Abstract
Parasitological assessment of marine fishes at Sodwana Bay in the iSimangaliso Marine Protected Area on the east coast of South Africa revealed a new species of cryptogonimid trematode infecting the pyloric caeca of the Dory Snapper, Lutjanus fulviflamma (Forsskål) (Lutjanidae). The new species is morphologically consistent with the concept of the large genus Siphoderina Manter, 1934; its phylogenetic position within this genus was validated through molecular sequencing of the ITS2 and partial 28S ribosomal DNA sub-regions. We name this species Siphoderina nana n. sp. and comment on the current state of understanding for this genus of cryptogonimids.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Research on the fish helminth fauna of South Africa throughout most of the twentieth century has been patchy, with studies being few and far between (Fantham, 1938; Prudhoe, 1956). In the 1970s and 1980s, key work by the likes of Bray (1974, 1978, 1984, 1985, 1987), Gibson (1983) and Gavrilyuk-Tkachuk and Nikolaeva (Gavrilyuk-Tkachuk, 1979; Nikolaeva & Tkachuk, 1979) greatly expanded knowledge on the helminth fauna of South Africa, but research, especially from within South Africa, continued to be relatively rare. As of 2022, just over 70 species of trematode were known from South African waters; this number compares unfavourably with regions of similar latitudes, such as the south coast of Australia (190 recorded species) and subtropical eastern China (529 recorded species) (Bray et al., 2016; Cribb et al., 2016). Most recent aquatic helminthological work in the country has primarily focused on trematodes in freshwater systems (Dos Santos et al., 2021; Hoogendoorn et al., 2020; Malatji & Mukaratirwa, 2019; Rakgole et al., 2019), and then mostly on intermediate stages, with a minority of studies concerning trematodes in marine systems (Huston et al., 2021; Vermaak et al., 2021). Just two new species of fish-infecting trematodes have been described from South Africa in the twenty-first century, one from freshwater (Dos Santos et al., 2021) and one from the marine environment (Vermaak et al., 2023).
Parasitological assessment of fishes from Sodwana Bay on the east Coast of South Africa yielded samples of several trematode species, including a species of cryptogonimid from the Dory Snapper, Lutjanus fulviflamma (Forsskål) (Lutjanidae). The Cryptogonimidae Ward, 1917, is a large trematode family with 79 recognised genera and over 280 recognised species (WoRMS, 2023), which, as sexual adults, infect a wide range of (primarily demersal, predatory) freshwater and marine fishes, crocodilians and some snakes and amphibians (Martin et al., 2023; Miller & Cribb, 2008b). Despite their widespread distribution and expansive host range, this is the first cryptogonimid species to be reported from South Africa.
Methods
Dory Snapper (L. fulviflamma) were collected by rod and reel from off Jesser Point, Sodwana Bay in the iSimangaliso Marine Protected Area National Park, South Africa. Following capture, fish were transported in aerated water from the collection site to the nearby field station where they were humanely killed by percussive stunning followed by cranial pithing, and dissected per the protocols of Cribb & Bray (2010). Any worms found were killed and heat-fixed in near-boiling saline, and preserved in 70% ethanol. Specimens for morphological analysis were prepared using standard protocols for Mayer’s haematoxylin preparations, per Yong et al. (2016). All measurements were made using an Olympus SC50 digital microscope camera, mounted on an Olympus BX-53 light microscope and cellSens™ version 1.13 imaging software. All measurements, unless otherwise stated, are in micrometres (μm) and given as a range with the mean in parentheses. Drawings were made with the aid of a camera lucida attachment mounted on the same Olympus BX-53 light microscope and digitised in Adobe Illustrator 6.0. Specimens for scanning electron microscopy were prepared by chemical dehydration, first in a graded ethanol series followed by a graded hexamethyldisilazane (HMDS) series, then sputter-coated with gold and photographed using a Phenom Pro Desktop scanning electron microscope (ThermoScientific, Waltham, Massachusetts, USA). Type-specimens have been deposited in the National Museum Parasite Collection of the National Museum, Bloemfontein, South Africa (NMB).
Genetic sequence data were generated from the novel material for cytochrome c oxidase subunit 1 mitochondrial marker (cox1 mtDNA), the non-coding second internal transcribed spacer unit of ribosomal DNA (ITS2 rDNA), and the large ribosomal subunit gene (28S rDNA). Genomic DNA was extracted using a DNeasy blood and tissue extraction kit (QIAGEN, Hilden, Germany) as per the manufacturer’s instructions, although extractions were eluted once only with 60 µl (vs recommend once to twice with 200 µl). The three target marker regions were amplified by conventional PCR, with 25 µl reaction volumes, comprising 2 µl unquantified extracted DNA, 1 µl per primer at 10 µM, 6 µl cresol-red visualisation dye, 9.3 µl purified injection water, 1 µl dNTPs at 4 mM, 2 µl MgCl2 at 25 mM and 0.2 µl Taq. DNA polymerase at 5.5 units/µl with 2.5 µl 10× reaction buffer (Fisher Biotec Australia), using the primers and cycle schedules provided in Yong et al. (2016) for rDNA and Wee et al. (2017) for mtDNA. These protocols yield a partial read for cox1 mtDNA, a full read for ITS2 rDNA with partial flanking 5.8S and 28S, and a partial read for 28S rDNA including the variable domains D1–D3.
Amplicons were visualised via electrophoresis using 1.5% agarose gels stained with SYBR Safe (Invitrogen, California, USA) and purified using Agencourt AMPure magnetic bead purification system (Beckman Coulter, California, USA). Sanger sequencing was completed by the Western Australian State Agricultural Biotechnology Centre (SABC) at Murdoch University, using an ABI Prism™ BigDye v3.1 Cycle Sequencing Kit (Applied Biosystems, California, USA) and an ABI 3730 96 capillary machine. Forward and reverse DNA strands were sequenced using the amplification primers for ITS2 rDNA and cox1 mtDNA, whereas the internal primers 300F and 1200R were used for sequencing the 28S rDNA marker; these primers are also detailed in Yong et al. (2016). Contiguous sequences were assembled, in Geneious™ version 10.2.2 (Kearse et al., 2012). The ITS2 region was determined by annotation through the ITS2 Database using the ‘Metazoa’ model (Ankenbrand et al., 2015).
