Abstract
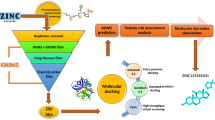
Among the neurodegenerative diseases responsible for cases of dementia, Alzheimer’s disease (AD) is responsible for 60–80% of cases. Caspase-3 is an enzyme essential in the synaptic degeneration caused by this disease, as it acts on the cleavage of tau and APP proteins, which leads to the formation of β-amyloid plaques in the brain. Therefore, the inhibition of caspase-3 may be an effective treatment method. In this work, QSAR studies based on descriptors derived from SMILES, similarity search, toxicity, and ADME studies were performed, including an evaluation of the applicability domain based on 71 1,2-benzisothiazol-3-ones capable of inhibiting caspase-3. Among the three hits identified, H2 proved to be the most interesting hit to be explored. Therefore, this hit was selected for future biological assay studies to confirm its activity. It may indicate new and original caspase inhibitors with drug-like properties suitable for AD therapy.




Similar content being viewed by others
Availability of data and materials
The study data were made available in the body of the article and in the supplementary material. If you have any questions, please contact the main author.
References
Sereniki A, Vital MABF (2008) Rev Psiquiatr Rio Gd Sul 30:(suppl 1). https://doi.org/10.1590/S0101-81082008000200002
Smith MDAC (1999) Rev Bras Neurol Psiquiatr 20(Suppl 2):3–7
Costa AS, Martins JPA, de Melo EB (2022) Struct Chem 33:1691–1706. https://doi.org/10.1007/s11224-022-02008-9
Forlenza OV (2005) Arch Clin Psychiatry (S Paulo) 32:137–148. https://doi.org/10.1590/S0101-60832005000300006
Mansur LL, Carthery MT, Caramelli P, Nitrini R (2005) Psicol Reflex Crit 18:300–307. https://doi.org/10.1590/S0102-79722005000300002
Porter AG, Jänicke RU (1999) Cell Death Differ 6:99–104. https://doi.org/10.1038/sj.cdd.4400476
Troy CM, Akpan N, Jean YY (2011) Prog Mol Biol Transl Sci 99:265–305. https://doi.org/10.1016/B978-0-12-385504-6.00007-5
Su JH, Zhao M, Anderson AJ, Srinivasan A, Cotman CW (2001) Brain Res 898:350–357. https://doi.org/10.1016/S0006-8993(01)02018-2
Roy K, Kar S, Das RN (2015) Understanding the basics of QSAR for applications in pharmaceutical sciences and risk assessment. 1st ed. Academic Press
Ferreira MMC (2002) J Braz Chem Soc 13:742–753. https://doi.org/10.1590/S0103-50532002000600004
Liu D, Tian Z, Yan Z, Wu L, Ma Y, Wang Q et al (2013) Bioorg Med Chem 21:2960–2967. https://doi.org/10.1016/j.bmc.2013.03.075
Guo Z, Yan Z, Zhou X, Wang Q, Lu M, Liu W et al (2015) Med Chem Res 24:1814–1829. https://doi.org/10.1007/s00044-014-1259-7
Wermuth CG (2011) The practice of medicinal chemistry. Academic Press
Stumpfe D, Hu Y, Dimova D, Bajorath J (2014) J Med Chem 57:18–28. https://doi.org/10.1021/jm401120g
Todeschini R, Consonni V (2008) Handbook of molecular descriptors. John Wiley & Sons
Puzyn T, Leszczynski J, Cronin MTD (2010) Recent advances in QSAR studies methods and applications. Springer
Martins JPA, Ferreira MMC (2013) Quim Nova 36:554–560. https://doi.org/10.1590/S0100-40422013000400013
Teófilo RF, Martins JP, Ferreira MMC (2009) J Chemom 23:32–48. https://doi.org/10.1002/cem.1192
Veerasamy R, Rajak H, Jain A, Sivadasan S, Varghese CP, Agrawal RK (2011) Int J Drug Des Discov 3:511–519
Calabrò M, Rinaldi C, Santoro G, Crisafulli C (2021) AIMS Neurosci 8:86–132. https://doi.org/10.3934/Neuroscience.2021005
Tropsha A (2010) Mol Inform 29:476–488. https://doi.org/10.1002/minf.201000061
Gramatica P (2007) QSAR Comb Sci 26:694–701. https://doi.org/10.1002/qsar.200610151
Tropsha A, Golbraikh A (2010) Predictive quantitative structure–activity relationships modeling: development and validation of QSAR models. Handbook of chemoinformatics algorithms. Chapman Hall/CRC p. 223–244
Shi LM, Fang H, Tong W, Wu J, Perkins R, Blair RM et al (2001) J Chem Inf Comput Sci 41:186–195. https://doi.org/10.1021/ci000066d
Roy K, Ambure P (2016) Chemom Intell Lab Syst 159:108–126. https://doi.org/10.1016/j.chemolab.2016.10.009
Rodrigues RP, Mantoani SP, de Almeida JR, Pinsetta FR, Semighini EP, da Silva VB et al (2012) Rev Virtual Quim 4:739–776. https://doi.org/10.5935/1984-6835.20120055
Zoete V, Daina A, Bovigny C, Michielin O (2016) J Chem Inf Model 56:1399–1404. https://doi.org/10.1021/acs.jcim.6b00174
Benfenati E, Manganaro A, Gini GC (2013) VEGA-QSAR. AI inside a platform for predictive toxicology. Italy: Politecnico di Mano. p. 21–28
Sahigara F, Mansouri K, Ballabio D, Mauri A, Consonni V, Todeschini R (2012) Molecules 17:4791–4810. https://doi.org/10.3390/molecules17054791
Minovski N, Župerl Š, Drgan V, Novič M (2013) Anal Chim Acta 759:28–42. https://doi.org/10.1016/j.aca.2012.11.002
Daina A, Zoete V (2016) ChemMedChem 11:1117–1121. https://doi.org/10.1002/cmdc.201600182
Daina A, Michielin O, Zoete V (2017) Sci Rep 7:42717. https://doi.org/10.1038/srep42717
Toropov AA, Benfenati E (2007) Curr Drug Discov Technol 4:77–116. https://doi.org/10.2174/157016307781483432
Kiralj R, Ferreira MMC (2009) J Braz Chem Soc 20:770–787. https://doi.org/10.1590/S0103-50532009000400021
Golbraikh A, Tropsha A (2002) J Mol Graph Model 20:269–276. https://doi.org/10.1016/S1093-3263(01)00123-1
Africa JGG, Arturo HCP, Bernardo LJM, Ching JKAR, de la Cruz OCE, Hernandez JBE et al (2022) Philipp J Sci 151:35–158. https://doi.org/10.56899/151.01.04
Manikandan P, Nagini S (2017) Curr Drug Targets 19:38–54. https://doi.org/10.2174/1389450118666170125144557
Orrling KM, Jansen C, Vu XL, Balmer V, Bregy P, Shanmugham A et al (2012) J Med Chem 55:8745–8756. https://doi.org/10.1021/jm301059b
Funding
The authors wish to thank Fundação Araucária (grant 2010/7354), PROAP/CAPES, and CNPq (grant 311048/2018–8).
Author information
Authors and Affiliations
Contributions
All authors contributed equally to the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Santos, P.B.J., de Melo, E.B. A QSAR and similarity search based on 1,2-benzisothiazol-3-ones to identify potential new inhibitors of caspase-3. Struct Chem (2024). https://doi.org/10.1007/s11224-024-02280-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11224-024-02280-x