Abstract
This paper aims to explore a potential connection between two hypotheses recently put forward in the context of language evolution. One hypothesis argues that some human-specific change(s) in the hominin brain developmental program habilitated the neuronal workspace that enabled “cognitive modernity” to unfold, also resulting in our globularized braincase. The other argues that the cultural niche resulting from our self-domestication favored the emergence of natural languages. In this article we document numerous links between the genetic changes we have claimed may have brought about globularization and neural crest cells, which have been claimed to explain the constellation of distinctive traits (physical, cognitive, and behavioral) found in all domesticated mammals. If these links turn out to be as robust as we think they are, globularization and self-domestication may well be closely related phenomena in the context of human evolution.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction: Linking Two Hypotheses
This paper aims to explore a potential connection between two hypotheses recently put forward linking the language phenotype and genotype. Both are ultimately aimed to cast light on how our species’ ability to develop mental rule systems that are put to use in thought and communication appeared. One (Boeckx 2013; Boeckx and Benítez-Burraco 2014a; Theofanopoulou 2015) is the idea that the globularization of the braincase that characterizes our species (Lieberman et al. 2002; Bruner 2004) is the reflex of a genetically regulated specific brain growth pattern that provided the neural scaffolding for “cognitive modernity”, most distinctively our ‘language-readiness’. This species-specific brain growth trajectory has been well characterized by several authors (Neubauer et al. 2010; Gunz et al. 2010, 2012; Bastir et al. 2011; Scott et al. 2014; Hublin et al. 2015). They have highlighted the differential growth of the parietal lobe, the cerebellum, and the frontal pole, as well as the olfactory bulb and quite possibly the temporal lobe. The logic of correlated growth also leads us to expect differential growth at the level of the dorsal thalamus (Boeckx and Benítez-Burraco 2014a) and the corpus callosum (Theofanopoulou 2015). On the basis of this information, Boeckx and Benítez-Burraco (2014a, b, 2015) and Benítez-Burraco and Boeckx (2015a) have put forward a potential candidate gene set underlying this brain and skull growth trajectory, many of which show sign of positive selection in anatomically modern humans (AMHs).
The other idea at the heart of this paper concerns to (self-)domestication. As Thomas (2014) discussed extensively, self-domestication in our species can prove extremely valuable in understanding our cognitive behavior, and in particular what we will call below the grammaticalization of our mind. Interestingly, Wilkins et al. (2014) have recently put forth domestication usually entails a constellation of distinctive traits (the “domestication syndrome”) many of them we believe related to the brain/cognitive changes that we think important for the evolution of language, and particularly, of globularity. According to Wilkins et al., the domestication syndrome, being triggered by different external stimuli, ultimately results from the hypofunction of the neural crest cells (NCCs) during embryonic development.
Our hypothesis is that the genetic changes we have claimed led to globularization are intimately related to those affected the NCCs, fueling (or being the main rationale for) the emergence of the (self-)domestication syndrome in our species, and ultimately, of full-fledged languages.
2 Domestication and the Grammaticalization of Mind
It is now clear that a rich “Universal Grammar” is not necessary (nor adequate) to capture properly and (biologically speaking) realistically all the complexity that the grammars of the world’s languages exhibit (Boeckx 2014). Rather, the very nature and context of cultural learning appear to be key in capturing all this grammatical paraphernalia (Hurford 2011; Kirby 2012; Smith and Kirby 2008; Thompson et al. 2016). But as Thomas (2014) points out, a major problem facing any attempt to account for language structure through a cultural mechanism is that the processes required by such a mechanism are only possible if we assume the existence of a range of preconditions, which we may call the “cultural niche”. The special social, behavioral, and neural conditions of human culture have been highlighted by Tomasello over a decade (see Tomasello 2009, 2014): we are indeed a super-social primate.
Thomas (2014) thinks that the behavioural, cognitive and temperamental outcomes of self-domestication might have formed the preconditions or the niche for the emergence of structured language through a cultural process. Thomas is not alone in this. Recently, Deacon (2009) and Hare and Tomasello (2005) have appealed to the idea of self-domestication and relaxed selective pressures to capture key aspects of our behavioral modernity, pointing out that sensitivity to communicative intent, at the heart of our communicative/pragmatic competence, characterize domesticated species in general. To add to this, Okanoya and colleagues have for several years now documented phenotypic differences between two vocal learning birds: the white-rumped munia and its domesticated variant, the Bengalese finch, and argued that domestication masked various selection pressures, particularly species recognition, thereby permitting increased variation and complexity (specifically in the syntax of their songs), and that this process was then reinforced by female preferences (Okanoya 2012; Kagawa et al. 2012, 2014; Takahasi and Okanoya 2010).
The intuition that our species, steeped as it is in culture, shares some aspects of domestication is indeed a venerable one. As Thomas (2014) observes, already the ancient Greeks had a single term “hemeros” to refer to domesticated animals and civilized people (Greeks); and another term “agrios” to refer to both wild animals and ‘savages’. Darwin himself in 1871 referred to “civilized men” as “in one sense highly domesticated”. But it was the observation that humans also share many of these typical characteristics of domesticated species that motivated people like Boas in 1938 to suggest that humans might also be thought of as domesticated. Even a cursory look at the fossil record highlight changes in anatomically modern humans that closely resemble the ones attested in domesticated animals: reduced skeletal and cranial robusticity, changes in dentition, retention of juvenile characteristics, changes in temperament (compared to other living primates), reduction of sexual dimorphism, etc. All these features suggest that domestication may have taken place as part of the emergence of AMHs, although, of course, once in place, it may have intensified in our recent history. This is how we suggest to reinterpret evidence like the decrease in brain size in the last 50,000 years (Bednarik 2012, 2014), or the reduction of brow ridge projection and the shortening of the upper facial skeleton in humans from the Middle Pleistocene to recent times, plausibly resulting from a reduction in androgen reactivity (Cieri et al. 2014).
As is characteristic of the literature on domestication, Thomas focuses on selectionist considerations when it comes to examining a number of the possible accounts of how humans might have become self-domesticated. Specifically, Thomas considers three, not necessarily mutually exclusive accounts: (a) an account that links self-domestication to a process of adaptation to the human-made environment (an environment in which humans have lived longer than any other species), (b) a second account that links self-domestication to humans having undergone selection against aggression (linked to the idea of domestication as selection on temperament), and (c) a process with effects analogous to artificial selection may have also been in operation in humans; in particular, the idea that culturally mediated constructs of mate-choice may have resulted in humans applying something akin to artificial selection to themselves (an idea that others, such as Vendramini 2014, if somewhat fancifully, have linked to the context of AMHs/Neanderthals interactions).
3 Domestication and Neural Crest Cells
It is a well-established fact that domesticated animals possess a distinctive an unusual suite of heritable traits not seen in their wild progenitors (what Wilkins et al. 2014 refer to as the “domestication syndrome”). Remarkably, the suite of characteristics that typifies domestication is consistent across most species, and include depigmentation, floppy ears, reduced ears, shorter muzzles, smaller teeth, docility, smaller brain or cranial capacity, neotenous (juvenile) behavior, curly tails, and affect reproductive cycles (more frequent estrous cycles). Of course, not all of these characteristics are found in all domesticates, but nearly all domesticated species exhibit a significant subset of these characteristics (Price 2002). Indeed, some of the hard tissue characteristics are actually used by archaeologists as diagnostic of domestication having occurred (Zeder et al. 2006).
As Wilkins et al. (2014) point out, most explanations focus on particular traits, while neglecting others, or on the possible selective factors involved in domestication rather than the underlying developmental and genetic causes of these traits. To correct this, Wilkins et al. (2014) put forth the idea that the domestication syndrome results predominantly from mild NCCs deficits during embryonic development. NCCs are the vertebrate-specific class of stem cells that first appear during early embryogenesis at the dorsal edge (hence the term “crest”) of the neural tube and then migrate ventrally throughout the body in both the cranium and the trunk, giving rise to the cellular precursors of many cell and tissue types and indirectly promoting the development of others. As noted above, the motivation behind Wilkins et al.’s (2014) hypothesis is that the development of the diverse traits characteristic of the domestication syndrome is closely linked to NCCs. This is especially true in the case of size and function of the adrenal glands, which play a central role in the physiology of both fear and stress responses. Adrenal hypofunction and reduced stress hormone levels are well documented in domesticated species and have been induced experimentally by selection for tameness during experimental domestication of foxes and rats. Accordingly, Wilkins et al. (2014) conjecture that initial selection for tameness leads to reduction of neural-crest-derived tissues of behavioral relevance, via multiple preexisting genetic variants that affect NCC numbers at the final sites, and that this neural crest hypofunction produces, as an unselected byproduct, the morphological changes in pigmentation, jaws, teeth, ears, etc. exhibited in the domestication syndrome.
Wilkins et al. (2014) go on to examine the underlying genetic basis for the NCC hypofunction, and include a list of genes that will be the center of our attention below. Specifically, our aim in the remainder of this paper is to show that many connections exist between the set of genes we have highlighted in the context of globularization in our past and ongoing research, and NCCs. We think that these connections are well worth taking into account given the idea that AMH is a self-domesticated species. We close with a few promising avenues of research this paper opens up.
