Abstract
Ocular swabs from canine distemper virus (CDV) suspected live or brain tissue from dead dogs were tested for the presence of CDV nucleoprotein (N) gene using reverse transcriptase polymerase chain reaction (RT-PCR). Partial “N” gene sequencing of the RT-PCR-positive samples and the local vaccine virus revealed that the Ind/Andaman 01/07 virus was highly divergent from the rest of the CDV isolates and from the vaccine strain. Quantitative real-time PCR (qRT-PCR) using SYBR Green I chemistry for CDV haemagglutinin “H” gene quantification showed C t values ranging from 29.76–30.67 in the RT-PCR-positive samples. Two of the positive samples, designated Ind/TN 01/07 and Ind/Andaman 01/07 were used for virus isolation in B95a cell line. Characteristic cytopathic changes such as rounding of cells, syncytia formation, and ballooning were seen from the first passage onwards. Specific cytoplasmic fluorescence was seen in infected cells with a commercial reference serum against CDV. To the best of our knowledge, this is the first report of CDV isolation from clinical cases in India.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Canine distemper virus (CDV), a Morbillivirus of the family Paramyxoviridae causes a highly systemic infection with prominent respiratory, gastrointestinal, and nervous signs in dogs (Patel and Heldensb 2009). The disease is characterized by a rapid onset of severe leucopenia and loss of lymphocyte proliferation ability. The resulting immunosuppression increases the host’s susceptibility to opportunistic infections, which are a main cause of distemper-associated deaths. This disease is widely prevalent in the tropical countries such as India.
The genome of CDV consists of single-stranded negative-sense RNA of more than 15,000 nucleotides, tightly encapsidated by the nucleocapsid protein (N). In addition, mature ribonucleoprotein complexes also contain copies of the phosphoprotein and large protein. The host cell-derived lipid envelope is spiked with transmembrane haemagglutinin (H) and fusion glycoproteins and, internally, the envelope is stabilized by the matrix protein (von Messling et al. 2003).
The disease has been controlled by the use of attenuated live virus vaccines. However, several CDV-vaccinated dogs also come down with CDV worldwide (Blixenkrone-Møller et al. 1992). These findings suggested that there are genomic/virulence differences between the wild-type CDV and vaccine strains (Harder and Osterhaus 1997). Analysis of viral genes provides insights into the understanding of the variations of CDV. The N protein gene constitutes an important structure of CDV comprising the conserved region, middle part, and variable region, N- and C-terminal parts. Some degree of variation of the N gene among CDV isolates had been demonstrated (Keawcharoen et al. 2005).
Sequence analysis of CDV strains from different geographic locations and from different animal species has revealed that the H gene undergoes genetic drift (Martella et al. 2007). Based on phylogenetic analysis of H gene sequences, most CDV field strains cluster into seven major genetic lineages, namely, America-1 (most vaccine strains), America-2, Asia-1, Asia-2, Europe, Artic-like, and European wildlife (Kapil et al. 2008; McCarthy et al. 2007).
Some CDV strains such as Onderstepoort vaccine strain are adapted to cell culture while CDV from clinical samples has been successfully isolated in MDCK or B95 cells derived from marmoset B lymphoblastoid cells or in Vero cells stably expressing canine signaling lymphocyte activation molecule (SLAM) (Seki et al. 2003).
In India, CDV infections are quite common and serological surveys using dot ELISA have been done (Parthiban et al. 2000; Ramadass and Latha, 2001). Latha et al. (2007) screened 160 conjunctival samples collected from dogs with clinical symptoms suggestive of canine distemper using dot enzyme-linked immunosorbant assay (ELISA) and 112 (70%) were positive. However, no virus isolation has been done from India. This study aimed at isolation and characterization of CDV in B95 cells from reverse transcription polymerase chain reaction (RT-PCR)-positive swab/tissue samples collected from CDV-infected dogs. To the best of our knowledge, CDV has not been isolated and characterized from India.
Materials and methods
Samples
Ocular swab samples collected from suspected dogs admitted to the Madras Veterinary College Teaching hospital and one brain tissue sample sent to us from the Andaman and Nicobar Islands were tested in RT-PCR for CDV N gene.
B95a cell line
Epstein–Barr virus transformed B cell line (B95a) procured from Indian Veterinary Research Institute, Mukteshwar, Uttaranchal and maintained in our laboratory was used for isolation of CDV.
