Abstract
Endemic (range restricted or precinctive) plant species are frequently observed to exhibit polyploidy (chromosome set duplication), which can drive shifts in ecology for angiosperms, but whether endemism is generally associated with polyploidy throughout the flowering plants has not been determined. We tested the hypothesis that polyploidy is more frequent and more pronounced (higher evident ploidy levels) for recently evolved endemic angiosperms. Chromosome count data, molecular dating and distribution for 4210 species (representing all major clades of angiosperms and including the largest families) were mined from literature-based databases. Upper boundary regression was used to investigate the relationship between the maximum number of chromosomes and time since taxon divergence, across clades and separately for families, comparing endemic with non-endemic species. A significant negative exponential relationship between maximum number of chromosomes and taxon age was evident across angiosperms (R2adj = 0.48 for all species, R2adj = 0.49 for endemics; R2adj = 0.44 for non-endemics; p always < 0.0001), recent endemics demonstrating greater maximum chromosome numbers (y intercept = 164 cf. 111) declining more rapidly with taxon age (decay constant = 0.12, cf. 0.04) with respect to non-endemics. The majority of families exhibited this relationship, with a steeper regression slope for endemic Campanulaceae, Asteraceae, Fabaceae, Poaceae, Caryophyllaceae and Rosaceae, cf. non-endemics. Chromosome set duplication is more frequent and extensive in recent angiosperms, particularly young endemics, supporting the hypothesis of recent polyploidy as a key explanation of range restriction. However, as young endemics may also be diploid, polyploidy is not an exclusive driver of endemism.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Why endemic species exhibit restricted geographical ranges is a key question in biogeography and ecology (Olivieri et al. 2015). Endemics may occupy a limited ecological niche (Williams et al. 2009), or originate by hybridization occurring only in the area of contact between progenitor species (Grünig et al. 2021), or they may be relics from a wider past range (‘paleo-endemics’; Favarger and Contandriopoulos 1961; Petrova et al. 2015) or recent species (‘neo-endemics’, sensu Stebbins and Major 1965) yet to disperse (Behroozian et al. 2020). Alternatively, genome duplication (polyploidy), effectively representing ‘instantaneous’ sympatric speciation (Mayr 1963), may be a major driver of plant evolution (Levin 1983; Otto and Whitton 2000; Soltis et al. 2014; Van de Peer et al. 2017). Polyploidy occurs when the cell cycle includes chromosome duplication but lacks the subsequent stages of cell component separation, resulting in cells with multiple chromosome complements. This can occur during mitosis (somatic doubling) or meiosis (non-reduction during sporogenesis), either within populations of a single species (autoploidy), or subsequent to interspecific hybridization (allopolyploidy; Ramsey and Schemske 1998). Polyploidy is often accompanied by larger nuclei and cells (genome size and nuclear and cell volumes are directly related; Cavalier-Smith 2005; Hodgson et al. 2010), altering physiological and morphological traits of offspring (see Van de Peer et al. 2020) that effect fitness (e.g. a greater tendency towards vegetative reproduction; Herben et al. 2017; see Soltis and Soltis 2016). Altered phenotypes change the ecology of polyploids with respect to diploid progenitors: e.g. larger cells result in larger organs (the ‘giga effect’), producing larger flowers of different colours and scents favouring different pollinator species (see Rezende et al. 2020). With different sets of chromosomes, polyploids are often reproductively isolated from mother plants (Husband and Sabara 2004; Laport et al. 2016; Lavania 2020). Ancient events of genome doubling are often associated with increased rates of speciation (Husband et al. 2013; Soltis and Soltis 2016; Landis et al 2018). Polyploidy occurs frequently in plants, particularly in angiosperms (Soltis and Soltis 2016; Lavania 2013, 2020): all angiosperms show evidence of multiple polyploidy events, except for Amborella trichopoda, the sister to all other living angiosperms which possesses only the ancestral ancient polyploidy event shared by all living flowering plants (Soltis et al. 2009; Jiao et al. 2011; Amborella Genome Project 2013; Lavania 2020).
However, despite being widely recognised as an important process for plant evolution and ecology, the extent to which polyploidy represents a general mechanism in the emergence of endemic species throughout the angiosperms has yet to be investigated.
Understanding the extent to which polyploidy is associated with endemism is complicated by the fact that ‘endemic’ is an ill-defined term. In part this is due to the historical use of the word, the original meaning being ‘a constant background presence in a particular area’ (e.g. “yellow fever is endemic to tropical Africa”), the opposite of ‘epidemic’ (i.e. spreading out of control). This meaning does not preclude the species being found elsewhere. In biology and ecology ‘endemic’ has taken on a meaning similar to ‘precinctive’, i.e. restricted to a precinct or place and found nowhere else. However, the precinct is often delimited on a case-by-case basis using artificial criteria such as geopolitical boundaries, which are variable in extent and often biologically irrelevant. The term ‘endemic’ is context dependent and applied over various scales (i.e. ‘continental endemic’, ‘local endemic’ or ‘narrow endemic’; Lavergne et al. 2004; Coelho et al. 2020), and may be considered to include the ecological requirements of the species and degree of habitat specificity (Boakes et al. 2010; Beck et al. 2014; Fithian et al. 2014). Indeed, terms such as ‘narrow endemism’ or ‘micro-endemism’ are typically qualified with information on the number of populations, degree of isolation and the genetic structure, environmental requirements and dispersal capacity of the taxon (Médail and Baumel 2018)—specific information that does not exist for many species. For the purposes of the present analysis the basic phenomenon under study is that of geographical range restriction, and to avoid the problems of scale surrounding the word ‘endemic’, here we explicitly define species as endemic (precinctive or range restricted) vs. non-endemic (relatively cosmopolitan) based on a number of criteria, including geographical distribution but also recognition as ‘endemic’ or ‘sub-endemic’ in national floras, in many cases combined with a specific epithet of the Latin binomial name that suggests belonging to a particular geographical location. We do not attempt to distinguish micro-endemics from more generally range-restricted endemics.
A further complication arises because ploidy level is not always easy to define, even when the basic number of chromosomes is known. This can result from ancient autopolyploidy or allopolyploidy (Parisod et al. 2010; Lavania 2013; Zozomová-Lihová et al. 2014), sometimes followed by diploidization and occasionally chromosome number reduction (i.e. diploidized paleo-polyploids; Tamayo-Ordóñez et al. 2016; Qiao et al. 2019). Moreover, some taxa show intraspecific variability, with multiple chromosome counts (different ploidy levels) arising from relatively frequent polyploidy events (Husband et al. 2013; Vimala et al. 2021). While polyploidy multiplies sets of chromosomes (and thus has striking effects during karyotype evolution), a range of processes can subtly rearrange single chromosomes, altering chromosome number or characteristics such as DNA content. These include insertion, deletion or duplication, inversion and intra- or inter-chromosomal reciprocal translocation, particularly evident for paleo-species that have accumulated changes over time (see Schubert and Lysak 2011; Vimala et al. 2021). As polyploidy drives change in the overall set of chromosomes, but complicating processes can further alter the number of chromosomes, the relationship between the number of chromosomes and polyploidy is not necessarily straightforward. With this caveat in mind, in the present study it is assumed that entire genome multiplication in the sporophyte generation (2n) is principally affected by polyploidy, and the number of chromosomes is used as a quantitative measure to represent the net result of karyotype evolution.
