Abstract
Plant mating systems may reflect an adaptation to a habitat type, with self-pollination being potentially common in unstable and disturbed conditions. We investigated the reproductive ecology of an Alpine, narrow-range toadflax, Linaria tonzigii Lona (Plantaginaceae), occurring in steep and dynamic mountain screes. We explored self-compatibility and spontaneous autogamy, seed viability, daily nectar production, pollinator behaviour and pollen transfer in wild populations, using hand pollination treatments, quantification of nectar volume, viability Tetrazolium assay, active pollinator sampling and video recordings, and UV-bright dust for pollen substitution. After ex novo sequencing of several genetic regions of L. tonzigii, we performed a multi-marker phylogenetic analysis of 140 Linaria species and tracked the occurrence of the self-compatibility trait. Our results showed that this species is self-compatible, pollinated mostly via spontaneous autogamy and pollinator-mediated geitonogamy, and self-pollinated seeds are as viable as cross-pollinated ones. Selfing could be due to pollinator rarity because, despite the studied species providing a high nectar volume, wild bees, moths and small beetles infrequently visited its flowers in the sparsely vegetated scree slopes. In addition, a preliminary survey showed a low genetic haplotype diversity in the study plant. Moreover, the phylogeny shows that self-compatibility is scattered in the tree, suggesting the adaptive nature of this reproductive trait in the genus Linaria. This study supports for theories that high prevalence of selfing is an adaptation to environments unfavourable for cross-pollination, and particularly where pollinators are rare, such as in perturbed, poorly vegetated high-elevation habitats.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants can reproduce in several ways, either asexually through vegetative propagation, or through sexual reproduction by means of self- or cross-pollination (Yang and Kim 2016). On one hand, self-pollination is expected to ensure progeny in condition of low pollinator activity, i.e. the hypothesis of “reproductive assurance” (Baker 1955; Stebbins, 1957). It is theorized that selfing holds a higher transmission rate compared to outcrossing, i.e. the “automatic selection” or “transmission advantage of selfing” hypothesis, (Fisher 1941; Stone et al. 2014) and it would assure the preservation of successful phenotypes that are particularly adapted to a local environment (Schmitt and Gamble 1990; Massol and Cheptou 2011). On the other hand, cross-pollination increases genetic diversity and therefore allows for a high potential of adaptation from a long-term perspective (Busch and Delph 2011; Cheptou 2019).
Although selfing bears all the risks related to inbreeding, theories state that it can occur when the benefits of transferring two copies of the maternal genes outweigh the risks of inbreeding depression (Lande and Schemske 1985; Charlesworth and Charlesworth 1987; Busch and Delph 2011). By selfing, the incidence of deleterious alleles is expected to rise, but this can be overcome through DNA purging, for example due to bottlenecks after extreme environmental events (Cooper and Cooper 1995). Furthermore, plant phylogenies clearly showed that plants transitioned from self-incompatibility to self-compatibility multiple times, independently (Igic et al. 2008; Goldberg et al. 2010), and this is even confirmed by cases of recent acquisition of the selfing (Voillemot et al. 2018). These aspects indicate the adaptive nature of the selfing trait, that could be advantageous at least in the short term (Cheptou 2019).
Despite the theories presented above describing how selfing could be advantageous for plant fitness, it seems less clear if selfing can be triggered by the type of habitat a plant lives in (Barrett et al. 2014). In general, it is believed that stable habitats are inhabited by highly outcrossing species, whilst unstable and disturbed habitats are usually colonized by selfing or even clonal plants (Lloyd 1980). Plausibly, this observation could be explained by associating self-pollination to environments where pollinators and/or mates are rare (Baker 1955). However, numerous exceptions to that observation have been noted and convincing experimental evidence is still needed (Barrett et al. 2014). In particular, when considering the incidence of selfing within speciose genera, there seems to be less consistency between reproductive strategy and habitat type. For example, both selfing and out crossing species of the genus Senecio occur in Australian unstable habitats (Lawrence 1985). Likewise, instances of selfing have been recorded in Linaria species both in stable habitats (e.g., grasslands and rock cliffs) and in unstable habitats (e.g., sand dunes and arable land) (Carrió et al. 2013). When comparing selfing rates between different populations, it was found that outcrossing predominates in areas where populations are frequently visited by pollinators, whilst in populations where visitation to flowers does not occur, selfing is the most common way to reproduce (Ness et al. 2010). This phenomenon shows that “reproductive assurance” could often interplay with “automatic transmission” in determining the presence and maintenance of the selfing trait in a given species (Barrett et al. 2014). In addition, it was shown that selfing carries an advantage where populations are fragmented and/or have low density, and when repeated colonization/extinction events have occurred (Good-Avila et al. 2008). These conditions link the selfing trait to taxon history, and not only to the type of habitat. Therefore, understanding how the plant reproductive strategies relate to the habitat is still a key question for understanding the reproductive biology of plants.
A wide variety of environments are colonized by species of the genus Linaria (Plantaginaceae), therefore it is a suitable model for studying mating systems. Whist some Linaria species have a large distribution range, a number of taxa occur in small areas, and many species grow in peculiar environmental conditions, e.g. rocks, cliffs or sand dunes (Segarra-Moragues and Mateu-Andrés 2007). A remarkable case is Linaria tonzigii Lona, calcicole toadflax with a narrow biogeographical range, that is endangered at the global scale according to the IUCN, and occurs in an endangered habitat listed in the EU Habitats Directive (European Commission 2009; Mangili and Rinaldi 2011). In addition, L. tonzigii has a high local cultural value and forms the logo of the botanical association Flora Alpina Bergamasca (F.A.B.). In addition, its reproductive biology is unknown and this can jeopardize even the simplest conservation measures (Falk 1992; Janečková et al. 2019). The species occurs on mountainous screes, a peculiar, disturbed habitat characterized by a dynamic and incoherent substrate with low vegetation cover (Fisher 1952; Cannone and Gerdol 2003). The ground instability is mainly caused by the frequent debris supply from the neighbouring cliffs, high material flow on the scree, absence of cementification in the rocky debris, and the instability is correlated with slope angle and other topological factors (Statham 1973; Cannone and Gerdol 2003). As a result of the low vegetation cover, plants occurring on the screes are exposed to wind and other harsh weather conditions that are typical for high altitudes (Fisher 1952). Such extreme conditions impact not only plant life (Fragnière et al. 2020) but also pollinator abundance and consequently also outcrossing rates (Mathews and Collins 2014).