RT-PCR for CDV “N” gene
Total RNA was extracted using TriZol (Invitrogen, USA) from the suspected swabs, commercial vaccine and one brain tissue sample following manufacturer’s instructions. cDNA was synthesized from 2 μg of total RNA using the High capacity cDNA archive kit (Applied Biosystems Inc, USA) with oligo-dT primers according to the manufacturer’s instructions. The primers used for the amplification of CDV N gene both the first round and nested PCR have been described by Youn et al. (2001) resulting in 549 and 419 bp products, respectively. PCR was performed using outer or nested primers with the RedDye mix (Bangalore Genei, Bangalore, India) in 25 μl reaction volumes. PCR and nested PCR conditions were as follows, one cycle of 95°C for 2 min followed by 30 cycles of 30 s at 95°C, 58°C for 1 min and 72°C for 2 min followed by one cycle at 70°C for 10 min using a Gradient Master Cycler (Eppendorf, Germany). The amplified products were analyzed by 2% agarose (Sigma, Poole, UK) gel electrophoresis in 1× TAE buffer at 50 mA for 1 h and products visualized by staining with ethidium bromide (BioRad, USA) using 100 bp ladder as a DNA size marker.
CDV “N” gene sequencing and analysis
The CDV “N” gene-positive RT-PCR products were purified using the Gel extraction kit (Qiagen, USA) following the manufacturer’s conditions and subjected to sequencing in a Genetic Analyzer (ABI Biosystems, USA). The nucleotide sequences of “N” gene were compared with the other CDV “N” gene sequences available in the GenBank using the MegAlign program of DNAStar software. A phylogenetic tree was also constructed with these sequences.
Quantitative real-time PCR for CDV H mRNA using SYBR Green I chemistry
RNA was extracted and cDNA synthesized from CDV suspected swabs and brain samples as described earlier. The sequence of primers for the CDV H gene and the endogenous control actin gene are described in Table 1. The PCR mix for SYBR Green I assay was prepared using 10 μl of the SYBR Green I PCR master mix (Applied Biosystems Inc, USA), 1 μl (1 pmol) of CDV H/actin, sense primers, 1 μl (1 pmol) of CDV H/actin anti-sense primers and 100 ng cDNA adjusted in DEPC water in 8 μl to give a total reaction volume of 20 μl. The real-time PCR (Applied Biosystems Inc., USA, Model 7500) was run under the universal cycling condition consisting of one cycle at 50°C for 2 min, one cycle at 95°C for 10 min and 40 cycles at 95°C for 15 s and 60°C for 1 min. The C t values were recorded for both the target and endogenous controls. The data were accepted only when the NTC had no amplification indicated by C t values above 40. In addition to C t values, the melting curve was also recorded to assess the specificity of amplification. All the C t values are mean of triplicate samples tested. The ∆C t values indicate the difference in the C t values between the target gene and the endogenous gene. The ∆∆C t value indicates the difference between the ∆C t of sample and the ∆C t of calibrator. The fold change in the gene expression is calculated as \( {{2}^{{ - \Delta \Delta {C_{\text{t}}}}}} \).
Virus isolation and identification using fluorescent antibody test
One RT-PCR-positive ocular swab sample and one positive brain tissue sample was infected into B95a cells and serially passaged five times using conventional methods. At each passage, cytopathic effects (CPE) produced by the virus were recorded. Using a commercially available anti-CDV FITC direct conjugate (VMRD, USA) virus replication was assessed in these cells using fluorescent antibody test (FAT). The presence of CDV in the passaged samples was also assessed by RT-PCR.
Results
The suspected dogs from which the samples were collected had conjunctival discharge, rise in temperature, anorexia, diarrhea and nasal discharge.
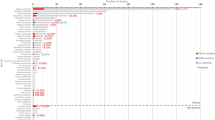
Two samples of ocular swabs from CDV suspected dogs (n = 13; Ind/TN 01/07 and Ind/TN 02/07) and one brain sample (Ind/Andaman 01/07) were positive in CDV “N” gene RT-PCR and nested PCR (Fig. 1). The nucleotide sequences of the three Indian CDV isolates had a percent homology up to 89% to 98% with other CDV isolated from other countries. A phylogenetic tree drawn with the aligned sequences revealed a distinct clade for two Indian viruses while the Ind/And/01/07 was found to be more divergent from other CDV (Fig. 2). The commercial vaccine grouped with other CDV from USA, Taiwan, Switzerland and China and was different from the Onderstepoort vaccine strain.
