Abstract
Like many peat wetlands around the world, Temperate Highland Peat Swamps on Sandstone (THPSS), located in the Sydney Basin, Australia, have been impacted by urban development. In this paper, we used Illumina 16S rRNA DNA amplicon sequencing to characterise and compare the bacterial communities of surface (top 0–2 cm) and deep (50 cm) sediments in peat swamps that occur in both urbanised and non-urbanised catchments. Proteobacteria (32.2% of reads), Acidobacteria (23.6%) and Chloroflexi (10.7%) were the most common phyla of the dataset. There were significant differences in the bacterial community structure between catchment types and depths apparent at the phyla level. Proteobacteria, Bacteroidetes, Actinobacteria and Verrucomicrobia made up a greater proportion of the reads in the surface sediments than the deeper sediments, while Chloroflexi and Nitrospirae were relatively more common in the deeper than the surface sediment. By catchment type, Acidobacteria were more common in swamps occurring in non-urbanised catchments, while Nitrospirae, Bacteroidetes and Actinobacteria were more common in those in urbanised catchments. Microbial community structure was significantly correlated with sediment pH, as was the relative abundance of several phyla, including Acidobacteria (negative correlation) and Bacteroidetes (positive correlation). As an indicator of trophic shift from oligotrophic to copiotrophic conditions associated with urbanised catchment, we found significant differences ratios of β-Proteobacteria to Acidobacteria and Bacteriodetes to Acidobacteria between the catchment types. Based on SIMPER results we suggest the relative abundance of Nitrosomonadaceae family as a potential indicator of urban degradation. As the first study to analyse the bacterial community structure of THPSS using sequencing of 16S rDNA, we reveal the utility of such analyses and show that urbanisation in the Blue Mountains is impacting the microbial ecology of these important peatland ecosystems.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Wetlands have the potential to mitigate surface water pollution and carbon emissions, which are among the most pressing and widespread anthropogenic disturbances (Mitsch and Gosselink 2015). Yet, wetlands globally are threatened by a multitude of pressures (Asselen et al. 2013). Urbanisation is a major threat to the physical and biological structure and function of wetlands. It is well known that changes to the imperviousness of surfaces within a catchment negatively affects the ecology of wetlands (Arnold and Gibbons 1996) by altering the hydrology, augmenting nutrient and pollutant concentrations and changing physicochemical conditions leading to a loss of ecosystem services (Walsh et al. 2005; Tippler et al. 2014). For example, stormwater runoff can channelize wetlands, lower water tables, and impact water quality (Tippler et al. 2014; Cowley et al. 2016).
Wetland microbial communities are sensitive and responsive to changing environmental conditions and shifts in microbial assemblage have the potential to alter ecosystem function (Urakawa and Bernhard 2017). Given that the sediment microbial community is fundamental to carbon and nitrogen cycling in wetlands (Mitsch and Gosselink 2015), understanding the implications of urbanisation to wetland microbial function is key, as is the ability to track and assess degradation associated with urbanisation, to ensure that ecosystem services and ecosystem integrity are maintained.
Since European colonisation and ensuing urbanisation, wetlands such as the endangered Temperate Highland Peat Swamps on Sandstone (THPSS) that occur in the Sydney Basin in south-eastern Australia, have undergone habitat loss and degradation (Kohlhagen et al. 2013). The THPSS, particularly those in the developed areas of the Blue Mountains, are increasingly under pressure from the effects of urbanisation (Belmer et al. 2015, 2018; Fryirs et al. 2016; Lane 2016). Impervious catchment area and the number of stormwater drains entering these swamps have significant negative impacts on the hydrological and geomorphic condition of these wetlands (Fryirs et al. 2016). THPSS in urbanised catchments have altered macroinvertebrate communities (Belmer et al. 2018; Hardwick 2019) and chemical properties (Belmer et al. 2015). Urbanisation is also associated with increased carbon exports and depletion of carbon stocks in these swamps at an ecologically and economically significant scale (Cowley et al. 2018a, b; Cowley and Fryirs 2020). As peat formation and carbon sequestration processes rely heavily on slow microbial decomposition of accumulated organic matter under saturated, anaerobic and naturally acidic conditions (Moore 1989), it is important to understand the microbial composition and processes of the ecosystem to improve management and protect these systems.
Surprisingly, there have been few comparative studies investigating the impacts of urbanisation on wetland microbial assemblages. A number of studies undertaken in streams and rivers have recently reported impacts from urbanisation on microbial assemblage composition in the water column and/or sediments (Ibekwe et al. 2016; Wang et al. 2016, 2018; Hosen et al. 2017; Jani et al. 2018; Roberto et al. 2018; Simonin et al. 2019). In tidal wetlands, microbial assemblages responded to urbanisation stressors by shifting function toward increased nitrification and xenobiotic degradation (Gonzalez Mateu et al. 2019). We previously reported differences in the community fingerprints between THPSS in urbanised and non-urbanised catchments, and increased likelihood of finding the microbial functional genes for methanogen and methnotrophs (mcrA & pmoA) in the urbanised swamps (Christiansen et al. 2019), but an in-depth taxonomic analysis is lacking.
The aim of this study was to characterise bacterial assemblages in sediments in THPSS (hereafter called swamps) by high-throughput 16S rRNA amplicon sequencing, and to compare the assemblages in swamps within urbanised and non-urbanised catchments. Further, we sought to identify taxa indicative of urban-impacted conditions and microbially-mediated ecosystem community functions that may also be affected. These swamps are saturated peat-forming systems that are naturally nutrient poor and acidic (Pemberton 2005). Consequently, we expected that undisturbed swamps would harbor taxa with oligotrophic traits of slow carbon mineralisation. The ratio of copiotrophic to oligotrophic taxa has been suggested as a reliable indicator of trophic status and ecological condition in wetlands and soils (Smit et al. 2001; Sims et al. 2013), which we apply in this study using the ratio of Proteobacteria to Acidobacteria (Smit et al. 2001; Hartman et al. 2008). Bacteroidetes and β-Proteobacteria are more consistently copiotrophic (Fierer et al. 2007) compared with the large and diverse phyla of Proteobacteria, so we also considered the ratios of β-Proteobacteria to Acidobacteria and Bacteroidetes to Acidobacteria as indicators of disturbance to trophic status. Our hypothesis was that greater relative abundances of oligotrophic taxa would be found in the non-urbanised catchment swamps and conversely a greater relative abundance of copiotrophic taxa would be found in the urbanised catchment swamps. We anticipated taxa associated with an urbanised catchment would include those with nitrification or bioremediation capabilities, indicative of pollution associated with stormwater runoff. We consider potential indicator taxa and the applicability for microbial biomonitoring in these ecosystems and more broadly.
Materials and methods
Study sites
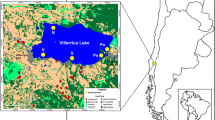
The study area is located in the Blue Mountains approximately 100 km west of Sydney (Fig. 1). The swamps are situated in shallow depressions atop the sandstone plateau of the Sydney Basin (Fryirs et al. 2019). The area experiences a temperate climate with mean maximum temperatures between 9.4 °C (July) to 23.3 °C (January) and mean minimum temperatures between 2.6 °C (July) to 12.9 °C (February), with mean annual rainfall of 1400 mm (BOM 2017). The swamps are generally covered by dense native vegetation that includes sedge, Melaleuca, Grevillea and Banksia species (Keith and Myerscough 1993) and have a characteristic stacking of sediment layers that comprise the valley fill (Freidman and Fryirs 2014; Fryirs et al. 2014a, b; Cowley et al. 2016). Typically these swamps lack standing water. The water table is (generally) about 20 cm below ground level, however the highly organic sediments are generally saturated (Hose et al. 2014; Cowley et al. 2018a, b).