Novel ITS2 and partial 28S rDNA sequences were aligned and analysed against comparable sequence data for other cryptogonimid taxa available on GenBank (see Table 1). As verified cox1 mtDNA sequence data are only publicly available for five cryptogonimid taxa, no alignment or analysis of the cox1 region was performed. Analyses of the ITS2 rDNA dataset were carried out using only sequences for species of Cryptogonimidae to verify the generic affiliation of our new taxon. Selected species of Heterophyidae Leiper, 1909 + Opisthorchiidae Looss, 1899 were used as outgroup taxa in alignments of the partial 28S rDNA dataset, as these are the closest families to the Cryptogonimidae, as indicated by large-scale molecular phylogenetic analyses by Olson et al. (2003) and Pérez-Ponce de León & Hernández-Mena (2019). Both alignments were performed using MUSCLE 3.7 (Edgar, 2004) with ClustalW sequence weighting and UPGMA clustering for iterations 1 and 2. The resultant alignments were refined by eye using MESQUITE (Maddison & Maddison, 2017) with the ends trimmed to a length matching over 75% of sequences per Yong et al. (2021) and indels greater than three consecutive characters and affecting more than 5% of sequences removed.
The ITS2 and partial 28S rDNA alignments were subjected to Bayesian inference and maximum likelihood phylogenetic analyses to explore affinities of the novel material using MrBayes v3.2.2 (Ronquist et al., 2012) and RAxML v7.2.8 (Stamatakis, 2014), respectively, via the CIPRES portal (Miller et al., 2010a). The GTR+I+Γ model of nucleotide substitution evolution was assumed in analyses for both datasets, based on the Akaike Information Criterion (AIC) in jModelTest v2.1.10 (Darriba et al., 2012). Each Bayesian inference analysis was run over 10,000,000 generations (ngen = 10,000,000) with two runs each containing four simultaneous Markov Chain Monte Carlo (MCMC) chains (nchains = 4) and every 1,000th tree saved (samplefreq = 1000). Bayesian analyses used the following parameters: nst = 6, rates = invgamma, ngammacat = 4, and the priors parameters of the combined dataset were set to ratepr = variable. Samples of substitution model parameters, and tree and branch lengths were summarised using the parameters ‘sump burnin = 3000’ and ‘sumt burnin = 3000’. Maximum likelihood analyses ran 100 bootstrap pseudoreplicates for both datasets.
Results
Molecular phylogenetic analyses
Alignment of the novel ITS2 rDNA sequence data with those of other cryptogonimid taxa available on GenBank yielded 491 characters for analysis. Bayesian inference and maximum likelihood analyses of the ITS2 rDNA alignment produced trees of essentially identical topologies, with lower nodal support in the latter; these trees are not presented. Alignment of the novel partial 28S rDNA sequence data with those of other cryptogonimid taxa available on GenBank yielded 861 characters for comparative analysis. Bayesian inference and maximum likelihood analyses of the partial 28S alignment produced trees with near-identical topologies (Fig. 1).
Molecular phylogeny of the Cryptogonimidae, including Siphoderina nana n. sp. and representatives of other genera for which molecular data were available, based on Bayesian inference and maximum likelihood analyses of the partial 28S rDNA dataset generated by this study (presented topology based on the Bayesian inference analysis), with posterior probabilities from the Bayesian inference analysis shown above the nodes and bootstrap support values from the maximum likelihood analysis below. Nodal support of less than 75 has been excluded.
In both ITS2 and partial 28S analyses, the novel sequences formed a clade with those of species of Siphoderina Manter, 1934, with high nodal support. Our trees largely resemble those for the family produced by Martin & Cutmore (2022) and Miller et al. (2018), with the clade formed by species of Siphoderina sister to that formed by species of Latuterus Miller & Cribb, 2007, but with disagreement regarding the position of Caulanus thomasi Miller & Cribb, 2007. In the ITS2 tree, our new species formed a clade with Siphoderina hirastricta (Manter, 1963), sister to Siphoderina poulini Miller & Cribb, 2008, but with low confidence at all nodes. In the partial 28S tree, our new species was basal to a clade formed by Siphoderina territans Miller & Cribb, 2008 + S. virga Miller & Cribb, 2008 + S. hirastricta + S. grunnitus Miller & Cribb, 2008.
Taxonomy
Cryptogonimidae Ward, 1917
Siphoderina Manter, 1934
Type-species: Siphoderina brotulae Manter, 1934, by original designation
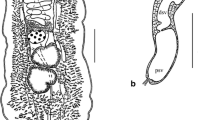
Siphoderina nana n. sp. Fig. 2A–D
Siphoderina nana n. sp. A Whole-body mount, ventral view of holotype specimen XXX; B–D scanning electron micrographs of the B oral sucker, C tegumental spines; and D ventrogenital sac opening. C caecum, CSR canalicular seminal receptacle, EP excretory pore, EV excretory vesicle, GP genital pore (opens internally within ventrogenital sac), OG oesophageal glands, Oo oötype, OP optical pigment, OS oral sucker, Ph pharynx, SV seminal vesicle, Tt testis, Ut uterus, Vit vitelline follicles, VS ventral sucker (within ventrogenital sac). Scale-bars: A 150 µm, B 25 µm, C 10 µm, D 15 µm.
Type-host: Lutjanus fulviflamma (Forsskål) (Lutjaniformes: Lutjanidae), Dory Snapper.
Infection site: Pyloric caeca.
Type-locality: Jesser Point, Sodwana Bay, (27° 32′ 19′′ S, 32° 40′ 48′′ E), iSimangaliso Marine Protected Area, KwaZulu-Natal Province, South Africa.
Prevalence and intensity: Three of nine L. fulviflamma infected with 3–30 worms.
Material examined: Holotype (NMB P 970) and 21 paratypes (NMB P 971–990), all slide-mounted.
Representative DNA sequences: Three identical replicates of ITS2 rDNA, 466 nucleotides, one submitted to GenBank (OR593670); two identical replicates of partial 28S rDNA, 1,121 nucleotides, one submitted to GenBank (OR593669); three sequences of cox1 mtDNA comprising two genotypes varying at 1 of 491 nucleotides (OR590623 representative of two replicates, and OR590624 representative of one replicate).
ZooBank registration: The species Siphoderina nana is registered in ZooBank under urn:lsid:zoobank.org:act:C29020A4-6535-4F31-B4A1-B623AC1FE964.
Etymology: The specific name nana is a feminine Latin noun, itself derived from the Ancient Greek “nânos”, meaning “dwarf”, as it is the smallest known species of the genus Siphoderina.
