Abstract
Discrete hazard models are widely applied for the analysis of time-to-event outcomes that are intrinsically discrete or grouped versions of continuous event times. Commonly, one assumes that the effect of explanatory variables on the hazard can be described by a linear predictor function. This, however, may be not appropriate when non-linear effects or interactions between the explanatory variables occur in the data. To address this issue, we propose a novel class of discrete hazard models that utilizes recursive partitioning techniques and allows to include the effects of explanatory variables in a flexible data-driven way. We introduce a tree-building algorithm that inherently performs variable selection and facilitates the inclusion of non-linear effects and interactions, while the favorable additive form of the predictor function is kept. In a simulation study, the proposed class of models is shown to be competitive with alternative approaches, including a penalized parametric model and Bayesian additive regression trees, in terms of predictive performance and the ability to detect informative variables. The modeling approach is illustrated by two real-world applications analyzing data of patients with odontogenic infection and lymphatic filariasis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The terms time-to-event data or survival data refer to data sets where the outcome variable corresponds to the time to the occurrence of a certain event of interest. In clinical research, for example, the time to death, the time to disease progression or the duration of hospitalization are widely applied time-to-event outcomes (Klein and Moeschberger 2003).
When building a regression model that relates the event time T to a set of explanatory variables \(\varvec{x} = (x_1,\ldots ,x_p)\) one must account for certain characteristics that are unique to time-to-event data. In particular, event times are usually subject to censoring, that is, the event of interest is not observed for all individuals under study. An appropriate approach for modeling time-to-event outcomes is to consider the hazard for the occurrence of an event at time t given by \(\xi (t) = \lim _{\Delta t \rightarrow 0}\{P(t<T\le t + \Delta t\vert \ T>t, \varvec{x})/ \Delta t\}\). The most popular hazard model is the Cox proportional hazards model by Cox (1972), which assumes that the effects of the explanatory variables on the hazard are constant over time. An alternative that does not require the proportional hazards assumption are accelerated failure time models (Kalbfleisch and Prentice 2002). This class of models is not dealt with here, but we refer to Kuss and Hoyer (2021) for recent developments in this field.
The classical Cox model assumes that the effects of the explanatory variables can be described by a parametric term (that is, by a linear function of \(\varvec{x}\)). This assumption, which is too restrictive in many applications, can be relaxed by the specification of smooth, non-linear functions (Sleeper and Harrington 1990; LeBlanc and Crowley 2004) or the use of recursive partitioning techniques also called tree-structured modeling, see, for example, Segal (1997), Zhang and Singer (1999), and Bou-Hamad et al. (2011a). Tree-structured models are strong tools that relate the explanatory variables to the outcome in a non-linear way and automatically detect relevant interactions between the explanatory variables if they are present. The visualization as a hierarchical tree makes the model easily accessible for practitioners and simulatable (Murdoch et al. 2019). In addition, tree-structured models inherently perform variable selection, which is particularly useful in high-dimensional settings. Within the scope of hazard models, Gordon and Olshen (1985) formerly extended classification and regression trees (CART, Breiman et al. 1984) and proposed to partition time-to-event data based on different measures of distance between two survival curves. Segal (1995) illustrated the application of tree-structured modeling for the analysis of HIV patient data, and Carmelli et al. (1997) applied tree-structured time-to-event analysis to investigate the relationship between obesity and mortality from coronary heart disease and cancer. More recently, Wallace (2014) applied conditional inference trees (Hothorn et al. 2006) to model time-to-event data, and Rancoita et al. (2016) proposed to use a Bayesian network for survival tree analysis with missing data.
All of the aforementioned methods have in common that they assume time to be measured or approximated by a continuous scale. In many applications, however, the event times are intrinsically discrete, or the exact continuous event times were not recorded and it is only known that the events occurred in a certain time interval (this is also referred to as the case of grouped event times). Grouped event times are typically observed in clinical and epidemiological studies with fixed prespecified follow-up times. Then time is recorded on a discrete time scale \(t = 1,\ldots ,k\). Two examples, which will be dealt with here, are days of hospitalization after jaw surgery in patients with acute odontogenic infection and time to an acute attack caused by infections through skin lesions in patients with lymphatic filariasis measured in months. In these cases, in which grouping effects are present, the application of statistical models designed for continuous time-to-event data is considered inappropriate by many authors (e.g. Tutz and Schmid (2016) and Berger and Schmid (2018)). Therefore, we consider the class of discrete hazard models that has prevailed for the analysis of discrete time-to-event outcomes (Willet and Singer 1993; Hashimoto et al. 2011; Tutz and Schmid 2016). A comprehensive introduction to parametric discrete hazard models and semi-parametric extensions was recently given by Berger and Schmid (2018). In this article, we propose novel alternatives to the widely used class of parametric discrete hazard models. More specifically, we propose different non-parametric extensions, where (part of) the parametric term is replaced by a tree structure while the common additive form of the predictor function is kept. As will be illustrated, the models are highly flexible and combine the advantages of classical parametric and tree-structured discrete hazard models.
