Abstract
Silver birch (Betula pendula Roth.) is an ecologically and economically important deciduous tree species in Northern Europe. In vitro culture initiation and shoot rejuvenation from birches in their reproductive phase is difficult and often unsuccessful. Currently, markers to evaluate the degree of juvenility of birch in vitro shoots, which could be used to track the rejuvenation process and to determine factors affecting it, have not been developed. The aims of this study were to assess expression of juvenility related microRNAs and target genes during micropropagation of silver birch, and to investigate factors affecting juvenility of birch genotypes with different in vitro morphogenic ability. Potential precursor sequences and target genes of the microRNAs miR156 and miR172 were determined by alignment to the birch unigene set. Ten potential miR156 and miR172 precursor sequences were identified, and were tested for the ability to form the required stem-loop structure. Based on precursor sequences, primers were designed for real time PCR analysis of precursor miRNA expression. Expression patterns of two miR156 family precursors (miR156_511 and miR156_789) and one miR172 precursor (miR172_1931) and two target genes (BpSPL1 and BpAP2) had the best correlation with juvenility/maturity in the analysed in vitro propagated silver birch samples. Expression patterns of these miRNA precursors and target genes were also investigated in samples cultured under different in vitro conditions. This study provides an initial survey of molecular markers for assessment of phase change in birch in vitro micropropagation.
Key message
This study investigated expression of juvenility related micro RNAs and target genes in Betula pendula micropropagated material with differing juvenility and in vitro culture conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Silver birch (Betula pendula Roth.) is one of the most ecologically and economically important tree species in northern boreal forests. Silver birch breeding programs, producing improved reproductive material for planting highly productive and qualitative birch stands have been implemented in many northern European countries (Hynynen et al. 2010; Gailis et al. 2020). This increases the added value of timber and contributes to ensuring the competitiveness and sustainable development of the forest industry. In addition, fast-growing forest stands provide an active carbon dioxide capture sink, which is important for mitigating climate change impacts.
Vegetative propagation (including micropropagation) of mature birch trees enables capture of genetic gain more rapidly than by sexual reproduction, due to selection and maintenance of both additive and non-additive gene effects (George et al. 2008). Vegetatively propagated material can then be used for further breeding and establishment of seed orchards, from which genetically improved nursery stock is obtained. Clonal propagation of high quality, selected silver birch germplasm can increase the productivity of birch stands, compared to conventional plantations (Zeltinš et al. 2018). In addition, vegetative propagation of selected genotypes, for use in controlled crosses, can shorten the breeding cycle by 10–15 years (Ewald et al. 2001). Plants, including woody plants undergo several development phases—embryonic, vegetative juvenile phase, vegetative mature phase and reproductive phase (Poethig 2013; Díaz-Sala 2016). In order to obtain the most accurate tree productivity and quality assessment, evaluation of birch breeding material according to phenotypic parameters is carried out when trees have reached their mature phase.
At maturity, vegetative propagation capacity declines abruptly and silver birch, as well as other tree species with high breeding value, become recalcitrant to propagation methods such as rooting of cuttings, therefore in vitro methods have been investigated (Welander 1993; Jain and Häggman 2007; Bonga 2016). Due to the effect of the in vitro culture environment, it is possible for initially mature plant material to undergo a rejuvenation process via axillary or adventitious shoot proliferation (Fig. 1), resulting in juvenile characteristics including high multiplication and rooting rate (Sánchez et al. 1997; Renau-Morata et al. 2005). Explants obtained from mature trees are often characterized by slow growth and reduced responsiveness to cytokinin and auxin treatment, resulting to low morphogenic ability (McCown 2000; Vinoth and Ravindhran 2018). Therefore, various in vitro methods, including medium composition and other cultivation conditions are used to achieve successful rejuvenation of mature shoots in vitro. Some birch genotypes show better responsiveness in vitro and can be successfully rejuvenated and regenerated, while others remain in a mature state and gradually die (Ewald et al. 2001; O’Dowd 2004). This responsiveness to in vitro regeneration is largely influenced by plant juvenility state (Fig. 1). However, the molecular mechanisms and stability of juvenility have not yet been fully elucidated (Basheer-Salimia 2007; Matsoukas et al. 2013; Jia et al. 2017).
Markers indicating juvenility would increase the effectiveness of research of this phenomenon in birch in vitro shoots. To date, no morphological, physiological and molecular markers have been developed to evaluate juvenility state or the rejuvenation process in silver birch in vitro shoots. Rejuvenation methods of the mother plant prior to cultivation as well as during cultivation can be used to achieve successful rejuvenation of mature shoots in vitro. To date, phase-change markers for other species have been based on morphological, anatomical and physiological traits, such as differences in foliar and stem morphology, reduced regeneration potential, cellular and subcellular anatomy, plant hormone types and concentrations and other chemicals. However, in some species, traits associated with the juvenile-to-adult phases of development involve rather subtle differences and are hard to recognize (Wang et al. 2011; Feng et al. 2016; Xu et al. 2016). It has been reported that phase change signals in the apical meristem come from leaves or leaf primordia, which also enables the acquisition of juvenile material by isolating the apical meristem at the correct time (Orkwiszewski and Poethig 2000).
Recent studies with annual model plant species indicate that the main endogenous regulators of juvenility are the evolutionary highly conserved microRNAs (miRNAs; miRs), miR156, miR172 and their target genes (Wu et al. 2009; Wang et al. 2011; Xing et al. 2014; Jia et al. 2017; Ahsan et al. 2019b, a). MiRNAs are a class of non-protein coding small RNAs (sRNAs) of ~ 20 to 24 nucleotides in length that play an important role in a variety of biological and metabolic processes, primarily through the coordinated action on the post-transcriptional control of gene expression (Carrington and Ambros 2003). MiRNAs control essential aspects of plant growth and development, and are also important in responses to stress and pathogens (Bartel 2004; Borges and Martienssen 2015; Wang et al. 2016). MiRNAs bind to complementary messenger RNA (mRNA) sequences, causing translational repression (inhibition of translation) or by direct cleavage of mRNA (Carrington and Ambros 2003). MiRNAs are initially synthesized as primary microRNA transcripts (pri-RNAs) one to several hundred nucleotides long, containing at least one hairpin stem loop structure. Several enzymes (Dicer-like enzyme 1, Hasty and others) are involved in the generation of ∼ 70 nt long miRNA precursors (pre-miRNAs) and subsequently mature miRNA molecules (Neutelings et al. 2012). MiRNAs are categorised into families based on sequence homology of the mature miRNAs. However, the primary and precursor sequences that encode identical or highly similar mature miRNAs can differ in sequence, and may have differing expression patterns, e.g. being expressed at different developmental stages or tissues, or in response to different abiotic or biotic conditions (Bielewicz et al. 2012).