Study site locations. Outlines show swamps that are identified by name and catchment type as detailed in Table 1. Modified from (Christiansen et al. 2019). Source: Basemaps produced with ArcGIS® software by Esri. Sources: Esri, DigitalGlobe, GeoEye, Earthstar Geographics, CNES/Airbus DS, USDA, USGS, AeroGRID, IGN, and the GIS User Community. Swamp outlines taken from the Macquarie University, Fryirs et al. (2016), THPSS mapping layer. 6 maps showing the spatial distribution of THPSS were produced for the following areas: Blue Mountains–VIS_ID 4480 Budderoo–VIS_ID 4481 Gosford–VIS_ID 4482 Newnes–VIS_ID 4483 Woronora–VIS_ID 4484 Penrose–VIS_ID 4485. Creative Commons license at: https://data.nsw.gov.au/data/dataset/temperate-highland-peat-swamps-on-sandstone-thpss-vegetation-maps-vis-ids-4480-to-4485. Satellite image providing context from © Google Earth 2019
Twelve swamps were sampled in this study. Six ‘urbanised catchment swamps’ were located within the towns of Katoomba, Wentworth Falls, Leura or Blackheath. The urbanised sites had impervious catchment area > 30% and between zero and eleven stormwater drains discharging into them (Fryirs et al. 2016) (Fig. 1, Table 1). Six ‘non-urbanised catchment swamps’ were located nearby in catchments that were dominated by natural vegetation, had < 15% impervious catchment area and no stormwater drains discharging into them. Sediment physico-chemistry has been described in Christiansen et al. (2019) and summarised for each site in Table S1. Most notably, sediment pH and surface ammonium concentrations were higher in urbanised than in non-urbanised catchment swamps. The electrical conductivity, moisture and organic content of sediments were similar between the swamps in different catchment types (Table S1) (Christiansen et al. 2019).
Sample collection
Sampling was conducted in September 2015. Sediment was collected at three representative random locations along the central axis of each swamp using a Russian D-corer. From each core, sediment was sampled from the top 1–2 cm and at 50 cm depth. These depths target contrasting sediment composition and conditions (Cowley et al. 2016). The surface is comprised of surface organic fines of living and decomposing organic matter and fine silt and sand and is likely to favour aerobic microbial metabolic processes. Sediments at 50 cm depth were generally alternating organic sands, which include layers of peat and mineral sand. These deeper sediments had lower electrical conductivity, ammonium, moisture and organic content than the surface sediments (Table S1). Sediment pH was similar between the two depths (Table S1) (Christiansen et al. 2019). Samples for molecular analysis were placed directly into separate sterile tubes (2 mL Eppendorf tubes) with a stainless steel spatula sterilised with ethanol in between samples. All sediment samples were snap-frozen immediately in the field using dry ice and stored upon return to the lab at –80 °C.
Molecular analysis
Extraction
Environmental DNA was extracted from sediment samples using the PowerSoil Total RNA Isolation Kit and DNA Elution Accessory Kit (MoBio) according to the manufacturer’s protocol. In this study only the DNA extraction was utilised. The purity and yield of the DNA was quantified using a NanoDrop 2000 spectrophotometer (Thermo Scientific Inc.). Eluted DNA was diluted with sterile MilliQ water (1:150) to avoid PCR inhibitors based on qPCR efficiency curves of serial dilutions.
Sequencing and pipeline processing
Amplicon sequencing was performed on the V1-V3 region of the 16S rRNA bacteria gene via the Illumina MiSeq platform using the primers 27f and 519r (Lane et al. 1985; Lane 1991). PCR amplification and sequencing were completed by Ramaciotti Centre for Genomics, University of New South Wales, Sydney (ISO/IEC 17,025 accredited laboratories).
Sequencing reads were processed using the EBI genomics project processing pipeline Version 4.0 (Mitchell et al. 2017). In summary, paired overlapping raw reads were merged to form longer reads using SeqPrep (v 1.1). For quality control, low quality ends and sequences (> 10% undetermined nucleotides) were removed using Trimmomatic (v 0.35), and sequences less than 100 nucleotides were removed using Biopython (v 1.65). Infernal (v 1.1.2) was used to identify ncRNA and lower quality overlaps removed by Cmsearch deoverlap script (v 1.0). Taxonomy and OTU classifications were assigned using MAPseq (v 1.2) with SILVAs SSU/LSU (v 128) database. Data are publicly available and stored in the European Nucleotide Archive (ENA) https://www.ebi.ac.uk/ena/browser/view/PRJEB23135.
Statistical analysis
Diversity, measured with Shannon diversity index, richness, estimated by Chao1 index, the relative abundances of phyla, and ratios of taxa were compared using a 3-factor repeated measures analysis of variance (ANOVA). In these analyses, catchment type (between-subject factor) and depth (within-subject factor) were considered fixed factors and site was considered a random factor and nested within catchment type. Sphericity was tested using Mauchley’s Test and the Geisser-Greenhouse Adjustments used where the assumption was not met. Tukey’s post hoc pair-wise comparisons were used to test for differences between levels where there was a significant interaction. Relationships between phyla relative proportions, diversity and richness were explored using Pearson’s correlation.
Bacterial assemblages were analysed at the operational taxonomic unit (OUT) level. Assemblages were compared between catchment types and depth using a 3-factor PERMANOVA, replicating the ANOVA analysis described above. The ‘catchment type’ and ‘depth’ were considered fixed factors with ‘site’ nested within catchment type as a random factor. Profiles at the OTU level were visualised using non-metric multidimensional scaling (NMDS). The OTU reads were standardised by total number of reads and square root transformed (Hellinger transformation) and made into a resemblance matrix using the Bray–Curtis similarity index (Clarke et al. 2006). Where differences were found, we followed up with SIMPER analysis to determine the OTU(s) that contributed most to the dissimilarities between groups.
Relationships between bacterial community assemblages and sediment conditions at the time of sampling (from Christiansen et al. 2019, Table S1) were analysed at each depth using distance based linear models (DistLM) with stepwise selection based on Bray–Curtis similarity. Sediment variables that were significant in explaining the community variation were further correlated (Pearson’s correlation) with the relative abundance of key phyla and Proteobacteria classes.
Analysis of taxonomic data was based on relative abundance (number of reads per OTU over the total number of reads for that sample). The phyla level dataset was limited to those contributing at least 1% of the total, and the OTU level dataset was limited to those contributing at least 0.1% of the total dataset. The diversity and richness index calculations were based on the complete dataset. The significance level (α) for all analyses was 0.05. The 3-factor ANOVA and Pearson’s correlation analyses were performed using Minitab (Minitab Inc, PA, USA). The PERMANOVA, SIMPER and DistLM analyses were performed with PRIMER 6 & PERMANOVA + add on version 1.0.8 (PRIMER-E Ltd., Plymouth, UK).
Results
Sample reads
The mean number of sequence reads per sample was 97,430, ranging from 25,877 to 175,294. A total of 2754 OTU were found. The mean number of OTUs per sample was 544 OTU and ranged from 128 to 1098. There were 179 OTUs that individually made up at least 0.1% of the dataset. The greatest number of those were found in the surface of the urbanised catchments (154 OTU), while 79 OTU were found at both depths and catchment types.