Tree-structured models for discrete time-to-event analysis were so far proposed by Bou-Hamad et al. (2009) and Bou-Hamad et al. (2011b). Their method first grows a tree and then fits a covariate-free discrete hazard model in each terminal node separately. Schmid et al. (2016) suggested a CART approach where both the explanatory variables and the time t are considered as candidates for tree building. Sparapani et al. (2016) expanded the Bayesian additive regression tree (Chipman et al. 2010) for binary outcomes to time-to-event outcomes considering grouped survival times. In addition, Tiendrébéogo et al. (2019) applied a model-based recursive partitioning approach based on the algorithm by Hothorn et al. (2006) to HIV patient data to identify characteristics that are associated with risk of death. For overviews on existing tree-structured methods for discrete-time hazard modeling and extensions for dynamic predictions, see also Kretowska (2019) and Moradian et al. (2021).
Our proposed approach differs from previous methods, as we do not apply a traditional recursive partitioning algorithm, but fitting and tree building is performed within the classical framework of additive discrete hazard models. When growing the trees, in each step the best split is selected among all current non-internal and non-terminal nodes, yielding a sequence of nested subtrees of different size.
The remainder of this article is structured as follows: In Sect. 2 the notation and general methodology for the analysis of discrete time-to-event data are described. In Sect. 3 we propose three different novel tree-structured regression models for time-to-event outcomes. The performance of the different models was assessed in a simulation study, which is presented in Sect. 4. In Sect. 5, we show the results of two applications using the proposed models to analyze data of patients with odontogenic infection and lymphatic filariasis. Finally, our findings and conceptual aspects are discussed in Sect. 6.
2 Notation and methodology
In the following, let \(T_i\) denote the event time and \(C_i\) denote the censoring time of individual i, \(i=1,\ldots ,n\). We assume that \(T_i\) and \(C_i\) are independent and take discrete values in \(\{1,\ldots ,k\}\). It is further assumed that the censoring mechanism is non-informative for \(T_i\), in the sense that \(C_i\) does not depend on any parameters used to model the event time. Considering right-censored data, the observation time for individual i is given by \({\tilde{T}}_i = \min (T_i, C_i)\), and \(\Delta _i:=I(T_i \le C_i)\) indicates whether the event of individual i was observed (\(\Delta _i =1\)) or not (\(\Delta _i=0\)). In case continuous time-to-event data has been grouped, the discrete event times \(t=1,\ldots ,k\) refer to time intervals \([0,a_1), [a_1, a_2),\ldots ,[a_{k-1}, \infty )\), where \(T_i = t\) means that the event occurred in time interval \([a_{t-1}, a_t)\).
An essential tool for modeling discrete time-to-event data is the discrete hazard function. For time-constant explanatory variables \(\varvec{x_{i}}=(x_{i1},\ldots ,x_{ip})\) it is given by
which is the conditional probability for the occurrence of an event at time t given that the event has not occurred until t. From (1) it follows that the discrete survival function, which denotes the probability that an event occurs after time t, can be written as
Based on the definition of the hazard function in (1), for a fixed time t, the discrete hazard represents a binary variable that specifies whether an event occurs at time t or not, given that \(T_i \ge t\). Hence, strategies for modeling binary outcome data can be adapted to discrete hazard modeling.
A class of regression models that relates the discrete hazard function to the explanatory variables \(\varvec{x_i}\) is defined by
where \(h(\cdot )\) is a strictly increasing distribution function and \(\eta (\cdot )\) is a real-valued predictor function depending on the explanatory variables and time. A commonly assumed semi-parametric form of the predictor function is given by
where \(\gamma _0 (t)\) describes the hazard over time (for any given values of \(\varvec{x_i}\), called baseline hazard) usually by the use of dummy variables for each time point or a smooth, non-linear function. The effects \(\varvec{\gamma }\in {\mathbb {R}}^p\) of the explanatory variables on the hazard are assumed to be linear and independent of time. Using the logistic distribution function for \(h(\cdot )\), Equation (3) yields the logistic discrete hazard model
which is also called the proportional continuation ratio model (cf. Tutz and Schmid 2016, Chapter 3). The continuation ratio at time t is given by
and denotes the ratio comparing the probability of an event at time t to the probability of an event after t. Comparing the continuation ratios of two individuals u and v at time t yields
Hence, proportionality is given in the sense that the comparison of two individuals with regard to their continuation ratios is independent of time. This facilitates interpretation of the estimated effects (see our applications in Sect. 5). For the remainder of this article, we will focus on the model using the logistic distribution function and refer to Sect. 6 for a discussion on the characteristics of models with other link functions.
