Abstract
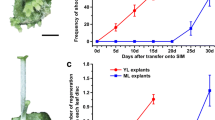
Arabidopsis plant that is doubly mutant for the de novo methylases DRM1 and DRM2 (drm1/drm2) produce larger and heavier callus than wild type (WT) plants when cultivated on callus induce medium (CIM) for a given period of time. This trend is observed both in seedlings and explants, which indicates that both DRM1 and DRM2 are involved in callus formation. Furthermore, the gene expression analysis revealed that the BBM expression was continuously increased with CIM cultivation, and the expression level was significantly higher in the drm1/drm2 mutants than WT. And the anti-stress response genes have altered in the drm1/drm2 mutants compared with WT, These data suggest that DNA methylation and corresponding genes play an important role in callus formation .







Similar content being viewed by others
References
Atta R et al (2009) Pluripotency of Arabidopsis xylem pericycle underlies shoot regeneration from root and hypocotyl explants grown in vitro. Plant J 57:626–644. doi:10.1111/j.1365-313X.2008.03715.x
Banno H, Ikeda Y, Niu QW, Chua NH (2001) Overexpression of Arabidopsis ESR1 induces initiation of shoot regeneration. Plant Cell 13:2609–2618
Beeckman T, De Smet I (2014) Pericycle. Curr Biol 24:R378–R379. doi:10.1016/j.cub.2014.03.031
Berdasco M et al (2008) Promoter DNA hypermethylation and gene repression in undifferentiated Arabidopsis cells. PLoS ONE 3:e3306. doi:10.1371/journal.pone.0003306
Boutilier K et al (2002) Ectopic expression of BABY BOOM triggers a conversion from vegetative to embryonic growth. Plant Cell 14:1737–1749
Cao X, Jacobsen SE (2002) Role of the arabidopsis DRM methyltransferases in de novo DNA methylation and gene silencing. Curr Biol 12:1138–1144
Chan SW, Henderson IR, Jacobsen SE (2005) Gardening the genome: DNA methylation in Arabidopsis thaliana. Nat Rev Genet 6:351–360. doi:10.1038/nrg1601
Chanvivattana Y, Bishopp A, Schubert D, Stock C, Moon YH, Sung ZR, Goodrich J (2004) Interaction of Polycomb-group proteins controlling flowering in Arabidopsis. Development 131:5263–5276. doi:10.1242/dev.01400
Desvoyes B, Sanchez MP, Ramirez-Parra E, Gutierrez C (2010) Impact of nucleosome dynamics and histone modifications on cell proliferation during Arabidopsis development. Heredity 105:80–91. doi:10.1038/hdy.2010.50
Dubrovsky JG, Doerner PW, Colon-Carmona A, Rost TL (2000) Pericycle cell proliferation and lateral root initiation in Arabidopsis. Plant Physiol 124:1648–1657
Fan M, Xu C, Xu K, Hu Y (2012) LATERAL ORGAN BOUNDARIES DOMAIN transcription factors direct callus formation in Arabidopsis regeneration. Cell Res 22:1169–1180. doi:10.1038/cr.2012.63
Feng Z, Zhu J, Du X, Cui X (2012) Effects of three auxin-inducible LBD members on lateral root formation in Arabidopsis thaliana. Planta 236:1227–1237. doi:10.1007/s00425-012-1673-3
Grafi G, Ben-Meir H, Avivi Y, Moshe M, Dahan Y, Zemach A (2007) Histone methylation controls telomerase-independent telomere lengthening in cells undergoing dedifferentiation. Dev Biol 306:838–846. doi:10.1016/j.ydbio.2007.03.023
Grafi G, Chalifa-Caspi V, Nagar T, Plaschkes I, Barak S, Ransbotyn V (2011) Plant response to stress meets dedifferentiation. Planta 233:433–438. doi:10.1007/s00425-011-1366-3
Heidmann I, de Lange B, Lambalk J, Angenent GC, Boutilier K (2011) Efficient sweet pepper transformation mediated by the BABY BOOM transcription factor. Plant Cell Rep 30:1107–1115. doi:10.1007/s00299-011-1018-x
Ikeuchi M, Sugimoto K, Iwase A (2013) Plant callus: mechanisms of induction and repression. Plant Cell 25:3159–3173. doi:10.1105/tpc.113.116053
