Torralbo et al. 2015) performed by our group were used. Sampling of the study was carried out along 2012 in a slaughterhouse located near Malaga (Spain). In this abattoir about 60,000 chickens are slaughtered and quartered each day. It is divided in six areas/stages: loading dock, scalding, evisceration, classified, quartering and final meat product/packing. Only DNA extracts from samples obtained in the first (cloacae from broilers in the loading dock) and the last stage (quartered carcasses surfaces (breast, wing, leg and back)) were used in this study.
After sampling, swabs were cultured in Campylobacter Free Blood Agar (Oxoid®) with CCDA supplement (Oxoid®). Bulk colonies for each isolate in blood agar plate were taken with a loopful of 10 µl to carry out DNA extraction according to the method described by Sambrook and Russell (2006). Phenotypic characterization using catalase and oxidase test, Gram stain for shape observation and mobility test under dark light microscopy, were performed in a first stage. Compatible Campylobacter isolates were confirmed by Multiplex PCR (Torralbo et al. 2015). Fifty-five from 86 C. jejuni isolates and 64 from 80 C. coli isolates were randomly selected for this study. Forty-four isolates (31 C. jejuni and 13 C. coli) were obtained from the initial stage (cloacae from broilers in the loading dock) and 75 (24 C. jejuni and 51 C. coli) from the final stage (meat ready for retail). The difference in the number of isolates selected from the initial and final stage is due to the greater number of isolates obtained in the final stage in the previous study from which Campylobacter was isolated.
Conventional PCR was used to identify the genes related to environmental persistence htrA, htrB and ppk1. The strain of C. coli DSMZ 4689 was used as positive control. The PCR protocols described by Nierop Groot et al. (2014), Ghunaim et al. (2015), and Candon et al. (2007) were used to identify the htrA, htrB and ppk1 genes, respectively. Information about the molecular weight of the PCR products and sequences of primers is shown in the Table 1. The polymerase kit MyTaq Red Mix®, from Bioline, which includes the polymerase, the nucleotides, the magnesium chloride and the load buffer was used to perform the PCR.
SPSS software (v. 22.0) was used to perform the statistical analyses. The initial, final and global prevalence of each gene was calculated for C. jejuni and C. coli. In addition, a Chi Squared test (χ2) was performed to establish the existence of significant differences between the prevalence of the genes in the initial and the final stage (p < 0.05). The same test was used to determine the existence of significant differences, at 95% confidence level, between the prevalence of C. jejuni and C. coli.
The procedures employed were ethically reviewed and approved by the bioethical committee of the University of Cordoba. The procedures employed in the submitted manuscript were performed only at laboratory level, in spaces approved by the biosecurity committee of the University of Cordoba.
Results
The overall prevalence of the htrA, htrB and ppk1 genes was 40.3% (48/119), 93.3% (111/119) and 68.9% (82/119), respectively.
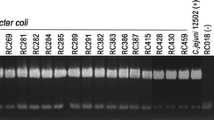
Table 2 shows the prevalence of the different genes in the initial and final stage for both Campylobacter species, C. jejuni and C. coli. Prevalence of htrA and htrB is significantly higher (p < 0.05) in the final stage when both species where included together. Significant differences are appreciated for htrA and htrB genes in C. jejuni, but not for C. coli. PCR examples for detection of C. jejuni and C. coli can be found in Figs. 1 and 2, respectively.
In Table 3 the prevalence of htrA, htrB and ppk1 genes in C. jejuni and C. coli is compared, showing that the prevalence of htrA and htrB genes is significantly higher in C. coli while prevalence of ppk1 gene is significantly higher in C. jejuni.
Discussion
In the present study we tried to support the hypothesis that htrA, htrB and ppk1 genes are involved in the environmental persistence of Campylobacter species. To the best of our knowledge, their role had only been evaluated by mean of laboratorial studies, but no previous field studies have been performed.
The main hypothesis in our study was that if these genes are involved in environmental persistence, their prevalence should increase since the strains having these genes would be positively selected during the stressful conditions existing along the processing at the abattoir. It only could be confirmed for htrA and htrB in C. jejuni at a confidence level of 95%. Differences in ppk1 for C. jejuni would be significant decreasing the signification level (p = 0.055).
Our results show that the three genes probably play a role in the environmental persistence of C. jejuni, but not in C. coli (Table 2). In contrast, a similar study elaborated by Hanning et al. (2010) with different genes related to the bacterial survival (racR, sobB, htrA and clpA), no association with actual survival was observed.