Additionally, the distinction between ‘neo-endemic’ and ‘paleo-endemic’ is also ambiguous. Despite the considerable attention given to the classification of endemic species in terms of when they originated (Favarger and Contandriopoulos 1961; Stebbins and Major 1965; Maers and Giller 2013), an absolute age threshold differentiating paleo- from neo-endemics remains undefined. While species younger than 1 million years are clearly neo-endemics (Kraft et al. 2010), the issue becomes complex for less recent species. Indeed, the term paleo-endemic refers more to a process than to a particular time or period per se (i.e. endemism by restriction or fragmentation of a previously extensive range). Ferreira and Boldrini (2011), for example, addressed the problem by suggesting the combination of a dated phylogeny (i.e. estimated age and degree of systematic isolation) with environmental context (based on stratigraphy and the age of underlying rocks). Lazarina et al. (2019) considered reproductive and geographical isolation, while Mishler et al. (2014) proposed a method based on their relative phylogenetic endemism index, to distinguish centres of neo- and paleo-endemism. Unfortunately, these methods are too unwieldy to be used for a prompt distinction between neo- and paleo-endemics in large datasets.
Particular relevance has been given to apo-endemics, or polyploids diverged from diploid progenitors (Favarger and Contandriopoulos 1961). However, the origin of a polyploid and divergence in the case of sympatric speciation is not always possible to date (Doyle and Egan 2010). Indeed, dating polyploidy events and their role in creating new taxa has so far been limited to the timing of major clade emergences (Wood et al. 2009), lacking sufficient detail to compare particular species within families or genera. In the present study, rather than entering into the debate regarding what constitutes a neo- or a paleo-endemic, we refer simply to the time period elapsed since the divergence of the taxon. Thus, the absolute timescale (in millions of years) is used here as a framework, and from hereon we explicitly avoid referring to arbitrary ‘neo-’ and ‘paleo-’ classes.
In summary, the comparison of estimated taxon age (Ma since divergence) against the number of chromosomes will test whether the chromosome complement is highest in recent species; information on the occurrence range of species will allow assessment of whether the phenomenon is general within the angiosperms or relatively prevalent in endemics. Based on these data, the principal objective of the present study is to assess whether polyploidization events are principal drivers of the emergence of new endemic species. Specifically, it is hypothesized that, despite a prevalence of diploid taxa throughout evolutionary time: (1) the highest chromosome counts are evident for angiosperm taxa that have diverged recently, (2) higher numbers of chromosomes are particularly evident for recent endemic (cf. non-endemic) taxa, and (3) the character of the ploidy level/divergence time relationship is consistent throughout the angiosperms, from ancient to recently diverged clades.
Materials and methods
Data mining
The relationship between genome duplication and the timing of speciation for endemic angiosperms used sporophyte chromosome count data, in the context of ‘time since divergence’ and geographical presence data. These data were collated from databases containing values from the scientific literature, and directly from the literature itself, aiming to broadly represent both endemic and non-endemic taxa across the angiosperms. The recent phylogeny of Leebens-Mack et al. (2019) was used, and the dataset specifically aimed to represent all major angiosperm clades and the largest families, starting with ANA-grade taxa (represented by Nymphaeaceae—other families in this clade are too under-represented in terms of both chromosome number and taxon age data), and including the monocots (represented by Poaceae and Orchidaceae), Magnoliids (Magnoliaceae), Ranunculales (Ranunculaceae), Caryophyllales (Caryophyllaceae), Asterids (Apiaceae, Campanulaceae, Asteraceae, Ericaceae, Primulaceae), Saxifragales (Saxifragaceae) and Core Rosids (Brassicaceae, Euphorbiaceae, Fabaceae, Rosaceae, Violaceae). Taxonomic name standardization was ensured using data from The Chromosome Counts Database (CCDB, v1.47: ccdb.tau.ac.il/browse), based on the automatic taxonomic name resolution software Taxonome (Kluyver and Osborne 2013) and The Plant List (v1.1: www.plantlist.org). Species with both ‘accepted’ and ‘unresolved’ taxonomic status were included: subspecies and varieties were discarded.
The diploid number of chromosomes for the sporophyte generation was attained from the CCDB (last access: October 2020), as a quantitative proxy of ploidy level. Only one count per species was included, except when different counts were equally reported in the database. When multiple counts were reported for a species, the modal value was retained; for species exhibiting multiple modal values, all were retained (e.g. 2n = 25, 2n = 30, 2n = 36 for Paphiopedilum victoria-mariae; Orchidaceae). Missing sporophyte values were calculated by doubling the gametophyte counts. B chromosomes were not considered. Negative values, 0 and 1 were considered errors (being biologically improbable) and discarded. When possible, the source material used for the counting was checked: usually mitotic counts were made using the root-tip squash method (Miller 1961), while meiotic counts were made from floral buds (see Windham et al. 2020). Chromosome counts for each taxon are presented in Table S1.
The estimated taxon age was obtained from the public database TimeTree: The Timescale of Life (TTOL, www.timetree.org; last access: September 2021). Molecular dating has been applied to an increasing number of species (the largest dated phylogenetic tree for the angiosperms comprises more than 36,000 species, belonging to ~ 8400 genera, 426 families and all orders; Janssens et al. 2020), but heterogeneity in datasets, sequences, calibrations and the software used can yield different estimates for the same species, often hindering comparison between the results of different studies (Pulquério and Nichols 2007). TTOL provides a comprehensive synthesis of data published between 1987 and 2013 (3998 studies; www.timetree.org/references) for 50,632 species, of which 14,465 angiosperms, and offers data uniformity, rapid data access and a robust foundation in the scientific literature (Hedges et al. 2006, 2015; Kumar and Hedges 2011; Kumar et al 2017). Divergence time between taxa is estimated through a hierarchical average linkage method (Hedges et al. 2015). Note that while confidence intervals for average divergence time estimates are not reported for all taxa in the present study, among-study variance does occur due a variety of factors, including differences in calibrations and gene and taxon sampling between studies. These interval estimates are calculated and reported by TTOL (for more details, see www.timetree.org/faqs#q2). Thus, in the present study, it is implicit that taxon age values represent estimated means based on a range of sampling methods employed across the scientific literature. A preliminary check of data included in the TTOL estimates was made from original chronograms in specific published papers, cited by TTOL. The discretional value of 0.001 Ma was attributed to extremely recent nodes, when a specific “estimated time” was not indicated [e.g. Adenocarpus hispanicus (Fabaceae), Anemone hepatica, Anemonastrum narcissiflora (Ranunculaceae), Magnolia coco, Magnolia obovata and Magnolia officinalis (Magnoliaceae), Table S1 presents estimated divergence times for all study species].