Description
[Based on 22 whole-mounted, unflattened specimens]. Body usually sub-globular, occasionally pyriform when oral sucker extended, 385–692 × 239–458 (547 × 357), 1.2–1.8 (1.5) times longer than wide. Tegumental spines flattened, broadly lanceolate, with serrated edges, arranged in the manner of overlapping scales, evenly covering whole of body except oral sucker, 5 long (Fig. 2B). Forebody 140–254 (182) long, represents 27.7–38.3% (33.6%) of body length. Remnant eyespot pigment present in some specimens; dispersed in forebody. Oral sucker infundibuliform, 70–126 × 103–161 (98 × 128), 0.5–1.2 (0.8) times longer than wide. Oral spines 60–72 (67), straight-tipped, ringing periphery of oral sucker, 14–16 (15) long (Fig. 2C). Ventral sucker distinctly smaller than oral sucker, spherical, set medially on ventral body surface within ventrogenital sac [per Miller & Cribb (2008a)], 41–61 × 42–60 (51 × 52), 0.8–1.2 (1.0) times longer than wide. Ventrogenital sac opens medially; aperture 10–22 (15) across, circular; gonotyl or other distinct tegumental folding absent (Fig. 2D). Prepharynx 11–40 (26). Pharynx ellipsoidal, similarly sized to ventral sucker, 39–60 × 36–47 (48 × 41), 1.0–1.5 (1.2) times longer than wide. Oesophagus straight, 11–32 (19). Gland cells, probably associated with mouth or pharynx, profuse, pyriform, centred on oesophagus, distributed throughout forebody to level of caecal bifurcation. Intestinal bifurcation medial, in space between pharynx and ventral sucker. Intestinal caeca blind-ended, straight, extremities often greatly expanded, pass parallel to lateral body margins, 220–503 (366) long, span 52.0–80.0% (66.6%) of total body length, terminate 35–115 (68) from posterior extremity of body; post-caecal zone 14–88 (44) or 3.5–18.4% (8.3%) of total body length.
Testes two, spherical to sub-spherical, opposite to slightly oblique, in mid-hindbody, left testis 93–248 × 93–209 (152 × 151); right testis 77–242 × 108–208 (155 × 151); pre-testicular zone 201–368 (292) or 45.6–61.5% (51.7%) of total body length; post-testicular zone 59–202 (127) or 13.1–29.3% (22.8%) of total body length. Seminal vesicle medial, saccular, usually deflexed, highly contorted in some specimens, 77–194 × 24–75 (118 × 47). Ejaculatory duct passes dorsal to ventral sucker, path almost entirely obscured by uterine eggs. Genital atrium obscured by ventral sucker, varying from almost indiscernible to very short, simple. Genital pore medial, opens antero-dorsal to ventral sucker within ventro-genital sac, 148–246 (192) or 29.5–41.3% (35.4%) of total body length from anterior extremity of body and 214–474 (340) or 55.6–68.8% (61.8%) from posterior extremity.
Ovary an irregularly-shaped, highly lobulate mass, asymmetrically medial in anterior hindbody between ventral sucker and testes, posteriorly overlaps anterior margins of testes, 75–219 × 67–237 (117 × 158). Canalicular seminal receptacle sub-medial, oval to oblate trapezoid, antero-dorsal to ovary, size and volume varying with sperm content, 34–120 × 54–155 (61 × 82). Oötype not discernible in most specimens, medial between testes, immediately posterior to ovary. Mehlis’ gland not discernible. Laurer’s canal not detected. Uterine coils convoluted and densely packed with eggs, commence immediately posterior to oötype, pass posteriad to occupy much of post-ovarian space, then ascend antero-sinistrally to level of ventral sucker and then dextro-laterally to occupy most of space between intestinal bifurcation and testes; distal-most coil ascends antero-medially to meet genital atrium. Metraterm absent. Eggs ovoid, translucent and lightly tanned in proximal uterine coils, gradually darkening as they mature within uterus, darkest in distal/anterior-most uterine coils, 15–16 × 8–9 (16 × 8). Vitellarium comprises a pair of densely clumped lateral fields that extend anteriorly to level of pharynx up to 94–185 (140) from anterior extremity and posteriorly to level of ovary, occasionally medially confluent, dorsal to uterine coils, appear to surround caeca and excretory vesicle. Vitelline ducts extend postero-diagonally from vitelline fields, meet medially in mid-body posterior to ventral sucker. Excretory vesicle Y-shaped, bifurcates in mid-hindbody postero-dorsal to ovary; arms vary greatly in volume, extend to level of pharynx.
Differential diagnosis
A combination of ovoid-to-fusiform body shape, smooth-margined testes opposite (as opposed to distinctly oblique or in tandem) in the anterior hindbody, and uterine coils that are extensively distributed pre- and post-testicular, and reach but do not exceed the level of the ventral sucker, differentiates Siphoderina nana n. sp. from all but nine of the 49 recognised species of Siphoderina: S. asiatica Gu & Shen, 1979, S. ghanensis (Fischthal & Thomas, 1968), S. grunnitus, S. marangsi Machida, 2009, S. provitellosa (Durio & Manter, 1969), S. ramadani (Nahhas, Sey & Nishimoto, 1998), S. sootai (Hafeezullah, 1975), S. territans, and S. ulaula (Yamaguti, 1970). Of these nine, eight infect species of lutjanid fishes, the exception being S. grunnitus, which is known only from the haemulid Plectorhinchus gibbosus (Lacépède). Siphoderina nana is smaller than all other Siphoderina species, with a maximum length of only 692 µm, distinctly smaller than the minimum size of S. asiatica (952 µm), S. ghanensis (899 µm), S. marangsi (800 µm), S. provitellosa (992 µm), S. sootai (1,112 µm) and S. ulaula (2,200 µm). Siphoderina nana also has more oral sucker spines (60–72) than S. marangsi (20–28) and S. ulaula (42–58), and fewer than S. provitellosa (~135).