Diversity and richness index
Shannon Diversity index values differed significantly between catchment types (p = 0.022), depths (p < 0.001) and sites (nested within catchment type) (p = 0.041). Overall, mean (± SD) diversity was greater in the urban catchment surface sediments (4.53 ± 0.33) than non-urbanised catchment surface sediments (4.17 ± 0.21), which were greater than deep urban (3.84 ± 0.39) and non-urbanised catchment sediments (3.57 ± 0.36) (Fig. 2). Mean OTU richness differed significantly between catchment types (p = 0.032) and depths (p < 0.001), but not sites nested within catchment type (p > 0.05). Mean OTU richness (± SD) was greatest in the urban catchment surface sediments (1058 ± 233) followed by non-urbanised catchment surface sediments (782 ± 112), deep urban catchment sediments (574 ± 180) and deep non-urbanised catchment sediments (500 ± 143) (Fig. 2).
Bacterial community diversity and richness by site, depth and catchment type. Black triangles are for swamps in non-urbanised catchments and grey circles are for swamps in urbanised catchments. Solid symbols are for surface and open are for deep sediments. Error bars indicate 1 standard deviation. All varied significantly between the depths (p < 0.001), p values are displayed on the graphs of those that varied significantly between catchments. Swamp names as follows: FC Fortress Creek, GC Grand Canyon, ME Michael Eade’s Reserve, MtH Mt Hay Creek, TS Timmy’s Swamp, VS Vista Swamp, 5A Fifth Avenue, KF Katoomba Falls Road, Ma Marmion Road, PP Pitt Park, PG Popes Glen, WFL Wentworth Falls Lake
Phyla analysis
Proteobacteria was the most common phylum, comprising 32.2% of reads of the dataset, followed by Acidobacteria with 23.6% and Chloroflexi with 10.7%. Other phyla that comprised more than 1% of the total reads were Planctomycetes (9.0%), Nitrospirae (3.3%), Bacteroidetes (3.0%), Actinobacteria (2.5%), Verrucomicrobia (2.3%), Firmicutes (1.4%) and Spirochaetes (1.1%).
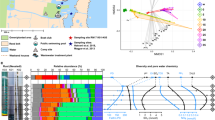
Several phyla were more abundant in either the deeper or surface sediments. Proteobacteria, Bacteroidetes, Actinobacteria and Verrucomicrobia made up a greater proportion of the reads in the surface sediments than the deeper sediments, while Chloroflexi and Nitrospirae were relatively more common in the deeper than the surface sediment (all p < 0.001). Acidobacteria had greater relative abundance in swamps in non-urbanised catchments (p = 0.046), while Bacteroidetes (p = 0.008), Nitrospirae (p = 0.008), and Actinobacteria (p = 0.007) were more abundant in swamps in urbanised catchments. The relative abundance of Proteobacteria as a whole did not differ significantly by catchment type, but there were differences of some Proteobacteria classes between catchment types. The proportion of α-Proteobacteria was greater in the swamps in non-urbanised catchments (p = 0.002) than the swamps in urbanised catchments, while the proportion of β-Proteobacteria (p = 0.004) and δ-Proteobacteria (p = 0.005) were greater in the swamps in urbanised than non-urbanised catchments. The proportion of each Proteobacteria class varied with depth. There was a significant interaction between catchment type and depth for Planctomycetes (p = 0.025), Nitrospirae (p = 0.012), Actinobacteria (p = 0.003) and Firmicutes (p = 0.002). In the swamps in non-urbanised catchments Planctomycetes was relatively more common in the deeper than surface sediments and Firmicutes made up a greater proportion of the surface sediment compared to the deep sediment assemblage but neither taxon differed markedly with depth in urbanised catchments (Fig. 3). In the urbanised catchments, Nitrospirae made up a greater proportion of the deep than the surface sediment assemblages and Actinobacteria made up a greater proportion of the surface compared to the deep sediment assemblages but neither taxon differed markedly with depth in non-urbanised catchments (Fig. 3).
Mean phylum composition (including Proteobacteria classes) for each swamp and overall mean by depth and arranged by catchment type (urbanised or non-urbanised). Asterisk (*/**) adjacent to phyla/class indicates significant differences (p < 0.05) between catchment types relative abundance in non-urbanised catchments. One (*) indicates greater relative abundance in non-urbanised catchment swamps and two (**) indicates greater relative abundance in urbanised catchment swamps. Swamp names as follows: FC Fortress Creek, GC Grand Canyon, ME Michael Eade’s Reserve, MtH Mt Hay Creek, TS Timmy’s Swamp, VS Vista Swamp, 5A Fifth Avenue, KF Katoomba Falls Road, Ma Marmion Road, PP Pitt Park, PG Popes Glen, WFL Wentworth Falls Lake
Bacterial community analysis
The ordination of the bacterial community shows a division among samples along the X axis that reflects catchment type (Fig. 4). There was further division along the Y axis, with surface sediments clustering toward the top of the plot and deep samples toward the bottom (Fig. 4). In general, the surface samples were clustered together more tightly, suggesting greater within group similarity, than the deep sediments (Fig. 4). This visualisation was supported by the PERMANOVA results. The bacterial community composition varied significantly by catchment type (p = 0.001) and depth (p = 0.001), site within catchment (p = 0.001) and there was an interaction between catchment type and depth (p = 0.037) and site and depth (p = 0.004) (Fig. 4). Pairwise tests of the sites revealed that Pitt Park (PP) and Popes Glen (PG) were significantly different from the other swamps in urbanised catchments, but not each other. Vista Swamp (VS) was different from all but Fortress Creek (FC), and Grand Canyon (GC) and Timmy’s Swamp (TS) were different.
NMDS of bacterial community based on OTU with greater than 0.1% relative abundance for the dataset. Black triangles are for swamps in non-urbanised catchments and grey circles are for swamps in urbanised catchments. Solid symbols are from surface sediment samples and open are from deep sediment (50 cm). Swamp names as follows: FC Fortress Creek, GC Grand Canyon, ME Michael Eade’s Reserve, MtH Mt Hay Creek, TS Timmy’s Swamp, VS Vista Swamp, 5A Fifth Avenue, KF Katoomba Falls Road, Ma Marmion Road, PP Pitt Park, PG Popes Glen, WFL Wentworth Falls Lake
Having found a significant difference between surface and deep sediments and a significant interaction between catchment type and depth, we ran a second PERMANOVA and SIMPER analysis on the surface and deep samples separately. For the surface sediments, the catchment type (p = 0.004) and sites (within catchment) (p = 0.036) were significantly different. The taxa that contributed most to the differences between catchment types were Candidatus_Koribacter (genus) and Acidobacteriales (order) from Acidobacteria, and Roseiarcus (genus) from α-Proteobacteria which were all more abundant in the swamps in non-urbanised catchments than the swamps in urbanised catchments. Nitrosomonadaceae (family) from β-Proteobacteria and Geobacter (genus) from δ-Proteobacteria were both more abundant in the swamps in urbanised than non-urbanised catchments. The assemblage composition at depth differed significantly with catchment type (p = 0.002) and site (within catchment type) (p = 0.002). The taxa that contributed most to the differences between the catchment types were Ktedonobacterales (order) from phylum Chloroflexi, Candidatus_Koribacter (genus) from Acidobacteria, Isosphaera (genus) and Isosphaeraceae (family) from Planctomycetes, and Acidobacteriales (order) from Acidobacteria, all of which had greater relative abundance in the swamps in non-urbanised than urbanised catchments (Fig. 5).