4 Globularization and Neural Crest Cells
The idea we would like to explore in this section is that genetic changes we claimed led to globularization may be closely linked to those that affect NCCs in domestic animals, given the well-attested roles of some of the core genes we have examined in the context of globularization at the level of the neural crest (as discussed in Boeckx and Benítez-Burraco 2014a, b; Benítez-Burraco and Boeckx 2015a, b core candidates for globularization fulfil the following criteria: they have experienced some evolutionary change after our split from Neanderthals/Denisovans; they play some role in brain growth, regionalization, and/or neural interconnection; they have been associated to conditions in which language, or cognitive properties often associated with language, are impaired, and they are candidates for craniosynostosis or some other similar condition affecting skull development and sutures). To test this possibility, we checked the literature to determine whether (and to what extent) (some of) the “domestication syndrome” genes highlighted by Wilkins et al. (2014) are also important for globularization and/or have changed in our lineage compared to Neanderthals and Denisovans. We also searched the literature via PubMed and OMIM (http://www.ncbi.nlm.nih.gov/omim/) to know more about the aberrant phenotypes linked to the mutation of these neural crest genes potentially implicated in globularization. We focused on clinical conditions and symptoms of interest for globularization (as described in Boeckx and Benítez-Burraco 2014a, b; Benítez-Burraco and Boeckx 2015a), including “language disorder”, “cognitive disorder”, “intellectual disability”, “syntactic deficit”, “semantic deficit”, “phonological deficit”, “speech deficit”, “dyslexia”, “schizophrenia”, “autism”, “autism spectrum disorder”, etc. We have also proceeded the other way around: we made extensive search of the literature to learn how many of (and to what extent) our candidates are involved in the development and function of the neural crest and could be also regarded as “neural crest genes”. We relied as well on computer tools to learn about the robustness of the links we posited, but also to uncover new links between the two sets of genes. For this, we mostly relied on String 10 (http://string-db.org/). String 10 predicts direct/physical and indirect/functional associations between proteins that derive from four sources: genomic context, high-throughput experiments, conserved coexpression, and the knowledge previously gained from text mining (Szklarczyk et al. 2011). As for phylogenetic changes, we determined the ancestral or derived nature of the genes of interest (compared to Neanderthals and Denisovans) and explored potential connections with other genes selected in AMHs. We also relied on available data on other molecular changes that occurred after our split from Neanderthals, including changes in promoter or enhancer activity, human-specific conserved deletions (hCONDELs), etc.
As far as Wilkins et al.’s (2014) candidates (their Table 2) are concerned, we found that all but one of them are related to (a) our putative globularization network, and/or (b) have changed recently in our species and/or (c) give rise to cognitive deficits that affect language and our mode of cognition when they are mutated. We will review these links briefly (see also Fig. 1).
The genes related to globularization and the genes related to the “domestication syndrome”. The network was generated by String 10 with the proteins discussed in Boeckx and Benítez-Burraco (2014a, b) and Benítez-Burraco and Boeckx (2015a) (see Table 1) and the candidate genes for domestication advanced by Wilkins et al. (2014). The medium confidence value was .0400 (this lower value enables to find a greater number of potential interactions among proteins; this lower value is then compensated for by checking whether the predicted interactions can be confirmed in the literature or in databases provided by functional assays). In this confidence view, stronger associations between proteins are represented by thicker lines. The figure does not represent a fully connected graph, but readers are asked to bear in mind that String 10 predicts associations between proteins that derive from a limited set of databases. The material discussed in the main text lead us to suspect connections between the globularization genes and the domestication genes that String does not generate (although we wish to note that just letting String add a few interactors, not discussed in this paper, yield a fully connected graph). It should be emphasized that the nature of String 10 is essentially predictive, and not explanatory. Although we have confirmed all the links we discuss here in the literature, they need to be confirmed at the brain level and in relation to language. Additionally, the diagram only represents the potential connectivity between the involved proteins, but this has to be mapped onto particular biochemical networks, signaling pathways, cellular properties, aspects of neuronal function, or cell-types of interest that can be confidently related to aspects of language development and function
To begin with, three SOX genes are highlighted by Wilkins et al. (2014): SOX10, SOX9, and SOX2. SOX10 is an interactor of PAX3 (another of Wilkins et al.’s 2014 candidates) and POU3F2 (Smit et al. 2000). Interestingly, POU3F2 interacts with FOXP2, a gene that has been consistently related to language and speech deficits (Maricic et al. 2013). AMHs show a derived allele of the binding site which is less efficient in activating transcription than the Neanderthal/Denisovan allele (Maricic et al. 2013). POU3F2 regulates neuronal migration and identity during the development of the neocortex (McEvilly et al. 2002; Sugitani et al. 2002) and has been linked to bipolar disorder (Mühleisen et al. 2014), and to developmental and language delays, intellectual disability, schizophrenia and autism spectrum disorders (ASD) (Huang et al. 2005; Lin et al. 2011; Potkin et al. 2009). Moreover, both PAX3 and SOX10 are robust candidates for Waardenburg syndrome, a complex condition in which sensorineural hearing loss is a prominent symptom (developmental delay is observed as well, but it may be subsidiary of hearing loss). Interestingly, SOX10 is hypermethylated in the brain of schizophrenics (Iwamoto et al. 2005).
We regard SOX9 of interest too because it mediates the retinoic acid-induced HES1 expression. Retinoic acid has proven to be important for brain plasticity (Luo et al. 2009), learning and memory (Etchamendy et al. 2003; Jiang et al. 2012). Importantly, it closely interacts with FOXP2 (Devanna et al. 2014) and with the genes related to globularization (Benítez-Burraco and Boeckx 2014). In turn, HES1 is a direct interactor of RUNX2, our core candidate underlying the changes that prompted globularization, because of its central role in suture ossification (Yoshida et al. 2003; Lattanzi et al. 2012), its involvement in the development of hippocampal GABAergic neurons (Pleasure et al. 2000; Benes et al. 2007) and of the thalamus (Reale et al. 2013); its involvement in the regulation of osteocalcin (Paredes et al. 2004) and osteopontin (Shen and Christakos 2005), which are important for both bone formation and brain organization (see Schroeter et al. 2006); and the fact that a selective sweep in this gene occurred after our split from Neanderthals (Green et al. 2010). The Hes1 pathway is related to language development and craniofacial development (reviewed in Boeckx and Benítez-Burraco 2014b). Additionally, Hes1 is involved in the development of both GABAergic neurons (Long et al. 2013) and dopaminergic neurons (Kameda et al. 2011). Finally, the transcriptional activation of Hes1 is a key step in the Slit/Robo signaling pathway that we believe important for the externalization of language (reviewed in Boeckx and Benítez-Burraco 2014b). We wish note as well that the phosphorylation of Sox9 is regulated downstream of Bmp signaling (Li et al. 2013). Importantly for us, two of the BMPs, namely BMP2 and BMP7, are core components of our gene network important for globularization (reviewed in Boeckx and Benítez-Burraco 2014a). Finally, we wish also note that sox9a expression is reduced in the pharyngeal arches of zebrafish in which dlx2a is knocked down (Sperber et al. 2008). DLX2, another of our core candidates, is important for tooth and craniofacial development (Jeong et al. 2008; Gordon et al. 2010), but also contributes to regulate thalamic development (Jones and Rubenstein 2004). According to McKinsey et al. (2013) Dlx2 (and Dlx1) controls some important steps of neuronal proliferation within the cortex via Zfhx1b, which is mutated in Mowat–Wilson syndrome (McKinsey et al. 2013), a condition characterized by speech delay, mental retardation, microcephaly, delayed motor development, hypertelorism, and cupped ears with fleshy, upturned lobules (Adam et al. 2006). Importantly, neural crest-specific removal of Zfhx1b in mouse gives rise to a wide range of neurocristopathies resembling Mowat-Wilson syndrome (Van de Putte et al. 2007). Moreover, the lack of Zfhx1b gives rise as well to multiple NCC defects causing Hirschsprung disease–mental retardation syndrome (Van de Putte et al. 2003). Interestingly, an hCODEL (not shared with Neanderthals) exists upstream SOX9 (McLean et al. 2011).