The last part of our study consisted of the comparison of the prevalence of these three genes between C. jejuni and C. coli at the first and last stage in the slaughterhouse. A significant increase in the prevalence of htrA (from 27.3 to 51.6%) and htrB (from 87.3 to 98.4%) was observed in C. coli (Table 3). The higher prevalence of these genes could explain the results reported by Torralbo et al. (2015), who described an increase in the relative prevalence of C. coli (compared with that one by C. jejuni) along the processing from the initial phase (41.5%) to the final stage (64.6%). There are also other studies describing a greater presence of C. coli at the end of processing at the slaughterhouse (Ma et al. 2014; Vinueza-Burgos et al. 2017).
On the other hand, the prevalence of ppk1 gene was significantly higher in C. jejuni (from 54.7 to 85.5%) (Table 3). This gene has been mainly related to the formation of biofilms, which has been extensively described in C. jejuni (Candon et al. 2007; Drozd et al. 2014), but scarcely for C. coli.
Resistance could be related not only to the presence of the gene but also with its level of expression. However, conventional PCR only provide information about the presence. Therefore, it would be convenient in the future to carry out novel studies using quantitative techniques (detection of DNA by qPCR or quantification of mRNA). It would be interesting also to continue the study including human isolations to evaluate if the presence of these genes is associated with an increase of human cases.
As conclusion, it has been observed that htrA, htrB and ppk1 genes are present in the isolates of C. jejuni and C. coli studied with medium to high prevalence. The prevalence of htrA and htrB significantly increased along the processing in C. jejuni, but not in C. coli. However, the prevalence of these two genes was significantly higher in C. coli. In contrast, the prevalence of ppk1 gene was significantly higher in C. jejuni.
Data Availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Baek KT, Vegge CS, Skórko-Glonek J, Brøndsted L (2011) Different contributions of HtrA protease and chaperone activities to Campylobacter jejuni stress tolerance and physiology. Appl Environ Microbiol 77(1):57–66. https://doi.org/10.1128/AEM.01603-10
Boehm M, Lind J, Backert S, Tegtmeyer N (2015) Campylobacter jejuni serine protease HtrA plays an important role in heat tolerance, oxygen resistance, host cell adhesion, invasion, and transmigration. European Journal of Microbiology & Immunology 5(1):68–80. https://doi.org/10.1556/EUJMI-D-15-00003
Bronowski C, James CE, Winstanley C (2014) Role of environmental survival in transmission of Campylobacter jejuni. FEMS Microbiol Lett 356(1):8–19. https://doi.org/10.1111/1574-6968.12488
Candon HL, Allan BJ, Fraley CD, Gaynor EC (2007) Polyphosphate kinase 1 is a pathogenesis determinant in Campylobacter jejuni. J Bacteriol 189(22):8099–8108. https://doi.org/10.1128/JB.01037-07
Drozd M, Chandrashekhar K, Rajashekara G (2014) Polyphosphate-mediated modulation of Campylobacter jejuni biofilm growth and stability. Virulence 5(6):680–690. https://doi.org/10.4161/viru.34348
EFSA (European Food Safety Authority) and ECDC (European Centre for Disease Prevention and Control) (2019). The European Union Summary Report on Trends and Sources of Zoonoses, Zoonotic Agents and Food-borne Outbreaks in 2018. EFSA Journal 17. https://doi.org/10.2903/j.efsa.2019.5926
Ghafir Y, China B, Dierick K, De Zutter L, Daube G (2007) A seven-year survey of Campylobacter contamination in meat at different production stages in Belgium. Int J Food Microbiol 116(1):111–120. https://doi.org/10.1016/j.ijfoodmicro.2006.12.012
Ghunaim H, Behnke JM, Aigha I, Sharma A, Doiphode SH, Deshmukh A, Abu-Madi MM (2015) Analysis of resistance to antimicrobials and presence of virulence/stress response genes in Campylobacter isolates from patients with severe diarrhoea. PLoS ONE 10(3):e0119268. https://doi.org/10.1371/journal.pone.0119268
Hanning I, Biswas D, Herrera P, Roesler M, Ricke SC (2010) Prevalence and characterization of Campylobacter jejuni isolated from pasture flock poultry. J Food Sci 75(7):M496-502. https://doi.org/10.1111/j.1750-3841.2010.01747.x