RT-PCR amplification of RNA extracted from ocular swabs of CDV-infected dog using outer and nested primers for CDV N gene. The electrophoresis was performed on agarose (2%) gel. Lane 1, 100 bp ladder; lane 2, CDV “N” gene with outer primers—549 bp product; lane 3, CDV “N” gene with nested primers—419 bp product
Phylogenetic tree of partial CDV “N” gene nucleotide sequences of Indian CDV isolates with respect to other available CDV sequences in GenBank. Alignment was done using MegAlign software of DNAStar program. The lengths of horizontal lines are proportional to the minimum number of nucleotide differences required to join nodes
Table 2 indicates the C t values of target (CDV H) and endogenous control (beta actin) genes. The C t of CDV “H” mRNA in one of the ocular swab samples were at least twofold higher than seen in the brain sample tested from a dead dog. The vaccine sample also had similar amounts of CDV H mRNA as that seen in an infected sample.
Two of the RT-PCR-positive samples, Ind/TN 01/07 and Ind/Andaman 01/07 were passaged in B95a cells. The CPE was noticed from the first passage itself and consisted of rounding of cells, clumping, ballooning, and large syncytia formation by 48 h post-infection (Fig. 3a, b). Using anti-CDV FITC conjugate, the identity of the isolated virus was established. Specific granular fluorescence was observed in the cytoplasm of infected cells (Fig. 4a, b). The cell culture passaged virus also gave positive amplification in RT-PCR for the CDV N gene in both outer and nested PCRs.
a Fluorescent antibody test performed on CDV-infected B95a cells at 48 h post-infection (×100). Note a typical apple green fluorescence indicating positive reaction in few infected cells. b Fluorescent antibody assay performed on CDV-infected B95a cells at 72 h post-infection (×100). Note a typical apple green fluorescence indicating positive reaction in several infected cells
Discussion
We have successfully isolated and identified CDV from clinical cases in India for the first time. Vero cells expressing canine SLAM is useful for isolation and passaging CDV without alteration of the viral genes (Lan et al. 2006). B95a cells have been recommended for primary isolation of CDV as it intrinsically expresses mammalian SLAM that has been shown to be a receptor for CDV. Amino acid residues at the SLAM interface appear to be important determinant the host–virus specificity of morbilliviruses (Ohishi et al. 2010). Hence, the CDV could show CPE even from the first passage itself and the typical morbilli virus changes could be observed. The isolated virus could be identified using the conventional FAT, which is the standard test used for assay of CDV antibodies also. B95a cells infected with this isolated Indian CDV could be used as a substrate for performance of FAT to assess CDV-specific antibodies in dog sera. The isolated virus can also be used in serum neutralization assays for CDV antibody quantification.
Ocular swabs would be ideal samples for RT-PCR-based confirmation of CDV genome since it was found that they have similar amounts of CDV “H” mRNA as that seen in the brain of a dead dog. Elia et al. (2006) also found that in addition to lymphoid tissues, conjunctival swab and urine were ideal clinical samples for detection of CDV by quantitative real-time PCR (qRT-PCR). The frontal lobe of the brain was also rich in virus.
Molecular epidemiology studies are largely based on the H protein gene that mediates receptor binding and is much more variable than other CDV proteins (Iwatsuki et al. 2000). However the primary aim of this study was to isolate CDV from India and to identify it. This was achieved using FAT and N gene based nested PCR of the original clinical sample. N gene PCR has been used largely for diagnosis of CDV also. Future studies should involve genotyping of CDV by sequencing the “H” genes of CDV from clinical cases. This study revealed variations in the N gene of CDV isolates that may have implications in the design of primers for detection of CDV genome targeting the N gene. The role of the variations in virus replication or host specificity needs further study.
qRT-PCR has been used earlier for quantifying CDV gene expression (Scagliarini et al. 2007; Elia et al. 2006). In this study, the qRT-PCR was used to quantify the H gene expression in the swabs and brain sample using the SYBR green chemistry. This could also be useful for diagnosis although it would be more expensive. The higher costs would be offset by the ease of performance, specificity of detection by melting curve analysis and no post-PCR processing requirement thereby reducing any potential laboratory based contamination. Thus, qRT-PCR could replace the conventional RT-PCR for clinical diagnosis of CDV in due course.