NMDS of bacterial assemblages in A surface (0–2 cm) and B deep (50 cm) sediments from swamps in unrbanised and non-urbanised catchments. Black triangles are for swamps in non-urbanised catchments and grey circles are for swamps in urbanised catchments. The vectors show the relationship of the relative abundance of the five OTUs that contributed the most variation between the catchment types based on SIMPER results. The OTUs are reported to the lowest taxonomic level possible and the taxonomic level of identification is indicated by the letter preceding the taxonomic name as follows: o order, f family, g genus. Swamp names as follows: FC Fortress Creek, GC Grand Canyon, ME Michael Eade’s Reserve, MtH Mt Hay Creek, TS Timmy’s Swamp, VS Vista Swamp, 5A Fifth Avenue, KF Katoomba Falls Road, Ma Marmion Road, PP Pitt Park, PG Popes Glen, WFL Wentworth Falls Lake
Indicator of trophic status
The ratios of Proteobacteria to Acidobacteria were not significantly higher in swamps in urbanised catchments (p = 0.063) than swamps in non-urbanised catchments (Fig. 6a). However, the ratios of β-Proteobacteria to Acidobacteria (p = 0.004) and the ratios of Bacteriodetes to Acidobacteria (p = 0.006) were (Fig. 6b, c, respectively). For each of these three ratios, the values were higher in the surface sediments than the deep sediments (p < 0.001 for all) and there were significant differences between sites within catchment types (p ≤ 0.001 for all). The Pitt Park and Popes Glen sites had higher values than other sites (Fig. 6).
Potential microbial indicators to assess swamp condition. Comparing swamps in urbanised catchments (grey circles) to those with swamps in non-urbanised catchments (black triangles). Surface sediment are represented by solid symbols, open symbols represent 50 cm deep. All varied significantly between the depths (p < 0.001), p values are displayed on the graphs of those that varied significantly between catchments. Ratios of Proteobacteria to Acidobacteria (a); β-Proteobacteria to Acidobacteria (b); Bacteriodetes to Acidobacteria (c) presented to represent ratios of copiotrophic to oligotrophic trait taxa; and the percentage of reads of family Nitrosomonadaceae (d), taxonomic group that contains ammonia oxidizing bacteria. Swamp names as follows: FC Fortress Creek, GC Grand Canyon, ME Michael Eade’s Reserve, MtH Mt Hay Creek, TS Timmy’s Swamp, VS Vista Swamp, 5A Fifth Avenue, KF Katoomba Falls Road, Ma Marmion Road, PP Pitt Park, PG Popes Glen, WFL Wentworth Falls Lake
Ammonia oxidizing bacteria (AOB) has been suggested as an indicator of wetland ecosystem condition (Sims et al. 2013). The Nitrosomonadaceae family, which was highlighted with the SIMPER results as a key taxon contributing to differences between the catchment types, includes AOB. The relative abundance of Nitrosomonadaceae was significantly higher in swamps in urbanised catchments (p = 0.002) than in swamps in non-urbanised catchments. Nitrosomonadaceae reads were also more common in the surface than in deep sediments (p < 0.001) and there were differences between the sites within the catchment types (p = 0.001) (Fig. 6).
Sediment environment and bacterial community
Sediment pH explained the greatest portion of the variation in the microbial community composition in the surface sediments (r2 = 0.51, p = 0.001). Ammonium (r2 = 0.03, p = 0.031) and moisture content (r2 = 0.03, p = 0.023) explained a small but statistically significant portion variation in the surface sediments (Fig. 7). By phyla, Acidobacteria (r = –0.820, p < 0.001) and Firmicutes (r = –0.625, p < 0.001) were negatively correlated with pH, as was class α-Proteobacteria (r = –0.771, p < 0.001). Bacteroidetes (r = 0.702, p < 0.001) and Nitrospirae (r = 0.544, p = 0.001), class β-Proteobacteria (r = 0.801, p < 0.001) and δ-Proteobacteria (r = 0.731, p < 0.001) were positively correlated with pH in the surface sediment. Actinobacteria (r = 0.404, p = 0.018) was positively correlated with ammonium concentrations.
Relationship between sediment conditions and bacterial community. Vectors of sediment conditions show Pearson’s correlation and only those were statistically significant (p < 0.05) from the DistLM analysis were included. Grey circles represent urbanised catchment swamps, black triangles represent in non-urbanised catchment swamps. Surface bacterial community a are represented by solid symbols, open symbols represent b 50 cm deep. EC = electrical conductivity. Swamp names as follows: FC Fortress Creek, GC Grand Canyon, ME Michael Eade’s Reserve, MtH Mt Hay Creek, TS Timmy’s Swamp, VS Vista Swamp, 5A Fifth Avenue, KF Katoomba Falls Road, Ma Marmion Road, PP Pitt Park, PG Popes Glen, WFL Wentworth Falls Lake
In the deep sediment, pH also explained the greatest proportion of the variation in assemblage structure (r2 = 0.33, p = 0.001) with electrical conductivity explaining a smaller amount of variation (r2 = 0.05, p = 0.012) (Fig. 7). Again, Acidobacteria (r = –0.469, p = 0.009) and class α-Proteobacteria (r = –0.670, p < 0.001) were positively correlated with pH, as was Planctomycetes (r = –0.534, p < 0.001). Bacteroidetes (r = 0.787, p < 0.001), Actinobacteria (r = 0.738, p < 0.001) and class β-Proteobacteria (r = 0.571, p < 0.001) were positively correlated with pH. Planctomycetes also had a weak positive correlation with electrical conductivity (r = 0.368, p = 0.035). The separation of bacterial assemblage samples in the nMDS (Fig. 7) along the same axis as the pH vector, supports the findings of the DistLM result that pH explained the greatest proportion of the variation in the bacterial assemblage structure. The few cases where an urbanised catchment swamp was clustered with those from the non-urbanised sites can be explained by the sample having a low sediment pH.
Discussion
Applying next generation sequencing techniques to the high conservation value Temperate Highland Peat Swamps on Sandstone of the Sydney Basin, we have found catchment urbanisation coincides with shifts in the bacterial assemblage and the relative abundance of functionally important taxa. Despite relatively small towns situated in the catchments of these swamps (e.g. Blackheath (population 5,000), Wentworth Falls (6,000), Katoomba (8,000) and Leura (4,600) (ABS 2019)), the patterns of change in the microbial assemblages were similar to those observed in highly urbanised or disturbed aquatic environments elsewhere (Hosen et al. 2017; Jani et al. 2018; Saxena et al. 2018; Wang et al. 2018). This highlights the sensitivity of bacteria and the potential of microbial measures as indicators of function, trophic status and ecosystem health (Fierer et al. 2007; Hartman et al. 2008; Sims et al. 2013; Ho et al. 2017; Urakawa and Bernhard 2017).