For the purposes of the analysis, we classified species as endemic (range-restricted) on a case-by-case basis using a combination of quantitative data (geographical range) guided by qualitative information such as designation as ‘endemic’ in national and regional floras. The geographical range for each species was obtained from public databases: Global Biodiversity Information Facilities (GBIF: https://www.gbif.org/), the Plants of the World Online portal (www.plantsoftheworldonline.org), taxon-specific databases (i.e. the Global Compositae Database—GCD: www.compositae.org; the Campanula portal: www.campanula.e-taxonomy.net) or specific papers (i.e. for Campanulaceae: Kandemir 2007 for Campanula coriacea; Crowl et al. 2015 for Catopsis delicatula; for Asteraceae: Zhang et al. 2011 for the genera Soroseris, Stebbinsia and Syncalathium). The highest richness of precinctive species is found in biodiversity hotspots (Cañadas et al. 2014; Noroozi et al. 2018), which have been identified in 36 areas around the globe (Conservation International, www.conservation.org; Critical Ecosystem Partnership Fund, www.cepf.net), and range from 18,972 km2 (New Caledonia) to 2,373,057 km2 (Indo-Burma region). Such heterogeneity often requires the identification of smaller, higher-concentration areas within these regions (“hotspots within hotspots”) with endemism again being considered across differing scales (Cañadas et al. 2014; Noroozi et al. 2018). Based on the geographical extent of these hotspots, a range not exceeding 600,000 km2 was one factor in the decision to classify a species as endemic. This threshold was chosen in order to include the remaining vegetation of the 36 Biodiversity Hotspots (see Table S2) according to Conservation International. Since the pre-industrialisation extension of some hotspots (i.e. Indo-Burma region, Brazil’s Cerrado, or Mediterranean Basin) exceeds 2,000,000 km2, and also includes urban areas, only the extension of the remaining vegetation (rather than the historical extent) was considered. Classification as endemic or not was also decided by determining whether species are recognised as endemic in national or local floras (for example: New Zealand Plant Conservation Network, http://www.nzpcn.org.nz; Cellinese et al. 2009 for endemic Campanulaceae of Crete; Brochmann et al. 1997 for endemics of Cape Verde; see Table S1 for details of each case, including flora languages), and when species epithets of Latin names indicated belonging to a geographical location (e.g. Amelanchier nantucketensis).
To check whether latitude affected data availability, for both the distribution of endemic species and chromosome counts, a control analysis was performed. Data records (Table S1) were randomised, and a subset of 500 species extracted and assigned a latitudinal zonation class: ‘Tropical’ (species occurring between the Tropics of Cancer and Capricorn, i.e. 23° 27′ N and 23° 27′ S), ‘Subtropical’ (between latitude 23° 27′ and 35° in each hemisphere, following www.globalbioclimatics.org), ‘Temperate’ (between latitude 35° and 66° 33′, the polar circles) and ‘Polar/Alpine’ (species at latitudes above the polar circles, or growing at elevations above 2000 metres above sea level, m a.s.l.). Species equally spread across two or more zones were considered as ‘Cosmopolitan’. To assess the distribution of endemic species according to latitude, the proportion of endemics with respect to the total number of species in each zone was calculated.
Data analysis
The coverage rate of the available data for each family was calculated as a percentage ratio between the number of species included in the analysis and the total number of species (both accepted and unresolved) reported in The Plant List. Three separate analyses were performed both on the totality of data collected (referred to as ‘angiosperms’) and on subsets for single families, further subdivided into analysis of all species (endemics and non-endemics), and endemics and non-endemics treated separately. For Nymphaeaceae, only one analysis was performed due to lack of data on endemic species, while for Magnoliaceae, Rosaceae, Saxifragaceae and Violaceae, analysis of endemic species was not performed, due to insufficient data (10 spp. or less). Analyses were performed using the statistical software R (v3.5.1; R Core Team 2018). Data were plotted according to the estimated time since divergence (x axis) and the number of chromosomes (y axis), using the ggplot2 package (Wickham et al. 2020).
To investigate the maximum number of chromosomes exhibited by taxa over geological time (time since taxon divergence), an upper boundary regression was applied, fitting the regression curve only to the highest values of the dataset. Boundary functions are widely used in ecology to highlight the maximum effects of processes, otherwise obscured by the weight of mean values (Pierce 2014, and references therein). To remove the effects of redundant chromosome counts within each family, age values along the x axis were divided into periods (‘bins’) of 1 million years, and regression fitted to the five highest y values within each bin. The function applied was an exponential decay with the formula: \(y=A{\mathrm{e}}^{\left(-kx\right)}-c\), where y is the sporophyte number of chromosomes, A is the initial quantity or y intercept (estimated y value for x = 0), k is the decay constant, x is the estimated time since divergence and c is the lowest y value for each family. The c parameter was introduced to obtain a horizontal asymptote equal to the lower chromosome count and avoid curves tending to zero, as zero chromosomes would be biologically unrealistic.
Finally, the percentage ratio between the number of polyploids (sensu Wood et al. 2009) and the total number of species for each family included in the analysis was calculated to test whether taxonomic groups are differentially predisposed to polyploidy, in terms of both formation and establishment. Data are available in Microsoft Excel spreadsheet format (Table S1).
Results
Analyses were performed on a total of 4530 records, representing 4210 species, 1344 (31.9%) of which were classified as endemic according to the combination of criteria used. For each family, the coverage rate of collected data generally did not exceed 4% of known species (Table S3), with the exception of Ranunculaceae (5.8%), Apiaceae (6.2%), Magnoliaceae (9.1%) and Campanulaceae (12.6%). According to the estimated crown age (Table S3), the oldest families (> 100 Ma) are Rosaceae (106 Ma, 95% CI 41–161 Ma), Orchidaceae (105 Ma, 95% CI 97–113 Ma) and Magnoliaceae (104 Ma, 95% CI 95–113 Ma), while Brassicaceae (46 Ma, 95% CI 19–71 Ma), Caryophyllaceae (52 Ma, 95% CI 39–62 Ma) and Apiaceae (54 Ma, 95% CI 29–57 Ma) are the most recent. However, uncertainty related to estimated crown age was sometimes substantial: in Rosaceae, the extreme case, the confidence interval differed by 120 million years. Fabaceae (71–80 Ma), Violaceae (67–77 Ma) and Ranunculaceae (75–88 Ma) exhibited the least variable estimates.
Recurrent numbers of chromosomes were evident in all families, which sometimes represent a high proportion of data, for example, 2n = 22 for Apiaceae (58%); 2n = 34, Campanulaceae (38.8%); 2n = 38, Magnoliaceae (67.7%); 2n = 32, Ranunculaceae (54.9%); and 2n = 48, Violaceae (39%). Model parameters (k = decay constant; A = y intercept) and statistics (R2adj and p-value) were extracted during model production in the R environment. Hereafter, the adjusted R2 values for each general analysis [total species (T) representing endemics (E) plus non-endemics (N)] are indicated by R2adjT, while for each analysis performed on endemics and non-endemics separately variance is indicated by R2adjE, and R2adjN, respectively. Upper boundary regression applied to the entire dataset (Fig. 1) showed a significant exponential decay trend (R2adjT = 0.48; R2adjE = 0.49; R2adjN = 0.44; p always < 0.0001) between the number of chromosomes and the estimated time since divergence, which was three times steeper in endemic species (k = 0.12) with respect to non-endemics (k = 0.04). The estimated y intercept was higher in endemics compared to non-endemics (A = 164 cf. 111, respectively; Fig. 1).