Lastly, SOX2 is one of the transcription factors that are differentially expressed in several postmitotic thalamic nuclei (Vue et al. 2007). SOX2 is one of the components of the SHH-GLI signaling pathway that we feel has played a significant role in the anatomical and physiological events leading to globularization (Boeckx and Benítez-Burraco 2015, Boeckx et al., manuscript), but also in NCC fate (more on SHH below). Specifically, Sox2 input and differential Gli-binding affinity have been shown to provide context and positional information in Shh-directed neural patterning. In the neural tube, GLI-bound cis-regulatory modules (CRMs) are enriched for SOX binding motifs, and SOX2 (among other SOX factors) acts as neural-specific GLI co-factors (Oosterveen et al. 2012, 2013; Peterson et al. 2012). Moreover, SOX2 regulates PQBP1, a gene linked to developmental delay, intellectual disability and microcephaly (Li et al. 2013), which also interacts with POU3F2 (Li et al. 2013). According to the Human Brain Transcriptome Atlas PQBP1 is expressed highly in several key regions of the brain in the context of globularization (cerebellum, hippocampus, and olfactory bulb). Interestingly, SOX2 is involved as well in the enhancer effect of human endogenous retroviruses (HERVs) on brain genes related to schizophrenia, specifically PRODH (Suntsova et al. 2013). And similarly to the other two SOX genes reviewed above, SOX2 is also regulated by RUNX2, some of the DLXs genes, and some of the BMPs. Hence, the transcriptional activity of Sox2 is downregulated by Runx2 and Dlx5 in response to interleukin-6 in bone marrow-derived mesenchymal multipotent cells (Yoon et al. 2014). In addition, SOX2 interacts with BMP signalling. For example, Li et al. (2015) have recently found that a BMP-Smad4-SHH-Gli1 signalling network provides a niche supporting transient Sox2+ dental epithelial stem cells in mouse molars. Importantly for our hypothesis, SOX2 is predicted (according to String 10) to interact specifically with BMP7 via NOG, an inhibitor of BMP signalling and a protein important for dopamine neuron production (Chiba et al. 2008). As we noted above, BMP7 is closely related to some of our core candidates (BMP2, RUNX2, DLX1, and DLX2 among others) and plays a pivotal main role in osteogenesis (Cheng et al. 2003) and skull and brain development (Yuge et al. 2011; Segklia et al. 2012). Mutations of BMP7 cause eye anomalies, deafness, developmental delay, and learning disabilities (Wyatt et al. 2010). Finally, together with OTX2 and PAX6, SOX2 plays a key role in controlling eye development, potentially affecting orbit size, a feature known to have changed in our species, compared to other extinct hominin species.
Among Wilkins et al.’s (2014) candidates we have found other genes that interact (according to Kulhwilm et al. 2013) with RUNX2, our key candidate for globularization. One of them is MTIF. Interestingly, an hCODEL (although shared with Neanderthals) exists upstream MTIF in AMHs (McLean et al. 2011). Also GDNF is differentially expressed after RUNX2 transfection. This gene is predominantly expressed in the pulvinar neostriatal interneuronal ensemble (Hidalgo-Figueroa et al. 2012), but also in the visual cortex (Pochon et al. 1997). GDNF is involved in dopaminergic differentiation (Christophersen et al. 2007) and also promotes formation of neuronal synapses by ligand-induced cell adhesion (Ledda et al. 2007). Interestingly for our hypothesis, GDNF is predicted (according to String 10) to interact with NCAM1, the carrier of polysialic acid (one of the neural crest stem markers) and a protein related to schizophrenia, bipolar disorder and Alzheimer’s disease (Atz et al. 2007), and to working memory performance (Bisaz et al. 2013). In turn, NCAM1 interacts with VCAM1, known to bear a fixed change (D414G) in AMHs compared to Neanderthals/Denisovans (Pääbo 2014, his table S1). NCAM1 is a potential target of RUNX2 too (Kuhlwilm et al. 2013), but also of FOXP2 (Konopka et al. 2009).
Additionally, we have found that four of Wilkins et al.’s (2014) candidates (Magoh, Fgf8, piebald (l), and Ret) are targets of FOXP2 (according to Spiteri et al. 2007; Vernes et al. 2011). We regard this an interesting link, provided that FOXP2 (together with ROBO1) is a core component of the machinery responsible for the two neural components often presented as critical for vocal learning: the direct cortico-laryngeal connection and the cortico-thalamo-basal ganglia pathway. As we review in Boeckx and Benítez-Burraco (2014b) FOXP2 and its interactome show strong links with our set of genes related to globularization and language-readiness. Magoh is involved in RNA splicing and its loss leads to a disorganized, microcephalic brain with fewer neurons (Silver et al. 2010). Interestingly, the splicing of MAGOH has a AMH-specific pattern (Lin et al. 2010). FGF8 encodes a protein involved in the regionalization of brain tissues in mammals (Fukuchi-Shimogori and Grove 2001). FGF8 interacts with RUNX2 via FGFR1, a candidate for craniosynostosis and Pfeiffer syndrome (Rossi et al. 2003), but also with DLX2 (according to String 10). The expression of FGF8 is also under the control of the SHH-GLI signaling pathway that we highlighted above as important for globularization (see Boeckx et al. manuscript for details). Specifically, both SHH and GLI3 modulates the FGF8/WNT signaling source in the forebrain (Kobayashi et al. 2010; Rash and Grove 2011). Interestingly, ≈98 % of Altaic Neanderthals and Denisovans gained a non-synonymous change in GLI3 that appears to be mildly disruptive (Castellano et al. 2014). FGF8 is a candidate for holoprosencephaly, a condition characterized by forebrain malformations, severe neurocognitive impairment, and midline facial anomalies (Sarnat and Flores-Sarnat 2001; Solomon et al. 2012). Concerning piebald (l) (also known as EDNRB), it is a candidate for Waardenburg syndrome (Read and Newton 1997), a condition involving sensorineural hearing loss and (subsidiary) developmental delay. Finally, RET is also found to be differentially expressed after RUNX2 transfection in neuroblastomic cells (Kuhlwilm et al. 2013). The RET regulatory function is conserved from human to zebrafish without sequence similarity (Fisher et al. 2006).
With regard to Tcof1, another of Wilkins et al.’s (2014) candidate, it contains a LIS1 motif (Emes and Ponting 2001). LIS1 (also known as PAFAH1B1) regulates neuronal migration and axonal growth (Smith et al. 2000; Toyo-oka et al. 2003; Shu et al. 2004). Mutations in LIS1 reduces brain size and give rise to different types of lissencephalies (Reiner et al. 1993; Lo Nigro et al. 1997) with different degrees of impact on linguistic abilities (Leventer et al. 2001). Individuals with LIS1 duplications had subtle brain defects, including microcephaly and dysgenesis of the corpus callosum. Importantly for our hypothesis, TCOF1 is also bound by RUNX2 based on ChIP analyses (Young et al. 2007). Tcof1 also acts as a modifier of Pax3 during enteric nervous system development (derived from vagal NCC) (Barlow et al. 2013). TCOF1 is a candidate for Treacher Collins syndrome, an autosomal-dominant condition involving mandibulofacial dysostosis. Heterozygous knockout of Tcof1 in mice causes severe craniofacial malformation (Dixon and Dixon 2004; Shows and Shiang 2008).
Additionally, we have found signs of positive selection in other genes highlighted by Wilkins et al. (2014) Hence, FoxD3, a transcriptional regulator essential for neural crest development, depends on Nodal (one of our candidates) for regulating zebrafish dorsal mesoderm development (Chang and Kessler 2010). FOXD3 is one of the enriched transcription factors in present-day human-specific differentially methylated regions (DMRs) (Gokhman et al. 2014). Similarly, a human-specific conserved deletion (hCONDEL) (although shared with Neanderthals) exists upstream CDH7 (McLean et al. 2011). Finally, the last two genes in Wilkins et al.’s (2014) list are also of interest for us. WSTF (also known as BAZ1B) plays a key role in chromatin remodeling and nucleosome repositioning (Kitagawa et al. 2003). BAZ1B belongs to the core 1.5 Mb region commonly deleted in Willliams syndrome (Somerville et al. 2005). Finally, piebald (s) (also known as EDN3) is significantly dysregulated in autistic children (Glatt et al. 2011). Like SOX10, EDN3 is also a candidate for Waardenburg syndrome type 4, a rare neural crest disorder combining sensorineural hearing loss and pigmentation defects (as in Waardenburg syndrome) and aganglionosis (as in Hirschsprung disease).
As we pointed our above, we made as well an extensive survey of the PubMed literature to know whether our candidates can be regarded as “neural crest genes”. The whole list of them, as resulting from our research on the molecular underpinnings of globularization and language-readiness (reviewed in Boeckx and Benítez-Burraco 2014a, b; Boeckx and Benítez-Burraco 2015a), currently encompasses 152 genes (see Table 1) Interestingly, we discovered that nearly 60 % of them play a role related to the neural crest. In the first part of this section we have focused on the genes that also belong to Wilkins et al.’s (2014) list. Below we provide an overview of the most promising connection between the neural crest and our set of genes (our selection was based on the volume of references currently available and degree of confidence in databases and computer tools like String 10.) (see Fig. 1).