Many studies have shown that overexpression of miR156 prolongs the juvenile phase and delays phase change for both herbaceous and woody plants (Wu and Poethig 2006; Wu et al. 2009; Wang et al. 2011; Zhang et al. 2015). Conversely, if miR156 expression decreases, phase change, and thus flowering, occurs more rapidly (Yang et al. 2013; Yu et al. 2013). MiR156 is highly expressed in the juvenile plant state and gradually decreases during maturation (Wu and Poethig 2006; Wu et al. 2009; Wang et al. 2011). MiR156 targets SQUAMOSA promoter binding protein-like (SBP/SPL) transcription factor genes, repressing the expression of SPL genes, which promote the transition to maturity by up-regulating key MADS-box genes, such as APETALA (AP1), LEAFY (LFY) and FRUITFULL (FUL). In Arabidopsis thaliana, 11 of 17 SPL genes have been identified as miR156 targets and are structurally divided into two groups: SPL3 (SPL3, SPL4, SPL5) and SPL9 (SPL2, SPL6, SPL9, SPL10, SPL11, SPL13, SPL15) (Xing et al. 2010). Each of these transcription factors is responsible for different phase-specific properties, including leaf shape, phylotaxis, rooting potential, flowering time, and leaf initiation (Bergonzi and Albani 2011). SPL1 is a birch homologue of the Arabidopsis SPL3 gene. SPL1 from Betula pendula is the first member of the SBP-box gene family characterized in trees (Lännenpää et al. 2004). MiR156 also suppresses expression of miR172 via SPL9 and SPL10, which leads to elevated levels of miR172 target genes belonging to the AP2-like transcription factor family genes: APETALA2 (AP2), TARGET OF EAT1 (TOE1), TOE2 and TOE3, SCHLAFMUTZE (SMZ), and SCHNARCHZAPFEN (SNZ).
Other physiological factors can also affect juvenility. One of the most important factors affecting juvenile state could be sugars. Arabidopsis mutants deficient in chlorophyll show delayed transition to maturity and photosynthesis products directly lower miR156A and miR156C gene expression (Yu et al. 2013). Plant nitrogen content also affects plant development and phase change. Nitrogen is an important factor influencing phase change. Nitrogen deficiency in A. thaliana plants has been observed to cause an increase in miR156 levels with a concomitant decrease in miR172 levels, which promotes the juvenile phase (Liang et al. 2012; Vidal et al. 2014).
Although a large number of miRNAs have been identified in various plant species, there are no reports on the regulatory miR156-SPL-miR172 network associated with phase change in Betula pendula. As the miR156-SPL-miR172 module is highly conservative among plant species, it could be used to determine the juvenility state of grafted plants, as well as to develop precise phenotypic and genetic markers that can be used to identify which explant types are the most juvenile and thus most suited for regeneration attempts. They would also be useful to monitor progress during rejuvenation to distinguish whether temporal variations in vegetative traits represent the same, or different, developmental processes. In studies on the change of plant development phases and their control mechanisms, it is important to use the most accurate markers characterizing the juvenile or mature phase. During rejuvenation, shoots switch from the vegetative mature phase to the vegetative juvenile phase. There is a lack of detailed studies on the physiological and molecular factors influencing the rejuvenation process. Depending on the plant species, each of these phases has morphological, anatomical and physiological characteristics that can be used as markers for a particular developmental phase (Zimmerman et al. 1985; Haffner et al. 1991; Preece 2008).
Utilising the latest molecular findings about miRNAs and their target genes, the main regulators of juvenility, the association of expression levels of miRNAs such as miR156, miR172 and their target genes with in vitro morphogenic ability of selected silver birch genotypes in Latvia was investigated. The aims of this study were to assess expression of juvenility related microRNAs and target genes during the micropropagation process of silver birch, and to investigate factors affecting juvenility of birch genotypes with different in vitro morphogenic ability. This can improve the propagation success rate for high quality selected birch genotypes, increasing the efficiency of breeding programs and the value of forest stands over the rotation period.
Materials and methods
Plant material and extraction of RNA
Plant material for miRNA and target gene expression analysis was collected for total RNA isolation and real time PCR analysis and stored at − 80 °C until extraction. Total RNA was extracted from leaves of rejuvenated in vitro shoots (REV). Two types of mature in vitro shoots were analysed, one of which (in vitro mature—IVM) exhibited typical signs of maturity (thick stems, large and thick leaves, inability to proliferate). The other type (in vitro mature rejuvenated—IVMR) was significantly different, and exhibited signs of a mature in vitro culture and then, after approximately 4 months, started growing in length, axillary or adventitious shoots appeared, and other signs of juvenility were observed (thin and long stems, thin leaves, ability to grow, high reproductive capacity). However, after approximately 6 months, this culture again showed signs of maturity. Leaves from a mature (approximately 20 years old) silver birch were used as a mature control (MAT), and 2 month old seedlings were used as a juvenile control (JUV). RNA was extracted using a standard phenol/chloroform/isoamyl alcohol protocol (Rubio-Piña and Zapata-Pérez 2011). Total extracted RNA from all samples was treated with the Turbo DNA-free kit (ThermoFisher Scientific, Cat. No. AM1907) following the manufacturer’s instructions.
RNA quality control
RNA concentration was measured with a Qubit and Quant-iT™ RNA BR Assay Kit (ThermoFisher, Cat. No. Q10210). RNA purity (DNA contamination) was tested by polymerase chain reaction (PCR) with an RNA stock solution as template and three birch genomic microsatellite locus primers L7.8, L7.4 and L1.10 (Kulju et al. 2004). Each forward primer was labelled with a different fluorophore (6-FAM, HEX, or TMR) to facilitate visualization using capillary electrophoresis. The PCR reactions for the microsatellite markers were carried out in a 10 μL solution containing a final concentration of 1 × HOT FIREPol® Blend Master Mix with 10 mM MgCl2 (Solis Biodyne, Cat. No. 04-27-00120), 0.3 mM of each primer, 1 μL RNA solution. PCR cycling conditions consisted of an initial denaturation step of 95 °C for 15 min; 35 cycles of 95 °C for 20 s, 55 °C for 30 s, and 72 °C for 45 s; followed by a final extension step of 72 °C for 10 min. PCR reaction were carried out in an Eppendorf Mastercycler gradient thermal cycler. Amplification fragments were separated on an ABI Prism 3130xl Genetic Analyzer (Life Technologies) and genotyped with GeneMapper 3.5. If no PCR amplification fragments were detected, RNA samples were considered to be free of DNA contamination. If PCR amplification fragments were detected, RNA samples were again treated with the Turbo DNA-free kit, and RNA concentrations purity were reanalysed prior to reverse transcription and real time PCR analysis.
Identification of potential miRNA precursors and target genes
Since mature birch miRNA sequences are not currently available, the mature miR156 and miR172 sequences and precursors were selected from other species (based on previous reports that these miRNAs and genes are the main regulators of juvenility) using the miRBase database version 22 (www.mirbase.org) and publicly available mature miRNA sequences from different plant species (Kozomara and Griffiths-Jones 2011, 2014). The CoGe database (https://genomevolution.org/coge/CoGeBlast.pl) was used to identify potential precursor miRNAs in the silver birch genome (Betula pendula (id 35079) scaffold assembly, vv1.2 scaffolds unmasked), containing the mature miRNA region. Precursor stem loop secondary structures were predicted using the Mfold program (http://mfold.rna.albany.edu/?q=mfold) web server (Zuker 2003). To distinguish miRNAs from other RNAs, minimum free-folding energy index (MFEI) (Zhang et al. 2006) was calculated to confirm that the precursor sequences conformed to the requirements for forming the miRNA precursor structures as reported previously (Zhang et al. 2006; Axtell and Meyers 2018; Krivmane et al. 2020). The location of the mature miRNA sequence on the stem-loop was taken into account to identify potential pre-miRNAs. As a result, ten predicted precursor miRNA stem loop sequences were found in the birch genome—seven for the miR156 and three for the miR172 family. The length of precursors varied from 82 to 134 nucleotides. Minimum folding free energy indexes ranged from 0.70 to 1.21, with most (nine precursor sequences) having a value of > 0.85.