Iwase A et al (2011) The AP2/ERF transcription factor WIND1 controls cell dedifferentiation in Arabidopsis. Curr bio 21:508–514. doi:10.1016/j.cub.2011.02.020
Knoester M, Pieterse CM, Bol JF, Van Loon LC (1999) Systemic resistance in Arabidopsis induced by rhizobacteria requires ethylene-dependent signaling at the site of application. Mol Plant Microbe Interact 12:720–727. doi:10.1094/MPMI.1999.12.8.720
Li H, Freeling M, Lisch D (2010) Epigenetic reprogramming during vegetative phase change in maize. Proc Nati Acad Sci USA 107:22184–22189. doi:10.1073/pnas.1016884108
Li W, Liu H, Cheng ZJ, Su YH, Han HN, Zhang Y, Zhang XS (2011) DNA methylation and histone modifications regulate de novo shoot regeneration in Arabidopsis by modulating WUSCHEL expression and auxin signaling. PLoS Genet 7:e1002243. doi:10.1371/journal.pgen.1002243
Liu J, Sheng L, Xu Y, Li J, Yang Z, Huang H, Xu L (2014) WOX11 and 12 are involved in the first-step cell fate transition during de novo root organogenesis in Arabidopsis. Plant Cell 26:1081–1093. doi:10.1105/tpc.114.122887
Lucas M, Guedon Y, Jay-Allemand C, Godin C, Laplaze L (2008) An auxin transport-based model of root branching in Arabidopsis thaliana. PLoS ONE 3:e3673. doi:10.1371/journal.pone.0003673
Marsch-Martinez N et al (2006) BOLITA, an Arabidopsis AP2/ERF-like transcription factor that affects cell expansion and proliferation/differentiation pathways. Plant mol bio 62:825–843. doi:10.1007/s11103-006-9059-1
Mikami T, Kinoshita T (1988) Genotypic effects on the callus formation from different explants of rice, Oryza sativa L. Plant Cell Tissue Organ Cult 1988, 12(3): 311−314.doi: 10.1007/BF00034374
Okushima Y et al (2005) Functional genomic analysis of the AUXIN RESPONSE FACTOR gene family members in Arabidopsis thaliana: unique and overlapping functions of ARF7 and ARF19. Plant Cell 17:444–463. doi:10.1105/tpc.104.028316
Prifti E, Zucker JD, Clement K, Henegar C (2008) FunNet: an integrative tool for exploring transcriptional interactions. Bioinformatics 24:2636–2638. doi:10.1093/bioinformatics/btn492
Sarkar AK et al (2007) Conserved factors regulate signalling in Arabidopsis thaliana shoot and root stem cell organizers. Nature 446:811–814. doi:10.1038/nature05703
Sen GL, Reuter JA, Webster DE, Zhu L, Khavari PA (2010) DNMT1 maintains progenitor function in self-renewing somatic tissue. Nature 463:563–567. doi:10.1038/nature08683
Srinivasan C et al (2007) Heterologous expression of the BABY BOOM AP2/ERF transcription factor enhances the regeneration capacity of tobacco (Nicotiana tabacum L.). Planta 225:341–351. doi:10.1007/s00425-006-0358-1
Sugimoto K, Gordon SP, Meyerowitz EM (2011) Regeneration in plants and animals: dedifferentiation, transdifferentiation, or just differentiation? Trends Cell Biol 21:212–218. doi:10.1016/j.tcb.2010.12.004
Te-chato S, Lim M (1999) Plant regeneration of mangosteen via nodular callus formation. Plant Cell Tissue Organ Cult 59:89–93. doi:10.1023/A:1006482513727
Tessadori F et al (2007) Large-scale dissociation and sequential reassembly of pericentric heterochromatin in dedifferentiated Arabidopsis cells. J Cell Sci 120:1200–1208. doi:10.1242/jcs.000026
Acknowledgments
This paper was supported by Natural Science Foundation of China (Grant No. 31370214).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
All of the authors have no conflicts of interests.
Additional information
Fangwei Jiang and Xiaofeng Xu have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jiang, F., Xu, X., Liu, H. et al. DRM1 and DRM2 are involved in Arabidopsis callus formation. Plant Cell Tiss Organ Cult 123, 221–228 (2015). https://doi.org/10.1007/s11240-015-0812-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-015-0812-5