In having 60–72 oral spines, Siphoderina nana is similar to S. asiatica (65–74), S. ghanensis (70–75), S. ramadani (up to 60) and S. territans (53–66); the number of oral spines in S. sootai is unknown. Of those, S. nana can be immediately differentiated from S. asiatica, S. ghanensis and S. sootai by body size. Unlike S. nana, all three of those species show no overlap between the ovary and the testes; the ovary of S. asiatica, in particular, appears stellate in shape rather than irregularly lobed. The former two species also show more elongate, fusiform body shapes, with a higher ratio of length to breadth: ~ 1.9 and 2.1 times longer than broad, respectively, versus 1.2–1.8 (mean 1.5) times for S. nana. S. territans overlaps with S. nana in body size and oral spine count, as well as having an irregularly lobed ovary which extensively overlaps the testes, extensive pre-testicular uterine coils and similar vitelline follicle distribution. It is, however, larger than S. nana, averaging 706 µm long (range 533–946 µm), has a greater body length/breadth ratio (1.6–2.4, mean 2.0) and larger eggs (18–23 × 8–11, mean 20 × 9, vs 15–16 × 8–9, mean 16 × 8). The host range of S. territans is so far limited to Lutjanus carponotatus (Richardson), whereas S. nana infects L. fulviflamma. The two species are also genetically distinct from one another, differing by 28 bp and 32 bp in the ITS2 and 28S rDNA regions respectively. Siphoderina nana most closely resembles S. ramadani and the two are known from the same host. However, S. ramadani is larger than S. nana, with a minimum body size of 691 µm (max 1408, mean 906) whereas the maximum size of S. nana is 692 µm, and the number of oral spines described for S. ramadani was “up to 60” (Nahhas et al., 1998) whereas the minimum number of oral spines for S. nana is 60. Obtaining molecular sequence data for S. ramadani to clarify its phylogenetic position, particularly in relation to S. nana n. sp., is desirable.
Discussion
Our description of Siphoderina nana n. sp. brings the number of species in this genus to 49. The taxonomic history of this genus has been complex since its original proposition by Manter (1934) for S. brotulae from the ophidiid Brotula barbata (Bloch & Schneider). Species of several genera (Metadena Linton, 1910, Neochasmus Van Cleave & Müller, 1932, Paracryptogonimus Yamaguti, 1934, Lappogonimus Oshmarin, Mamaev & Parukhin, 1961 and Pseudallacanthochasmus Velasquez, 1961) have been subsequently transferred to Siphoderina (Miller & Cribb, 2008a, b). The majority of species (35 of 49) infect lutjanids, five infect the ecologically-similar centropomids and a further two infect haemulids, which are phylogenetically most closely related to lutjanids (Betancur-R et al., 2017); two species, S. ghanensis and S. grandispinus (Velasquez, 1961), have been recorded from both lutjanids and haemulids (Fischthal & Thomas, 1968; Velasquez, 1961). This leaves a minority of species infecting ecologically and phylogenetically disparate hosts, e.g. species of Gobiidae [S. microvata (Tubangui, 1928)], Cirrhitidae [S. mexicana (Bravo-Hollis, 1953)] and Uranoscopidae (S. xenocephali Machida, 2009) (Bravo-Hollis, 1953; Machida, 2009; Tubangui, 1928).
Several species of Siphoderina have been recorded from multiple phylogenetically, geographically and ecologically disparate hosts, further complicating interpretations of true host range within the genus. The type-species, S. brotulae, for example, was described from a brotula (Ophidiidae), but was subsequently reported infecting snappers (Lutjanidae) and goosefishes (Lophiidae), a host range across three fish orders (Ophidiiformes, Lutjaniformes and Lophiiformes, respectively) (Dyer et al., 1992; Manter, 1934, 1947). Siphoderina americana (Manter, 1940) has been recorded from both eastern and western American coasts, in seven lutjanid species (Fischthal, 1977; Manter, 1940), a centropomid (Lamothe-Argumedo, 1969), a batrachoidid (Pearse, 1949) and a serranid (Fischthal, 1977), as well as lutjanids from off Karwar, Karnataka on the west coast of India (Hafeezullah & Siddiqi, 1970). Most remarkably, S. grandispinus, described by Velasquez (1961) from an unknown Lutjanus species from the Philippines and also known from the haemulid Pomadasys hasta (Bloch) from off Kochi (Cochin), Kerala in western India (Hafeezullah & Siddiqi, 1970), has also been reported from the freshwater characiform Hoplias malabaricus (Bloch) (Erythrinidae) from Porto Alegre, Brazil (Fortes et al., 1996).
In many cases, the accounts in these reports are no more than cursory entries in bigger lists of host-parasite combinations, the result of more general parasitological surveys, with no accompanying descriptions, illustrations or specimen accession information. The overly simple nature of some early descriptions and their accompanying illustrations, and frequent use of excessive flattening during specimen preparation that warps anatomical features, also hinders effective interpretation. Most of these reports pre-date molecular sequencing techniques. Studies which make use of molecular sequence data, like those conducted by Miller & Cribb (2008a), indicate a tendency among species of Cryptogonimidae, including Siphoderina, toward high host-specificity. This, in turn, is reflective of a wider trend among trematodes of tropical marine fishes, in which genuinely euryxenic species are comparatively rare, and species once regarded as having broad host ranges on the basis of morphology alone are revealed to be complexes of multiple species when scrutinised using molecular sequencing (Miller et al., 2011). We are therefore sceptical of many of these reports. The few reports with accompanying illustrations leave little doubt that the taxa observed were indeed cryptogonimids, and these taxa may even be closely related to the species they are purported to be. In the absence of available specimens and especially molecular sequence data, we cannot definitively rule on their actual identity. Nevertheless, it seems likely that at least some of these reports of species of Siphoderina will prove erroneous.
Siphoderina is the largest cryptogonimid genus and among the least distinctive concepts in the family. As such, it currently accommodates substantial morphological variability. For example, considered in isolation, it seems unlikely that the diminutive S. nana, reaching at most 692 µm long, should be considered a good congener of S. onaga (Yamaguti, 1970), which reportedly reaches 11,200–12,500 µm long (Machida, 2009). Nevertheless, in our new analyses, all represented species of Siphoderina continue to form a monophyletic clade. Re-collection and sequencing of other Siphoderina species, especially the type-species S. brotulae and those other than from lutjanids and haemulids, will be informative for determining and refining the composition and definition of the genus. Indeed, although it is possible for species of Cryptogonimidae from a single genus to greatly vary morphologically [see Miller & Cribb (2007a, 2013)], for at least a few of the most morphologically disparate taxa, we suspect they will prove to not be close relatives of “good” species of Siphoderina at all. Rather, we suspect that the genus Siphoderina might incorporate multiple disparate groups, currently united via an overly generalised concept, and that the highly morphologically divergent forms might represent distinct radiations, possibly corresponding to ecologically or phylogenetically related host groups. Such a pattern is not without precedent among trematodes, e.g. the blood fluke genus Cardicola Short, 1953 (Aporocotylidae) (see Yong et al., 2021).