In this study, we investigated the pollination biology of L. tonzigii, with reference to its habitat (mountain screes) and the evolutionary history of the genus. As L. tonzigii displays conspicuous zygomorphic flowers and a long nectar spur, we hypothesize a major contribution by long-tongued pollinators in both visitation and pollination (e.g. Vlašánková et al. 2017). In addition, the timing of daily nectar production could reveal the plant preference for a specific pollinator group (Zimmerman 1988), and thus we hypothesize that nectar production prior to dusk time would be connected to a preference for nocturnal pollination, whilst a production prior to and during the daylight could indicate a preference for diurnal ones (Zimmerman 1988). Therefore, in order to shed light on the pollination system of the plant, we focused on (a) its self-compatibility, spontaneous autogamy and cross-pollination; and on (b) the daily pattern of nectar allocation, the pollinator fauna and foraging behaviour. Furthermore, we aimed to (c) explore the genetic identity of the study species and place it in the phylogeny of the genus Linaria, information that was not previously available. The latter will be helpful for comparing the species' mating systems with that of related species.
Methods
The plant and the study area
Linaria tonzigii is a perennial hemicryptophyte plant that is endemic to a small area within the Bergamasque Orobie mountains, in the south-central part of the Alpine chain (area number 68 in the Alpenvereinseinteilung der Ostalpen classification and 29 in the SOUISA system). It occurs in an extremely restricted range (convex hull of about 10 km2) and is found in unstable and dynamic habitats characterized by incoherent accumulations of coarse gravel in particularly steep mountain screes (Giupponi and Giorgi 2019). It usually occurs in the alpine and subalpine altitudinal planes and exclusively on calcareous substrates (Orsenigo et al. 2016). The plant is about 6–12 cm tall, each stalk usually hosts 3–6 (rarely 10) single flowers, and the flowering phenology lasts from around mid-June to mid-July, with some individuals flowering earlier or later in the season.
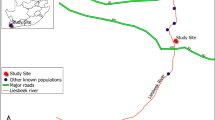
Most of the field work for this study took place in two locations on Mt. Arera, with little to no isolation, that can be considered as a unique population (N 45.933, E 9.804 at 2100 m a.s.l.; N 45.929, E 9.807 at 2000 m a.s.l., 500 m apart). An additional population located in Mt. Cavallo was included in the pollinator survey (N 46.034, E 9.695 at 2100 m a.s.l., 12 km from Mt. Arera). These populations occur on screes with a slope range of 53.5–60.4% (calculated as difference in elevation between top and bottom of the scree divided by their distance).
Pollination biology
In the field, the study plants in pre-flowering stage were covered with bags of fine mesh to exclude pollinators during the time of the investigation. Twenty plant individuals received four pollination treatments each, as follows: (1) Cross-pollination by hand (with pollen from various randomly chosen individuals), (2) Self-pollination by hand (with a flower own pollen), (3) Autogamy (autonomous self-pollination), (4) Emasculation (test of agamospermy). These treatments were carried out during the peak flowering period of the population and were applied during two consecutive days in the field. Single flowers were deprived of the anthers during the pre-flowering stage (except those of the autogamy treatment) and the flowers of a given individual plant were treated at the same time. To avoid an effect of the maternal plant on the results, all four treatments were applied to the flowers of each selected individual simultaneously, as in Hamilton and Mitchell-Olds (1994).
The seed set for each of these treatments was calculated as the number of seeds per capsule and compared to the seed set of plants exposed to open pollination. The seeds in each capsule of these treatments were weighed and the total weight was divided by the number of seeds relative to each fruit (Kern ABT 120-5DNM, 0.00001 g). The seed viability in open-, self- and cross- pollination treatments was tested with a tetrazolium assay at 1% concentration for 24 h after cutting the seeds in half for exposing the embryo to the solution.
The number and mean weight of seeds per capsule were analysed as response variables in linear mixed effect models, pollination treatment was a predictor and plant individual identity was a random intercept. The assumptions of normality were checked visually. The percentage of viable seeds per capsule was tested as a proportion of the total number of seeds (viable + non-viable seeds) in a generalized mixed effect model with binomial error distribution, with the pollination treatment as a predictor variable and the plant individual identity as a random intercept.
To measure the difference and significance of seed number, weight, or viability of each treatment in relation to open pollination, the regression slopes and the Z-statistic were used. For these and the above detailed regressions, the package glmmTMB in R was used (Brooks et al. 2017; R Core Team 2017). A post hoc test comparing the seed number associated to cross- and self- pollination was performed with the package emmeans (Lenth 2020).
Nectar production and sugar concentration
Flowers were emptied in each location and the sugar concentration in the nectar was measured with a field refractometer (Eclipse Handheld Refractometer − Sugar, 0–50% − Bellingham + Stanley, London), one flower per plant from randomly chosen individuals (different from those who received pollination treatment, see above). For recording the pattern of nectar production during the day, the nectar volume was measured with microcapillaries (0.5 μL microcapillary tube Drummond Microcaps, Sigma-Aldrich) by comparing the volume of 24 h with that produced after night and every 6 h during the day (i.e., collections at 9:00 A.M, 3:00 P.M., 9:00 P.M. from the same flowers, with the dusk time and the sun set usually happening before 9 PM in the sampling area). All flowers were covered in bags of fine mesh to exclude flower visitors, and corolla spurs were previously emptied to assure nectar production during the time frame indicated above. The nectar volume of a total of 48 flowers was measured, half of which were used for the 24 h-volume assessment, and the others for the 6-h production.
The volume of nectar was analysed as a response variable in generalized mixed effect models with Gamma error distribution and log-link function. Categorical predictor variables were whether nectar was taken after 24 h or during a phase of the day. Plant individual identity was the random intercept. The difference in estimated coefficients associated to the predictors and its significance was tested with the delta beta and the associated Z-statistic, using the package glmmTMB in R (Brooks et al. 2017; R Core Team 2017).