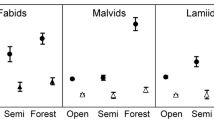
The relationship between time since taxon divergence and number of chromosomes (as a proxy for ploidy level) across all major clades of the Angiosperms, applied to: a the entire dataset, b endemic species and c non-endemic species. Squares represent endemic species, and circles represent non-endemic species. Broken lines represent ‘upper boundary regressions’, or 3-parameter Lorentzian regressions fitted to the five highest values in each ‘bin’ or 1 million year interval (bin value data points are filled in dark grey; points under the upper boundary curve are unfilled). Dotted lines represent the ± 95% confidence interval
In analyses performed on single families, results were varied but common patterns were evident. The overall negative relationship between the number of chromosomes and taxon age was confirmed in the majority of the families, with differing degrees of significance: Campanulaceae (R2adjT = 0.42; R2adjE = 0.22; R2adjN = 0.31, p < 0.0001; Fig. 2a–c), Asteraceae (R2adjT = 0.46; R2adjE = 0.45; R2adjN = 0.23; p < 0.0001; Fig. 2d–f), Fabaceae (Fig. 2g–i), Poaceae (Fig. 2j–l), Caryophyllaceae (Fig. S1a–c), Ranunculaceae (Fig. S1g–i) and Rosaceae (Fig. S1d–f). However, similar exponential decay patterns were only evident as non-significant trends for endemic Primulaceae (Fig. S1k; R2adjE = 0.065; p = 0.116). The relationship was stronger for endemics, as confirmed by the higher values of k (e.g. Asteraceae, k = 0.15 and 0.04 for endemics vs. non-endemics, respectively; Caryophyllaceae, k = 0.23 and 0.15 for endemics vs. non-endemics, respectively) and the entire family (e.g. k = 0.11 for Asteraceae, or k = 0.16 for Caryophyllaceae). The negative trend was of borderline significance for endemic Apiaceae (R2adjE = 0.06; Fig. S2b), not significant for endemic vs. non-endemic Orchidaceae (R2adjT = 0.19; R2adjE = − 0.02; R2adjN = 0.14; p = 0.51 in endemics and < 0.001 for the family and non-endemics; Fig. S2d–f) and non-significant for Violaceae (R2adjT = 0.05; R2adjN = 0.03; p always > 0.1; Fig. S2g–i) and Magnoliaceae (negative values for R2adjT and R2adjN; p always > 0.6; Fig. S2j–l), although data were lacking for endemics of these latter two families (Fig. S2h, k).
The relationship between time since taxon divergence and number of chromosomes for examples of angiosperm families exhibiting a declining upper boundary relationship: a–c Campanulaceae, all spp., endemic spp. and non-endemic spp., respectively, d–f Asteraceae, g–i Fabaceae and j–l Poaceae. Squares represent endemic species, and circles represent non-endemic species. Broken lines represent ‘upper boundary regressions’, or 3-parameter Lorentzian regressions fitted to the five highest values in each ‘bin’ or 1 million year interval (bin value data points are filled in dark grey; points under the upper boundary curve are unfilled). Dotted lines ± 95% CI Note that x and y data ranges are different for each family
For Ericaceae, regressions were not significant (R2adj always < 0.01; p always > 0.1; Fig. 3). In contrast, Brassicaceae (Fig. 4) and Euphorbiaceae (Fig. S3) exhibited statistically significant negative slopes for non-endemics and the families as a whole, while slopes for endemics were not significant (− R2adj, p ~ 0.7) with increasing tendencies with taxon age (k = − 0.03 and − 0.009, respectively). Finally, Nymphaeaceae (Fig. 5) showed a significant positive relationship, with k = − 0.04, R2adjT = 0.29 and p = 0.03. A similar, but non-significant, tendency was shown in Saxifragaceae (k = − 0.02 for the family and k = − 0.01 for non-endemics, Fig. S4a–c), with negative R2adj values and p always > 0.3.
Non-significant trends in the family of Ericaceae, with a positive slope in a the analysis of the whole family and c non-endemic species only, contrasted with a negative slope for b endemics. Squares represent endemic species, and circles represent non-endemic species. Broken lines represent ‘upper boundary regressions’ fitted to the five highest values in each ‘bin’ (points filled in dark grey; points under the upper boundary curve are unfilled). Dotted lines ± 95% CI
Significant negative relationship between the number of chromosomes and estimated taxon age in a the entire family of Brassicaceae, c non-endemic Brassicaceae and b non-significant positive trend in endemic Brassicaceae. Squares represent endemic species, and circles represent non-endemic species. Broken lines represent ‘upper boundary regressions’ fitted to the five highest values in each ‘bin’ (points filled in dark grey; points under the upper boundary curve are unfilled). Dotted lines ± 95% CI
The proportion of polyploid taxa within each family was found to differ substantially between families (Fig. S5): polyploidy was evident for the majority of taxa in Violaceae (92%), Primulaceae (76%) and Campanulaceae (68%), while it occurred in less than 10% of species of the Fabaceae (8%) and Orchidaceae (6%). Note that there was no statistically significant geographical bias (in terms of bioclimatic zones) for either the number of chromosomes (Fig. S6) or the proportion of the flora that were endemic species (Fig. S7).
Discussion
We demonstrated a negative exponential relationship between maximum number of chromosomes and time since divergence with a decay constant three times greater for endemic angiosperms with respect to non-endemics (k = 0.12 cf. 0.04, respectively; Fig. 1). Moreover, the estimated y intercept was much higher for endemics with respect to non-endemics (A = 164 cf. 111, respectively), indicating that recently diverged species with higher numbers of chromosomes are more likely to be range restricted (endemic). Indeed, the results broadly support the hypotheses that polyploidy is particularly evident in recently diverged angiosperms (Hypothesis 1), especially for recent endemic species (Hypothesis 2). While this phenomenon was evident for most of the families investigated, it was not always observed, and the hypothesis of a mechanism working consistently across the angiosperms (Hypothesis 3) was only partially supported. The distribution of chromosome counts with time (i.e. towards recent ages) highlighted the pattern of progressive multiplication of the chromosome set, with high concentrations of records corresponding to diploid, tetraploid and hexaploid counts (2n = 2x, 4x and 6x, respectively): this is particularly evident in Apiaceae, Caryophyllaceae, Ranunculaceae and Rosaceae. Thus, polyploidization appears to be an important mechanism for the emergence of new species generally.
Contrasting results for different families suggest that phylogenetic effects operate within each family (revealed by the direction, range and variability of patterns; Fig. S5). For Ericaceae, the regression analyses were not significant: it is likely that other prevailing mechanisms, such as adaptive radiation, drive the emergence of new ericaceous species. Despite showing an overall negative tendency between the number of chromosomes and taxon age, a similar interpretation could explain the weak significance for Orchidaceae, also indicated by the highly variable 95% confidence intervals.