The novel sequence data for S. nana are the first for a cryptogonimid from South Africa. Prior to the new analyses, sequence data existed for just 10 of the (now) 49 species of Siphoderina, all of which were generated from specimens collected in Australian waters (Martin & Cutmore, 2022; Miller & Cribb, 2008a), and indeed the majority of sequences available for cryptogonimids are from Australian taxa. In the new analyses, novel data for S. nana resolved deep within the Siphoderina clade, clearly demonstrating that the Australian taxa do not form a monophyletic clade, i.e. S. nana is more closely related to some Australian species than those species are to some other Australian congeners. Further collection and sequencing efforts from South Africa and elsewhere in the western Indian Ocean are required to begin to understand the extent to which cryptogonimid species might be shared across the Indo-West Pacific, although, given the considerable efforts in Australia, it would appear that S. nana at least should be considered unlikely to occur there.
The largest radiation of species of Siphoderina has occurred in lutjanid fishes, and several lutjanid species are reported hosts for multiple Siphoderina species. The host for Siphoderina nana, L. fulviflamma, is the reported host for six other species of Siphoderina. Gu & Shen (1979) described S. asiatica from this host, as well as L. sanguineus (Cuvier), from off Hainan Island, China. Saoud et al. (1988) recorded S. leilae (Nagaty, 1957) (as Metadena leilae) in L. fulviflamma from the Persian/Arabian Gulf off Qatar. Nahhas et al. (1998) described S. ramadani in L. fulviflamma from the Kuwaiti coast, also in the Persian/Arabian Gulf. Nahhas et al. (2003) reported Siphoderina acanthostomus (Yamaguti, 1934) (as Paracryptogonimus acanthostomus Yamaguti, 1934) from this fish host from Fiji. Most recently, Miller & Cribb (2008a) described two species in L. fulviflamma, S. jactus Miller & Cribb, 2008 and S. quasispina Miller & Cribb, 2008 from Heron Island in the southern Great Barrier Reef, eastern Australia, with S. quasispina also found in the same host off northwestern Australia. Thus, L. fulviflamma is host to the greatest number of Siphoderina species of any lutjanid, and indeed any fish. Our phylogenetic analyses do not indicate that all these species represent a single radiation; rather, we infer that species of Siphoderina have invaded L. fulviflamma at least twice. It is unclear why L. fulviflamma seemingly supports greater richness of Siphoderina, except that this fish is common across its broad range and so has likely been sampled more frequently than most other lutjanids. Further sampling of lutjanids across the Indo-West Pacific, and perhaps especially from the western coast of Africa, will almost certainly continue to uncover substantial further richness of Siphoderina species.
References
Ankenbrand, M. J., Keller, A., Wolf, M., Schultz, J., & Förster, F. (2015). ITS2 Database V: Twice as much. Molecular Biology and Evolution, 32, 3030–3032.
Betancur-R., R., Wiley, E. O., Arratia, G., Acero, A., Bailly, N., Miya, M., Lecointre, G., & Orti, G. (2017). Phylogenetic classification of bony fishes. BMC Evolutionary Biology, 17, e162.
Bravo-Hollis, M. (1953). Dos especies nuevas de Cryptogonimidae Ciurea, 1933, de Puerto Vallarta, Jalisco, Mexico. Anales del Instituto de Biología, Universidad Nacional Autónoma de México, Series Zoología, 24, 137–145.
Bray, R. A. (1974). Acanthocephala in the flatfish Solea bleekeri (Soleidae) from Cape Province, South Africa. Journal of Helminthology, 48, 179–185.
Bray, R. A. (1978). Two new species of Enenterum Linton, 1910 (Digenea) in the marine fish Neoscorpis lithophilus (Kyphosidae) from the south-western Indian Ocean. Journal of Helminthology, 52, 131–139.
Bray, R. A. (1984). Some helminth parasites of marine fishes and cephalopods of South Africa: Aspidogastrea and the digenean families Bucephalidae, Haplosplanchnidae, Mesometridae and Fellodistomidae. Journal of Natural History, 18, 271–292.
Bray, R. A. (1985). Some helminth parasites of marine fishes of South Africa: Families Gorgoderidae, Zoogonidae, Cephaloporidae, Acanthocolpidae and Lepocreadiidae (Digenea). Journal of Natural History, 19, 377–405.
Bray, R. A. (1987). Some helminth parasites of marine fishes of south Africa: family Opecoelidae (Digenea). Journal of Natural History, 21, 1049–1075.
Bray, R. A., Waeschenbach, A., Cribb, T. H., Weedall, G., Dyal, P., & Littlewood, D. T. (2009). The phylogeny of the Lepocreadioidea (Platyhelminthes, Digenea) inferred from nuclear and mitochondrial genes: Implications for their systematics and evolution. Acta Parasitologica, 54, 310–329.
Bray, R. A., Diaz, P. E., & Cribb, T. H. (2016). Knowledge of marine fish trematodes of Atlantic and Eastern Pacific Oceans. Systematic Parasitology, 93, 223–235.
Cribb, T. H., & Bray, R. A. (2010). Gut wash, body soak, blender and heat-fixation: approaches to the effective collection, fixation and preservation of trematodes of fishes. Systematic Parasitology, 76, 1–7.
Cribb, T. H., Bray, R. A., Diaz, P. E., Huston, D. C., Kudlai, O., Martin, S. B., Yong, R. Q-Y., & Cutmore, S. C. (2016). Trematodes of fishes of the Indo-West Pacific: told and untold richness. Systematic Parasitology, 93, 237–247.
Darriba, D., Taboada, G. L., Doallo, R., & Posada, D. (2012). jModeltest 2: more models, new heuristics and parallel computing. Nature Methods, 9, 772.
Dos Santos, Q. M., Gilbert, B. M., Avenant-Oldewage, A., & Dumbo, J. C. (2021). Morphological and molecular description of Allocreadium apokryfi sp. n. (Digenea: Allocreadiidae) from native Labeobarbus aeneus (Cyprinidae) in South Africa, including notes on its biology, evolutionary history and an updated key of African Allocreadium. Folia Parasitologica, 68, e013.