Pollinator monitoring, pollinator behaviour and pollen analysis
Flower visitors found inside or entering the corollas and touching the reproductive structures of the flowers were sampled with a mouth aspirator or by hand netting. Surveys consisted in walking repeatedly across the screes during day and evening, approximately between 8:00 A.M. and 11:00 P.M. for 20 days distributed in the two populations of Mt. Arera in 2018 and 2019 and a population of Mt. Cavallo in 2019. Additionally, video cameras with infrared (IR) night vision were placed both during day and night with continuous recording (SuperEye RJ0090-UK, OneThingCam™).
Pollen from the insects body was removed using fuchsin jelly (i.e., from the head of large bees and from the body of small beetles), which was subsequently melted onto glass slides and inspected with an optic microscope (Leitz Laborulux K). These slides were compared to a reference slide of pollen of L. tonzigii created by touching freshly opened L. tonzigii flowers with fuchsin gel (this reference slide is stored in MIB:ZPL herbarium Department of Biotechnology and Biosciences, University of Milano-Bicocca, Milan, Italy).
To quantify self-pollination and geitonogamy in wild populations, we tracked the flow of the pollen with pollen-substitutes. Flowers of 10 randomly chosen individuals were dusted with UV-reflective dust (RadGlo RC 33, Radiant Color, Europark 1046 B 3530 Houthalen) on their inner surface of the corolla near to and on the anthers, similarly to Wanderley et al. (2020). At night, a UV emitting torch (MorPilot, 51 Led) was used for checking the inner corolla of all flowers on the same stalk as the source UV flower and on all individual plants occurring within one meter radius from the dusted, source flower.
DNA extraction and phylogenetic analysis
One to five plants of L. tonzigii, L. vulgaris and L. alpina (the three Linaria species occurring in the same administrative province where the study was conducted) were sampled in several populations for subsequent laboratory analysis (Table S1 in Online Resource Appendix 1). These samples were vouchered following the protocol specified by the Global Registry of Biodiversity Repositories (http://grbio.org/) and the data standards for DNA barcode records (http://www.boldsystems.org/docs/dwg_data_standards-final.pdf). Young leaves were used for genetic screening (1–3 per plant) and they were stored immediately at − 20 °C until laboratory analysis. The specimens were stored in herbarium at the Department of Biotechnology and Biosciences, University of Milano-Bicocca, Milan, Italy (herbarium code MIB:ZPL).
One hundred milligrams of each sample were used for DNA extraction using Plant DNeasy Isolation and Purification kit (Qiagen, Hilden, Germany). Purified DNA concentration and quality of each sample were estimated fluorometrically with a NanoDrop™ 1000 Spectrophotometer (Thermo Scientific, USA), by measuring the absorbance (Abs) at 260, 280, and 230 nm. DNA-based characterization of L. tonzigii was performed by amplification and sequencing of four DNA barcode markers (i.e., the plastidial rbcL, matK, the intergenic spacer trnH-psbA, and the nuclear ITS) and other three genomic regions that showed variation in studies of other plant species of a range size similar to L. tonzigii (i.e., agt1, rpl32-trnL and trnL-trnF; Blanco-Pastor and Vargas 2013; Zecca et al. 2017). For each locus, PCR amplification was performed using puReTaq Ready-To-Go PCR beads (Amersham Bioscience, Freiburg, Germany) in a 25 μL reaction according to the manufacturer's instructions. PCR cycles consisted of an initial denaturation step for 7 min at 94 °C, 35 cycles of denaturation (45 s at 94 °C), annealing (30 s at 50 °C for rbcL and trnL-trnF, 53 °C for matK and trnH-psbA, 55 °C for ITS, agt1 and rpl32-trnL), and extension (1 min at 72 °C), and a final extension at 72 °C for 7 min. The primer pairs used were 1F/724R for rbcL (Fay et al. 1998), 390F/1326R for matK (Cuénoud et al. 2002), psbA/trnH for trnH-psbA (Newmaster et al. 2008), p5/u4 for ITS (Cheng et al. 2016), AGT1_F1/AGT1_R1 for agt1 (Kovacova et al. 2014), rpL32-F/trnL for rpl32-trnL (Shaw et al. 2007) and trnT-F_cF/trnT-F_fR for trnL-trnF (Taberlet et al. 1991).
PCR products were bidirectionally sequenced using an ABI 155 3730XL automated sequencer at Macrogen Inc., Korea. Sequence editing and alignment were performed using Bioedit (Hall 1999) and the sequence data were submitted to the European Bioinformatics Institute of the European Molecular Biology Laboratory (EMBL-EBI, see Table S1 in Online Resource Appendix S1). The variability among L. tonzigii samples was tested with the regions trnH-psbA, agt1, rpl32-trnL and trnL-trnF as in Blanco-Pastor and Vargas (2013) and Zecca et al. (2017). The genetic variation at these markers was described by calculating the number of haplotypes, the haplotype diversity and the nucleotide diversity with DnaSP v5.10.1 software (Librado and Rozas 2009).
To analyse the phylogeny, we downloaded sequences for nuclear (ITS and agt1) and plastidial (rbcL, matK, trnH-psbA, rpl32-trnL, and trnL-trnF) markers in all the Linaria species and subspecies available in Genbank using the rentrez package version 1.2.1 (Winter 2017) in R (R Core Team 2017). We also obtained sequences for these markers in Maurandya scandens, Chaenorhinum macropodum, Antirrhinum majus, and A. graniticum to be used as outgroups. Since the genetic identity of the analysed L. tonzigii specimens were very similar to each other (see section ‘Results’), only the sequence of the Mt. Arera haplotype was used in the phylogenetic analysis. Downloaded and newly sequenced markers were aligned using MAFFT version 7.307 (Katoh and Standley 2013) and the best evolutionary model and partition scheme was estimated using PartitionFinder version 2.1 (Lanfear et al. 2017). Phylogenetic trees were performed using both, Maximum Likelihood as implemented in RaxML version 8.2.9 (Stamatakis 2014) and Bayesian Inference as implemented in MrBayes version 3.2.6 (Ronquist and Huelsenbeck 2003) and visualized using the ape 5.1 library (Paradis and Schliep 2018) in R. Bayesian analyses were performed using 4 chains, 20 million generations sampled every 1000 generations each with 0.25 as relative burning value. For maximum likelihood analyses, branch support was estimated after 100 bootstrap replicates.