As hypothesized, in non-urbanised catchment swamps we found greater relative abundance of taxa associated with oligotrophic traits, such as Acidobacteria (Fierer et al. 2007). While Acidobacteria are a diverse and ubiquitous phylum (Janssen 2006; Jones et al. 2009; Kielak et al. 2016), they are generally competitively advantaged in low resource (Smit et al. 2001; Eichorst et al. 2007; Fierer et al. 2007; Ward et al. 2009; Ho et al. 2017) and low pH environments (Sait et al. 2006; Hartman et al. 2008). Along the same lines, studies elsewhere have found greater relative abundance of Acidobacteria in streams from forested catchments compared with urbanised catchments (Hosen et al. 2017) and in natural wetlands compared with artificial wetlands (Peralta et al. 2013). The taxa that were identified as key contributors to the dissimilarity between the catchment types and were relatively more abundant in the non-urbanised catchment swamps, also had oligotrophic traits. For example, Candidatus Koribacter (Acidobacteria) is a genus adapted to low resource conditions and slow growth (Ward et al. 2009). The genus Roseiarcus (class α-Proteobacteria) has been characterised as slow growing and moderately acidophilic (Kulichevskaya et al. 2014). The family Isosphaeraceae (phylum Planctomycetes) have been commonly found in boreal peatlands and are slow decomposers and includes members that may be capable of anaerobic ammonium oxidation (Youssef and Elshahed 2014; Fuerst 2017; Ivanova et al. 2017), which is associated with low nutrient levels (Burgin and Hamilton 2007; Urakawa and Bernhard 2017).
In contrast to the swamps in non-urbanised catchments, the swamps in urbanised catchments had higher proportions of taxa associated with copiotrophic life strategies, Bacteroidetes and β-Proteobacteria, (Fierer et al. 2007). While these taxa are common in freshwater environments, Bacteroidetes respond positively to pollution and β-Proteobacteria abundances are often associated with nutrient enriched environments (Wobus et al. 2003; Hartman et al. 2008; Zhang et al. 2012; Saxena et al. 2018). Members of Bacteroidetes can degrade high molecular weight organic compounds (Thomas et al. 2011; Martin et al. 2012) and have been reported to increase in disturbed urban waters including River Ganges, India (Jani et al. 2018), at industrial outfalls in the Nanxijiang River, China (Lu and Lu 2014) and in urban streams in Chicago, USA (Drury et al. 2013). Members of β-Proteobacteria are nitrogen cyclers and include ammonia oxidisers in the family Nitrosomonadaceae, (Garrity 2005), which was among the top contributors of dissimilarity of the swamp surface sediment assemblages between the catchment types in this study. High relative abundances of ammonia oxidising bacteria (AOB) indicate nutrient enrichment (Sims et al. 2013; Urakawa and Bernhard 2017; Korbel et al. 2022), which is consistent with the correlations between ammonia levels and AOB in the surface sediments of the swamps across an urbanisation gradient (Christiansen et al. 2019).
In the swamps from urbanised catchments, Actinobacteria, δ-Proteobacteria and Nitrospirae were also relatively more abundant. Species in the genus Geobacter (class δ-Proteobacteria) are capable of oxidising organic compounds and metals and are important for biogeochemical processes and bioremediation (Lovley et al. 2011; Cologgi et al. 2014). They had greatest relative abundance in the urbanised catchment swamps and were highlighted as a key taxon explaining the dissimilarity between the catchment types. Geobacter have been associated with contaminated environments (Lovley et al. 2011) including polluted river systems (Liao et al. 2019). Thus, the increased Geobacter abundance may indicate contamination is occurring in swamps found in urbanised catchments, perhaps caused by runoff from roads. Nitrospirae, which are capable of nitrite oxidation (Daims 2014) are favoured in higher nitrogen conditions and have been shown to increase in abundance around the outfalls of wastewater treatment plants (Drury et al. 2013). However, we found no correlation between ammonium levels and Nitrospirae. Alternatively, higher relative abundances of Nitrospirae may be a result of members from the genus Leptospirillum (Order Nitrospirae), which are capable of iron (II) oxidation (Daims 2014), and reflect conditions suitable for iron cycling. In nearby swamps, evidence of bacterial iron oxidation has been linked to groundwater inputs from the local iron-rich sandstone and historical disturbance of the sediment profile through excavation (Hardwick 2019). Supporting this, Katoomba Falls Road (KF) swamp had the highest relative abundance of Nitrospirae, and we observed iron flocs indicating active iron cycling at the time of sampling.
In accordance with previous studies (Fierer and Jackson 2006; Hartman et al. 2008; Lauber et al. 2009), sediment pH was again important in determining the composition of the bacterial assemblage, with significant correlations with several phyla and to the assemblage overall. The swamps in this study are naturally acidic and nutrient poor environments (Keith and Myerscough 1993; Pemberton 2005), while the impacts of urbanisation are locally associated with augmented nutrients and higher pH (Belmer et al. 2015; Lane 2016; Christiansen et al. 2019). It is not surprising that we found greater relative abundance of Acidobacteria in the swamps in non-urbanised catchments and a negative correlation with pH. Conversely, relative abundance of β-Proteobacteria and Bacteroidetes were most strongly positively correlated with pH. The higher pH in the urbanised catchment swamps is likely due to high alkalinity inputs from concrete stormwater infrastructure and the impervious catchment area (Walsh et al. 2005; Christiansen et al. 2019; Carroll et al. 2020). Stormwater contains sediment, nutrients and other pollution and contaminants (Barbosa et al. 2012) to which bacteria may also be responding.
Given that we observed patterns of taxonomic shifts similar to other studies, we propose that microbial indicators can be applied to analysis and assessment of these swamp’s ecosystem health. The use of microbes as biological indicators for assessing ecosystem health and restoration efficacy has received increased attention lately (Watts et al. 2008; Sims et al. 2013; Aylagas et al. 2017; Urakawa and Bernhard 2017; Reumer et al. 2018; Simonin et al. 2019; Hilderbrand et al. 2020; Ritson et al. 2021). The ratio of Proteobacteria to Acidobacteria has been suggested as a microbial indicator using the principle that these taxa broadly represent copiotrophic and oligotrophic life strategies (respectively) and so a change in the ratio indicates a trophic shift (Smit et al. 2001; Hartman et al. 2008). Although we did not find the ratio of Proteobacteria to Acidobacteria to be significantly different between the catchment types, we found ratios of taxa with more reliable copiotrophic traits (Bacteriodetes or β-Proteobacteria) (Fierer et al. 2007) were significantly different between the catchment types. Given that Proteobacteria is a large and diverse phylum including taxa with varied traits (Ho et al. 2017), it may not be best suited to represent the copiotrophic taxonomic group. Indeed, in the current study, and in other nutrient poor, low pH systems, Proteobacteria often makes up the greatest overall proportion of the sequencing reads (Too et al. 2018). As discussed above, both Bacteriodetes and β-Proteobacteria made up a significantly higher proportion of the bacterial community in swamps in urbanised catchments than in swamps in non-urbanised catchments, whereas there was no clear relationship between Proteobacteria and the catchment types.
Several studies have also recommended that analysis and assessment of ecosystem health should focus on taxa that are responsible for important biogeochemical processes such as nitrogen cycling or measures of diversity and richness (Sims et al. 2013; Urakawa and Bernhard 2017). Sims et al. (2013) highlighted ammonia oxidizing bacteria (AOB) as indicative of disturbance and nutrient enrichment. In accordance with that, we have found that the Family Nitrosomonadaceae, which includes AOB (Garrity 2005), was a defining taxon in the surface sediments of swamps found in urbanised catchments. Potentially Nitrosomonadaceae could represent an indicator taxon for which qPCR assays could be developed. We also found measures of diversity and richness differed between catchment types. Importantly, bacterial diversity is related to functionality and ecosystem services (Bell et al. 2005; Delgado-Baquerizo et al. 2016). While high diversity is often associated with healthy ecosystems (Urakawa and Bernhard 2017), the swamps in non-urbanised catchments had lower diversity and richness. Contamination and disturbance can negatively influence diversity (Baker and Banfield 2003; Lenart-Boroń and Boroń 2014), however, in naturally acidic and nutrient poor systems, such as those in the THPSS in non-urbanised catchments, the inverse can be true (Griebler and Lueders 2009; Horton et al. 2019). Any shift in diversity associated with degradation, whether positive or negative, indicates a departure from the natural condition and a potential shift in function.