Primer design for amplification of precursor miRNAs and target genes
Primers for pre-miRNA amplification were designed and selected using predicted pre-miRNA sequences (Primer 3 version 0.4.0; Table 1). The main criterion was that these primers must amplify a 70–150 bp long stem loop region, including the mature miRNA sequence. Forward primers were placed as close as possible to the mature miRNA sequence, but the reverse primers were placed within or as close as possible to the stem loop sequence. Published sequences of SPL and AP2-like target genes in the NCBI database (https://www.ncbi.nlm.nih.gov/) were used for identification of similar sequences in the silver birch genome in the GoGe database and primers for target genes were designed using Primer 3 (Table 1). The nucleotide sequence of Betula pendula SPL1 is similar to Antirrhinum SBP2 and Arabidopsis SPL3, except that SPL1 in Betula pendula does not contain an intron typical to all other known SBP-box genes studied thus far (Lännenpää et al. 2004). Therefore, two SPL target gene primers (for BpSPL1 and BpSPL9) were designed. Ethylene-responsive transcription factor RAP sequences were identified in the GoGe database by homology with AP2-like genes and primers were designed as described previously (Table 1).
Expression analysis of precursor miRNAs and target genes using RT-PCR
The Taqman Reverse transcription kit (Thermo Fisher Scientific, Cat. no. 4304134) was used for reverse transcription of 1 μg RNA with Oligo d(T)16 (Thermo Fisher Scientific Cat. no. 4304134). The obtained cDNA was diluted to 10 ng/μl with nuclease-free water and 2 μl of cDNA was used for RT-PCR analysis.
Comparative Ct RT- PCR was performed with the Maxima SYBR Green/ROX qPCR Master Mix (2 ×) (Thermo Fisher Scientific, Cat. No. K0221) using a standard protocol on a StepOnePlus thermocycler (Thermo Fisher Scientific, PN 4376785). For each sample, three technical replications were done. Four reference genes (endogenous controls) were used: actin (Ruonala et al. 2006), α-tubulin (Keinänen et al. 2007; Žiarovská et al. 2013; Fernández-Fuego et al. 2017), Peptidyl-prolyl isomerase (cyclophilin) and transcription factor CBF1 (Žiarovská et al. 2013). After evaluation of the four reference genes, the actin (primers—Bp_Act F and R) and cyclophilin (Bp_Cycloph F and R) genes were selected for analysis of precursor miRNA and target gene expression. The other endogenous controls were not used for data normalization because large differences (Ct values between samples varied by 3 to 7) were observed between sample Ct values. Real-time PCR conditions were: 10 min at 95 °C, followed by 40 cycles of denaturation for 15 s at 95 °C, 15 s at 95 °C, 60 s at 60 °C, 15 s at 95 °C.
Analysis of RT-PCR data was done using the StepOne ™ Software v.2.3. The cycle threshold (Ct) is the cycle at which fluorescence exceeds a fixed threshold. The Ct values for the analysed precursor miRNAs, target genes and endogenous controls were obtained using the default threshold settings. Relative expression levels (relative target quantity—RQ) were determined using the 2−ΔΔCt method after normalization by comparison with the endogenous control Ct values. The minimum and maximum RQ values indicate the error associated with the RQ value for the analysed precursor miRNAs and target genes. These values were computed using RQmin = 2 – (RQ – SE), RQmax = 2 – (RQ + SE), where SE is the standard error for the RQ. Statistical significance of differences in the expression levels of the analysed precursor miRNAs and target genes was established using one-way analysis of variance (ANOVA) and t tests, with a p-value threshold of 0.05.
Effect of in vitro culture conditions on juvenility
In addition, the effect of subculture duration, growth media and sucrose concentrations, and plant growth regulators on the rejuvenation of stabilized in vitro birch shoots was determined by analysing the expression of miR156, miR172 precursors and their target genes. For this study, a birch shoot culture obtained in 2016 from clone L29 and maintained on woody plant medium (WPM) (Lloyd and McCown 1980) supplemented with zeatin at a concentration of 0.5 mg L−1 was used. The effect of three different subculture durations was determined. Shoots were cultured on WPM medium supplemented with zeatin 0.5 mg L−1 and were transferred to new medium every 10 days, 30 days or not transferred. After 60 days, shoots were harvested for miRNA and target gene expression analysis.
The effect of growth media was determined by culturing the shoots for 60 days in two macronutrient media supplemented with zeatin 0.5 mg L−1—Murashige–Skoog (MS) (Murashige and Skoog 1962) macronutrient medium (total nitrogen content 840 mg L−1), and woody plant media (WPM) (Lloyd and McCown 1980) (total nitrogen content 206 mg L−1). Both media had 6 g L−1 agar and 20 g L−1 sucrose. Shoots were subcultured onto fresh media after 30 days, and samples collected for analysis after a total of 60 days.
The effect of sucrose concentration was determined by culturing the shoots for 60 days in WPM medium with sucrose concentrations of 20 and 40 g L−1. The effect of different plant growth regulators was determined in three variants by culturing the shoots for 60 days on WPM medium supplemented with: (1) 6-benzylaminopurine (BAP) at a concentration of 1.0 mg L−1; (2) indole-3-acetic acid (IAA) at a concentration of 0.5 mg L−1; (3) zeatin at a concentration of 0.5 mg L−1. All samples were subcultured onto fresh media after 30 days, and samples were collected for RNA extraction after 60 days.
The in vitro cultures were initiated by removing 1.5 cm long shoot tips were from shoots of the stabilized cultures in a laminar box. Shoot tips were placed vertically in medium (30 mL) in a glass jar (200 mL). Each variant was represented by 10 jars of 10 shoots each. The pH of all study media was adjusted to 5.8 before autoclaving using 0.1 M HCl and 1 M NaOH. Cultures were kept in the growth room at 25 ± 3 °C for 16/8 h light/dark photoperiod, white, fluorescent light (photosynthetically active radiation (PAR) 140–160*mol*m−2*s−1).
Results
Expression analysis of precursor miRNAs and target genes in tissue culture and control samples of differing juvenility
Relative expression of the seven miR156 precursors was highest in the juvenile control (JUV), except for miR156_801, where the expression was highest in the REV and IVM samples. In all cases, relative expression was lowest in the mature (MAT) controls. The expected patterns of expression (upregulated in JUV, REV and IVMR samples, downregulated in IVM and MAT samples) was observed with five of the seven analysed pre-miR156 primer pairs (miR156_801 and miR156_51 did not conform to this pattern) (Supplementary file 1). In general, changes in relative expression levels were low between all sample types, except for miR156_237 and miR156_237v2, where relative expression in JUV samples was over 60- and 30-fold higher than the other samples, respectively, including the rejuvenated in vitro shoot samples (REV). Relative expression of the miR156_789, miR156_511 and miR156_374 precursors had the expected expression patterns with reference to juvenility/maturity over all analysed samples, including the in vitro samples (Fig. 2).