Data on self-compatibility for Linaria species were taken from Valdés (1970), Docherty (1982), Carrió et al. (2013), Voillemot et al. (2018).
Results
Pollination biology
Pollination treatment was a significant categorical predictor of the seed set (χ2 = 38.893, df = 1, P ≤ 0.001; Fig. 1a). Flowers exposed to open pollination produced less seeds than the hand cross- or self-pollinated ones, and open pollination produced a similar number of seeds as spontaneous autogamy (Table 1). The numbers of cross- and self-pollinated seeds were not significantly different in the post hoc test (estimated difference cross-self = − 6.35, SE = − 3.38, P = 0.07). Testing for agamospermy did not yield any seeds.
Boxplots of seed set (a), seed weight (b) and seed viability (c) of L. tonzigii from the pollination treatments of hand cross-pollination (HCP), hand self-pollination (HSP), autonomous self-pollination (ASP), agamospermy (AGP) and control open pollination (OP). Significant differences to open pollination are indicated with “*” and statistical details are reported in Table 1
The seed weight was significantly related to the pollination treatment (χ2 = 47.669, df = 1, P ≤ 0.001; Fig. 1b), however, there was no significant difference between open pollination and either hand cross- or self-pollination or spontaneous autogamy (Table 1). The rate of seed viability was not significantly predicted by the pollination treatment (χ2 = 0.296, df = 1, P = 0.862; Table 1, Fig. 1c).
Daily nectar production and sugar concentration
Sugar concentration in the nectar averaged around 35.4%. The nectar volume produced varied during the day: the volume produced in 24 h was similar to that produced in the morning, but it was significantly higher than that produced around noon and evening (Table 2).
Pollinator monitoring and behaviour
In total, 141 h of video were recorded and about 240 h of active sampling carried out. Despite the high amount of time spent in sampling (actively or with videocameras), few insects were found.
The active sampling of pollinators yielded insects of different functional groups and taxonomical orders that were either inside the flowers or foraging on flower resources, and scattered in different flowers along the surveyed screes (total sums indicated): 32 small thrips (Thysanoptera), 41 small beetles of Coleoptera: Staphylinidae (Staphylinidae: Eusphalerum sp.), a large moth of Lepidoptera: Noctuidae, three large wild bees of Hymenoptera: Apidae that were two queens of the bumblebee Bombus monticola and a female bee of Anthophora quadrimaculata. All these insects are stored as voucher specimens in PB collection. Video recordings yielded additional flower visitors and their behaviour, namely: a large moth of Lepidoptera: Noctuidae and a butterfly of Erebia sp. (Lepidoptera: Nymphalidae) collecting nectar, a queen of the bumblebee Bombus monticola collecting nectar and a worker of Bombus sp. collecting pollen. In addition, an Andrena cf. rogenhoferi bee was recorded and landed on the corolla of a L. tonzigii flowers carrying a lot of light-yellow pollen, which is a colour very similar to that of L. tonzigii, but it was not seen entering the flower. Videos showed very clearly that whilst foraging, the bumblebees visited in fast succession a relatively high number of L. tonzigii flowers (i.e., 19, 7, 6 flowers), whilst the moth and the butterfly visited only a few flowers (1 and 2 flowers, respectively). Instead, the vegetated meadows growing at the sides of the screes host a high activity of pollinating insects both during the day and during the night. For instance, eight moths were found visiting flowers of different families during a transect walk of one hour, but none on L. tonzigii during that time.
The captured specimens were examined in the laboratory and pollen of very similar morphology to L. tonzigii was found on the body of both small beetles and of the large bees, and the latter carried a higher quantity of pollen on their body (i.e. their head) than the small beetles. 99% of the pollen taken with the fuchsin jelly was identified as belonging to L. tonzigii. Pollen was not found on the one moth collected.
UV-reflective dust indicated that most of the investigated flowers did not receive dust (47.92%), whilst flowers of the dust-treated stalks received more dust than those of surrounding stalks (29.86% and 22.22%, respectively).
Some flowers had a hole in the spur, a particular feature compatible with nectar robbing, especially in those flowers growing closer to the highly vegetated meadows.
Phylogenetic analysis and genetic identity
The genetic diversity among the analysed specimens was very low, with three poorly differentiated haplotypes at agt1, trnH-psbA and rpl32-trnL (see Table S2 in Online Resource Appendix S1).
A total of 140 species and subspecies of the genus Linaria and Nuttallanthus (Linaria s.l.) were included in the phylogenetic analysis. The best models for the different partitions were SYM + I + G (ITS), GTR + G (matK), JC (rbcL) and HKY (psbA-trnH). L. tonzigii clustered within the Clade E (sensu Fernández-Mazuecos et al. 2013b), together with species of the Section Speciosae: L. purpurea, L. repens, and L. capraria (Fig. 2). This position in the tree is consistently observed when nuclear and plastidial markers are used separately (see Figs. S1–S4 in Online Resource Appendix S2).
Phylogenetic analysis of the genus Linaria s.l. based on concatenated genetic markers (ITS, agt1, rbcL, matK, trnH-psbA, rpl32-trnL, trnL-trnF). Species sequences newly generated for this study are indicated in bold. Species mating system data compiled from the literature are indicated as SI (self-incompatible), SC (self-compatible) and SI/SC (both self-incompatible and self-compatible depending on the original study), and L. tonzigii is SC (this study). The branches of the outgroup species are in black at the bottom of the tree (descending vertically Maurandya scandens, Chaenorhinum macropodum, Antirrhinum majus, A. graniticum)
Discussion
In this study, we investigated several aspects of the pollination ecology and genetic identity of a toadflax species, L. tonzigii, inhabiting Mediterranean screes, a threatened habitat according to the IUCN (European Commission 2009). The target species is a rare and steno-endemic plant, threatened of extinction (“Endangered” category in the IUCN Red List, Mangili and Rinaldi 2011). Based on the literature records of other Linaria species and on the flower morphology (i.e., zygomorphic flowers with a nectar spur), we previously hypothesized that the pollinating fauna would be composed uniquely of long-tongued insects. In detail, other Linaria species are visited by large bees, bee flies of the Bombyliidae family and both diurnal and nocturnal Lepidopterans (Sutton 1988; Stout et al. 2000; Fernández-Mazuecos et al. 2013a), which are insects with long mouthparts. In addition, since the flowers of L. tonzigii have a very long spur, we hypothesised that if nectar is produced in the evening, a preference for nocturnal pollination could be expected (Zimmerman 1988).