Here, we have shown bacteria assemblage shifted in ways that are consistent with ecosystem degradation. A change from greater oligotrophic taxa to copiotrophic taxa under urbanised catchment conditions suggests carbon is being metabolised and exported from the system providing a microbial mechanism for the increased carbon exports reported by Cowley and Fryirs (2020). Altered processes threaten the peat formation processes and therefore the viability of these unique ecosystems in the landscape. The relationship between sediment conditions and bacterial assemblages found in this study highlights the importance of natural ecosystem pH levels for the maintenance of microbial function. Given that THPSS, wetland and peatland monitoring programs are investigating new and more innovative ways of assessing ecosystem function, this study demonstrates the potential for microbial indicators as a meaningful biomonitoring tool.
Availability of data and material
Sequencing data is publicly accessible through the European Nucleotide Archive (ENA) https://www.ebi.ac.uk/ena/browser/view/PRJEB23135
References
Arnold CL, Gibbons CJ (1996) Impervious surface coverage: The emergence of a key environmental indicator. J Am Plann Assoc 62(2):243–258
Asselen S, Verburg P, Vermaat J, Janse J (2013) Drivers of wetland conversion: A global meta-analysis. PLoS One 8(11)
Aylagas E, Borja Á, Tangherlini M, Dell’Anno A, Corinaldesi C, Michell CT, Irigoien X, Danovaro R, Rodríguez-Ezpeleta N (2017) A bacterial community-based index to assess the ecological status of estuarine and coastal environments. Mar Pollut Bull 114(2):679–688
Baker BJ, Banfield JF (2003) Microbial communities in acid mine drainage. FEMS Microbiol Ecol 44(2):139–152
Barbosa AE, Fernandes JN, David LM (2012) Key issues for sustainable urban stormwater management. Water Res 46(20):6787–6798
Bell T, Newman JA, Silverman BW, Turner SL, Lilley AK (2005) The contribution of species richness and composition to bacterial services. Nature 436(7054):1157–1160
Belmer N, Tippler C, Wright IA (2018) Aquatic ecosystem degradation of high conservation value upland swamps, Blue Mountains Australia. Water Air Soil Pollut 229(3):98
Belmer N, Wright I, Tippler C (2015) Urban geochemical contamination of high conservation value upland swamps, Blue Mountains Australia. Water Air Soil Pollut 226(10):1–5
Burgin AJ, Hamilton SK (2007) Have we overemphasized the role of denitrification in aquatic ecosystems? A review of nitrate removal pathways. Frontiers Ecol Environ 5:89–96
Carroll R, Reynolds JK, Wright IA (2020) Geochemical signature of urbanisation in Blue Mountains upland swamps. Sci Total Environ 699:134393
Christiansen NA, Fryirs KA, Green TJ, Hose GC (2019) The impact of urbanisation on community structure, gene abundance and transcription rates of microbes in upland swamps of eastern Australia. PLoS One 14(3):e0213275
Clarke KR, Gorley RN (2006) Primer v6: User manual/tutorial. PRIMER-E Ltd
Cologgi DL, Speers AM, Bullard BA, Kelly SD, Reguera G (2014) Enhanced uranium immobilization and reduction by geobacter sulfurreducens biofilms. Appl Environ Microbiol 80(21):6638–6646
Cowley K, Looman A, Maher DT, Fryirs K (2018a) Geomorphic controls on fluvial carbon exports and emissions from upland swamps in eastern Australia. Sci Total Environ 618:765–776
Cowley KL, Fryirs KA (2020) Forgotten peatlands of eastern australia: An unaccounted carbon capture and storage system. Sci Total Environ 730:139067
Cowley KL, Fryirs KA, Hose GC (2016) Identifying key sedimentary indicators of geomorphic structure and function of upland swamps in the blue mountains for use in condition assessment and monitoring. Catena 147:564–577
Cowley KL, Fryirs KA, Hose GC (2018b) The hydrological function of upland swamps in eastern australia: The role of geomorphic condition in regulating water storage and discharge. Geomorphology 310:29–44
Daims H (2014) The family Nitrospiraceae. The prokaryotes: Other major lineages of bacteria and the archaea. Rosenberg E, DeLong EF, Lory S, Stackebrandt E and Thompson F. Berlin, Heidelberg, Springer Berlin Heidelberg 733–749
Delgado-Baquerizo M, Maestre FT, Reich PB, Jeffries TC, Gaitan JJ, Encinar D, Berdugo M, Campbell CD, Singh BK (2016) Microbial diversity drives multifunctionality in terrestrial ecosystems. Nat Commun 7:10541
Drury B, Rosi-Marshall E, Kelly JJ (2013) Wastewater treatment effluent reduces the abundance and diversity of benthic bacterial communities in urban and suburban rivers. Appl Environ Microbiol 79(6):1897
Eichorst SA, Breznak JA, Schmidt TM (2007) Isolation and characterization of soil bacteria that define Terriglobus gen. Nov., in the phylum Acidobacteria. Appl Environ Microbiol 73(8):2708
Fierer N, Bradford MA, Jackson RB (2007) Toward an ecological classification of soil bacteria. Ecology 88(6):1354–1364
Fierer N, Jackson RB (2006) The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci USA 103(3):626–631
Freidman BL, Fryirs KA (2014) Rehabilitating upland swamps using environmental histories: A case study of the Blue Mountains peat swamps, eastern Australia. Geografiska Annaler Ser Phys Geogr 97:337–353
Fryirs K, Freidman B, Williams R, Jacobsen G (2014a) Peatlands in eastern australia? Sedimentology and age structure of Temperate Highland Peat Swamps on Sandstone (THPSS) in the Southern Highlands and Blue Mountains of NSW. Australia Holocene 24(11):1527–1538
Fryirs K, Gough J, Hose GC (2014b) The geomorphic character and hydrological function of an upland swamp, Budderoo Plateau, Southern Highlands, NSW. Australia Phys Geogr 35(4):313–334
Fryirs KA, Cowley K, Hose GC (2016) Intrinsic and extrinsic controls on the geomorphic condition of upland swamps in eastern NSW. CATENA 137:100–112
Fryirs KA, Farebrother W, Hose GH (2019) Understanding the spatial distribution and physical attributes of upland swamps in the Sydney Basin as a template for their conservation and management. Aust Geogr 50(1):91–110. https://doi.org/10.1080/00049182.2018.1449710
Fuerst JA (2017) Chapter 1 - Planctomycetes—new models for microbial cells and activities. Academic Press, Microbial resources. Kurtböke I, pp 1–27
Garrity GM (2005) Bergey’s manual® of systematic bacteriology: Volume two the Proteobacteria part c the alpha-, beta-, delta-, and epsilonproteobacteria. Boston, MA, Boston, MA: Springer US
Gonzalez Mateu M, Park CE, McAskill CP, Baldwin AH, Yarwood SA (2019) Urbanization altered bacterial and archaeal composition in tidal freshwater wetlands near Washington DC, USA, and Buenos Aires, Argentina. Microorganisms 7(3)
Griebler C, Lueders T (2009) Microbial biodiversity in groundwater ecosystems. Freshw Biol 54(4):649–677