In Saxifragaceae, Magnoliaceae, Nymphaeaceae and Violaceae, results were probably affected by limited data availability. This could also explain the lower significance of the analyses for endemics and non-endemics with respect to the family-level analysis in Primulaceae, and the atypical trend in endemic Euphorbiaceae and Brassicaceae, characterised by wide and irregular 95% confidence intervals. In particular, the estimated age for endemic Brassicaceae did not exceed 9 Ma, with only seven species older than 3 Ma (i.e. Cochlearia aragonensis, Draba hederifolia, Streptanthus glandulosus, Vella asperum, Vella bourgaeana, Vella pseudocytisus and Vella spinosa). In these situations, the nature of the data requires careful consideration: four out of these species belonging to the same genus, Vella, are endemics of Spain and were dated relying on a single study (i.e. Simón-Porcar et al. 2015); independence of observations could not be assured, since V. asperum, V. bourgaeana and V. pseudocytisus are closely related (see also Siljak-Yakovlev and Peruzzi 2012). This is likely to have effects on species distribution: these species are also found in the same habitat (disturbed xerophytic shrublands on gypsum substrate; Gómez Campo 1993). With regard to karyotypes, the three species share the same basic chromosome number (x = 17), but while V. bourgaeana is diploid (2n = 2x = 34), V. pseudocytisus is mainly tetraploid (2n = 4x = 68), and V. asperum is hexaploid (2n = 6x = 102) (Simón-Porcar et al. 2015), further confirming the role of polyploidy in speciation events. For these reasons, detailed here for a single prominent case, all analyses based on a restricted number of records should be interpreted with caution.
Additionally, weaker regressions or positive trends were generally evident for older families: Orchidaceae (105 Ma), Magnoliaceae (104 Ma), Nymphaeaceae (89 Ma), Saxifragaceae (88 Ma) and Ericaceae (82 Ma; Table S3). This suggests that for certain ancient clades polyploidization may not be the main driver of speciation, although Rosaceae, the oldest clade (106 Ma), agreed with the hypothesis of higher polyploidy occurrence in recently diverged species. It is noticeable that some of these ancient families (i.e. Nymphaeaceae, Saxifragaceae, Rosaceae) showed high percentages of polyploid taxa, sensu Wood et al. 2009 (higher than 45%, Fig. S5). In Saxifragaceae, a high number of chromosomes in older taxa but a lack of a relationship over time could indicate an initial burst of speciation via polyploidy followed by a lesser involvement of polyploidy in speciation. Therefore, polyploidy could be relatively widespread even across ancient families, but it is evidently not the main driver of speciation for these families.
Why recent polyploids are relegated to limited ranges is not immediately evident from our dataset. Polyploids are often adaptable species able to survive in harsh environmental contexts, and thus advantaged when colonizing new habitats (Flovik 1940; Brochmann et al. 2004; Manzaneda et al. 2012; te Beest et al. 2012; Mas de Xaxars et al. 2016; Paule et al. 2018; Stevens et al. 2020). Intriguingly, alteration of phenotype by polyploidy suggests that plant functioning and fitness may be fundamentally changed. However, a preliminary classification of the species in our dataset according to Grime’s CSR ecological strategies (method of Pierce et al. 2017) showed that no particular strategy class was associated with polyploidy: species adapted to survive competition, stress or disturbance all exhibited an extremely wide range of chromosome numbers (data not shown). Rather than reflecting fitness and adaptation, the high incidence of endemic polyploids could depend mainly on limited time for dispersal and colonizing new areas, evident for certain species (Behroozian et al. 2020). It has been determined that polyploid plant species rely mainly on vegetative reproduction (Herben et al. 2017), which is usually ineffective for wide or rapid dispersal (Winkler and Fischer 2001; Herben et al. 2016). Indeed, the rearrangements of the chromosome set and the encumbrance created by multiple chromosomes in polyploid cells hinder meiosis, imposing disadvantages for sexual reproduction (Herben et al. 2017). Another polyploid trait impacting dispersal, related to the ‘giga effect’, is that of larger and heavier seeds (Stevens et al. 2020), of particular importance to wind-dispersed species. Moreover, some studies (e.g. Corneillie et al. 2019; Mo et al. 2020) have shown slower growth rates for polyploids compared to diploids. Together, these traits may limit dispersal, colonisation and seedling recruitment processes for polyploids.
The present study has a number of limitations that should be considered in interpreting these results. For example, the available data cannot indicate the extent to which anagenesis (speciation via evolutionary change directly within a single lineage) occurs, and speciation by cladogenesis is assumed. Increased availability of data regarding phylogenies (inclusion of greater numbers of species), tropical species (which are less well studied; Prance 1977; Prance and Campbell 1988; Sosef et al. 2017), species molecular dating, chromosome counts and distributions will provide further support to this analysis. This particularly applies to families with less complete records (e.g. Ericaceae, Magnoliaceae, Nymphaeaceae), in order to determine whether contrasting results for these families are indicative of truly different patterns or are data artefacts.
Conclusion
Chromosome duplication is more prevalent in recent angiosperms, in particular young endemics, confirming the role of polyploidy as a key driver of recent endemism throughout the flowering plants. This pattern was generally evident across flowering plant families, but some cases in which patterns were lacking may reflect insufficient data availability. Dispersal limitation is more likely for polyploid taxa, potentially explaining why recently evolved polyploids tend to remain relegated to small ranges compared to diploids. However, the majority of young species (both endemics and non-endemics) are diploid, and thus polyploidy is not an exclusive driver of endemism.
Data availability
The dataset is available in Microsoft Excel format as Table S1 (Online Resource 1).
Code availability
N/A.