This field study of L. tonzigii only partly confirmed these expectations. Our data showed a prevalence of diurnal pollinators, which also matched the higher production of nectar volume during morning hours. This result hints that the study plant allocates resources to diurnal pollinators (Zimmerman 1988), and this conclusion could potentially be strengthened in future by recording the daily dynamics of the scent production. This is in congruence with a general pattern that pollinators usually visit flowers when the nectar resources are the highest (Real and Rathcke 1991; Aguiar and Pansarin 2013; Biella et al. 2019). In addition, our survey indicated that not all flower visitors were long-tongued insects, and not all the pollinators were equal in terms of abundance and visitation behaviour. Large bees with long mouthparts were rarely found in the flowers, although they visited several flowers per foraging bout, whilst small beetles with short mouthparts were more abundant visitors, but apparently very sedentary in the flowers. Moreover, the fact that open pollinated flowers produced similar seeds number as the autogamic flowers, together with the pollinator behaviour recorded in the videos and the pollen tracking suggest that the majority of the pollen flow is within the same flower (spontaneous autogamy) or between flowers of the same stalk (geitonogamous pollination). Thus, these evidences suggest for a prevalence of self-pollination in L. tonzigii. The reason why large and mobile pollinators rarely visited L. tonzigii may be due to low vegetation cover in this habitat (Giupponi and Giorgi 2019), and that highly mobile winged pollinators would prefer foraging in patches with generally higher plant abundance (Redhead et al. 2016). We observed seemingly higher pollinator activity in nearby vegetated meadows than in the screes habitat, although this was not quantified. Rare visits to flowers agrees with the evidence that the plant is pollen limited, because the open pollination flowers produced less seeds than the hand pollination treatments, that is a clear indication of low amount of pollen delivered by pollinators (Kalisz and Vogler 2003). This agrees with the “reproductive assurance” hypothesis. Furthermore, it could be expected that this plant spread also vegetatively, as it grows with underground radicated stalks (see Fig. S5 in Online Resource Appendix S1). These features are compatible with a vegetative reproduction (Pfeiffer et al. 2006), but future studies will need to provide empirical evidence confirming the role of asexual reproduction in this species.
Self- and cross- pollination experiments indicated that the species is self-compatible and that spontaneous autogamy produced as many seeds as open pollinated ones. Moreover, the self-pollinated seeds were equally as viable as cross-pollinated ones. All these results strongly indicate no avoidance of self-pollination and that fruiting is more important than the source of the fecundating pollen (i.e., whether the pollen is from the same flower/individual or from different ones). Self-pollination bears implications for genetic diversity. The hypothesis of automatic advantage of selfing implies that self-pollination is a sustainable strategy if the level of genetic diversity is low, as after DNA purging and bottlenecks (Busch and Delph 2011). We preliminary checked for genetic differences between individuals of L. tonzigii collected both in the same population and in different parts of the distribution range using similar genetic markers as other studies focused on geographic ranges comparable to ours (Zecca et al. 2017; Blanco-Pastor et al. 2019). Although the loci used here are less variable than what would be obtainable with microsatellites, DD-Rad or other genomic approaches based on high-troughput sequencing, we still found a very low genetic diversity amongst individuals. Even if preliminary, this result is not contradicting a scenario of severe diversity reduction during the history of the species or due to selfing, similar to reports in other cases of plants with low genetic diversity (Szczecińska et al. 2016; Blambert et al. 2016; Zecca et al. 2017).
Previous studies supposed that the majority of Linaria species are self-incompatible (Bruun 1937; Valdés 1970). However, the species pool considered in those studies was poor. In the multi-marker phylogenetic analysis in our study, the high frequency of the missing information on self-compatibility limits the drawing of clear conclusions on the evolutionary patterns of this character at the level of the entire genus, until additional data are made available. Yet, the fact that the self-compatibility trait occurs in different clades indicate that in Linaria spp. the acquisition of this trait arose multiple times and it may be interpreted as an adaptation to local environments, as reported in other plant genera as well (Goodwillie 1999; Goldberg et al. 2010). This seems to be corroborated by another species of this genus that showed different levels of self-compatibility in different parts of its range, varying from self-incompatible to self-compatible populations (Voillemot et al. 2018). Some similar cases are occasionally reported for other plant genera (Busch et al. 2011). The advantage of self-pollination is that it can assure reproduction when pollen flow levels are low (Cheptou 2019), such as in unstable environments with unreliability of pollinator presence and activity, as is the case in dynamic mountain screes inhabited by L. tonzigii. Autonomous selfing interplays with pollinator-mediated selfing (geitonogamy), that is considered to assure reproduction in populations where flowering conspecifics are scarce and sparsely distributed (Lloyd 1979; Kropf and Renner 2008). In the dynamic and sparsely vegetated screes of this study, individuals of L. tonzigii are often far apart and geitonogamy could benefit reproduction in this challenging environment, in combination with other ways of reproduction. On the long-term, however, it is believed that self-compatibility could become disadvantageous as it could decrease the potential for adaptability and even increase the extinction risk (Goldberg et al. 2010). As indicated by Cheptou (2019), only by tracking the demographic history of the species and by quantifying the roles of inbreeding depression, DNA purging and rearrangement, and the need of reproductive assurance will clarify if L. tonzigii populations will be likely to persist over time.