To begin with, BMP7 differentiates cortical neural stem cells into a variety of neural crest lineages (Gajavelli et al. 2004). Similarly, BMP2 (also mentioned above) is required for regulating the migration of NCCs (Correia et al. 2007) and also recruits multipotent neural crest progenitors (Glejzer et al. 2011). Also CDC42 controls neural crest stem cell proliferation (Fuchs et al. 2009). Inactivation of Cdc42 in NCCs causes craniofacial and cardiovascular morphogenesis defects (Liu et al. 2013). Concerning CITED2, this is an important neural crest gene (Yin et al. 2002). According to Bamforth et al. 2001 mice lacking Cited2 exhibit adrenal agenesis, neural crest defects and exencephaly. Moreover, CITED2 has been implicated in a new disease pathway involved in Hirschsprung’s disease (Tang et al. 2015). Regarding its phylogenetic relevance, we wish note that Prüfer et al. (2014) highlight a highly disruptive intergenic change near CITED2 that is 99 % derived in AMHs and ancestral in both Altai Neanderthals and Denisovans. Moreover, CITED show strong links with genes important for language and globularization. To begin with, it is a regulatory target of FOXP2 (Nelson et al. 2013). Additionally, it interacts, via LHX2 (another of our candidates for globularization), with ROBO1 (Marcos-Mondéjar et al. 2012), one of the best-known candidate genes for dyslexia (Hannula-Jouppi et al. 2005), and crucially involved in the neural establishment of vocal learning abilities (Wang et al. 2015). Regarding CTNNB1, this gene temporally controls, as part of the Wnt/β-catenin signalling pathway, multiple steps of neural crest development, from neural crest induction, lineage decisions, to differentiation (Hari et al. 2012). Dlx1 and Dlx2 are robust neural crest markers too (Ishii et al. 2012) and are involved in patterning and morphogenetic processes in the neural crest-derived mesenchyme (Mallo 2001). Both Dlx5 and Dlx6 are crucial for aspects of neural crest development (Ruest et al. 2003). Specifically, DLX5 positions the neural crest at the border of the neural plate (McLarren et al. 2003). FGF genes (FGF7 and FGF8) are known to act in combination with BMP genes to generate the cranial neural crest (Endo et al. 2012). In turn, NCCs regulate the amount of Fgf8 produced by the anterior neural ridge (ANR) and the isthmus (the two brain organizers) via the secretion of anti-BMP signalling molecules (Le Douarin and Dupin 2012). Also FGFR1 (together with MAPK signalling) is responsible for neural crest induction (Stuhlmiller and García-Castro 2012). Moreover, Fgfr1 patterns the pharyngeal region to create a permissive environment for neural crest cell migration (Trokovic et al. 2003). Concerning Pax3, it is among the earliest genes activated in neural crest progenitors and cooperates in neural crest patterning (Maczkowiak et al. 2010; Bae et al. 2014; Plouhinec et al. 2014). In turn, Pax6 is involved in regulating the migration of NCCs from the anterior midbrain (Matsuo et al. 1993). SLITs and ROBOs (as part of the Slit/Robo signalling) are necessary to confine early NCCs to the ventral migratory pathway in the trunk (Jia et al. 2005). Moreover, Slits affect the timely migration of NCCs via the Robo receptor (Giovannone et al. 2012). Concerning SHH, we have already pointed out the important role played by the SHH signalling pathway in NCC fate. Specifically, Shh promotes the survival of NCCs (Delloye-Bourgeois et al. 2015) and the development of multipotent neural crest progenitors (endowed with both mesenchymal and neural potentials) (Calloni et al. 2007). Importantly, Shh signalling is required for cranial neural crest morphogenesis and chondrogenesis (Wada et al. 2005). Moreover, NCCs receive both Bmp and Shh signalling to induce Satb2 expression, which is another of our candidates (Sheehan-Rooney et al. 2013). Interestingly as well, a neural crest deficit in Down syndrome mice is associated with deficient mitotic response to Shh (Roper et al. 2009). Finally, we wish highlight VCAN, which encodes versican-1, a protein that guide migratory NCCs (Dutt et al. 2006) and which shows a fixed N3042D change in AMHs (Pääbo 2014; table S1).
Finally, we wish note that the recent epigenomic profiling of human and chimpanzee NCCs (Prescott et al. 2015) has revealed significant divergence across species in craniofacial cis-regulatory landscapes, resulting in expression differences of crucial NCC regulators and ultimately, in human facial variation. Some of these enhancers with a species-bias overlap with human accelerated regions (HARs). Among the genes that are flanked by species-biased enhancers and that show species-biased expression profile one finds some of our candidates (e.g. PAX3). Moreover, these human-biased enhancers have been found to be clustered in super-enhancers. Interestingly, among the genes potentially regulated by these super-enhancers we have found some of our candidates too (e.g. SRGAP2, and MECP2). Importantly, although Prescott and colleagues found that Denisovans and Neanderthals primarily share the human-like variants of some key regulatory motifs, they found as well a small set of changes that are unique to modern humans compared to these extinct hominins. Interestingly, this divergence largely results from cis-sequence changes rather than changes in the trans regulatory environment, and to some extent from the activity of retroelements (many of them belonging to ERV1, ERVL-MaLR, and ERVK subclasses), which have been hypothesised to have played a key role in the genomic changes that brought about modern language (Benitez-Burraco and Uriagereka 2016).
5 Discussion
In all fairness we are still not in a position to prove the functional connections we have sought to highlight in this paper. Because most genes involved in development are pleiotropic, playing a role in different tissues and at different times, the connections found in silico or inferred from the literature need to be validated in vitro and in vivo in tissues of interest to achieve a more satisfactory level of certainty. In particular, we still do not understand our globularization network well enough to be able to demonstrate that changes in the expression of (some of) the genes we have examined reduce the input to the neural crest, giving rise to the sort of hyporegulation that Wilkins et al. (2014) highlight in their work. But we would like to list two promising lines of research that we think could reinforce the connection we have tried to establish here.
Firstly, we find it particularly interesting that Wilkins et al. (2014) characterize the reduced neural crest input at the heart of the “domestication syndrome” as an instance of “mild neurocristopathy”. Interestingly, mutations in some of our candidates for language-readiness result in neurocristopathy (plausibly via the interaction with the neural crest genes). In this context, Hirschsprung disease strikes us again as a good starting point for future exploration, considering that the list of genes related to the disease includes many of our candidates, like ASCL1, DLX1, MAPT, NRG1, ROBO1, SLIT2, SOX2, SOX10, and ZFHX1B (Jiang et al. 2011; Tang et al. 2015). Additionally, there appears to exist comorbidity of neurocristopathy and skull/brain/face anomalies (Sjamsudin et al. 2001; Jian et al. 1995; Parry et al. 2013). Interestingly, in the case of Parry et al. (2013), the gene they focus on, GSC, interacts with some of our core candidates, including DLX genes and MEF2 genes (Miller et al. 2007; Fossat et al. 2012). Exploring this comorbidity should help validate some of the links we posited above.
The constellation of symptoms observed in ASD could lend support to the view that the link between globularization and neural crest function (and between language-readiness and language change via domestication) is a robust one, and particularly, that the changes in the globularization network and the NCC-derived network of the sort we have highlighted in this paper resulted in a distinctive modification of the cognitive phenotype of ancient hominins. Accordingly, ASD entails anomalies in the skull and the brain, cognitive and language deficits, and an abnormal presentation of the “domestication syndrome” (see Benítez-Burraco et al. 2016; Theofanopoulou and Boeckx 2016 for reviews from different angles). Although the involvement of the neural crest in the aetiopathogenesis of ASD has not been documented in detail, we wish note that neurocristopathies commonly involve autistic features (Fernell et al. 1999). Interestingly too, ASD candidates are overrepresented among the genes known to be involved in the “domestication syndrome” and the set of genes important for language-readiness and many of these genes show altered expression profiles in the brain of autists (Benítez-Burraco and Murphy 2016; Benítez-Burraco et al. 2016).
6 Conclusion
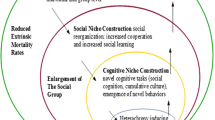
The globularization of the brain and braincase seems to be related to emergence of other traits that we believe important for modern language, including the refinement of the neural substrates for ‘externalization’ (via the ROBO/SLIT/FOXP2 interactomes) (see Boeckx and Benítez-Burraco 2014b for details) or new inter- and intrahemispheric connections underlying modern cognition (via the SHH-GLI pathway, the AUTS2 interactome, etc.) (see Benítez-Burraco and Boeckx 2015a, b; Boeckx et al. manuscript for details). The evidence we have reviewed in this paper suggests to us that a globularized brain and brain case may also be linked to the developmental/genetic context for a domestic phenotype, which could then have been selected for the reasons Thomas (2014) discussed (see Fig. 2). Put another way, globularization may be part of Deacon’s (2009) discussion of ‘relaxation of selection’, leading to a niche favoring cultural selection.
This is not to say that globularization and self-domestication are one and the same. It is well established that globularization led to a critical change at the level of the parietal lobe and associated parietal bone (Bruner 2004, 2010; Bruner et al. 2015). Crucially, though, calvarial bones arise from two embryonic tissues, namely, the neural crest and the mesoderm. Unlike the frontal bone, which is neural crest-derived, the parietal bone is paraxial mesodermal-derived (Quarto et al. 2010). If posterior braincase changes were driving globularization, as Bruner contends, these changes cannot be entirely reduced to NCC-related (i.e. domestication-related) changes.