Relative expression (RQ) of the miR156 family miRNA precursors. REV rejuvenated in vitro shoots, IVM typical mature in vitro shoots, IVMR mature in vitro shoots with signs of juvenility, MAT mature control, JUV juvenile control. Bars indicate minimum and maximum RQ values, samples labelled with different letters are statistically different (p < 0.05)
Expression patterns of two of the miR172 precursors was as expected, with relative expression levels highest in the mature controls, and low in the other samples. Compared to the reference rejuvenated sample, expression of miR172_1931 was over 160-fold higher, and expression of miR172_42 was over 50-fold higher (Fig. 3). However, the expression of miR172_365 was similar in all samples (Supplementary file 2).
Relative expression (RQ) of the miR172 family miRNA precursors. REV rejuvenated in vitro shoots, IVM typical mature in vitro shoots, IVMR mature in vitro shoots with signs of juvenility, MAT mature control, JUV juvenile control. Bars indicate minimum and maximum RQ values, samples labelled with different letters are statistically different (p < 0.05)
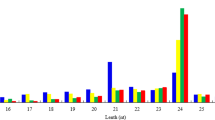
Two potential target genes of miR156 were analysed—BpSPL1 and BpSPL9. As expected, the highest expression of both target genes was observed in the mature controls (Supplementary file 3). However, the relative changes in expression were much higher for BpSPL1 compared to BpSPL9 (over 150-fold and fourfold, respectively, compared to the rejuvenated in vitro shoots). For BpSPL1, the lowest expression was observed in the juvenile control, and apart from the mature control, expression of BpSPL1 was highest in the in vitro sample that displayed typical signs of maturity (IVM) (Fig. 4). Expression of the BpSPL9 gene did not follow the expected patterns, as expression in the juvenile control was higher than all in vitro culture samples, and was lowest in the in vitro sample that displayed typical signs of maturity (IVM).
Relative expression (RQ) of the miR156 and miR172 family target genes BpSPL1 and BpAP2. REV rejuvenated in vitro shoots, IVM typical mature in vitro shoots, IVMR—mature in vitro shoots with signs of juvenility, MAT mature control, JUV juvenile control. Bars indicate minimum and maximum RQ values, samples labelled with different letters are statistically different (p < 0.05)
Three potential target genes of miR172 were analysed—BpAP2, BpRAP_488, BpRAP_288, which are all expected to be more highly expressed in juvenile tissues (Supplementary file 3). Changes in relative expression values were comparatively small—the largest was a less than fivefold increase of expression in the mature control for the BpRAP_488 gene (compared to rejuvenated in vitro shoots). Expression patterns of the BpAP2 gene were as expected, with the highest expression in the juvenile samples—REV, IVMR and JUV, while lower expression was observed in the mature samples—IVMR and MAT (Fig. 4). Expression patterns of the BpRAP_488 and BpRAP_288 genes were not as expected, with the highest relative expression levels observed in the IVM sample for BpRAP_288, and lower relative expression for the REV sample compared to the MAT control sample for BpRAP_488.
In summary, analysing the expression patterns of the precursor miRNAs and their target genes in tissue culture and control samples of differing juvenility, two miR156 precursors (miR156_511 and miR156_789) and their target gene BpSPLI as well as mir172_1931 and the target gene BpAP2 exhibited the expected expression patterns in the analysed samples and showed the best correlation with juvenility/maturity state. The expression of these miRNA precursors and their target genes was further investigated in samples incubated under differing culture conditions.
Effect of in vitro culture conditions on the expression of selected precursor miRNAs and target genes
Stabilized birch in vitro shoots were used to investigate the effect of subculture duration, growth media and sucrose content and different plant growth regulators on the expression of the selected precursor miRNAs and their target genes.
Three subculture durations were tested over a total period of 60 days—subculture every 10 days, every 30 days, and no subculturing. Samples were collected for RNA extraction at the end of the 60 day period, and relative expression values of the selected miRNA precursors and target genes determined (Figs. 5 and 6, Supplementary file 4). Expression of miR156_511 increased slightly with increasing time between subcultures. Expression of miR156_789 was almost equal in the samples subcultured every 10 days, and the samples not subcultured for 60 days, while in the samples subcultured every 30 days, expression levels were almost five times lower. Subculturing after 30 days is regarded as the optimum for rejuvenation of plant tissues, and expression of the miR156 precursors was expected to be highest in the 30 day subculture samples. However, expression of the miR156 target gene BpSPL1 was as expected, with similar levels in 10 day and 60 subculture samples, and two times lower in the 30 day samples. The expression of miR172_1931 was highest in the 10 day subculture samples, and approximately 20 times lower in the 30 and 60 day samples. There was not a corresponding change in expression of the target gene BpAP2, which was similar in 10 and 60 day samples, and two times lower in the 30 day samples.
Expression of these precursors and target genes was analysed on WPM supplemented with IAA, BAP, and twice the concentration of sucrose. In addition, samples cultured on MS media were analysed. The standard WPM was used as a reference sample for calculation of relative expression levels (Figs. 5 and 6, Supplementary file 5). For the miR156 precursors, the highest relative expression was found in samples cultured on the WPM reference and the MS media. On the standard WPM, miR156_511 had the highest expression, while miR156_789 was most highly expressed on MS media. Expression of miR156 precursors was more than two times lower in samples cultured on WPM supplemented with IAA, BAP, and sucrose, and relative expression values in these samples were similar to each other. However, expression of the BpSPL1 target gene was similar in all WPM samples. Expression of BpSPL1 was not detected in the sample cultured on MS media. Expression of miR172_1931 was highest in the WPM + IAA sample, and approximately ten times lower in the other samples. However, expression of the target gene BpAP2 was also highest in the WPM + IAA sample, followed by the MS media sample, and then the standard WPM sample.
The relative expression patterns of the selected miRNA precursors and target genes in samples cultured in differing conditions were not as well correlated with the expected patterns, in particular the expression directions of the precursor miRNAs and their corresponding target genes.
Discussion
Expression of miR156, miR72 and their target genes control plant development phase change and floral induction in both herbaceous and woody plants (Wu and Poethig 2006; Wu et al. 2009; Wang et al. 2011). The expression pattern of miR156 is consistent with its role in promoting juvenile development, as it is expressed at high levels in young seedlings and at lower levels in older plants. An analysis of miR156 levels in fully expanded leaves of species that undergo significant changes in leaf morphology during vegetative phase change demonstrated that juvenile leaves have significantly higher levels of miR156 than adult leaves (Wang et al. 2011), which is consistent with our results. The results from this study indicate that of seven miR156 miRNA precursors, six were expressed at high levels in young seedlings and at lower levels (or were undetected) in mature birch samples. The expression of miR156_511 and miR156_789 were similar to the expected expression patterns over all analysed tissues types, but miR156_237 had the highest expression levels in juvenile samples. miR156 expression decreases with plant age, and correspondingly the target gene SPL expression in juvenile plants is low and increases with age (Wu and Poethig 2006).