Availability of data
References
Aguiar JMRBV, Pansarin ER (2013) Does Oeceoclades maculata (Orchidaceae) reabsorb nectar? Eur J Environ Sci 3:113–118
Baker HG (1955) Self-compatibility and establishment after “long-distance” dispersal. Evolution 9:347–349. https://doi.org/10.1111/j.1558-5646.1955.tb01544.x
Barrett SCH, Arunkumar R, Wright SI (2014) The demography and population genomics of evolutionary transitions to self-fertilization in plants. Philos Trans R Soc B 369:20130344. https://doi.org/10.1098/rstb.2013.0344
Biella P, Akter A, Ollerton J, Tarrant S, Janeček Š, Jersáková J, Klecka J (2019) Experimental loss of generalist plants reveals alterations in plant-pollinator interactions and a constrained flexibility of foraging. Sci Rep 9:7376. https://doi.org/10.1038/s41598-019-43553-4
Blambert L, Mallet B, Humeau L, Pailler T (2016) Reproductive patterns, genetic diversity and inbreeding depression in two closely related Jumellea species with contrasting patterns of commonness and distribution. Ann Bot 118:93–103. https://doi.org/10.1093/aob/mcw014
Blanco-Pastor JL, Vargas P (2013) Autecological traits determined two evolutionary strategies in Mediterranean plants during the Quaternary: low differentiation and range expansion versus geographical speciation in Linaria. Mol Ecol 22:5651–5668. https://doi.org/10.1111/mec.12518
Blanco-Pastor JL, Fernández-Mazuecos M, Coello AJ, Pastor J, Vargas P (2019) Topography explains the distribution of genetic diversity in one of the most fragile European hotspots. Divers Distrib 25:74–89. https://doi.org/10.1111/ddi.12836
Brooks ME, Kristensen K, van Benthem KJ, Magnusson A, Berg CW, Nielsen A, Skaug HJ, Machler M, Bolker BM (2017) glmmTMB balances speed and flexibility among packages for zero-inflated generalized linear mixed modeling. R J 9:378–400. https://doi.org/10.32614/RJ-2017-066
Bruun HG (1937) Genetical notes on Linaria, I—II. Hereditas 22:395–400. https://doi.org/10.1111/j.1601-5223.1937.tb02657.x
Busch JW, Delph LF (2011) The relative importance of reproductive assurance and automatic selection as hypotheses for the evolution of self-fertilization. Ann Bot 109:553–562. https://doi.org/10.1093/aob/mcr219
Busch JW, Joly S, Schoen DJ (2011) Demographic signatures accompanying the evolution of selfing in Leavenworthia alabamica. Mol Biol Evol 28:1717–1729. https://doi.org/10.1093/molbev/msq352
Cannone N, Gerdol R (2003) Vegetation as an ecological indicator of surface instability in rock glaciers. Arct Antarct Alp Res 35:384–390. https://doi.org/10.1657/1523-0430(2003)035[0384:VAAEIO]2.0.CO;2
Carrió E, Güemes J, Herreros R (2013) Pollination biology in an endangered rocky mountain toadflax (Linaria cavanillesii). Plant Biosyst Int J Deal Asp Plant Biol 147:354–363. https://doi.org/10.1080/11263504.2012.736424
Charlesworth D, Charlesworth B (1987) Inbreeding depression and its evolutionary consequences. Annu Rev Ecol Syst 18:237–268. https://doi.org/10.1146/annurev.es.18.110187.001321
Cheng T, Xu C, Lei L, Li C, Zhang Y, Zhou S (2016) Barcoding the kingdom Plantae: new PCR primers for ITS regions of plants with improved universality and specificity. Mol Ecol Resour 16:138–149. https://doi.org/10.1111/1755-0998.12438
Cheptou P-O (2019) Does the evolution of self-fertilization rescue populations or increase the risk of extinction? Ann Bot 123:337–345. https://doi.org/10.1093/aob/mcy144
Cooper A, Cooper RA (1995) The Oligocene bottleneck and New Zealand biota: genetic record of a past environmental crisis. Proc R Soc Lond B 261:293–302. https://doi.org/10.1098/rspb.1995.0150
Cuénoud P, Savolainen V, Chatrou LW, Powell M, Grayer RJ, Chase MW (2002) Molecular phylogenetics of Caryophyllales based on nuclear 18S rDNA and plastid rbcL, atpB, and matK DNA sequences. Am J Bot 89:132–144. https://doi.org/10.3732/ajb.89.1.132
Docherty Z (1982) Self-incompatibility in Linaria. Heredity 49:349–352. https://doi.org/10.1038/hdy.1982.108
European Commission (2009) Composite report on the conservation status of habitat types and species as required under Article 17 of the Habitats Directive—report from the Commission to the Council and the European Parliament COM 358. European Commission, Brussels
Falk DA (1992) From conservation biology to conservation practice: strategies for protecting plant diversity. In: Fiedler PL, Jain SK (eds) Conservation biology: the theory and practice of nature conservation, preservation and management. Springer, Boston, pp 397–431
Fay MF, Bayer C, Alverson WS, de Bruijn AY, Chase MW (1998) Plastid rbcL sequence data indicate a close affinity between Diegodendron and Bixa. Taxon 47:43–50. https://doi.org/10.2307/1224017
Fernández-Mazuecos M, Blanco-Pastor JL, Gómez JM, Vargas P (2013a) Corolla morphology influences diversification rates in bifid toadflaxes (Linaria sect. Versicolores). Ann Bot 112:1705–1722. https://doi.org/10.1093/aob/mct214
Fernández-Mazuecos M, Blanco-Pastor JL, Vargas P (2013b) A phylogeny of toadflaxes (Linaria Mill.) based on nuclear internal transcribed spacer sequences: systematic and evolutionary consequences. Int J Plant Sci 174:234–249. https://doi.org/10.1086/668790
Fisher RA (1941) Average excess and average effect of a gene substitution. Ann Eugen 11:53–63. https://doi.org/10.1111/j.1469-1809.1941.tb02272.x
Fisher FJF (1952) Observations on the vegetation of screes in Canterbury, New Zealand. J Ecol 40:156–167. https://doi.org/10.2307/2258027
Fragnière Y, Pittet L, Clément B, Bétrisey S, Gerber E, Ronikier M, Parisod C, Kozlowski G (2020) Climate change and alpine screes: no future for glacial relict Papaver occidentale (Papaveraceae) in Western Prealps. Diversity 12:346. https://doi.org/10.3390/d12090346
Giupponi L, Giorgi A (2019) A contribution to the knowledge of Linaria tonzigii Lona, a steno-endemic species of the Orobie Bergamasche Regional Park (Italian Alps). Eco Mont J Prot Mt Areas Res 11:16–24. https://doi.org/10.1553/eco.mont-11-1s16