References
Adam MP, Schelley S, Gallagher R, Brady AN, Barr K, Blumberg B et al (2006) Clinical features and management issues in Mowat–Wilson syndrome. Am J Med Genet A 140:2730–2741
Atz ME, Rollins B, Vawter MP (2007) NCAM1 association study of bipolar disorder and schizophrenia: polymorphisms and alternatively spliced isoforms lead to similarities and differences. Psychiatr Genet 17:55–67
Bae CJ, Park BY, Lee YH, Tobias JW, Hong CS, Saint-Jeannet JP (2014) Identification of Pax3 and Zic1 targets in the developing neural crest. Dev Biol 386:473–483
Bamforth SD, Bragança J, Eloranta JJ, Murdoch JN, Marques FI, Kranc KR et al (2001) Cardiac malformations, adrenal agenesis, neural crest defects and exencephaly in mice lacking Cited2, a new Tfap2 co-activator. Nat Genet 4:469–474
Barlow AJ, Dixon J, Dixon M, Trainor PA (2013) Tcof1 acts as a modifier of Pax3 during enteric nervous system development and in the pathogenesis of colonic aganglionosis. Hum Mol Genet 22:1206–1217
Bastir M, Rosas A, Gunz P, Peña-Melian A, Manzi G, Harvati K et al (2011) Evolution of the base of the brain in highly encephalized human species. Nat Commun 2:588
Bednarik RG (2012) An aetiology of hominin behavior. HOMO J 63:319–335
Bednarik RG (2014) Doing with less: hominin brain atrophy. HOMO J 6:433–449
Benes FM, Lim B, Matzilevich D, Walsh JP, Subburaju S, Minns M (2007) Regulation of the GABA cell phenotype in hippocampus of schizophrenics and bipolars. Proc Natl Acad Sci USA 104:10164–10169
Benítez-Burraco A, Boeckx C (2014) FOXP2, retinoic acid, and language: a promising direction. Front Cell Neurosci 8:387
Benítez-Burraco A, Boeckx C (2015a) Possible functional links among brain- and skull-related genes selected in modern humans. Front Psychol 6:794
Benítez-Burraco A, Boeckx C (2015b) Approaching motor and language deficits in autism from below: a biolinguistic perspective. Front Integr Neurosci 9:25
Benítez-Burraco A, Murphy E (2016) The oscillopatic nature of language deficits in autism: from genes to language evolution. Front Hum Neurosci 10:120
Benitez-Burraco A, Uriagereka J (2016) The immune syntax revisited: opening new windows on language evolution. Front Mol Neurosci 8:84
Benítez-Burraco A, Lattanzi W, Murphy E (2016) Language impairment in ASD resulting from a failed domestication of the human brain: a hypothesis. BioRxiv. doi:10.1101/046037
Bisaz R, Boadas-Vaello P, Genoux D, Sandi C (2013) Age-related cognitive impairments in mice with a conditional ablation of the neural cell adhesion molecule. Learn Mem 20:183–193
Boas F (1938) The mind of primitive man. Macmillan, New York
Boeckx C (2013) Biolinguistics: Forays into human cognitive biology. J Anthropol Sci 91:63–89
Boeckx C (2014) Elementary syntactic structures. Cambridge University Press, Cambridge
Boeckx C, Benítez-Burraco A (2014a) The shape of the human language-ready brain. Front Psychol 5:282
Boeckx C, Benítez-Burraco A (2014b) Globularity and language-readiness: generating new predictions by expanding the set of genes of interest. Front Psychol 5:1324
Boeckx C, Benítez-Burraco A (2015) Osteogenesis and neurogenesis: a robust link also for language evolution. Front Cell Neurosci 9:291
Boeckx C, Theofanopoulou C, Benítez-Burraco A (manuscript) GLI transcription factors and globularization of the human brain
Bruner E (2004) Geometric morphometrics and paleoneurology: brain shape evolution in the genus Homo. J Hum Evol 47:279–303
Bruner E (2010) Morphological differences in the parietal lobes within the human genus. Curr Anthropol 51:S77–S88
Bruner E, Amano H, de la Cuétara JM, Ogihara N (2015) The brain and the braincase: a spatial analysis on the midsagittal profile in adult humans. J Anat 227:268–276
Calloni GW, Glavieux-Pardanaud C, Le Douarin NM, Dupin E (2007) Sonic Hedgehog promotes the development of multipotent neural crest progenitors endowed with both mesenchymal and neural potentials. Proc Natl Acad Sci USA 104:19879–19884
Castellano S, Parra G, Sánchez-Quinto FA, Racimo F, Kuhlwilm M, Kircher M et al (2014) Patterns of coding variation in the complete exomes of three Neandertals. Proc Natl Acad Sci USA 111:6666–6671
Chang L, Kessler DS (2010) Foxd3 is an essential Nodal-dependent regulator of zebrafish dorsal mesoderm development. Dev Biol 342:39–50
Chiba S, Lee YM, Zhou W, Freed CR (2008) Noggin enhances dopamine neuron production from human embryonic stem cells and improves behavioral outcome after transplantation into Parkinsonian rats. Stem Cells 26:2810–2820
Christophersen NS, Grønborg M, Petersen TN, Fjord-Larsen L, Jørgensen JR, Juliusson B et al (2007) Midbrain expression of Delta-like 1 homologue is regulated by GDNF and is associated with dopaminergic differentiation. Exp Neurol 204:791–801
Cieri RL, Churchill SE, Franciscus RG, Tan J, Hare B (2014) Craniofacial feminization, social tolerance and the origins of behavioral modernity. Curr Anthropol 55:419–443
Correia AC, Costa M, Moraes F, Bom J, Nóvoa A, Mallo M (2007) Bmp2 is required for migration but not for induction of neural crest cells in the mouse. Dev Dyn 236:2493–2501
Darwin C (1871) The descent of man, and selection in relation to sex. John Murray, London
Deacon TW (2009) Relaxed selection and the role of epigenesis in the evolution of language. In: Blumberg M, Freeman J, Robinson S (eds) Oxford handbook of developmental behavioral neuroscience. Oxford University Press, New York
Delloye-Bourgeois C, Jacquier A, Charoy C, Reynaud F, Nawabi H, Thoinet K et al (2015) PlexinA1 is a new Slit receptor and mediates axon guidance function of Slit C-terminal fragments. Nat Neurosci 1:36–45
Devanna P, Middelbeek J, Vernes SC (2014) FOXP2 drives neuronal differentiation by interacting with retinoic acid signaling pathways. Front Cell Neurosci 8:305
Dixon J, Dixon MJ (2004) Genetic background has a major effect on the penetrance and severity of craniofacial defects in mice heterozygous for the gene encoding the nucleolar protein Treacle. Dev Dyn 229:907–914
Dutt S, Kléber M, Matasci M, Sommer L, Zimmermann DR (2006) Versican V0 and V1 guide migratory neural crest cells. J Biol Chem 17:12123–12131
Emes RD, Ponting CP (2001) A new sequence motif linking lissencephaly, Treacher Collins and oral–facial–digital type 1 syndromes, microtubule dynamics and cell migration. Hum Mol Genet 10:2813–2820
Endo Y, Ishiwata-Endo H, Yamada KM (2012) Extracellular matrix protein anosmin promotes neural crest formation and regulates FGF, BMP, and WNT activities. Dev Cell 23:305–316
Etchamendy N, Enderlin V, Marighetto A, Pallet V, Higueret P, Jaffard R (2003) Vitamin A deficiency and relational memory deficit in adult mice: relationships with changes in brain retinoid signalling. Behav Brain Res 145:37–49
Fernell E, Olsson VA, Karlgren-Leitner C, Norlin B, Hagberg B, Gillberg C (1999) Autistic disorders in children with CHARGE association. Dev Med Child Neurol 41:270–272
Fisher S, Grice EA, Vinton RM, Bessling SL, McCallion AS (2006) Conservation of RET regulatory function from human to zebrafish without sequence similarity. Science 312:276–279
Fossat N, Jones V, Garcia-Garcia MJ, Tam PP (2012) Modulation of WNT signaling activity is key to the formation of the embryonic head. Cell Cycle 11:26–32
Fuchs S, Herzog D, Sumara G, Büchmann-Møller S, Civenni G, Wu X et al (2009) Stage-specific control of neural crest stem cell proliferation by the small rho GTPases Cdc42 and Rac1. Cell Stem Cell 4:236–247
Fukuchi-Shimogori T, Grove EA (2001) Neocortex patterning by the secreted signaling molecule FGF8. Science 294:1071–1074
Gajavelli S, Wood PM, Pennica D, Whittemore SR, Tsoulfas P (2004) BMP signaling initiates a neural crest differentiation program in embryonic rat CNS stem cells. Exp Neurol 188:205–223
Giovannone D, Reyes M, Reyes R, Correa L, Martinez D, Ra H et al (2012) Slits affect the timely migration of neural crest cells via robo receptor. Dev Dyn 241:1274–1288