Dyer, W. G., Williams, E. H., & Bunkley-Williams, L. (1992). Homalometron dowgialloi sp. n. (Homalometridae) from Haemulon flavolineatum and additional records of digenetic trematodes of marine fishes in the West Indies. Journal of the Helminthological Society of Washington, 59, 182–189.
Edgar, R. C. (2004). MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Research, 32, 1792–1797.
Fantham, H. B. (1938). Lecithostaphylus spondyliosomae n. sp., a trematode parasite of the Hottentot fish, Spondyliosoma blochii, found in South African waters. Transactions of the Royal Society of South Africa, 26, 387–393.
Fischthal, J. H. (1977). Some digenetic trematodes of marine fishes from the Barrier Reef and Reef Lagoon of Belize. Zoologica Scripta, 6, 81–88.
Fischthal, J. H., & Thomas, J. D. (1968). Digenetic trematodes of some freshwater and marine fishes from Ghana. Proceedings of the Helminthological Society of Washington, 35, 126–140.
Fortes, E., Hoffmann, R. P., & Scariot, J. (1996). Trematódeos digenéticos de Hoplias malabaricus (Bloch, 1794) do Lago Guaíba, Porto Alegre, RS, Brasil. Revista Brasileira de Medicina Veterinária, 18, 68–70.
Gavrilyuk-Tkachuk, L. P. (1979). New species of trematodes from commercial fishes of the Indian Ocean. Biologiya Morya, Vladivostok, 3, 83–86.
Gibson, D. I. (1983). Kenmackenzia gen. nov. and Kenmackenziinae subfam. nov. (Digenea, Sclerodistomidae): new taxa to accommodate the giant trematode Distoma gigas Nardo. Journal of Natural History, 17, 189–202.
Gu, C., & Shen, J. (1979). Ten new species of digenetic trematodes of marine fishes. Acta Zoologica Sinica, 4, 342–355.
Hafeezullah, M., & Siddiqi, A. H. (1970). Digenetic trematodes of marine fishes of India. Part 1. Bucephalidae and Cryptogonimidae. Indian Journal of Helminthology, 12, 1–22.
Hoogendoorn, C., Smit, N. J., & Kudlai, O. (2020). Resolution of the identity of three species of Diplostomum (Digenea: Diplostomidae) parasitising freshwater fishes in South Africa, combining molecular and morphological evidence. International Journal for Parasitology: Parasites and Wildlife, 11, 50–61.
Huston, D. C., Cutmore, S. C., Miller, T. L., Sasal, P., Smit, N. J., & Cribb, T. H. (2021). Gorgocephalidae (Digenea: Lepocreadioidea) in the Indo-West Pacific: new species, life-cycle data and perspectives on species delineation over geographic range. Zoological Journal of the Linnean Society, 193, 1416–1455.
Jayawardena, U. A., Tkach, V. V., Navaratne, A. N., Amerasinghe, P. H., & Rajakaruna, R. S. (2013). Malformations and mortality in the Asian Common Toad induced by exposure to pleurolophocercous cercariae (Trematoda: Cryptogonimidae). Parasitology International, 62, 246–252.
Kearse, M., Moir, R., Wilson, A., Stones-Havas, S., Cheung, M., Sturrock, S., Buxton, S., Cooper, A., Markowitz, S., Duran, C., Thierer, T., Ashton, B., Meintjes, P., & Drummond, A. (2012). Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics, 28, 1647–1649.
Kvach, Y., Bryjová, A., Sasal, P., & Winkler, H. M. (2017). A revision of the genus Aphalloides (Digenea: Cryptogonimidae), parasites of European brackish water fishes. Parasitology Research, 116, 1973–1980.
Lamothe-Argumedo, R. (1969). Trematodos de peces. IV. Registro de cuatro especies de trematodos de peces marinos de la costa del Pacifico Mexicano. Anales de Instituto de Biología, Universidad Nacional Autónoma de México, Serie Zoologia, 40, 179–194.
Le, T. H., Nguyen, K. T., Nguyen, N. T., Doan, H. T., Dung, D. T., & Blair, D. (2017). The ribosomal transcription units of Haplorchis pumilio and H. taichui and the use of 28S rDNA sequences for phylogenetic identification of common heterophyids in Vietnam. Parasites & Vectors, 10, e17.
Machida, M. (2009). Cryptogonimidae (Trematoda, Digenea) from fishes of Japanese and adjacent waters. Bulletin of the National Science Museum, Series A (Zoology), 35, 137–155.
Maddison, W. P., & Maddison, D. R. (2017). Mesquite: a modular system for evolutionary analysis. Version 3.2. Retrieved from http://mesquiteproject.org.
Malatji, M. P., & Mukaratirwa, S. (2019). Molecular detection of natural infection of Lymnaea (Pseudosuccinea) columella (Gastropoda: Lymnaeidae) with Fasciola gigantica (Digenea: Fasciolidae) from two provinces of South Africa. Journal of Helminthology, 94, e38.
Manter, H. W. (1934). Some digenetic trematodes from deep-water fish of Tortugas, Florida. Papers from Tortugas Laboratory, 28, 257–345.
Manter, H. W. (1940). Digenetic trematodes of fishes from the Galapagos Islands and the neighbouring Pacific. Reports of the Allan Hancock Pacific Expeditions, 2, 329–497.
Manter, H. W. (1947). The digenetic trematodes of marine fishes of Tortugas. American Midland Naturalist, 38, 257–416.
Martin, S. B., & Cutmore, S. C. (2022). Siphoderina hustoni n. sp. (Platyhelminthes: Trematoda: Cryptogonimidae) from the Maori snapper Lutjanus rivulatus (Cuvier) on the Great Barrier Reef. Systematic Parasitology, 99, 403–417.
Martin, S. B., De Silva, M. L. I., Pathirana, E., & Rajapakse, R. P. V. J. (2023). Polyphyly of the Dinurinae Looss, 1907 (Digenea: Hemiuridae) and resurrection of the Mecoderinae Skrjabin & Guschanskaja, 1954 based on novel collection of Tubulovesicula laticaudi Parukhin, 1969 from marine elapid snakes in Sri Lanka. Parasitology International, 97, 102776.
Martínez-Aquino, A., Vidal-Martínez, V. M., & Aguirre-Macedo, M. L. (2017). A molecular phylogenetic appraisal of the acanthostomines Acanthostomum and Timoniella and their position within Cryptogonimidae (Trematoda: Opisthorchioidea). PeerJ, 5, e4158.