Goldberg EE, Kohn JR, Lande R, Robertson KA, Smith SA, Igić B (2010) Species selection maintains self-incompatibility. Science 330:493–495. https://doi.org/10.1126/science.1194513
Good-Avila SV, Mena-Alí JI, Stephenson AG (2008) Genetic and environmental causes and evolutionary consequences of variations in self-fertility in self incompatible species. In: Franklin-Tong VE (ed) Self-incompatibility in flowering plants. Springer, Berlin, Heidelberg, pp 33–51
Goodwillie C (1999) Multiple origins of self-compatibility in Linanthus Section Leptosiphon (Polemoniaceae): phylogenetic evidence from Internal-Transcribed-Spacer sequence data. Evolution 53:1387–1395. https://doi.org/10.1111/j.1558-5646.1999.tb05403.x
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98. https://doi.org/10.14601/Phytopathol_Mediterr-14998u1.29
Hamilton MB, Mitchell-Olds T (1994) The mating system and relative performance of selfed and outcrossed progeny in Arabis fecunda (Brassicaceae). Am J Bot 81:1252–1256. https://doi.org/10.2307/2445400
Igic B, Lande R, Kohn JR (2008) Loss of self-incompatibility and its evolutionary consequences. Int J Plant Sci 169:93–104. https://doi.org/10.1086/523362
Janečková P, Janeček Š, Bartoš M, Hrázskỳ Z (2019) Reproductive system of the critically endangered taxon Gentianella praecox subsp. bohemica. Preslia 91:77–92. https://doi.org/10.23855/preslia.2019.077
Kalisz S, Vogler DW (2003) Benefits of autonomous selfing under unpredictable pollinator environments. Ecology 84:2928–2942. https://doi.org/10.1890/02-0519
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Kovacova V, Zluvova J, Janousek B, Talianova M, Vyskot B (2014) The evolutionary fate of the horizontally transferred Agrobacterial Mikimopine Synthase gene in the genera Nicotiana and Linaria. PLoS ONE 9:e113872. https://doi.org/10.1371/journal.pone.0113872
Kropf M, Renner SS (2008) Pollinator-mediated selfing in two deceptive orchids and a review of pollinium tracking studies addressing geitonogamy. Oecologia 155:497–508. https://doi.org/10.1007/s00442-007-0919-4
Lande R, Schemske DW (1985) The evolution of self-fertilization and inbreeding depression in plants. I. Genetic models. Evolution 39:24–40. https://doi.org/10.1111/j.1558-5646.1985.tb04077.x
Lanfear R, Frandsen PB, Wright AM, Senfeld T, Calcott B (2017) PartitionFinder 2: new methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Mol Biol Evol 34:772–773. https://doi.org/10.1093/molbev/msw260
Lawrence ME (1985) Senecio L. (Asteraceae) in Australia: reproductive biology of a genus found primarily in unstable environments. Aust J Bot 33:197–208. https://doi.org/10.1071/bt9850197
Lenth R (2020) emmeans: estimated marginal means, aka least-squares means. 2018. R Package Version 144 HttpsCRANR-Proj
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinform Oxf Engl 25:1451–1452. https://doi.org/10.1093/bioinformatics/btp187
Lloyd DG (1979) Some reproductive factors affecting the selection of self-fertilization in plants. Am Nat 113:67–79. https://doi.org/10.1086/283365
Lloyd DG (1980) Demographic factors and mating patterns in angiosperms. In: Solbrig OT (ed) Demography and evolution in plant populations. Blackwell, Oxford, pp 67–88
Mangili F, Rinaldi G (2011) Linaria tonzigii. IUCN Red List Threat Species 2011 e.T162172A5553011. https://doi.org/10.2305/IUCN.UK.2011-1.RLTS.T162172A5553011.en
Massol F, Cheptou P-O (2011) Evolutionary syndromes linking dispersal and mating system: the effect of autocorrelation in pollination conditions. Evolution 65:591–598. https://doi.org/10.1111/j.1558-5646.2010.01134.x
Mathews K, Collins B (2014) Plant and pollinator communities of high elevation rock outcrops. Nat Areas J 34:300–309. https://doi.org/10.3375/043.034.0306
Ness RW, Wright SI, Barrett SC (2010) Mating-system variation, demographic history and patterns of nucleotide diversity in the tristylous plant Eichhornia paniculata. Genetics 184:381–392. https://doi.org/10.1534/genetics.109.110130
Newmaster SG, Fazekas AJ, Steeves RAD, Janovec J (2008) Testing candidate plant barcode regions in the Myristicaceae. Mol Ecol Resour 8:480–490. https://doi.org/10.1111/j.1471-8286.2007.02002.x
Orsenigo S, Bacchetta G, Calevo J, Castello M, Cogoni D, Gennai M, Licht W, Montagnani C, Perrino EV, Pinna SM, Silletti G, Vela E, Viciani D, Vidali M, Wagensommer RP, Zappa E, Fenu G (2016) Global and regional IUCN Red List assessments: 1. Ital Bot 1:61–85. https://doi.org/10.3897/italianbotanist.1.8647
Paradis E, Schliep K (2018) ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35:526–528
Pfeiffer T, Fritz S, Stech M, Frey W (2006) Vegetative reproduction and clonal diversity in Rhytidium rugosum (Rhytidiaceae, Bryopsida) inferred by morpho-anatomical and molecular analyses. J Plant Res 119:125–135. https://doi.org/10.1007/s10265-005-0255-x
R Core Team (2017) R: a language and environment for statistical computing. R Foundation for Statistical Computing. ISBN 3-900051-07-0, Vienna, Austria
Real LA, Rathcke BJ (1991) Individual variation in nectar production and its effect on fitness in Kalmia latifolia. Ecology 72:149–155. https://doi.org/10.2307/1938910
Redhead JW, Dreier S, Bourke AFG, Heard MS, Jordan WC, Sumner S, Wang J, Carvell C (2016) Effects of habitat composition and landscape structure on worker foraging distances of five bumble bee species. Ecol Appl 26:726–739. https://doi.org/10.1890/15-0546
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574. https://doi.org/10.1093/bioinformatics/btg180
Schmitt J, Gamble SE (1990) The effect of distance from the parental site on offspring performance and inbreeding depression in Impatiens capensis: a test of the local adaptation hypothesis. Evolution 44:2022–2030. https://doi.org/10.1111/j.1558-5646.1990.tb04308.x