Glatt SJ, Cohen OS, Faraone SV, Tsuang MT (2011) Dysfunctional gene splicing as a potential contributor to neuropsychiatric disorders. Am J Med Genet B Neuropsychiatr Genet 156:382–392
Glejzer A, Laudet E, Leprince P, Hennuy B, Poulet C, Shakhova O et al (2011) Wnt1 and BMP2: two factors recruiting multipotent neural crest progenitors isolated from adult bone marrow. Cell Mol Life Sci 68:2101–2114
Gokhman D, Lavi E, Prüfer K, Fraga MF, Riancho JA, Kelso J et al (2014) Reconstructing the DNA methylation maps of the Neandertal and the Denisovan. Science 344:523–527
Gordon CT, Brinas IM, Rodda FA, Bendall AJ, Farlie PG (2010) Role of Dlx genes in craniofacial morphogenesis: Dlx2 influences skeletal patterning by inducing ectomesenchymal aggregation in ovo. Evol Dev 12:459–473
Green RE, Krause J, Briggs AW, Maricic T, Stenzel U, Kircher M et al (2010) A draft sequence of the neanderthal genome. Science 328:710–722
Gunz P, Neubauer S, Maureille B, Hublin JJ (2010) Brain development after birth differs between Neanderthals and modern humans. Curr Biol 20:R921–R922
Gunz P, Neubauer S, Golovanova L, Doronichev V, Maureille B, Hublin JJ (2012) A uniquely modern human pattern of endocranial development. Insights from a new cranial reconstruction of the Neandertal newborn from Mezmaiskaya. J Hum Evol 62:300–313
Hannula-Jouppi K, Kaminen-Ahola N, Taipale M, Eklund R, Nopola-Hemmi J, Kääriäinen H et al (2005) The axon guidance receptor gene ROBO1 is a candidate gene for developmental dyslexia. PLoS Genet 1:e50
Hare B, Tomasello M (2005) Human-like social skills in dogs? Trends Cogn Sci 9:439–444
Hari L, Miescher I, Shakhova O, Suter U, Chin L, Taketo M et al (2012) Temporal control of neural crest lineage generation by Wnt/β-catenin signaling. Development 139:2107–2117
Hidalgo-Figueroa M, Bonilla S, Gutiérrez F, Pascual A, López-Barneo J (2012) GDNF is predominantly expressed in the PV+ neostriatal interneuronal ensemble in normal mouse and after injury of the nigrostriatal pathway. J Neurosci 32:864–872
Huang YT, Iwamoto K, Kurosaki T, Nasu M, Ueda S (2005) The neuronal POU transcription factor Brn-2 interacts with Jab1, a gene involved in the onset of neurodegenerative diseases. Neurosci Lett 382:175–178
Hublin JJ, Neubauer S, Gunz P (2015) Brain ontogeny and life history in Pleistocene hominins. Philos Trans R Soc 370:20140062
Hurford JR (2011) The origins of grammar: language in the light of evolution, vol 2. Oxford University Press, Oxford
Ishii M, Arias AC, Liu L, Chen YB, Bronner ME, Maxson RE (2012) A stable cranial neural crest cell line from mouse. Stem Cells Dev 21:3069–3080
Iwamoto K, Bundo M, Yamada K, Takao H, Iwayama-Shigeno Y, Yoshikawa T et al (2005) DNA methylation status of SOX10 correlates with its downregulation and oligodendrocyte dysfunction in schizophrenia. J Neurosci 25:5376–5381
Jeong J, Li X, McEvilly RJ, Rosenfeld MG, Lufkin T, Rubenstein JL (2008) Dlx genes pattern mammalian jaw primordium by regulating both lower jaw-specific and upper jaw-specific genetic programs. Development 135:2905–2916
Jia L, Cheng L, Raper J (2005) Slit/Robo signaling is necessary to confine early neural crest cells to the ventral migratory pathway in the trunk. Dev Biol 282:411–421
Jian XC, Chen XQ, Hunan C (1995) Neurocristopathy that manifests right facial cleft and right maxillary duplication. Oral Surg Oral Med Oral Pathol Oral Radiol Endod 79:546–550
Jiang Q, Ho YY, Hao L, Nichols Berrios C, Chakravarti A (2011) Copy number variants in candidate genes are genetic modifiers of Hirschsprung disease. PLoS One 6:e21219
Jiang W, Yu Q, Gong M, Chen L, Wen EY, Bi Y et al (2012) Vitamin A deficiency impairs postnatal cognitive function via inhibition of neuronal calcium excitability in hippocampus. J Neurochem 121:932–943
Jones EG, Rubenstein JLR (2004) Expression of regulatory genes during differentiation of thalamic nuclei in mouse and monkey. J Comp Neurol 477:55–80
Kagawa H, Yamada H, Lin R-S, Mizuta T, Hasegawa T, Okanoya K (2012) Ecological correlates of song complexity in white-rumped munias: the implication of relaxation of selection as a cause for signal variation in birdsong. Interact Stud 13:263–284
Kagawa H, Suzuki K, Takahasi M, Okanoya K (2014) Domestication changes innate constraints for birdsong learning. Behav Process 106:91–97
Kameda Y, Saitoh T, Fujimura T (2011) Hes1 regulates the number and anterior–posterior patterning of mesencephalic dopaminergic neurons at the mid/hindbrain boundary (isthmus). Dev Biol 358:91–101
Kirby S (2012) Language is an adaptive system: the role of cultural evolution in the origins of structure. In: Tallerman M, Gibson KR (eds) The Oxford handbook of language evolution. Oxford University Press, Oxford
Kitagawa H, Fujiki R, Yoshimura K, Mezaki Y, Uematsu Y, Matsui D et al (2003) The chromatin-remodeling complex WINAC targets a nuclear receptor to promoters and is impaired in Williams syndrome. Cell 113:905–917
Kobayashi T, Yasuda K, Araki M (2010) Coordinated regulation of dorsal bone morphogenetic protein 4 and ventral Sonic hedgehog signaling specifies the dorso-ventral polarity in the optic vesicle and governs ocular morphogenesis through fibroblast growth factor 8 upregulation. Dev Growth Differ 52:351–363
Konopka G, Bomar JM, Winden K, Coppola G, Jonsson ZO, Gao F et al (2009) Human-specific transcriptional regulation of CNS development genes by FOXP2. Nature 462:213–217
Kuhlwilm M, Davierwala A, Pääbo S (2013) Identification of putative target genes of the transcription factor RUNX2. PLoS One 8:e83218
Lattanzi W, Bukvic N, Barba M, Tamburrini G, Bernardini C, Michetti F et al (2012) Genetic basis of single-suture synostoses: genes, chromosomes and clinical implications. Childs Nerv Syst 9:1301–1310
Le Douarin NM, Dupin E (2012) The neural crest in vertebrate evolution. Curr Opin Genet Dev 22:381–389
Ledda F, Paratcha G, Sandoval-Guzman T, Ibanez CF (2007) GDNF and GFR alpha 1 promote formation of neuronal synapses by ligand-induced cell adhesion. Nat Neurosci 10:293–300
Leventer RJ, Cardoso C, Ledbetter DH, Dobyns WB (2001) LIS1 missense mutations cause milder lissencephaly phenotypes including a child with normal IQ. Neurology 57:416–422
Li C, Xu S, Gotlieb AI (2013) The progression of calcific aortic valve disease through injury, cell dysfunction, and disruptive biologic and physical force feedback loops. Cardiovasc Pathol 22:1–8
Li J, Feng J, Liu Y, Ho TV, Grimes W, Ho HA et al (2015) BMP-SHH signaling network controls epithelial stem cell fate via regulation of its niche in the developing tooth. Dev Cell 33:125–135
Lieberman DE, McBratney BM, Krovitz G (2002) The evolution and development of cranial form in Homo sapiens. Proc Natl Acad Sci USA 99:1134–1139
Lin L, Shen S, Jiang P, Sato S, Davidson BL, Xing Y (2010) Evolution of alternative splicing in primate brain transcriptomes. Hum Mol Genet 19:2958–2973
Lin M, Pedrosa E, Shah A, Hrabovsky A, Maqbool S, Zheng D et al (2011) RNA-Seq of human neurons derived from iPS cells reveals candidate long non-coding RNAs involved in neurogenesis and neuropsychiatric disorders. PLoS ONE 6:e23356
Liu JA, Wu MH, Yan CH, Chau BK, So H (2013) Phosphorylation of Sox9 is required for neural crest delamination and is regulated downstream of BMP and canonical Wnt signalling. Proc Natl Acad Sci USA 110:2882–2887
Lo Nigro C, Chong CS, Smith AC, Dobyns WB, Carrozzo R, Ledbetter DH (1997) Point mutations and an intragenic deletion in LIS1, the lissencephaly causative gene in isolated lissencephaly sequence and Miller-Dieker syndrome. Hum Mol Genet 6:157–164
Long Z, Medlock C, Dzemidzic M, Shin YW, Goddard AW, Dydak U (2013) Decreased GABA levels in anterior cingulate cortex/medial prefrontal cortex in panic disorder. Prog Neuropsychopharmacol Biol Psychiatry 44:131–135
Luo T, Wagner E, Dräger UC (2009) Integrating retinoic acid signaling with brain function. Dev Psychol 45:139–150