One of the ways in which miR156 represses flowering is through its effect on miR172, a miRNA that represses several AP2-like transcription factors that inhibit flowering. miR172 expression is low in juvenile plants, but increases with age (Wang et al. 2011). High levels of miR172 promote flowering by repressing the expression of these floral repressors. This higher expression of miR172 as plants mature is complementary to the decrease in miR156 expression, and is thought to be a consequence of this decrease because plants over-expressing miR156 have reduced levels of miR172, whereas plants with reduced levels of miR156 have elevated levels of miR172 (Wu et al. 2009; Jung et al. 2011). In addition to being directly repressed by miR156, SPL3 genes are transcriptionally repressed by the AP2-like proteins targeted by miR172, and are thus indirectly repressed by miR156 through its effect on miR172 expression. Our results indicate that a similar mechanism is present in silver birch shoots. According to the obtained results, the best correlation was shown by miR156_511, miR156_789 and their target gene BpSPL1. The largest difference in expression between juvenile, four-week-old shoot, and mature, 20-year-old birch samples was shown by BpSPL1, indicating that it may be a more accurate marker of juvenility than BpSPL9. BpSPL1, which is homologous to the SPL3 gene in Arabidopsis thaliana, is one of the major target genes for miR156 (Lännenpää et al. 2004; Gandikota et al. 2007). The increased (and expected) expression of BpSPL1, but not BpSPL9, in mature tissues found in this study is in concordance with results from avocado, macadamia and mango trees, where the SPL3/4/5 family genes were upregulated in mature tissues, but not the SPL9 genes (Ahsan et al. 2019b). Similar results were reported in Betula luminifera, where most of the miR156—targeted SPL genes, especially SLP1, SPL3 and SPL6, were dramatically up-regulated with age (Li et al. 2018). Mature miR156 was highly accumulated in younger seedlings (5- and 10-day-old), and dramatically down-regulated in older, 1.5-year-old and 4-year-old plants, suggesting important roles of these miR156-targeted SPL genes in vegetative phase change of Betula luminifera as well as in Betula pendula. SPL genes have been reported to be targeted by miR156 in many species, including Arabidopsis (Rhoades et al. 2002), rice (Xie et al. 2006), Populus (Li and Lu 2014), Citrus (Shalom et al. 2015) and Ziziphus jujube (Shao et al. 2017). However, differing SPLI expression patterns were reported in B. pendula (Lännenpää et al. 2004), where expression was even higher in juvenile shoots than in the reproductive stage leaves, and no differences between leaves at different distances from inflorescences were observed, and therefore the authors concluded that Betula pendula SPL1 is not involved in phase transition. However, our results indicated that BpSPL1 is highly expressed in mature samples in comparison to the juvenile and rejuvenated samples, therefore further investigation of expression of BpSPL1 in various tissues, developmental stages and genetic backgrounds is needed to resolve this issue. Results from this study indicated that expression of the BpAP2 gene, but not the RAP_488, RAP_288 genes, was higher in tissues with juvenile characteristics. However, the differences in expression levels were much smaller between mature and juvenile tissues for the BpAP2 gene, compared to the BpSPL1 gene. This is similar to results from avocado, macadamia and mango, where the expression of the AP2-like genes did not correspond to miR172 abundance or tree maturity (Ahsan et al. 2019b).
Expression patterns of some of the miRNA precursors were not as expected, or were similar in all analysed sample types. These unexpected expression patterns could be due to the differential expression of precursor miRNAs encoding identical mature miRNAs in different tissues or different biotic or abiotic conditions (Bielewicz et al. 2012). Further investigation of the expression patterns of these precursor miRNAs in a range of tissue types, growth conditions, and genetic backgrounds is needed to determine the function of these precursors in birch. In addition, the relative expression levels of the precursors where the expected expression patterns were observed may also differ in different tissues, growth conditions and genetic backgrounds.
Shoots, obtained from mature trees, during stabilization and after in vitro initiation, gradually rejuvenate, which is associated with the return of morphological, anatomical and physiological signs of juvenility (Mullins et al. 1979; Struve and Lineberger 1988; Brand and Lineberger 1992; Read and Bavougian 2012; McCown 2013). However, in this study, gene expression changes are less pronounced in in vitro culture than in juvenile plants, indicating that rejuvenation might only occur partially. This was also reported by several other molecular studies on in vitro plant rejuvenation (Li et al. 2012; Bastías et al. 2016; Sgamma et al. 2016; Jia et al. 2017). Rejuvenation, similarly to maturation, occurs in the apical meristem of the shoot. Culture environment influences the apical meristem where changes of gene expression occur, leading to rejuvenation (Sgamma et al. 2016). The expression of precursor miRNA and their target genes in stabilized in vitro shoots of birch cultured in the optimal WPM + zeatin medium with expression in samples cultured on WPM media supplemented with BAP was similar, with the samples cultured with BAP tending to have expression patterns more resembling mature tissues compared to the samples cultured with zeatin. The samples cultured on WPM + IAA had expression patterns most resembling mature tissues, particularly the expression of miR172_1931, and the target gene BpAP2. IAA is not optimal for long-term in vitro cultivation of birch, and the tendency towards maturation of samples cultivated on WPM + IAA media could be a reflection of this.
In vitro shoots, that are not formed de novo, are incompletely rejuvenated, and rejuvenation only persists while specific culture conditions are maintained (Bastías et al. 2016). Nitrogen and sucrose levels and plant growth regulators have a significant effect on plant juvenility in both intact plants (Hackett 1985; Liang et al. 2012; Yang et al. 2013) and in vitro plants (Franclet et al. 1987). The increased amount of nitrogen in the MS media promoted maturation, with miR156_511 expression lower than in the standard WPM media, and the expression of BpSPL1 in this variant was slightly increased. Rapid growth induced by increased nitrogen content accelerated the development of plants and promoted maturation and the transition to the reproductive phase, similar to the influence of growth-promoting factors in nature (Robinson and Wareing 1969). Interestingly, according to most physiological markers, these shoots appeared as juvenile (Girgžde unpublished results), however, molecular analysis indicated that they were maturing, suggesting that molecular markers may be more sensitive than physiological markers, or may indicate transition to maturity at an earlier stage than detected by physiological markers.
We expected that both subculturing every 10 days and no subculturing for 60 days would induce more rapid maturation of shoots and therefore lower miR156 expression compared to the standard 30 day subculturing. Subculturing every 10 days could increase cytokinin levels, while subculturing every 60 days could increase stress on plant tissues due to nutrient deficiencies. However, the expression patterns of the precursor miRNAs and their target genes were not consistent with this hypothesis. The expression of miR156_511 was lowest in the samples transferred every 10 days, but was highest in the samples not transferred for 60 days, while the expression of miR156_789 was highest in the 10 and 60 day samples, compared to the standard 30 day subcultured samples. However, these relative expression levels of the miRNAs did not correspond with the expression levels of the target genes, with BpSPL1 expression also being highest in the 10 and 60 day samples. The expression patterns of the BpSPL1 gene suggests that the standard 30 day subculture period is most effective at maintaining juvenility in in vitro culture. Subculturing every 10 days, caused plants to become vitrified/hyperhydric (results not shown), indicated by water-soaked shoots with a glassy appearance. This condition is mostly caused by an excess of ammonium and cytokinin (zeatin) (Ziv and Chen 2008). Hyperhydricity causes reduction in the gas exchange between cells and environment leading to hypoxic stress, and the increase of reactive oxygen species (Rojas-Martínez et al. 2010). Diler et al. (2016) observed that expression of miR156 was four fold lower in hyperhydric in vitro shoots, compared to normal shoots. However, Khraiwesh et al. (2012) reported that miR156 expression increased in a hypoxic environment.
miR156_511 and miR156_789 expression was decreased in WPM media with elevated sucrose concentrations, which is consistent with information on the inhibitory effect of sucrose on miR156 expression (Yang et al. 2013; Yu et al. 2013). However, no corresponding changes in BpSPL1 expression were observed, which suggests that the observed morphological, anatomical and physiological features, such as reduced growth and periderm formation are more related to the specific in vitro factor physiological reactions and are less closely related to maturation at the molecular level.