References
Amborella Genome Project (2013) The Amborella genome and the evolution of flowering plants. Science 342:1241089. https://doi.org/10.1126/science.1241089
Beck J, Böller M, Erhardt A, Schwanghart W (2014) Spatial bias in the GBIF database and its effect on modeling species’ geographic distributions. Ecol Inform 19:10–15. https://doi.org/10.1016/j.ecoinf.2013.11.002
Behroozian M, Ejtehadi H, Memariani F, Pierce S, Mesdaghi M (2020) Are endemic species necessarily ecological specialists? Functional variability and niche differentiation of two threatened Dianthus species in the montane steppes of northeastern Iran. Sci Rep 10:11774. https://doi.org/10.1038/s41598-020-68618-7
Boakes EB, McGowan PJK, Fuller RA, Chang-qing D, Clark NE, O’Connor K, Mace GM (2010) Distorted views of biodiversity: spatial and temporal bias in species occurrence data. PLoS Biol 8(6):e1000385. https://doi.org/10.1371/journal.pbio.1000385
Brochmann C, Rustan ØH, Lobin W, Kilian N (1997) The endemic vascular plants of the Cape Verde Islands, W Africa. Sommerfeltia 24:1–356
Brochmann C, Brysting AK, Alsos IG, Borgen L, Grundt HH, Scheen A-C, Elven R (2004) Polyploidy in Arctic plants. Biol J Linn Soc Lond 82:521–536. https://doi.org/10.1111/j.1095-8312.2004.00337.x
Cañadas EM, Fenu G, Peñas J, Lorite J, Mattana E, Bacchetta G (2014) Hotspots within hotspots: endemic plant richness, environmental drivers, and implications for conservation. Biol Conserv 170:282–291. https://doi.org/10.1016/j.biocon.2013.12.007
Cavalier-Smith T (2005) Economy, speed and size matter: evolutionary forces driving nuclear genome miniaturisation and expansion. Ann Bot 95:147–175
Cellinese N, Smith SA, Edwards EJ, Kim S-T, Haberle RC, Avramakis M, Donoghue MJ (2009) Historical biogeography of the endemic Campanulaceae of Crete. J Biogeogr 36:1253–1269. https://doi.org/10.1111/j.1365-2699.2008.02077.x
Coelho N, Gonçalves S, Romano A (2020) Endemic plant species conservation: biotechnological approaches. Plants 9(3):345. https://doi.org/10.3390/plants9030345
Corneillie S, De Storme N, Van Acker R, Fangel JU, De Bruyne M, De Rycke R, Geelen D, Willats WGT, Vanholme B, Boerjan W (2019) Polyploidy affects plant growth and alters cell wall composition. Plant Physiol 179(1):74–87. https://doi.org/10.1104/pp.18.00967
Crowl AA, Visger CJ, Mansion G, Hand R, Wu H-H, Kamari G, Phitos D, Cellinese N (2015) Evolution and biogeography of the endemic Roucela complex (Campanulaceae: Campanula) in the Eastern Mediterranean. Ecol Evol 5(22):5329–5343. https://doi.org/10.1002/ece3.1791
Doyle JJ, Egan AN (2010) Dating the origins of polyploidy events. N Phytol 186:73–85. https://doi.org/10.1111/j.1469-8137.2009.03118.x
Favarger C, Contandriopoulos J (1961) Essai sur l’endemisme. Bull Soc Bot Suisse 71:384–408 ((in the French language))
Ferreira PMA, Boldrini II (2011) Potential reflection of distinct ecological units in plant endemism categories. Conserv Biol 25(4):672–679. https://doi.org/10.1111/j.1523-1739.2011.01675.x
Fithian W, Elith J, Hastie T, Keith DA (2014) Bias correction in species distribution models: pooling survey and collection data for multiple species. Methods Ecol Evol 6(4):424–438. https://doi.org/10.1111/2041-210X.12242
Flovik K (1940) Chromosome numbers and polyploidy within the flora of Spitzbergen. Hereditas 26(3–4):430–440. https://doi.org/10.1111/j.1601-5223.1940.tb03246.x
Gómez Campo C (1993) Vella L. In: Castroviejo S et al (eds) Flora Iberica, vol 4. Real Jardín Botánico, CSIC, Madrid, pp 414–417
Grünig S, Fischer M, Parisod C (2021) Recent hybrid speciation at the origin of the narrow endemic Pulmonaria helvetica. Ann Bot 127:21–31. https://doi.org/10.1093/aob/mcaa145
Hedges SB, Dudley J, Kumar S (2006) TimeTree: a public knowledge-base of divergence times among organisms. Bioinformatics 22(23):2971–2972. https://doi.org/10.1093/bioinformatics/btl505
Hedges SB, Marin J, Suleski M, Paymer M, Kumar S (2015) Tree of life reveals clock-like speciation and diversification. Mol Biol Evol 32(4):835–845. https://doi.org/10.1093/molbev/msv037
Herben T, Tackenberg O, Klimešová J (2016) Reproduction by seed and clonality in plants: correlated syndromes or independent strategies? J Ecol 104:1696–1706. https://doi.org/10.1111/1365-2745.12646
Herben T, Suda J, Klimešová J (2017) Polyploid species rely on vegetative reproduction more than diploids: a re-examination of the old hypothesis. Ann Bot 120(2):341–349. https://doi.org/10.1093/aob/mcx009
Hodgson JG, Sharafi M, Jalili A, Díaz S et al (2010) Stomatal vs. genome size in angiosperms: the somatic tail wagging the genomic dog? Ann Bot 105:573–584
Husband BC, Sabara HA (2004) Reproductive isolation between autotetraploids and their diploid progenitors in fireweed, Chamerion angustifolium (Onagraceae). N Phytol 161(3):703–713. https://www.jstor.org/stable/1514534. Accessed 12 Jan 2022
Husband BC, Baldwin SJ, Suda J (2013) The incidence of polyploidy in natural plant populations: major patterns and evolutionary processes. In: Leitch IJ et al (eds) Plant genome diversity, vol 2. Springer, Wien, pp 255–276
Janssens SB, Couvreur TLP, Mertens A, Dauby G, Dagallier L-PMJ, Abeele SM, Vandelook F, Mascarello M, Beeckman H, Sosef M, Droissart V, van der Bank M, Maurin O, Hawthorne W, Marshall C, Réjou-Méchain M, Beina D, Baya F, Merckx V, Verstraete B, Hardy O (2020) A large-scale species level dated angiosperm phylogeny for evolutionary and ecological analyses. Biodivers Data J 8:e39677. https://doi.org/10.3897/BDJ.8.e39677
Jiao Y, Wickett NJ, Ayyampalayam S, Chanderbali AS, Landherr L, Ralph PE, Tomsho LP, Hu Y, Liang H, Soltis PS, Soltis DE, Clifton SW, Schlarbaum SE, Schuster SC, Ma H, Leebens-Mack J, dePamphilis CW (2011) Ancestral polyploidy in seed plants and Angiosperms. Nature 473:97–100. https://doi.org/10.1038/nature09916
Kandemir A (2007) A new Campanula (Campanulaceae) from east Anatolia, Turkey. Nord J Bot 25:53–57. https://doi.org/10.1111/j.0107-055X.2007.00091_14.x
Kluyver TA, Osborne CP (2013) Taxonome: a software package for linking biological species data. Ecol Evol 3(5):1262–1265. https://doi.org/10.1002/ece3.529
Kraft NJB, Baldwin BG, Ackerly DD (2010) Range size, taxon age and hotspots of neoendemism in the California flora. Divers Distrib 16:403–413. https://doi.org/10.1111/j.1472-4642.2010.00640.x