Hofreuter D (2014) Defining the metabolic requirements for the growth and colonization capacity of Campylobacter jejuni. Front Cell Infect Microbiol 4:137. https://doi.org/10.3389/fcimb.2014.00137
Josefsen MH, Bhunia AK, Engvall EO, Fachmann MS, Hoorfar J (2015) Monitoring Campylobacter in the poultry production chain—from culture to genes and beyond. J Microbiol Methods 112:118–125. https://doi.org/10.1016/j.mimet.2015.03.007
Ma L, Wang Y, Shen J, Zhang Q, Wu C (2014) Tracking Campylobacter contamination along a broiler chicken production chain from the farm level to retail in China. Int J Food Microbiol 181:77–84. https://doi.org/10.1016/j.ijfoodmicro.2014.04.023
Mossong J, Mughini-Gras L, Penny C, Devaux A, Olinger C, Losch S, Cauchie H, van Pelt W, Ragimbeau C (2016) Human Campylobacteriosis in Luxembourg, 2010–2013: A Case-Control Study Combined with Multilocus Sequence Typing for Source Attribution and Risk Factor Analysis. Sci Rep 6:20939. https://doi.org/10.1038/srep20939
Nierop Groot MN, de Boer AG, van Pelt W, van der Hulst-van Arkel MC, de Leeuw P, Widjaja HC, Smits MA, van der Wal FJ (2014) A SpoT polymorphism correlates with chill stress survival and is prevalent in clinical isolates of Campylobacter jejuni. Poult Sci 93(11), 2900-2909. https://doi.org/10.3382/ps.2014-04055
Oh E, McMullen L, Jeon B (2015) Impact of oxidative stress defense on bacterial survival and morphological change in Campylobacter jejuni under aerobic conditions. Front Microbiol 6:295. https://doi.org/10.3389/fmicb.2015.00295
Phongsisay V, Perera VN, Fry BN (2007) Expression of the htrB gene is essential for responsiveness of Salmonella typhimurium and Campylobacter jejuni to harsh environments. Microbiology 153(Pt 1):254–262. https://doi.org/10.1099/mic.0.29230-0
Sambrook J, Russell DW (2006) Purification of nucleic acids by extraction with phenol:chloroform. CSH Protoc. 2006(1). https://doi.org/10.1101/pdb.prot4455
Seliwiorstow T, Baré J, Berkvens D, Van Damme I, Uyttendaele M, De Zutter L (2016) Identification of risk factors for Campylobacter contamination levels on broiler carcasses during the slaughter process. Int J Food Microbiol 226:26–32. https://doi.org/10.1016/j.ijfoodmicro.2016.03.010
Silva J, Leite D, Fernandes M, Mena C, Gibbs PA, Teixeira P (2011) Campylobacter spp as a Foodborne Pathogen: A Review. Frontiers in Microbiology 2:200. https://doi.org/10.3389/fmicb.2011.00200
Torralbo A, Borge C, García-Bocanegra I, Méric G, Perea A, Carbonero A (2015) Higher resistance of Campylobacter coli compared to Campylobacter jejuni at chicken slaughterhouse. Comp Immunol Microbiol Infect Dis. https://doi.org/10.1016/j.cimid.2015.02.003
Vinueza-Burgos C, Wautier M, Martiny D, Cisneros M, Van Damme I, De Zutter L (2017) Prevalence, antimicrobial resistance and genetic diversity of Campylobacter coli and Campylobacter jejuni in Ecuadorian broilers at slaughter age. Poult Sci 96(7):2366–2374. https://doi.org/10.3382/ps/pew487
Wagenaar JA, French NP, Havelaar AH (2013) Preventing Campylobacter at the source: why is it so difficult? Clin Infect Dis 57(11):1600–1606. https://doi.org/10.1093/cid/cit555
Acknowledgements
The authors thank to Alicia Torralbo Montoro and UVESA Company for their help during sampling.
Funding
Open Access funding provided thanks to the CRUE-CSIC agreement with Springer Nature. This research was funded by a grant of the University of Cordoba for Research Groups (sub-modality 1.2 UCO-Activa).
Author information
Authors and Affiliations
Contributions
Study conception and design was performed by A. Carbonero and J. Perea. Preparation of the material and laboratory analyses were performed by C. Borge, Y. Trujillo and A. Maldonado-Iniesta. Statistical analyses were performed by A. Carbonero an M. Riofrio. The first draft of the manuscript was written by I. Garcia-Bocanegra. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that there are no conflicts of interest regarding the results presented in this study.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Carbonero, A., Maldonado-Iniesta, A., Trujillo, Y. et al. Identification of genes associated with environmental persistence in Campylobacter jejuni and Campylobacter coli isolates from processing in a broiler abattoir. Vet Res Commun 46, 1325–1330 (2022). https://doi.org/10.1007/s11259-022-09981-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11259-022-09981-w