The most significant decrease in miR156 expression in shoots was induced by WPM media with 0.5 mg L−1 IAA. BpSPL1 gene expression was also increased in samples cultured on this media, suggesting that it may be one of the major contributors to in vitro maturation. Based on miR156 and BpSPL1 expression data, supplementation of media with 1.0 mg L−1 BAP also slightly contributed to the maturation of shoots. Excess auxin and cytokinin levels cause growth arrest and maturation of the stabilized birch in vitro shoots.
The relative expression patterns of the miRNA precursors in the samples with differing subculture duration and media composition were not as clear as in the samples with differing juvenility/maturity. The miRNA precursor and target gene expression was not as clearly in opposite directions, as was particularly observed for the miR156 precursors and the BpSP1 gene. This may be due to the smaller changes in relative expression in in vitro samples compared to the mature and juvenile controls. Further studies of the effect of tissue culture conditions on miRNA and target gene expression are needed, using a range of different genotypes, and analysing a larger number of miRNA precursors, as in vitro conditions may induce differential expression of genes and miRNA precursors compared to in vivo plant tissues.
Conclusions
This study identified that expression of two miR156 precursors (miR156_511 and miR156_789) and the target gene BpSPL1 were the best indicators of juvenility state in B. pendula. The miR172 precursor (miR172_1931) and target gene BpAP2 also had the expected expression patterns in juvenile, rejuvenated and mature birch samples, however, the relative differences in expression levels were smaller than for the miR156 precursors and BpSPL1. The obtained results indicate that changes in expression of BpSPL1 are associated with phase change in silver birch. Expression patterns of the miR156 precursors and BpSPL1 were in opposite directions as expected, particularly in the samples with differing juvenility/maturity, and these opposite expression patterns were most pronounced in the juvenile and mature control samples. The expression patterns of the miRNA precursors and target genes was not as clear cut in the samples cultured in differing conditions. However, the relative expression of the miRNA precursors was indicative of expected responses to different in vitro culture conditions. The molecular markers developed in this study can be used to assess juvenility of birch tissues, and provide a foundation for further studies of miRNA and target gene expression studies in relation to juvenility and in vitro culture conditions.
Data availability
All data generated or analysed during this study are included in this published article [and its supplementary information files].
References
Ahsan MU, Hayward A, Alam M et al (2019a) Scion control of miRNA abundance and tree maturity in grafted avocado. BMC Plant Biol 19:382. https://doi.org/10.1186/s12870-019-1994-5
Ahsan MU, Hayward A, Irihimovitch V et al (2019b) Juvenility and vegetative phase transition in tropical/subtropical tree crops. Front Plant Sci 10:729. https://doi.org/10.3389/fpls.2019.00729
Axtell MJ, Meyers BC (2018) Revisiting criteria for plant microRNA annotation in the era of big data. Plant Cell 30:272–284. https://doi.org/10.1105/TPC.17.00851
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297. https://doi.org/10.1016/S0092-8674(04)00045-5
Basheer-Salimia R (2007) Juvenility, maturity. and rejuvenation woody plants. Hebron Univ Res J A 3:17–43. https://doi.org/10.1186/s13007-018-0280-0
Bastías A, Almada R, Rojas P et al (2016) Aging gene pathway of microRNAs 156/157 and 172 is altered in juvenile and adult plants from in vitro propagated Prunus sp. Cienc e Investig Agrar 43:429–441. https://doi.org/10.4067/S0718-16202016000300009
Bergonzi S, Albani MC (2011) Reproductive competence from an annual and a perennial perspective. J Exp Bot 62:4415–4422. https://doi.org/10.1093/JXB/ERR192
Bielewicz D, Dolata J, Zielezinski A et al (2012) mirEX: a platform for comparative exploration of plant pri-miRNA expression data. Nucleic Acids Res 40:D191–D197. https://doi.org/10.1093/NAR/GKR878
Bonga JM (2016) Conifer clonal propagation in tree improvement programs. In: Park Y, Bonga J, Moon H (eds) Vegetative propagation of forest trees. National Institute of Forest Science (NiFos), Seoul, pp 3–31
Borges F, Martienssen RA (2015) The expanding world of small RNAs in plants. Nat Rev Mol Cell Biol 16:727–741. https://doi.org/10.1038/nrm4085
Brand MH, Lineberger RD (1992) In vitro rejuvenation of Betula (Betulaceae): morphological evaluation. Am J Bot 79:626–635. https://doi.org/10.1002/j.1537-2197.1992.tb14604.x
Carrington JC, Ambros V (2003) Role of microRNAs in plant and animal development. Science (80-) 301:336–338. https://doi.org/10.1126/SCIENCE.1085242
Díaz-Sala C (2016) Physiological, cellular, molecular and genomic analysis of the effect of maturation on propagation capacity. In: Park Y, Bonga JM, Moon H-K (eds) Vegetative propagation of forest trees. National Institute of Forest Science (NiFos), Seoul, pp 75–96
Diler E, Unver T, Karakülah G (2016) Differential expression of hyperhydricity responsive peach miRNAs. J Integr Bioinform 13:308. https://doi.org/10.2390/biecoll-jib-2016-308
Ewald D, Naujoks G, Welander M, et al (2001) Micropropagation and birch field trials. In: Proceedings of the workshop on high quality birch: clonal propagation and wood properties. Swedish University of Agricultural Sciences, Department of Crop Science, Ronneby, Sweden, pp 37–46
Feng S, Xu Y, Guo C et al (2016) Modulation of miR156 to identify traits associated with vegetative phase change in tobacco (Nicotiana tabacum). J Exp Bot 67:1493–1504. https://doi.org/10.1093/JXB/ERV551
Fernández-Fuego D, Keunen E, Cuypers A et al (2017) Mycorrhization protects Betula pubescens Ehr. from metal-induced oxidative stress increasing its tolerance to grow in an industrial polluted soil. J Hazard Mater 336:119–127. https://doi.org/10.1016/J.JHAZMAT.2017.04.065
Franclet A, Boulay M, Bekkaoui F, et al (1987) Rejuvenation. Springer, Dordrecht pp 232–248
Gailis A, Karklina A, Purvinš A et al (2020) Effect of breeding on income at first commercial thinning in silver birch plantations. Forests. https://doi.org/10.3390/f11030327