Kumar S, Hedges SB (2011) TimeTree2: species divergence times on the iPhone. Bioinformatics 27(14):2023–2024. https://doi.org/10.1093/bioinformatics/btr315
Kumar S, Stecher G, Suleski M, Hedges SB (2017) TimeTree: a resource for timelines, timetrees, and divergence times. Mol Biol Evol 34(7):1812–1819. https://doi.org/10.1093/molbev/msx116
Landis JB, Soltis DE, Li Z, Marx HE, Barker MS, Tank DC, Soltis PS (2018) Impact of whole genome duplication events on diversification rates in angiosperms. Am J Bot 105(3):433–444. https://doi.org/10.1002/ajb2.1060
Laport RG, Minckley RL, Ramsey J (2016) Ecological distributions, phenological isolation, and genetic structure in sympatric and parapatric populations of the Larrea tridentata polyploid complex. Am J Bot 103(7):1358–1374. https://doi.org/10.3732/ajb.1600105
Lavania UC (2013) Polyploidy, body size, and opportunities for genetic enhancement and fixation of heterozygosity in plants. Nucleus (April) 56(1):1–6. https://doi.org/10.1007/s13237-013-0075-7
Lavania UC (2020) Plant speciation and polyploidy: in habitat divergence and environmental perspective. Nucleus 63:1–5. https://doi.org/10.1007/s13237-020-00311-6
Lavergne S, Thompson JD, Garnier E, Debussche M (2004) The biology and ecology of narrow endemic and widespread plants: a comparative study of trait variation in 20 congeneric pairs. Oikos 107(3):505–518. https://www.jstor.org/stable/3548237. Accessed 12 Jan 2022
Lazarina M, Kallimanis AS, Dimopoulos P, Psaralexi M, Michailidou D-E, Sgardelis SP (2019) Patterns and drivers of species richness and turnover of neo-endemic and palaeo-endemic vascular plants in a Mediterranean hotspot: the case of Crete, Greece. J Biol Res Thessalon 26:12. https://doi.org/10.1186/s40709-019-0106-x
Leebens-Mack JH, Barker MS et al (2019) One thousand plant transcriptomes and the phylogenomics of green plants. Nature 574:679–685. https://doi.org/10.1038/s41586-019-1693-2
Levin DA (1983) Polyploidy and novelty in flowering plants. Am Nat 122(1):1–25. https://www.jstor.org/stable/2461003. Accessed 12 Jan 2022
Maers AA, Giller P (2013) Analytical biogeography: an integrated approach to the study of animal and plant distributions. Springer, Dordrecht
Manzaneda AJ, Rey PJ, Bastida JM, Weiss-Lehman C, Raskin E, Mitchell-Olds T (2012) Environmental aridity is associated with cytotype segregation and polyploidy occurrence in Brachypodium distachyon (Poaceae). N Phytol 193(3):797–805. https://doi.org/10.1111/j.1469-8137.2011.03988.x
Mas de Xaxars G, Garnatje T, Pellicer J, Siljak-Yakovlev S, Vallès J, Garcia S (2016) Impact of dysploidy and polyploidy on the diversification of high mountain Artemisia (Asteraceae) and allies. Alp Bot 126:35–48. https://doi.org/10.1007/s00035-015-0159-x
Mayr E (1963) Animal species and evolution. Harvard University Press, Cambridge
Médail F, Baumel A (2018) Using phylogeography to define conservation priorities: the case of narrow endemic plants in the Mediterranean Basin hotspot. Biol Conserv 224:258–266. https://doi.org/10.1016/j.biocon.2018.05.028
Miller WH (1961) Root tip cell squashes for the study of cell and chromosome morphology. Am Biol Teach 23(8):513–515. https://doi.org/10.2307/4439736
Mishler BD, Knerr N, Gonzáles-Orozco CE, Thornhill AH, Laffan SW, Miller JT (2014) Phylogenetic measures of biodiversity and neo- and paleo-endemism in Australian Acacia. Nat Commun 5:4473. https://doi.org/10.1038/ncomms5473
Mo L, Chen J, Chen F, Xu Q, Tong Z, Huang H, Dong R, Lou X, Lin E (2020) Induction and characterization of polyploids from seeds of Rhododendron fortunei Lindl. J Integr Agric 19(8):2016–2026. https://doi.org/10.1016/S2095-3119(20)63210-5
Noroozi J, Talebi A, Doostmohammadi M, Rumpf SB, Linder HP, Schneeweiss GM (2018) Hotspots within a global biodiversity hotspot—areas of endemism are associated with high mountain ranges. Sci Rep 8:10345. https://doi.org/10.1038/s41598-018-28504-9
Olivieri I, Tonnabel J, Ronce O, Mignot A (2015) Why evolution matters for species conservation: perspectives from three case studies of plant metapopulations. Evol Appl 9(1):196–211. https://doi.org/10.1111/eva.12336
Otto SP, Whitton J (2000) Polyploid incidence and evolution. Annu Rev Genet 34:401–437. https://doi.org/10.1146/annurev.genet.34.1.401
Parisod C, Holderegger R, Brochmann C (2010) Evolutionary consequences of autopolyploidy. N Phytol 186:5–17. https://doi.org/10.1111/j.1469-8137.2009.03142.x
Paule J, Dunkel FG, Schmidt M, Gregor T (2018) Climatic differentiation in polyploidy apomictic Ranunculus auricomus complex in Europe. BMC Ecol 18:16. https://doi.org/10.1186/s12898-018-0172-1
Petrova G, Moyankova D, Nishii K, Forrest L, Tsiripidis I, Drouzas AD, Djilianov D, Möller M (2015) The European Paleoendemic Haberlea rhodopensis (Gesneriaceae) has an Oligocene origin and a Pleistocene diversification and occurs in a long-persisting refugial area in Southeastern Europe. Int J Plant Sci 176(6):499–514. https://doi.org/10.1086/681990
Pierce S (2014) Implications for biodiversity conservation of the lack of consensus regarding the humped-back model of species richness and biomass production. Funct Ecol 28:253–257. https://doi.org/10.1111/1365-2435.12147
Pierce S, Negreiros D, Cerabolini BEL, Kattge J, Díaz S, Kleyer M, Shipley B, Wright SJ, Soudzilovskaia NA, Onipchenko VG, van Bodegom PM, Frenette-Dussault C, Weiher E, Pinho BX, Cornelissen JHC, Grime JP, Thompson K, Hunt R, Wilson PJ, Buffa G, Nyakunga OC, Reich PB, Caccianiga M, Mangili F, Ceriani RM, Luzzaro A, Brusa G, Siefert A, Barbosa NPU, Chapin FS III, Cornwell WK, Fang J, Fernandes GW, Garnier E, Le Stradic S, Peñuelas J, Melo FPL, Slaviero A, Tabarelli M, Tampucci D (2017) A global method for calculating plant CSR ecological strategies applied across biomes worldwide. Funct Ecol 31(2):444–457. https://doi.org/10.1111/1365-2435.12722
Prance GT (1977) Floristic inventory of the Tropics: where do we stand? Ann Mo Bot Gard 64(4):659–684. https://www.jstor.org/stable/2395293. Accessed 12 Jan 2022
Prance GT, Campbell DG (1988) The present state of tropical floristics. Taxon 37(3):519–548. https://doi.org/10.2307/1221097
Pulquério MJF, Nichols RA (2007) Dates from the molecular clock: how wrong can we be? Trends Ecol Evol 22(4):180–184. https://doi.org/10.1016/j.tree.2006.11.013
Qiao X, Li Q, Yin H, Qi K, Li L, Wang R, Zhang S, Paterson AH (2019) Gene duplication and evolution in recurring polyploidization–diploidization cycles in plants. Genome Biol 20:38. https://doi.org/10.1186/s13059-019-1650-2