Miller, M. A., Pfeiler, E., & Schwartz, T. (2010a). Creating the CIPRES Science Gateway for inference of large phylogenetic trees. Proceedings of the Gateway Computing Environments Workshop (GCE), 14 Nov. 2010, New Orleans, LA. New Orleans, LA., p. 1–8.
Miller, T. L., Adlard, R. D., Bray, R. A., Justine, J.-L., & Cribb, T. H. (2010b). Cryptic species of Euryakaina n. g. (Digenea: Cryptogonimidae) from sympatric lutjanids in the Indo-West Pacific. Systematic Parasitology, 77, 185–204.
Miller, T. L., Bray, R. A., & Cribb, T. H. (2011). Taxonomic approaches to and interpretation of host specificity of trematodes of fishes: lessons from the Great Barrier Reef. Parasitology, 138, 1710–1722.
Miller, T. L., Bray, R. A., Justine, J-L., & Cribb, T. H. (2010c). Varialvus gen. nov. (Digenea, Cryptogonimidae), from species of Lutjanidae (Perciformes) off the Great Barrier Reef, New Caledonia and the Maldives. Acta Parasitologica, 55, 327–339.
Miller, T. L., & Cribb, T. H. (2007a). Coevolution of Retrovarium n. gen. (Digenea: Cryptogonimidae) in Lutjanidae and Haemulidae (Perciformes) in the Indo-west Pacific. International Journal of Parasitology, 37, 1023–1045.
Miller, T. L., & Cribb, T. H. (2007b). Two new cryptogonimid genera (Digenea, Cryptogonimidae) from Lutjanus bohar (Perciformes, Lutjanidae): analyses of ribosomal DNA reveals wide geographic distribution and presence of cryptic species. Acta Parasitologica, 52, 104–113.
Miller, T. L., & Cribb, T. H. (2007c). Two new cryptogonimid genera Beluesca n. gen. and Chelediadema n. gen. (Digenea: Cryptogonimidae) from tropical Indo-West Pacific Haemulidae (Perciformes). Zootaxa, 1543, 45–60.
Miller, T. L., & Cribb, T. H. (2008a). Eight new species of Siphoderina Manter, 1934 (Digenea, Cryptogonimidae) infecting Lutjanidae and Haemulidae (Perciformes) off Australia. Acta Parasitologica, 53, 344–364.
Miller, T. L., & Cribb, T. H. (2008b). Family Cryptogonimidae Ward, 1917. In: Gibson, D. I., Bray, R. A. & Jones, A. (Eds) Keys to the Trematoda. Wallingford, UK: CABI Publishing, pp. 51–112.
Miller, T. L., & Cribb, T. H. (2009). Gynichthys diakidnus n. g., n. sp. (Digenea: Cryptogonimidae) from the grunt Plectorhinchus gibbosus (Lacépède, 1802) (Perciformes: Haemulidae) off the Great Barrier Reef, Australia. Systematic Parasitology, 74, 103–112.
Miller, T. L., & Cribb, T. H. (2013). Dramatic phenotypic plasticity within species of Siphomutabilus n. gen. (Digenea: Cryptogonimidae) from Indo-Pacific caesionines (Perciformes: Lutjanidae). Systematic Parasitology, 86, 101–112.
Miller, T. L., Cutmore, S. C., & Cribb, T. H. (2018). Two species of Neometadena Hafeezullah & Siddiqi, 1970 (Digenea: Cryptogonimidae) from Moreton Bay, Australia, including the description of Neometadena paucispina n. sp. from Australian Lutjanidae. Systematic Parasitology, 65, 655–664.
Miller, T. L., Downie, A. J., & Cribb, T. H. (2009). Morphological disparity despite genetic similarity; new species of Lobosorchis Miller & Cribb, 2005 (Digenea: Cryptogonimidae) from the Great Barrier Reef and the Maldives. Zootaxa, 1992, 37–52.
Nahhas, F. M., Sey, O., & Nishimoto, R. (1998). Digenetic trematodes of marine fishes from the Kuwaiti Coast of the Arabian Gulf: Families Pleorchiidae, Fellodistomidae, and Cryptogonimidae, with a description of two new species, Neoparacryptogonimus sphericus and Paracryptogonimus ramadani. Journal of the Helminthological Society of Washington, 65, 129–140.
Nahhas, F. M., Tran, M., & Nguyen, T. P. (2003). Digenetic trematodes of marine fishes from Suva, Fiji (Cryptogonimidae) including description of a new species. Acta Parasitologica, 48, 176–181.
Nikolaeva, V. M., & Tkachuk, L. P. (1979). A new trematode genus (Didymozoidae) from the common mackerel of the Indian Ocean. Parazitologiya, 13, 552–555.
Olson, P. D., Cribb, T. H., Tkach, V. V., Bray, R. A., & Littlewood, D. T. (2003). Phylogeny and classification of the Digenea (Platyhelminthes: Trematoda). International Journal of Parasitology, 33, 733–755.
Pantoja, C. S., Hernández-Mena, D. I., Pérez-Ponce de León, G., & Luque, J. L. (2018). Phylogenetic position of Pseudosellacotyla lutzi (Freitas, 1941) (Digenea: Cryptogonimidae), a parasite of Hoplias malabaricus (Bloch) in South America, through 28S rDNA sequences, and new observations of the ultrastructure of their tegument. Journal of Parasitology, 104, 530–538.
Pearse, A. S. (1949). Observations on flatworms and nemerteans collected at Beaufort, N.C. Proceedings of the U.S. National Museum, 100, 25–38.
Pérez-Ponce de León, G., & Hernández-Mena, D. I. (2019). Testing the higher-level phylogenetic classification of Digenea (Platyhelminthes, Trematoda) based on nuclear rDNA sequences before entering the age of the ‘next-generation’ Tree of Life. Journal of Helminthology, 93, 260–276.
Prudhoe, S. (1956). On a new trematode from South African fishes. Annals and Magazine of Natural History, 12, 72–75.
Rakgole, J., Moema, E. B., & King, P. H. (2019). Phylogenetic studies of larval digenean trematodes from freshwater snails and fish species in the proximity of Tshwane metropolitan, South Africa. Onderstepoort Journal of Veterinary Research, 86, 1–7.