Gandikota M, Birkenbihl RP, Höhmann S et al (2007) The miRNA156/157 recognition element in the 3’ UTR of the Arabidopsis SBP box gene SPL3 prevents early flowering by translational inhibition in seedlings. Plant J 49:683–693. https://doi.org/10.1111/J.1365-313X.2006.02983.X
George EF, Hall MA, Klerk GJ (2008) Plant tissue culture procedure-background. Plant Propag Tissue Cult 1:1–28. https://doi.org/10.1007/978-1-4020-5005-3_1
Hackett WP (1985) Juvenility, maturation, and rejuvenation in woody plants. In: Janick J (ed) Horticultural reviews, vol 7. AVI Publishing Company, Connecticut, pp 109–155
Haffner V, Enjalric F, Lardet L et al (1991) Maturation of woody plants: a review of metabolic and genomic aspects. Ann Des Sci for INRA/EDP Sci 48:615–630. https://doi.org/10.1051/forest:19910601
Hynynen J, Niemistö P, Viherä-Aarnio A et al (2010) Silviculture of birch (Betula pendula Roth and Betula pubescens Ehrh.) in Northern Europe. Forestry 83:103–119. https://doi.org/10.1093/forestry/cpp035
Jain SM, Häggman H (2007) Protocols for micropropagation of woody trees and fruits. Protoc Micropropag Woody Trees Fruits. https://doi.org/10.1007/978-1-4020-6352-7
Jia XL, Chen YK, Xu XZ et al (2017) miR156 switches on vegetative phase change under the regulation of redox signals in apple seedlings. Sci Rep 71(7):1–13. https://doi.org/10.1038/s41598-017-14671-8
Jung JH, Seo PJ, Kang SK (2011) miR172 signals are incorporated into the miR156 signaling pathway at the SPL3/4/5 genes in Arabidopsis developmental transitions. Plant Mol Biol 761(76):35–45. https://doi.org/10.1007/S11103-011-9759-Z
Keinänen SI, Hassinen VH, Kärenlampi SO, Tervahauta AI (2007) Isolation of genes up-regulated by copper in a copper-tolerant birch (Betula pendula) clone. Tree Physiol 27:1243–1252. https://doi.org/10.1093/treephys/27.9.1243
Khraiwesh B, Zhu JK, Zhu J (2012) Role of miRNAs and siRNAs in biotic and abiotic stress responses of plants. Biochim Biophys Acta Gene Regul Mech 1819:137–148. https://doi.org/10.1016/j.bbagrm.2011.05.001
Kozomara A, Griffiths-Jones S (2011) miRBase: integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res. https://doi.org/10.1093/NAR/GKQ1027
Kozomara A, Griffiths-Jones S (2014) miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. https://doi.org/10.1093/NAR/GKT1181
Krivmane B, Šņepste I, Škipars V et al (2020) Identification and in silico characterization of novel and conserved microRNAs in methyl jasmonate-stimulated scots pine (Pinus sylvestris L.) needles. Forests 11:384. https://doi.org/10.3390/F11040384
Kulju KKM, Pekkinen M, Varvio S (2004) Twenty-three microsatellite primer pairs for Betula pendula (Betulaceae). Mol Ecol Notes 4:471–473. https://doi.org/10.1111/j.1471-8286.2004.00704.x
Lännenpää M, Jänönen I, Hölttä-Vuori M et al (2004) A new SBP-box gene BpSPL1 in silver birch (Betula pendula). Physiol Plant 120:491–500. https://doi.org/10.1111/j.0031-9317.2004.00254.x
Li C, Lu S (2014) Molecular characterization of the SPL gene family in Populus trichocarpa. BMC Plant Biol 14:1–15. https://doi.org/10.1186/1471-2229-14-131/FIGURES/10
Li H, Zhao X, Dai H et al (2012) Tissue culture responsive MicroRNAs in strawberry. Plant Mol Biol Report 30:1047–1054. https://doi.org/10.1007/S11105-011-0406-2
Li XY, Lin EP, Huang HH et al (2018) Molecular characterization of squamosa promoter binding protein-like (SPL) gene family in Betula luminifera. Front Plant Sci 9:608. https://doi.org/10.3389/FPLS.2018.00608/BIBTEX
Liang G, He H, Yu D (2012) Identification of nitrogen starvation-responsive microRNAs in Arabidopsis thaliana. PLoS ONE 7:e48951. https://doi.org/10.1371/JOURNAL.PONE.0048951
Lloyd G, McCown B (1980) Commercially-feasible micropropagation of mountain laurel Kalmia latifolia, by use of shoot-tip culture. CABI 30:421–427
Matsoukas IG, Massiah AJ, Thomas B (2013) Starch metabolism and antiflorigenic signals modulate the juvenile-to-adult phase transition in Arabidopsis. Plant Cell Environ 36:1802–1811. https://doi.org/10.1111/PCE.12088
McCown BH (2000) Special symposium: in vitro plant recalcitrance recalcitrance of woody and herbaceous perennial plants: dealing with genetic predeterminism. Vitr Cell Dev Biol Plant 363(36):149–154. https://doi.org/10.1007/S11627-000-0030-6
McCown BH (2013) Woody shrubs and trees. In: Smith RH (ed) Plant tissue culture. Techniques and experiments, 3rd edn. Elsevier, San Diego, pp 93–101
Mullins MG, Nair Y, Sampet P (1979) Rejuvenation in vitro: induction of juvenile characters in an adult clone of Vitis vinifera L. Ann Bot 44:623–627
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15:473–497. https://doi.org/10.1111/J.1399-3054.1962.TB08052.X
Neutelings G, Fénart S, Lucau-Danila A, Hawkins S (2012) Identification and characterization of miRNAs and their potential targets in flax. J Plant Physiol 169:1754–1766. https://doi.org/10.1016/j.jplph.2012.06.011
O’Dowd N (2004) The improvement of Irish birch. Phase 1: selection of individuals and populations. COFORD, Dublin
Orkwiszewski JAJ, Poethig RS (2000) Phase identity of the maize leaf is determined after leaf initiation. Proc Natl Acad Sci USA 97:10631–10636. https://doi.org/10.1073/PNAS.180301597
Poethig RS (2013) Vegetative phase change and shoot maturation in plants. Curr Top Dev Biol 105:125–152. https://doi.org/10.1016/B978-0-12-396968-2.00005-1
Preece J (2008) Stock plant physiological factors affecting growth and morphogenesis. In: George E, Hall M, De Klerk J (eds) Plant propagation by tissue culture, 3rd edn. Springer, Dordrecht, pp 403–422
Read PE, Bavougian CM (2012) In vitro rejuvenation of woody species. Humana Press, Totowa pp 383–395
Renau-Morata B, Ollero J, Arrillaga I, Segura J (2005) Factors influencing axillary shoot proliferation and adventitious budding in cedar. Tree Physiol 25:477–486. https://doi.org/10.1093/TREEPHYS/25.4.477
Rhoades MW, Reinhart BJ, Lim LP et al (2002) Prediction of plant microRNA targets. Cell 110:513–520. https://doi.org/10.1016/S0092-8674(02)00863-2
Robinson LW, Wareing PF (1969) Experiments on the juvenile-adult phase change in some woody species. New Phytol 68:67–78. https://doi.org/10.1111/J.1469-8137.1969.TB06420.X
Rojas-Martínez L, Visser RGF, de Klerk GJ (2010) The hyperhydricity syndrome: waterlogging of plant tissues as a major cause. Propag Ornam Plants 10:169–175