Segarra-Moragues JG, Mateu-Andrés I (2007) Levels of allozyme diversity in closely related toadflaxes (Linaria, Plantaginaceae) and their correspondence with the breeding systems of the species. Conserv Genet 8:373–383. https://doi.org/10.1007/s10592-006-9176-z
Shaw J, Lickey EB, Schilling EE, Small RL (2007) Comparison of whole chloroplast genome sequences to choose noncoding regions for phylogenetic studies in angiosperms: the tortoise and the hare III. Am J Bot 94:275–288. https://doi.org/10.3732/ajb.94.3.275
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinform Oxf Engl 30:1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Statham I (1973) Scree slope development under conditions of surface particle movement. Trans Inst Br Geogr. https://doi.org/10.2307/621711
Stebbins GL (1957) Self fertilization and population variability in the higher plants. Am Nat 91:337–354
Stone JL, VanWyk EJ, Hale JR (2014) Transmission advantage favors selfing allele in experimental populations of self-incompatible Witheringia solanacea (Solanaceae). Evolution 68:1845–1855. https://doi.org/10.1111/evo.12419
Stout JC, Allen JA, Goulson D (2000) Nectar robbing, forager efficiency and seed set: bumblebees foraging on the self incompatible plant Linaria vulgaris (Scrophulariaceae). Acta Oecol 21:277–283. https://doi.org/10.1016/S1146-609X(00)01085-7
Sutton DA (1988) A revision of the tribe Antirrhineae. Oxford University Press, Oxford
Szczecińska M, Sramko G, Wołosz K, Sawicki J (2016) Genetic diversity and population structure of the rare and endangered plant species Pulsatilla patens (L.) Mill in East Central Europe. PLoS ONE. https://doi.org/10.1371/journal.pone.0151730
Taberlet P, Gielly L, Pautou G, Bouvet J (1991) Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Mol Biol 17:1105–1109. https://doi.org/10.1007/BF00037152
Valdés B (1970) Taxonomía experimental del género Linaria IV. Reproducción sexual: autogamia y polinización intraespecífica. Bol R Soc Esp Hist Nat Biol 68:79–89
Vlašánková A, Padyšáková E, Bartoš M, Mengual X, Janečková P, Janeček Š (2017) The nectar spur is not only a simple specialization for long-proboscid pollinators. New Phytol 215:1574–1581. https://doi.org/10.1111/nph.14677
Voillemot M, Rougemont Q, Roux C, Pannell JR (2018) The divergence history of the perennial plant Linaria cavanillesii confirms a recent loss of self-incompatibility. J Evol Biol 31:136–147. https://doi.org/10.1111/jeb.13209
Wanderley AM, dos Santos EKR, Galetto L, Benko-Iseppon AM, Machado ICS (2020) Pollen flow within and among isolated populations of two rare, self-compatible plant species from inselbergs of Northeast Brazil. Plant Ecol. https://doi.org/10.1007/s11258-020-01004-5
Winter DJ (2017) rentrez: an R package for the NCBI eUtils API. R J 9:520–526. https://doi.org/10.32614/RJ-2017-058
Yang YY, Kim JG (2016) The optimal balance between sexual and asexual reproduction in variable environments: a systematic review. J Ecol Environ 40:1–18. https://doi.org/10.1186/s41610-016-0013-0
Zecca G, Casazza G, Piscopo S, Minuto L, Grassi F (2017) Are the responses of plant species to Quaternary climatic changes idiosyncratic? A demographic perspective from the Western Alps. Plant Ecol Divers 10:273–281. https://doi.org/10.1080/17550874.2017.1393702
Zimmerman M (1988) Nectar production, flowering phenology, and strategies for pollination. In: Doust JL, Doust LL (eds) Plant reproductive ecology: patterns and strategies. Oxford University Press, Oxford, pp 157–178
Acknowledgements
The authors thank the botanist association Flora Alpina Bergamasca (F.A.B.) for technical help, and the administration of the protected area, Parco delle Orobie Bergamasche, for sampling permits (No. 1428 and 1283MV/RC). The authors thank Kathryn Harrold for text revision and Adriana Nurtaza, Davide Magnani and Andrea Tapparo for their contribution to the laboratory work. AJMP was supported by a post-doctoral fellowship funded by the Portuguese Foundation for Science and Technology (SFRH/BPD/111015/2015).
Funding
Open access funding provided by Università degli Studi di Milano - Bicocca within the CRUI-CARE Agreement. AJMP supported by the Portuguese Foundation for Science and Technology (SFRH/BPD/111015/2015).
Author information
Authors and Affiliations
Contributions
Conceptualization: PB, JJ, LM; Data collection: PB, AA, NT; Laboratory analyses: PB, AA, NT; Statistical analyses: PB, AJMP, AG; Writing: all authors.
Corresponding author
Ethics declarations
Conflict of interest
The author declare that there is no conflict of interest.
Ethical approval
This study was approved by the authorities of the Parco delle Orobie Bergamasche (permits no. 1428 and no. 1283MV/RC).
Additional information
Communicated by Marjana Westergren.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
11258_2021_1123_MOESM1_ESM.pdf
Supplementary file1 (PDF 894 KB) Appendix S1—Additional details on Linaria samples used for genetic analyses, additional results on the genetic identity and variation in L. tonzigii (Table S1 and S2), figure S5 with an example of L. tonzigii rooting system.
11258_2021_1123_MOESM2_ESM.pdf
Supplementary file2 (PDF 725 KB) Appendix S2—Figures S1–S4 with the phylogenetic analyses of concatenated sequences for nuclear and for plastidial genetic markers of the genus Linaria s.l., inferred separately with RaxML and MrBayes
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Biella, P., Akter, A., Muñoz-Pajares, A.J. et al. Investigating pollination strategies in disturbed habitats: the case of the narrow-endemic toadflax Linaria tonzigii (Plantaginaceae) on mountain screes. Plant Ecol 222, 511–523 (2021). https://doi.org/10.1007/s11258-021-01123-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11258-021-01123-7