Ramsey J, Schemske DW (1998) Pathways, mechanisms, and rates of polyploidy formation in flowering plants. Annu Rev Ecol Syst 29:467–501. https://doi.org/10.1146/annurev.ecolsys.29.1.467
R Core Team (2018) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna. http://www.R-project.org/. Accessed 12 Jan 2022
Rezende L, Suzigan J, Amorim FW, Moraes AP (2020) Can plant hybridization and polyploidy lead to pollinator shift? Acta Bot Bras 34(2):229–242. https://doi.org/10.1590/0102-33062020abb0025
Schubert I, Lysak MA (2011) Interpretation of karyotype evolution should consider chromosome structural constraints. Trends Genet 27(6):207–216. https://doi.org/10.1016/j.tig.2011.03.004
Siljak-Yakovlev S, Peruzzi L (2012) Cytogenetic characterization of endemics: past and future. Plant Biosyst 146(3):694–702
Simón-Porcar VI, Pérez-Collazos E, Catalán P (2015) Phylogeny and systematics of the western Mediterranean Vella pseudocytisus–V. aspera complex (Brassicaceae). Turk J Bot 39:472–486. https://doi.org/10.3906/bot-1406-87
Soltis PS, Soltis DE (2016) Ancient WGD events as drivers of key innovations in Angiosperms. Curr Opin Plant Biol 30:159–165. https://doi.org/10.1016/j.pbi.2016.03.015
Soltis DE, Albert VA, Leebens-Mack J, Bell CD, Paterson AH, Zheng C, Sankoff D, dePamphilis CW, Wall PK, Soltis PS (2009) Polyploidy and Angiosperm diversification. Am J Bot 96(1):336–348. https://doi.org/10.3732/ajb.0800079
Soltis DE, Visger CJ, Soltis PS (2014) The polyploidy revolution then…and now: Stebbins revisited. Am J Bot 101(7):1057–1078. https://doi.org/10.3732/ajb.1400178
Sosef MSM, Dauby G, Blach-Overgaard A et al (2017) Exploring the floristic diversity of tropical Africa. BMC Biol 15:art.15. https://doi.org/10.1186/s12915-017-0356-8
Stebbins GL, Major J (1965) Endemism and speciation in the California flora. Ecol Monogr 35(1):1–35. https://doi.org/10.2307/1942216
Stevens AV, Nicotra AB, Godfree RC, Guja LK (2020) Polyploidy affects the seed, dormancy and seedling characteristics of a perennial grass, conferring an advantage in stressful climates. Plant Biol 22:500–513. https://doi.org/10.1111/plb.13094
Tamayo-Ordóñez MC, Espinosa-Barrera LA, Tamayo-Ordóñez YJ, Ayil-Gutiérrez B, Sánchez-Teyer LF (2016) Advances and perspectives in the generation of polyploid plant species. Euphytica 209:1–22. https://doi.org/10.1007/s10681-016-1646-x
te Beest M, Le Roux JJ, Richardson DM, Brysting AK, Suda J, Kubešová M, Pyšek P (2012) The more the better? The role of polyploidy in facilitating plant invasions. Ann Bot 109:19–45. https://doi.org/10.1093/aob/mcr277
Van de Peer Y, Mizrachi E, Marchal K (2017) The evolutionary significance of polyploidy. Nat Rev Genet 18:411–424. https://doi.org/10.1038/nrg.2017.26
Van de Peer Y, Ashman T-L, Soltis PS, Soltis DE (2020) Polyploidy: an evolutionary and ecological force in stressful times. Plant Cell 33(1):11–26. https://doi.org/10.1093/plcell/koaa015
Vimala Y, Lavania S, Lavania UC (2021) Chromosome change and karyotype differentiation—implications in speciation and plant systematics. Nucleus 64:33–54. https://doi.org/10.1007/s13237-020-00343-y
Wickham H, Chang W, Henry L, Pedersen TL, Takahashi K, Wilke C, Woo K, Yutani H, Dunnington D (2020) RStudio: create elegant data visualisations using the grammar of graphics. R package version 3.3.3
Williams SE, Williams YM, Van Der Wal J, Isaac JL, Shoo LP, Johnson CN (2009) Ecological specialization and population size in a biodiversity hotspot: how rare species avoid extinction. Proc Natl Acad Sci USA 106(2):19737–19741. https://doi.org/10.1073/pnas.0901640106
Windham MD, Pryer KM, Poindexter DB, Li F-W, Rothfels CJ, Beck JB (2020) A step-by-step protocol for meiotic chromosome counts in flowering plants: a powerful and economical technique revisited. Appl Plant Sci 8(4):e11342. https://doi.org/10.1002/aps3.11342
Winkler E, Fischer M (2001) The role of vegetative spread and seed dispersal for optimal life histories of clonal plants: a simulation study. Evol Ecol 15:281–301. https://doi.org/10.1023/A:1016080714200
Wood TE, Takebayashic N, Barker MS, Mayrosee I, Greenspoond PB, Rieseberg LH (2009) The frequency of polyploid speciation in vascular plants. Proc Natl Acad Sci USA 106(33):13875–13879. https://doi.org/10.1073/pnas.0811575106
Zhang J-W, Nie Z-L, Wen J, Sun H (2011) Molecular phylogeny and biogeography of three closely related genera, Soroseris, Stebbinsia, and Syncalathium (Asteraceae, Cichorieae), endemic to the Tibetan Plateau, SW China. Taxon 60(1):15–26. https://doi.org/10.1002/tax.601003
Zozomová-Lihová J, Krak K, Mandáková T, Shimizu KK, Španiel S, Vít P, Lysak MA (2014) Multiple hybridization events in Cardamine (Brassicaceae) during the last 150 years: revisiting a textbook example of neoallopolyploidy. Ann Bot 113(5):817–830. https://doi.org/10.1093/aob/mcu012
Acknowledgements
We thank Dr. Diego Fontaneto (CNR, Verbania Pallanza, Italy) for helpful comments on the statistical analysis, and two anonymous reviewers for comments that greatly improved the manuscript. The authors also thank Professor Daniele Bassi and the Department of Agricultural and Environmental Sciences for administrative help and organisation of the Postgraduate Programme, which made this study possible.
Funding
SV was supported by a Ph.D. Scholarship from the Department of Agricultural and Environmental Sciences, University of Milan, as part of the 35th round of funding in the Doctoral School of Agriculture, Environment and Bioenergy (Dottorato di Ricerca in Agricoltura, Ambiente e Bioenergia).
Author information
Authors and Affiliations
Contributions
SV and SP conceived the study. SV collated data and performed the analyses with support from SP and MM. SV wrote the manuscript and all authors were involved in manuscript editing.
Corresponding author
Ethics declarations
Conflict of interest
SP is an Editorial Board Member.
Additional information
Communicated by Marjana Westergren.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Sara Villa: early career author.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Villa, S., Montagna, M. & Pierce, S. Endemism in recently diverged angiosperms is associated with polyploidy. Plant Ecol 223, 479–492 (2022). https://doi.org/10.1007/s11258-022-01223-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11258-022-01223-y