Rubio-Piña JA, Zapata-Pérez O (2011) Isolation of total RNA from tissues rich in polyphenols and polysaccharides of mangrove plants. Electron J Biotechnol. https://doi.org/10.2225/VOL14-ISSUE5-FULLTEXT-10
Ruonala R, Rinne PLH, Baghour M et al (2006) Transitions in the functioning of the shoot apical meristem in birch (Betula pendula) involve ethylene. Plant J 46:628–640. https://doi.org/10.1111/J.1365-313X.2006.02722.X
Sánchez MC, Ballester A, Vieitez AM (1997) Reinvigoration treatments for the micropropagation of mature chestnut trees. Ann Des Sci for 54:359–370. https://doi.org/10.1051/FOREST:19970404
Sgamma T, Cirilli M, Caboni E et al (2016) In vitro plant culture system induces phase transition in fruit-bearing plants. Acta Hortic 1110:13–20. https://doi.org/10.17660/ACTAHORTIC.2016.1110.3
Shalom L, Shlizerman L, Zur N et al (2015) Molecular characterization of SQUAMOSA PROMOTER BINDING PROTEIN-LIKE (SPL) gene family from Citrus and the effect of fruit load on their expression. Front Plant Sci 6:1–10. https://doi.org/10.3389/FPLS.2015.00389/ABSTRACT
Shao F, Lu Q, Wilson IW, Qiu D (2017) Genome-wide identification and characterization of the SPL gene family in Ziziphus jujuba. Gene 627:315–321. https://doi.org/10.1016/J.GENE.2017.06.044
Struve DK, Lineberger RD (1988) Restoration of high adventitious root regeneration potential in mature Betula papyrifera Marsh, softwood stem cuttings. Can J for Res 18:265–269. https://doi.org/10.1139/x88-038
Vidal EA, Moyano TC, Canales J, Gutiérrez RA (2014) Nitrogen control of developmental phase transitions in Arabidopsis thaliana. J Exp Bot 65:5611–5618. https://doi.org/10.1093/JXB/ERU326
Vinoth A, Ravindhran R (2018) In vitro morphogenesis of woody plants using thidiazuron. Thidiazuron Urea Deriv Plant Growth Regul. https://doi.org/10.1007/978-981-10-8004-3_10
Wang JW, Park MY, Wang LJ et al (2011) MiRNA control of vegetative phase change in trees. PLOS Genet 7:e1002012. https://doi.org/10.1371/JOURNAL.PGEN.1002012
Wang H, Jiao X, Kong X et al (2016) A signaling cascade from miR444 to RDR1 in rice antiviral RNA silencing pathway. Plant Physiol 170:2365–2377. https://doi.org/10.1104/PP.15.01283
Welander M (1993) Micropropagation of birch. 223–246. https://doi.org/10.1007/978-94-015-8116-5_14
Wu G, Poethig RS (2006) Temporal regulation of shoot development in Arabidopsis thaliana by miR156 and its target SPL3. Development 133:3539–3547. https://doi.org/10.1242/DEV.02521
Wu G, Park MY, Conway SR et al (2009) The sequential action of miR156 and miR172 regulates developmental timing in Arabidopsis. Cell 138:750–759. https://doi.org/10.1016/J.CELL.2009.06.031
Xie K, Wu C, Xiong L (2006) Genomic organization, differential expression, and interaction of SQUAMOSA promoter-binding-like transcription factors and microRNA156 in rice. Plant Physiol 142:280–293. https://doi.org/10.1104/PP.106.084475
Xing S, Salinas M, Höhmann S et al (2010) miR156-targeted and nontargeted SBP-box transcription factors act in concert to secure male fertility in Arabidopsis. Plant Cell 22:3935–3950. https://doi.org/10.1105/TPC.110.079343
Xing L, Zhang D, Li Y et al (2014) Genome-wide identification of vegetative phase transition-associated microRNAs and target predictions using degradome sequencing in Malus hupehensis. BMC Genomics 15:1–22. https://doi.org/10.1186/1471-2164-15-1125/FIGURES/11
Xu Y, Guo C, Zhou B et al (2016) Regulation of vegetative phase change by SWI2/SNF2 chromatin remodeling ATPase BRAHMA. Plant Physiol 172:2416. https://doi.org/10.1104/PP.16.01588
Yang L, Xu M, Koo Y et al (2013) Sugar promotes vegetative phase change in Arabidopsis thaliana by repressing the expression of MIR156A and MIR156C. Elife. https://doi.org/10.7554/ELIFE.00260
Yu S, Li C, Zhou CM et al (2013) Sugar is an endogenous cue for juvenile-to-adult phase transition in plants. Elife. https://doi.org/10.7554/ELIFE.00269
Zeltinš P, Matisons R, Gailis A et al (2018) Genetic parameters of growth traits and stem quality of silver birch in a low-density clonal plantation. Forests 9:1–8. https://doi.org/10.3390/f9020052
Zhang B, Pan X, Wang Q et al (2006) Computational identification of microRNAs and their targets. Comput Biol Chem 30:395–407. https://doi.org/10.1016/J.COMPBIOLCHEM.2006.08.006
Zhang L, Hu YB, Wang H et al (2015) Involvement of miR156 in the regulation of vegetative phase change in plants. J Am Soc Hortic Sci 140:387–395. https://doi.org/10.21273/JASHS.140.5.387
Žiarovská J, Labajová M, Ražná K et al (2013) Changes in expression of BetV1 allergen of silver birch pollen in urbanized area of Ukraine. J Environ Sci Health A 48:1479–1484. https://doi.org/10.1080/10934529.2013.796788
Zimmerman RH, Hackett WP, Pharis RP (1985) Hormonal aspects of phase change and precocious flowering. Horm Regul Dev III:79–115. https://doi.org/10.1007/978-3-642-67734-2_4
Ziv M, Chen J (2008) The anatomy and morphology of tissue cultured plants. In: George EF, Hall MA, De Klerk G (eds) Plant propagation by tissue culture, 3rd edn. Springer, Wageningen
Zuker M (2003) Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res 31:3406–3415. https://doi.org/10.1093/NAR/GKG595
Acknowledgements
This project was funded by the Latvian Council of Science grant No. lzp-2019/1-0387 “Development of molecular markers for assessment of juvenility during micropropagation of silver birch (Betula pendula Roth)”.
Funding
This project was funded by the Latvian Council of Science Grant No. lzp-2019/1-0387 “Development of molecular markers for assessment of juvenility during micropropagation of silver birch (Betula pendula Roth)”.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and laboratory analyses were performed by BK, EG and IS. Data analyses were performed by BK and DR. The first draft of the manuscript was written by BK and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Communicated by Melekşen Akın.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Krivmane, B., Girgžde, E., Samsone, I. et al. Expression of juvenility related microRNAs and target genes during micropropagation of silver birch (Betula pendula Roth.). Plant Cell Tiss Organ Cult 152, 455–469 (2023). https://doi.org/10.1007/s11240-022-02419-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-022-02419-w