Maczkowiak F, Matéos S, Wang E, Roche D, Harland R, Monsoro-Burq AH (2010) The Pax3 and Pax7 paralogs cooperate in neural and neural crest patterning using distinct molecular mechanisms, in Xenopus laevis embryos. Dev Biol 340:381–396
Mallo M (2001) Formation of the middle ear: recent progress on the developmental and molecular mechanisms. Dev Biol 231:410–419
Marcos-Mondéjar P, Peregrín S, Li JY, Carlsson L, Tole S, López-Bendito G (2012) The lhx2 transcription factor controls thalamocortical axonal guidance by specific regulation of robo1 and robo2 receptors. J Neurosci 13:4372–4385
Maricic T, Günther V, Georgiev O, Gehre S, Curlin M, Schreiweis C et al (2013) A recent evolutionary change affects a regulatory element in the human FOXP2 gene. Mol Biol Evol 30:844–852
Matsuo T, Osumi-Yamashita N, Noji S, Ohuchi H, Koyama E, Myokai F et al (1993) A mutation in the Pax-6 gene in rat small eye is associated with impaired migration of midbrain. Nat Genet 3:299–304
McEvilly RJ, de Díaz MO, Schonemann MD, Hooshmand F, Rosenfeld MG (2002) Transcriptional regulation of cortical neuron migration by POU domain factors. Science 295:1528–1532
McKinsey G, Lindtner S, Trzcinski B, Visel A, Pennacchio LA, Huylebroeck D et al (2013) Dlx1&2-dependent expression of Zfhx1b (Sip1, Zeb2) regulates the fate switch between cortical and striatal interneurons. Neuron 77:83–98
McLarren KW, Litsiou A, Streit A (2003) DLX5 positions the neural crest and preplacode region at the border of the neural plate. Dev Biol 259:34–47
McLean CY, Reno PL, Pollen AA, Bassan AI, Capellini TD, Guenther C et al (2011) Human-specific loss of regulatory DNA and the evolution of human-specific traits. Nature 471:216–219
Miller CT, Swartz ME, Khuu PA, Walker MB, Eberhart JK, Kimmel CB (2007) mef2ca is required in cranial neural crest to effect Endothelin1 signaling in zebrafish. Dev Biol 308:144–157
Mühleisen TW, Leber M, Schulze TG, Strohmaier J, Degenhardt F, Treutlein J et al (2014) Genome-wide association study reveals two new risk loci for bipolar disorder. Nat Commun 5:3339
Nelson CS, Fuller CK, Fordyce PM, Greninger AL, Li H, DeRisi JL (2013) Microfluidic affinity and ChIP-seq analyses converge on a conserved FOXP2-binding motif in chimp and human, which enables the detection of evolutionarily novel targets. Nucleic Acids Res 41:5991–6004
Neubauer S, Gunz P, Hublin JJ (2010) Endocranial shape changes during growth in chimpanzees and humans: a morphometric analysis of unique and shared aspects. J Hum Evol 59:555–566
Okanoya K (2012) Behavioural factors governing song complexity in Bengalese finches. Int J Comp Psychol 25:44–59
Oosterveen T, Kurdija S, Alekseenko Z, Uhde CW, Bergsland M, Sandberg M et al (2012) Mechanistic differences in the transcriptional interpretation of local and long-range Shh morphogen signaling. Dev Cell 23:1006–1019
Oosterveen T, Kurdija S, Ensterö M, Uhde CW, Bergsland M, Sandberg M et al (2013) SoxB1-driven transcriptional network underlies neural-specific interpretation of morphogen signals. Proc Natl Acad Sci USA 110:7330–7335
Pääbo S (2014) The human condition—a molecular approach. Cell 157:216–226
Paredes R, Arriagada G, Cruzat F, Villagra A, Olate J, Zaidi K et al (2004) Bone-specific transcription factor Runx2 interacts with the 1α,25-dihydroxyvitamin D3 receptor to up-regulate rat osteocalcin gene expression in osteoblastic cells. Mol Cell Biol 24:8847–8861
Parry DA, Logan CV, Stegmann AP, Abdelhamed ZA, Calder A, Khan S et al (2013) SAMS, a syndrome of short stature, auditory-canal atresia, mandibular hypoplasia, and skeletal abnormalities is a unique neurocristopathy caused by mutations in Goosecoid. Am J Hum Genet 93:1135–1142
Peterson KA, Nishi Y, Ma W, Vedenko A, Shokri L, Zhang X et al (2012) Neural-specific Sox2 input and differential Gli-binding affinity provide context and positional information in Shh-directed neural patterning. Genes Dev 26:2802–2816
Pleasure SJ, Anderson S, Hevner R, Bagri A, Marin O, Lowenstein DH et al (2000) Cell migration from the ganglionic eminences is required for the development of hippocampal GABAergic interneurons. Neuron 28:727–740
Plouhinec JL, Roche DD, Pegoraro C, Figueiredo AL, Maczkowiak F, Brunet LJ et al (2014) Pax3 and Zic1 trigger the early neural crest gene regulatory network by the direct activation of multiple key neural crest specifiers. Dev Biol 386:461–472
Pochon NAM, Menoud A, Tseng JL, Zurn AD, Aebischer P (1997) Neuronal GDNF expression in the adult rat nervous system identified by in situ hybridization. Eur J Neurosci 9:463–471
Potkin SG, Turner JA, Guffanti G, Lakatos A, Fallon JH, Nguyen DD et al (2009) A genome-wide association study of schizophrenia using brain activation as a quantitative phenotype. Schizophr Bull 35:96–108
Prescott SL, Srinivasan R, Marchetto MC, Grishina I, Narvaiza I, Selleri L et al (2015) Enhancer divergence and cis-regulatory evolution in the human and chimp neural crest. Cell 163:68–83
Price EO (2002) Animal domestication and behavior. CABI Publishing, New York
Prüfer K, Racimo F, Patterson N, Jay F, Sankararaman S, Sawyer S et al (2014) The complete genome sequence of a Neanderthal from the Altai Mountains. Nature 505:43–49
Quarto N, Wan DC, Kwan MD, Panetta NJ, Li S, Longaker MT (2010) Origin matters: differences in embryonic tissue origin and Wnt signaling determine the osteogenic potential and healing capacity of frontal and parietal calvarial bones. J Bone Miner Res 25:1680–1694
Read AP, Newton VE (1997) Waardenburg syndrome. J Med Genet 34:656–665
Reale ME, Webb IC, Wang X, Baltazar RM, Coolen LM, Lehman MN (2013) The transcription factor Runx2 is under circadian control in the suprachiasmatic nucleus and functions in the control of rhythmic behavior. PLoS One 8:e54317
Reiner O, Carrozzo R, Shen Y, Wehnert M, Faustinella F, Dobyns WB et al (1993) Isolation of a Miller–Dieker lissencephaly gene containing G protein beta-subunit-like repeats. Nature 364:717–721
Roper RJ, VanHorn JF, Cain CC, Reeves RH (2009) A neural crest deficit in Down syndrome mice is associated with deficient mitotic response to Sonic hedgehog. Mech Dev 126:212–219
Rossi M, Jones RL, Norbury G, Bloch-Zupan A, Winter RM (2003) The appearance of the feet in Pfeiffer syndrome caused by FGFR1 P252R mutation. Clin Dysmorphol 12:269–274
Ruest LB, Hammer RE, Yanagisawa M, Clouthier DE (2003) Dlx5/6-enhancer directed expression of Cre recombinase in the pharyngeal arches and brain. Genesis 37:188–194
Sarnat HB, Flores-Sarnat L (2001) Neuropathologic research strategies in holoprosencephaly. J Child Neurol 16:918–931
Schroeter M, Zickler P, Denhardt DT, Hartung HP, Jander S (2006) Increased thalamic neurodegeneration following ischaemic cortical stroke in osteopontin-deficient mice. Brain 129:1426–1437
Scott N, Neubauer S, Hublin JJ, Gunz P (2014) A shared pattern of postnatal endocranial development in extant hominoids. Evol Biol 41:572–594
Segklia A, Seuntjens E, Elkouris M, Tsalavos S, Stappers E, Mitsiadis TA et al (2012) Bmp7 regulates the survival, proliferation, and neurogenic properties of neural progenitor cells during corticogenesis in the mouse. PLoS One 7:e34088
Sheehan-Rooney K, Swartz ME, Lovely CB, Dixon MJ, Eberhart JK (2013) Bmp and Shh signaling mediate the expression of satb2 in the pharyngeal arches. PLoS One 8:59533
Shen Q, Christakos S (2005) The vitamin D receptor, Runx2, and the Notch signaling pathway cooperate in the transcriptional regulation of osteopontin. J Biol Chem 280:40589–40598
Shows KH, Shiang R (2008) Regulation of the mouse Treacher Collins Syndrome homolog (Tcof1) promoter through differential repression of constitutive expression. DNA Cell Biol 27:589–600
Shu T, Ayala R, Nguyen MD, Xie Z, Gleeson JG, Tsai LH (2004) Ndel1 operates in a common pathway with LIS1 and cytoplasmic dynein to regulate cortical neuronal positioning. Neuron 44:263–277