Hardwick L (2019) Functional ecological processes in upland swamps and chain of ponds systems in the Blue Mountains and Southern Highlands of eastern New South Wales. PhD, Macquarie University, Australia
Hartman WH, Richardson CJ, Vilgalys R, Bruland GL (2008) Environmental and anthropogenic controls over bacterial communities in wetland soils. Proc Natl Acad Sci USA 105(46):17842–17847
Hilderbrand RH, Keller SR, Laperriere SM, Santoro AE, Cessna J, Trott R (2020) Microbial communities can predict the ecological condition of headwater streams. PLoS One 15(8):e0236932
Ho A, Di Lonardo DP, Bodelier PLE (2017) Revisiting life strategy concepts in environmental microbial ecology. FEMS Microbiol Ecol 93(3)
Horton DJ, Uzarski DG, Learman DR, Theis KR (2019) Microbial community structure and microbial networks correspond to nutrient gradients within coastal wetlands of the Laurentian Great Lakes. FEMS Microbiol Ecol 95(4)
Hose GC, Bailey J, Stumpp C, Fryirs K (2014) Groundwater depth and topography correlate with vegetation structure of an upland peat swamp, Budderoo Plateau, NSW, Australia. Ecohydrology 1392–1402
Hosen JD, Febria CM, Crump BC, Palmer MA (2017) Watershed urbanization linked to differences in stream bacterial community composition. Front Microbiol 8(1452)
Ibekwe AM, Ma J, Murinda SE (2016) Bacterial community composition and structure in an urban river impacted by different pollutant sources. Sci Total Environ 566–567:1176–1185
Ivanova AA, Naumoff DG, Miroshnikov KK, Liesack W, Dedysh SN (2017) Comparative genomics of four Isosphaeraceae Planctomycetes: A common pool of plasmids and glycoside hydrolase genes shared by Paludisphaera borealis px4t, Isosphaera pallida is1bt, Singulisphaera acidiphila dsm 18658t, and strain sh-pl62. Front Microbiol 8(412)
Jani K, Ghattargi V, Pawar S, Inamdar M, Shouche Y, Sharma A (2018) Anthropogenic activities induce depletion in microbial communities at urban sites of the River Ganges. Curr Microbiol 75(1):79–83
Janssen PH (2006) Identifying the dominant soil bacterial taxa in libraries of 16S rRNA and 16S rRNA genes. Appl Environ Microbiol 72(3):1719
Jones RT, Robeson MS, Lauber CL, Hamady M, Knight R, Fierer N (2009) A comprehensive survey of soil Acidobacterial diversity using pyrosequencing and clone library analyses. ISME 3(4):442
Keith DA, Myerscough PJ (1993) Floristics and soil relations of upland swamp vegetation near Sydney. Aust J Ecol 18(3):325–344
Kielak AM, Barreto CC, Kowalchuk GA, van Veen JA, Kuramae EE (2016) The ecology of Acidobacteria: Moving beyond genes and genomes. Front Microbiol 7:744–744
Kohlhagen T, Fryirs K, Semple AL (2013) Highlighting the need and potential for use of interdisciplinary science in adaptive environmental management: The case of endangered upland swamps in the Blue Mountains, NSW. Australia Geogr Res 51(4):439–453
Korbel KL, Greenfield P, Hose GC (2022) Agricultural practices linked to shifts in groundwater microbial structure and denitrifying bacteria. Sci Tot Environ 807:150870
Kulichevskaya IS, Danilova OV, Tereshina VM, Kevbrin VV, Dedysh SN (2014) Descriptions of Roseiarcus fermentans gen. Nov., sp. Nov., a bacteriochlorophyll a-containing fermentative bacterium related phylogenetically to alphaproteobacterial methanotrophs, and of the family Roseiarcaceae fam. Nov. Int J Syst Evol Microbiol 64(8):2558–2565
Lane A (2016) Decades of healing: Wasteland to wetland. Australia, Greater Sydney Landcare Network Inc, Penrith
Lane DJ (1991) 16S/23S rRNA sequencing. Nucleic acid techniques in bacterial systematics (modern microbiological methods). Stackebrandt E and Goodfellow M. New York, NY, John Wiley and Sons 31:115–175
Lane DJ, Pace B, Olsen GJ, Stahl DA, Sogin ML, Pace NR (1985) Rapid determination of 16S ribosomal RNA sequences for phylogenetic analyses Proc Natl Acad Sci USA 82(20):6955–6959
Lauber CL, Hamady M, Knight R, Fierer N (2009) Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl Environ Microbiol 75(15):5111–5120
Lenart-Boroń A, Boroń P (2014) The effect of industrial heavy metal pollution on microbial abundance and diversity in soils — a review. Environmental risk assessment of soil contamination. Hernandez-Soriano MC. https://www.intechopen.com/books/environmental-risk-assessment-of-soil-contamination/the-effect-of-industrial-heavy-metal-pollution-on-microbial-abundance-and-diversity-in-soils-a-revie, IntechOpen. Accessed 15 Oct 2019
Liao H, Yu K, Duan Y, Ning Z, Li B, He L, Liu C (2019) Profiling microbial communities in a watershed undergoing intensive anthropogenic activities. Sci Total Environ 647:1137–1147
Lovley DR, Ueki T, Zhang T, Malvankar NS, Shrestha PM, Flanagan KA, Aklujkar M, Butler JE, Giloteaux L, Rotaru A-E, Holmes DE, Franks AE, Orellana R, Risso C, Nevin KP (2011) Geobacter: The microbe electric’s physiology, ecology, and practical applications. Adv Microb Physiol Poole RK Acad Press 59:1–100
Lu X-M, Lu P-Z (2014) Characterization of bacterial communities in sediments receiving various wastewater effluents with high-throughput sequencing analysis. Microb Ecol 67(3):612–623
Martin F, Torelli S, Le Paslier D, Barbance A, Martin-Laurent F, Bru D, Geremia R, Blake G, Jouanneau Y (2012) Betaproteobacteria dominance and diversity shifts in the bacterial community of a pah-contaminated soil exposed to phenanthrene. Environ Pollut 162:345–353
Mitchell AL, Scheremetjew M, Denise H, Potter S, Tarkowska A, Qureshi M, Salazar GA, Pesseat S, Boland MA, Hunter Fiona MI, ten Hoopen P, Alako B, Amid C, Wilkinson DJ, Curtis TP, Cochrane G, Finn RD (2017) Ebi metagenomics in 2017: Enriching the analysis of microbial communities, from sequence reads to assemblies. Nucleic Acids Res 46(D1):D726–D735
Mitsch WJ, Gosselink JG (2015) Wetlands. John Wiley & Sons, Incorporated
Moore PD (1989) The ecology of peat-forming processes: A review. Int J Coal Geol 12(1):89–103
Pemberton M (2005) Australian peatlands: A brief consideration of their origin, distribution, natural values and threats. J R Soc West Aust 88(Part 3):81–89
Peralta RM, Ahn C, Gillevet PM (2013) Characterization of soil bacterial community structure and physicochemical properties in created and natural wetlands. Sci Total Environ 443:725–732
Reumer M, Harnisz M, Lee HJ, Reim A, Grunert O, Putkinen A, Fritze H, Bodelier PLE, Ho A (2018) Impact of peat mining and restoration on methane turnover potential and methane-cycling microorganisms in a northern bog. Appl Environ Microbiol 84(3):e02218-e12217