Abbreviations
- CDV:
-
Canine distemper virus
- RT-PCR:
-
Reverse transcription polymerase chain reaction
- qRT-PCR:
-
Quantitative real-time PCR
- FAT:
-
Fluorescent antibody test
- N:
-
Nucleoprotein
- H:
-
Haemagglutinin
References
Blixenkrone-Møller, M., Svansson, V., Appel, M., Krogsrud, J., Have, P. and Orvell, C., 1992. Antigenic relationship between wild-type isolates of morbilliviruses from different carnivores, Archives of Virology, 123, 279–294
Elia, G., Decaro, N., Martella, V., Cirone, F., Lucente, M.S., Lorusso, E., Trani, L. D. and Buonavoglia, C., 2006. Detection of canine distemper virus in dogs by real-time RT-PCR, Journal of Virological Methods, 136, 171 – 176
Harder, T.C. and Osterhaus, A.D., 1997. Canine distemper virus: a morbillivirus in search of new hosts? Trends in Microbiology, 5, 120–124
Iwatsuki, K., Tokiyoshi, S., Hirayama, N., Nakamura, K., Ohashi, K., Wasaka, C., Mikami, T. and Kai, C., 2000. Antigenic difference in the H proteins of canine distemper viruses, Veterinary Microbiology, 71, 281–286
Kapil, S., Allison, R.W., Johnston III, L., Murray, B. L., Holland, S., Meinkoth, J. and Johnson, B., 2008. Canine distemper virus strains circulating among North American dogs, Clinical and Vaccine Immunology, 15, 702–712
Keawcharoen, J., Theamboonlers, A., Jantaradsamee, P., Rungsipipat, A., Poovorawan, Y. and Oraveerakul, K., 2005. Nucleotide sequence analysis of nucleocapsid protein gene of canine distemper virus isolates in Thailand. Veterinary Microbiology, 105, 137–142
Lan, N.T., Yamaguchi, R., Kawabata, A., Uchida, K., Kai, K., Sugano, S. and Tateyama, S., 2006. Stability of canine distemper virus (CDV) after 20 passages in Vero-DST cells expressing the receptor protein for CDV, Veterinary Microbiology, 118, 177–188
Latha, D., Srinivasan, S.R., Thirunavukkarasu, P.S., Gunaselan, L., Ramadass, P. and Narayanan, R.B., 2007. Assessment of canine distemper virus infection in vaccinated and unvaccinated dogs, Indian Journal of Biotechnology, 6, 35–40
Martella, V., Elia, G., Lucente, M.S., Decaro, N., Lorusso, E., Banyai, K., Blixenkrone-Moller, M., Lan, N.T., Yamaguchi, R., Cirone, R., Carmichael, R. E. and Buonavoglia, C., 2007. Genotyping canine distemper virus (CDV) by a hemi-nested multiplex PCR provides a rapid approach for investigation of CDV outbreaks, Veterinary Microbiology, 122, 37–42
McCarthy, A.J., Shaw, M.A. and Goodman, S.J., 2007. Pathogen evolution and disease emergence in carnivores, Proceedings of the Royal Society of London, Series B, Biological Sciences, 274, 3165–3174
Ohishi, K., Ando, A., Suzuki, R., Takishita, K., Kawato, M., Katsumata, E., Ohtsu, D., Okutsu, K., Tokutake, K., Miyahara, H., Nakamura, H., Murayama, T. and Maruyama, T., 2010. Host–virus specificity of morbilliviruses predicted by structural modeling of the marine mammal SLAM, a receptor, Comparative Immunology, Microbiology and Infectious Diseases, 33, 227–241
Parthiban, M., Meenambigai, T.V., Manohar Paul, W. and Mahalinga Nainar, A., 2000. Usefulness of dot-ELISA in detection of canine distemper virus antigen, Indian Journal of Animal Sciences, 70, 265–266
Patel, J. R. and Heldensb, J. G. M., 2009. Review of companion animal viral diseases and immunoprophylaxis, Vaccine, 27, 491–504
Ramadass, P. and Latha, D., 2001. Dot Enzyme Immunoassay for detection of canine distemper virus, Indian Veterinary Journal, 78, 981–983
Scagliarini, A., Dal Pozzo, F., Gallina, L., Vaccari, F. and Morganti, L., 2007. TaqMan based real time PCR for the quantification of canine distemper virus. Veterinary Research Communications, 31, 261–263
Seki, F., Ono, N., Yamaguchi, K. and Yanagi, Y., 2003. Efficient isolation of wild strains of canine distemper virus in Vero cells expressing canine SLAM (CD 150) and their adaptability to marmoset B95a SLAM, Journal of Virology, 77, 9943–9950
von Messling, V., Springfeld, C., Devaux, P. and Cattaneo, R., 2003. A ferret model of canine distemper virus virulence and immunosuppression, Journal of Virology, 77, 12579–12591
Youn, H.Y., Yoo, H.S. and Han, H.R., 2001. Detection of canine distemper virus through one step RT-PCR combined with nested PCR, Journal of Veterinary Science, 2, 59–66
Acknowledgements
The authors thank TANUVAS for providing facilities for this work. This work was funded partly by the Department of Biotechnology, New Delhi.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pawar, R.M., Raj, G.D., Gopinath, V.P. et al. Isolation and molecular characterization of canine distemper virus from India. Trop Anim Health Prod 43, 1617–1622 (2011). https://doi.org/10.1007/s11250-011-9880-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11250-011-9880-7