Silver DL, Watkins-Chow DE, Schreck KC, Pierfelice TJ, Larson DM, Burnetti AJ et al (2010) The exon junction complex component Magoh controls brain size by regulating neural stem cell division. Nat Neurosci 13:551–558
Sjamsudin J, David DJ, Singh GD (2001) An Indonesian child with orofacial duplication and neurocristopathy anomalies: case report. J Craniomaxillofac Surg 29:195–197
Smit DJ, Smith AG, Parsons PG, Muscat GE, Sturm RA (2000) Domains of Brn2 that mediate homodimerization and interaction with general and melanocytic transcription factors. Eur J Biochem 267:6413–6422
Smith K, Kirby S (2008) Cultural evolution: implications for understanding the human language faculty and its evolution. Philos Trans R Soc Lond B Biol Sci 363:3591–3603
Smith DS, Niethammer M, Ayala R, Zhou Y, Gambello MJ, Wynshaw-Boris A et al (2000) Regulation of cytoplasmic dynein behaviour and microtubule organization by mammalian Lis1. Nat Cell Biol 2:767–775
Solomon BD, Pineda-Alvarez DE, Gropman AL, Willis MJ, Hadley DW, Muenke M (2012) High intellectual function in individuals with mutation-positive microform holoprosencephaly. Mol Syndrom 3:140–142
Somerville MJ, Mervis CB, Young EJ, Seo EJ, del Campo M, Bamforth S et al (2005) Severe expressive-language delay related to duplication of the Williams–Beuren locus. N Engl J Med 353:1694–1701
Sperber SM, Saxena V, Hatch G, Ekker M (2008) Zebrafish dlx2a contributes to hindbrain neural crest survival, is necessary for differentiation of sensory ganglia and functions with dlx1a in maturation of the arch cartilage elements. Dev Biol 314:59–70
Spiteri E, Konopka G, Coppola G, Bomar J, Oldham M, Ou J et al (2007) Identification of the transcriptional targets of FOXP2, a gene linked to speech and language, in developing human brain. Am J Hum Genet 81:1144–1157
Stuhlmiller TJ, García-Castro MI (2012) Current perspectives of the signaling pathways directing neural crest induction. Cell Mol Life Sci 69:3715–3737
Sugitani M, Sugai T, Onoda N (2002) Postsynaptic activity of metabotropic glutamate receptors in the piriform cortex. NeuroReport 13:1473–1476
Suntsova M, Gogvadze EV, Salozhin S, Gaifullin N, Eroshkin F, Dmitriev SE et al (2013) Human-specific endogenous retroviral insert serves as an enhancer for the schizophrenia-linked gene PRODH. Proc Natl Acad Sci USA 110:19472–19477
Szklarczyk D, Franceschini A, Kuhn M, Simonovic M, Roth A, Minguez P et al (2011) The STRING database in 2011: functional interaction networks of proteins, globally integrated and scored. Nucleic Acids Res 39:D561–D568
Takahasi M, Okanoya K (2010) Song learning in wild and domesticated strains of white-rumped munia, Lonchura striata, compared by cross-fostering procedures: domestication increases song variability by decreasing strain-specific bias. Ethology 116:396–405
Tang W, Tang J, He J, Zhou Z, Qin Y, Qin J et al (2015) SLIT2/ROBO1-miR-218-1-RET/PLAG1: a new disease pathway involved in Hirschsprung disease. J Cell Mol Med 19:1197–1207
Theofanopoulou C (2015) Brain asymmetry in the white matter making and globularity. Front Psychol 6:1355
Theofanopoulou, C and Boeckx, C (2016). Oxytocin pathways, language-related genes, and the emergence of modern cognition. 10th FENS Forum of Neuroscience, Bella Center, Copenhagen, Denmark, 2–6 July
Thomas J (2014) Self-domestication and language evolution. Ph.D. thesis, University of Edinburgh
Thompson B, Smith K, Kirby S (2016) Culture shapes the evolution of cognition. PNAS 113:4530–4535
Tomasello M (2009) The cultural origins of human cognition. Harvard University Press, Cambridge
Tomasello M (2014) A natural history of human thinking. Harvard University Press, Cambridge
Toyo-oka K, Shionoya A, Gambello MJ, Cardoso C, Leventer R, Ward HL et al (2003) 14-3-3epsilon is important for neuronal migration by binding to NUDEL: a molecular explanation for Miller–Dieker syndrome. Nat Genet 34:274–285
Trokovic N, Trokovic R, Mai P, Partanen J (2003) Fgfr1 regulates patterning of the pharyngeal region. Genes Dev 17:141–153
Van de Putte T, Maruhashi M, Francis A, Nelles L, Kondoh H, Huylebroeck D et al (2003) Mice lacking Zfhx1b, the gene that codes for Smad-interacting protein-1, reveal a role for multiple neural crest cell defects in the etiology of Hirschsprung disease–mental retardation syndrome. Am J Hum Genet 72:465–470
Van de Putte T, Francis A, Nelles L, van Grunsven LA, Huylebroeck D (2007) Neural crest-specific removal of Zfhx1b in mouse leads to a wide range of neurocristopathies reminiscent of Mowat-Wilson syndrome. Hum Mol Genet 16:1423–1436
Vendramini D (2014) Them and us: how Neanderthal predation created modern humans. Kardoorair Press, Armidale, p 2014
Vernes SC, Oliver PL, Spiteri E, Lockstone HE, Puliyadi R, Taylor JM et al (2011) Foxp2 regulates gene networks implicated in neurite outgrowth in the developing brain. PLoS Genet 7:e1002145
Vue TY, Aaker J, Taniguchi A, Kazemzadeh C, Skidmore JM, Martin DM et al (2007) Characterization of progenitor domains in the developing mouse thalamus. J Comp Neurol 505:73–91
Wada N, Javidan Y, Nelson S, Carney TJ, Kelsh RN, Schilling TF (2005) Hedgehog signaling is required for cranial neural crest morphogenesis and chondrogenesis at the midline in the zebrafish skull. Development 132:3977–3988
Wang R, Chen CC, Hara E, Rivas MV, Roulhac PL, Howard JT et al (2015) Convergent differential regulation of SLIT-ROBO axon guidance genes in the brains of vocal learners. J Comp Neurol 523:892–906
Wilkins AS, Wrangham RW, Fitch WT (2014) The “domestication syndrome” in mammals: a unified explanation based on neural crest cell behavior and genetics. Genetics 197:795–808
Wyatt AW, Osborne RJ, Stewart H, Ragge NK (2010) Bone morphogenetic protein 7 (BMP7) mutations are associated with variable ocular, brain, ear, palate, and skeletal anomalies. Hum Mutat 31:781–787
Yin Z, Haynie J, Yang X, Han B, Kiatchoosakun S, Restivo J et al (2002) The essential role of Cited2, a negative regulator for HIF-1alpha, in heart development and neurulation. Proc Natl Acad Sci USA 99:10488–10493
Yoon DS, Kim YH, Lee S, Lee KM, Park KH, Jang Y et al (2014) Interleukin-6 induces the lineage commitment of bone marrow-derived mesenchymal multipotent cells through down-regulation of Sox2 by osteogenic transcription factors. FASEB J 28:3273–3286
Yoshida T, Kanegane H, Osato M, Yanagida M, Miyawaki T, Ito Y et al (2003) Functional analysis of RUNX2 mutations in cleidocranial dysplasia: novel insights into genotype-phenotype correlations. Blood Cells Mol Dis 30:184–193
Young DW, Hassan MQ, Pratap J, Galindo M, Zaidi SK, Lee SH et al (2007) Mitotic occupancy and lineage-specific transcriptional control of rRNA genes by Runx2. Nature 445:442–446
Yuge K, Kataoka A, Yoshida AC, Itoh D, Aggarwal M, Mori S et al (2011) Region-specific gene expression in early postnatal mouse thalamus. J Comp Neurol 519:544–561
Zeder MA, Emshwiller E, Smith BD, Bradley DG (2006) Documenting domestication: the intersection of genetics and archaeology. Trends Genet 22:139–155
Acknowledgments
Preparation of this work was supported by funds from the Spanish Ministry of Economy and Competitiveness (Grants FFI2013-43823-P and FFI2014-61888-EXP), as well as funds from a Marie Curie International Reintegration Grant from the European Union (PIRG-GA-2009-256413), research funds from the Fundació Bosch i Gimpera, and from the Generalitat de Catalunya (2014-SGR-200, and FI-grant).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Human and Animal Rights
Their research did not involve human participants or animals.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Benítez-Burraco, A., Theofanopoulou, C. & Boeckx, C. Globularization and Domestication. Topoi 37, 265–278 (2018). https://doi.org/10.1007/s11245-016-9399-7
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11245-016-9399-7