Ritson JP, Alderson DM, Robinson CH, Burkitt AE, Heinemeyer A, Stimson AG, Gallego-Sala A, Harris A, Quillet A, Malik AA, Cole B, Robroek BJM, Heppell CM, Rivett DW, Chandler DM, Elliott DR, Shuttleworth EL, Lilleskov E, Cox F, Clay GD, Diack I, Rowson J, Pratscher J, Lloyd JR, Walker JS, Belyea LR, Dumont MG, Longden M, Bell NGA, Artz RRE, Bardgett RD, Griffiths RI, Andersen R, Chadburn SE, Hutchinson SM, Page SE, Thom T, Burn W, Evans MG (2021) Towards a microbial process-based understanding of the resilience of peatland ecosystem service provisioning – a research agenda. Sci Total Environ 759:143467
Roberto AA, Van Gray JB, Leff LG (2018) Sediment bacteria in an urban stream: Spatiotemporal patterns in community composition. Water Res 134:353–369
Sait M, Davis KER, Janssen PH (2006) Effect of pH on isolation and distribution of members of subdivision 1 of the phylum acidobacteria occurring in soil. Appl Environ Microbiol 72(3): 1852
Saxena G, Mitra S, Marzinelli EM, Xie C, Wei TJ, Steinberg PD, Williams RBH, Kjelleberg S, Lauro FM, Swarup S (2018) Metagenomics reveals the influence of land use and rain on the benthic microbial communities in a tropical urban waterway. mSystems 3(3):e00136–00117
Simonin M, Voss KA, Hassett BA, Rocca JD, Wang S-Y, Bier RL, Violin CR, Wright JP, Bernhardt ES (2019) In search of microbial indicator taxa: Shifts in stream bacterial communities along an urbanization gradient. Environ Microbiol 21(10):3653–3668
Sims A, Zhang Y, Gajaraj S, Brown PB, Hu Z (2013) Toward the development of microbial indicators for wetland assessment. Water Res 47(5):1711–1725
Smit E, Leeflang P, Gommans S, van den Broek J, van Mil S, Wernars K (2001) Diversity and seasonal fluctuations of the dominant members of the bacterial soil community in a wheat field as determined by cultivation and molecular methods. Appl Environ Microbiol 67(5):2284
Thomas F, Hehemann J-H, Rebuffet E, Czjzek M, Michel G (2011) Environmental and gut bacteroidetes: The food connection. Front Microbiol 2(93)
Tippler C, Wright IA, Davies PJ, Hanlon A (2014) The influence of concrete on the geochemical qualities of urban streams. Mar Freshw Res 65(11):1009–1017
Too CC, Sickel W, Keller A, Lee SM, Yule CM (2018) Microbial community structure in a Malaysian tropical peat swamp forest: the influence of tree species and depth. Front Microbiol 9:28–59
Urakawa H, Bernhard AE (2017) Wetland management using microbial indicators. Ecol Eng 108:456–476
Walsh CJ, Roy AH, Feminella JW, Cottingham PD, Groffman PM, Morgan RM (2005) The urban stream syndrome: current knowledge and the search for a cure. J North Am Benthol Soc 24(3):706–723. https://doi.org/10.1080/00049182.2018.1449710
Wang L, Zhang J, Li H, Yang H, Peng C, Peng Z, Lu L (2018) Shift in the microbial community composition of surface water and sediment along an urban river. Sci Total Environ 627:600–612
Wang P, Chen B, Yuan R, Li C, Li Y (2016) Characteristics of aquatic bacterial community and the influencing factors in an urban river. Sci Total Environ 569–570:382–389
Ward NL, Challacombe JF, Janssen PH, Henrissat B, Coutinho PM, Wu M, Xie G, Haft DH, Sait M, Badger J, Barabote RD, Bradley B, Brettin TS, Brinkac LM, Bruce D, Creasy T, Daugherty SC, Davidsen TM, DeBoy RT, Detter JC, Dodson RJ, Durkin AS, Ganapathy A, Gwinn-Giglio M, Han CS, Khouri H, Kiss H, Kothari SP, Madupu R, Nelson KE, Nelson WC, Paulsen I, Penn K, Ren Q, Rosovitz MJ, Selengut JD, Shrivastava S, Sullivan SA, Tapia R, Thompson LS, Watkins KL, Yang Q, Yu C, Zafar N, Zhou L, Kuske CR (2009) Three genomes from the phylum Acidobacteria provide insight into the lifestyles of these microorganisms in soils. Appl Environ Microbiol 75(7):2046
Watts CH, Vojvodic-Vukovic M, Arnold GC, Didham RK (2008) A comparison of restoration techniques to accelerate recovery of litter decomposition and microbial activity in an experimental peat bog restoration trial. Wetl Ecol Manag 16(3):199–217
Wobus A, Bleul C, Maassen S, Scheerer C, Schuppler M, Jacobs E, Röske I (2003) Microbial diversity and functional characterization of sediments from reservoirs of different trophic state. FEMS Microbiol Ecol 46(3):331–347
Youssef NH, Elshahed MS (2014) The phylum planctomycetes. The prokaryotes: Other major lineages of bacteria and the archaea. Rosenberg E, DeLong EF, Lory S, Stackebrandt E and Thompson F. Berlin, Heidelberg, Springer Berlin Heidelberg 759–810
Zhang D-C, Mörtelmaier C, Margesin R (2012) Characterization of the bacterial archaeal diversity in hydrocarbon-contaminated soil. Sci Total Environ 421–422:184–196
Acknowledgements
We thank the following volunteers who contributed their time collecting field data; Ari Edmonds, Lorraine Hardwick and Kirsten Cowley. Fieldwork was approved under NSW National Parks and Wildlife Service Scientific License No. SL101129, a Sydney Catchment Authority License (D2014/69166) and with permission of the Blue Mountains City Council. This work was supported by an Australian Research Council Linkage grant (LP130100120) with the Greater Sydney Local Land Services as industry partners, and a grant awarded under the Department of Sustainability, Environment, Water, Population and Communities (DSEWPaC) and Australian National University (ANU) Research Program on Temperate Highland Peat Swamps on Sandstone (THPSS) to GH and KF. NC held a Macquarie University Research Excellence Scholarship (MQRES) and received Higher Degree Research support.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions. This work was supported by an Australian Research Council Linkage grant (LP130100120) with the Greater Sydney Local Land Services as industry partners, and a grant awarded under the Department of Sustainability, Environment, Water, Population and Communities (DSEWPaC) and Australian National University (ANU) Research Program on Temperate Highland Peat Swamps on Sandstone (THPSS) to GH and KF. NC held a Macquarie University Research Excellence Scholarship (MQRES) and received Higher Degree Research support.
Author information
Authors and Affiliations
Contributions
Conceptualisation: N Christiansen; Methodology: N Christiansen, G Hose, T Green; Formal analysis and investigation: N Christiansen, G Hose; Writing—original draft preparation: N Christiansen; Writing—review and editing: G Hose, K Fryirs; Funding acquisition: K Fryirs, G Hose; Supervision: G Hose, T Green, K Fryirs.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Authors give consent for publication.
Conflicts of interest/competing interests
Authors have none to declare.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Christiansen, N.A., Green, T.J., Fryirs, K.A. et al. Bacterial communities in peat swamps reflect changes associated with catchment urbanisation. Urban Ecosyst 25, 1455–1468 (2022). https://doi.org/10.1007/s11252-022-01238-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11252-022-01238-3