Ronquist, F., Teslenko, M., van der Mark, P., Ayres, D. L., Darling, A., Hohna, S., Larget, B., Liu, L., Suchard, M. A., & Huelsenbeck, J. P. (2012). MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Systematic Biology, 61, 539–542.
Saoud, M. F. A., Ramadan, M. M., & Al Kawari, K. S. R. (1988). Helminth parasites of fishes from the Arabian Gulf. 4. On Allacanthochasmus lutjani n. sp. and Metadena leilae Nagaty, 1957 (Digenea: Cryptogonimidae). Qatar University Science Bullletin, 8, 161–172.
Stamatakis, A. (2014). RAxML Version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics, 30, 1312–1313.
Stoyanov, B., Neov, B., Pankov, P., Radoslavov, G., Hristov, P., & Georgiev, B. B. (2015). Redescription of Aphalloides coelomicola Dollfus, Chabaud & Golvan, 1957 (Digenea, Opisthorchioidea) based on specimens from Knipowitschia caucasica (Berg) (Actinopterygii, Gobionellidae) from a Black Sea lagoon, with comments on the systematic position of the genus. Systematic Parasitology, 91, 1–12.
Thaenkham, U., Nawa, Y., Blair, D., & Pakdee, W. (2011). Confirmation of the paraphyletic relationship between families Opisthorchiidae and Heterophyidae using small and large subunit ribosomal DNA sequences. Parasitology International, 60, 521–523.
Tubangui, M. A. (1928). Trematode parasites of Philippine vertebrates. Philippine Journal of Science, 36, 351–369.
Velasquez, C. C. (1961). Cryptogonimidae (Digenea: Trematoda) from Philippine food fishes. Journal of Parasitology, 47, 914–918.
Vermaak, A., Smit, N. J., & Kudlai, O. (2021). Molecular and morphological characterisation of the metacercariae of two species of Cardiocephaloides (Digenea: Strigeidae) infecting endemic South African klipfish (Perciformes: Clinidae). Folia Parasitologica, 68, e007.
Vermaak, A., Smit, N. J., & Kudlai, O. (2023). Molecular characterisation of three species of Coitocaecum (Digenea: Opecoelidae) infecting Clinus superciliosus (Clinidae) in South Africa, with description of Coitocaecum brayi sp. n. Folia Parasitologica, 70, e015.
Wee, N. Q-X., Cribb, T. H., Bray, R. A., & Cutmore, S. C. (2017). Two known and one new species of Proctoeces from Australian teleosts: Variable host-specificity for closely related species identified through multi-locus molecular data. Parasitology International, 66, 16–26. https://doi.org/10.1016/j.parint.2016.11.008.
WoRMS. (2023). WoRMS (World Register of Marine Species), World Wide Web electronic publication. Retrieved 3/2023 from https://www.marinespecies.org/index.php.
Yong, R. Q-Y., Cribb, T. H., & Cutmore, S. C. (2021). Molecular phylogenetic analysis of the problematic genus Cardicola (Digenea: Aporocotylidae) indicates massive polyphyly, dramatic morphological radiation and host-switching. Molecular Phylogenetics & Evolution, 164, e107290.
Yong, R. Q-Y., Cutmore, S. C., Wee, N. Q-X., & Cribb, T. H. (2016). A complex of Cardicola (Digenea: Aporocotylidae) species infecting the milkfish, Chanos chanos (Gonorynchiformes), with descriptions of two new species. Systematic Parasitology, 93, 831–846.
Acknowledgments
The collection work conducted in iSimangaliso Marine Protected Area was undertaken in part with funding from the National Research Foundation (NRF) of South Africa (NRF Project No. CPRR160429163437, Grant No. 105979, NJS as principal investigator); the opinions and conclusions expressed in this article are those of the authors and do not necessarily reflect those of the NRF. RQYY and SBM participated in this collection expedition with support from Holsworth Wildlife Research Endowments and Australian Government Research Training Program (RTP) Scholarships. RQYY gratefully acknowledges the current support of a North-West University (NWU) Postdoctoral Research Fellowship. SBM is currently supported by the Australian Biological Resources Study (ARBS) National Taxonomy Research Grants Program (NTRGP) grant G046WN7 and an Australia and Pacific Science Foundation (APSF) research grant APSF21048. The authors are very grateful to Hon. A/Prof. Tom Cribb of The University of Queensland for making available laboratory facilities to allow for morphological processing of specimens while RQYY was between laboratory placements. The authors also gratefully acknowledge Prof Paul Sikkel of the University of Miami for catching the host fishes while on expedition, Frances Brigg of the Western Australian State Agricultural Biotechnology Centre, Murdoch University for Sanger sequencing, Willie Landman of North-West University’s Unit of Environmental Sciences & Management for assistance and guidance with processing and photographing specimens for SEM, and Prof Gerardo Pérez Ponce de León of UNAM, Mexico, for sending copies of references which were unavailable to the authors. This is contribution number 792 from the NWU Water Research Group. This article is registered in ZooBank under the designation urn:lsid:zoobank.org:pub:66ACC844-7E82-4E44-978E-B735704AAC8D.
Funding
Open access funding provided by North-West University.
Author information
Authors and Affiliations
Contributions
RQYY, SBM and NJS were all members of the expedition on which the specimens used in this study were collected, and all parties were involved in the initial discovery and collection of these specimens. The expedition and all necessary permits were organised by NJS and attendance co-funded by all three parties. RQYY performed all morphological processing and analyses as well as phylogenetic analyses. SBM funded and performed all molecular sequencing. NJS provided funding for SEM processing. RQYY composed the manuscript, with substantial intellectual and editorial contributions from SBM and NJS.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no mutual conflicts of interest, competing financial interests, any secondary or funding bodies, and no personal interests or relationships that could affect the validity of this research.
Ethical approval
The authors declare that this work was performed in compliance with the ethical standards that legally prevail in their respective countries. Collection work was performed under the terms of a research agreement with iSimangaliso Wetland Park Authority allowing for collections to be made within a national park, and with ethics clearance granted under the permit number NWU-00440-16-A5-01.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yong, R.Q.Y., Martin, S.B. & Smit, N.J. A new species of Siphoderina Manter, 1934 (Digenea: Cryptogonimidae) infecting the Dory Snapper Lutjanus fulviflamma (Teleostei: Lutjanidae) from the east coast of South Africa. Syst Parasitol 100, 673–686 (2023). https://doi.org/10.1007/s11230-023-10116-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11230-023-10116-1