As mentioned above, discrete hazard models can be fitted by means of binary response models. This is because the log-likelihood of Model (3) can be expressed as
which is equivalent to the log-likelihood of a binomial model with independent observations \(y_{it}\). In order to derive coefficient estimates, one has to define the binary outcome values for each individual i as
Hence, before fitting the model with software for binary outcomes, the original time-to-event data has to be converted into an augmented data matrix that contains the binary outcome values. This results in a design matrix with \({\tilde{T}}_i\) rows for each individual i, where the vector of explanatory variables is repeated row-wise, and with a total number of \({\tilde{n}}=\sum _{i=1}^{n}{\tilde{T}}_i\) rows. Further details on data preparation and estimation of discrete hazard models can be found in Berger and Schmid (2018). Note that, in the following, the term individual will be used to refer to one row in the original, non-augmented data matrix, and the term observation to refer to one row of the augmented data matrix.
3 Tree-structured discrete hazard models
The logistic discrete hazard model with predictor function (4) assumes that the effect on the hazard can be described by a linear combination of the explanatory variables. This may be too restrictive, in particular when non-linear effects or interactions between the explanatory variables are present. To address this issue, we propose to incorporate tree-based splits into the predictor function of the discrete hazard model (3). Specifically, either the effects of the explanatory variables \(\varvec{x}\) or the effects of \(\varvec{x}\) as well as the effect of the time t on the hazard are replaced by a tree structure.
3.1 Smooth baseline coefficients
A tree-structured predictor function that allows for more flexibility, but still preserves the additive structure of the semi-parametric model (4) has the form
where the function \(tr(\cdot )\) is determined by a common tree structure. This means that the function \(tr(\cdot )\) sequentially partitions the observations into disjoint subsets \(N_m, m = 1,..,M\), based on the values of the explanatory variables and assigns a regression coefficient \(\gamma _m\) to each subset \(N_m\). Hence, this function can be written as
where \(I(\cdot )\) denotes the indicator function. When growing the tree, analogously to CART a binary split partitioning the observations of one parental node into two child nodes is performed in each step (cf. Hastie et al. 2009).
The function \(\gamma _0 (t)\) in Model (7) is determined by a P-spline (Eilers and Marx 1996). More precisely, a number of B-spline basis functions (de Boor 1978) are used, and a term to penalize differences between adjacent coefficients is included in the likelihood function. Then the coefficients are fitted by maximizing the penalized log-likelihood
where \(\varepsilon \in {\mathbb {R}}^{+}\) is a penalty parameter and \(J \in {\mathbb {R}}^{+}\) is the penalty term preventing the estimated function from becoming too rough. When using a P-spline, J is a difference penalty on adjacent B-spline coefficients. For details, see Wood (2011, 2017).
The tree \(tr(\cdot )\) is constructed in a stepwise procedure, starting from the model with baseline coefficients, only. Then the first split yields a model with predictor
where \(x_{j}\) is the explanatory variable selected for the first split, \(c_j\) is the corresponding split point, and \(\gamma _{1}^{[1]}\) is the effect for the first subset. Note that the second node defined by \(I(x_{ij} > c_j)\) needs to serve as reference node with \(\gamma _2^{[1]}:=0\) to ensure parameter identifiability. Secondly, a different or the same variable and a corresponding split point is selected to further split one of the current nodes. Assuming that the left node is split based on variable \(x_{k}\) with split point \(c_k\) yields the predictor
where \(\gamma _1 ^{[2]}\) and \(\gamma _2^{[2]}\) are the effects for the two new subsets built after the second split. Further splits are performed analogously until a predefined stopping criterion is met (see Sect. 3.4 for details). The design of the augmented data matrices for fitting the model with predictor (10) is illustrated in Online Resource Supplement 1, see Equations (S1) and (S2).
3.2 Piecewise constant baseline coefficients
Model (7) allows for non-linear effects of the explanatory variables as well as (higher-order) interactions between them. Moreover, the effects of the explanatory variables can easily be interpreted from the tree structure. Modeling the baseline coefficients by a smooth (P-spline) function may, however, not be adequate, as abrupt changes of the effect strength appear particularly plausible when considering discrete event times (Puth et al. 2020). Therefore, in the following, we assume that the baseline hazard does not vary over the whole range of t, but is constant within several time intervals.
An alternative to model (7) that also preserves the additive structure of parametric discrete hazard models, but allows to fit piecewise constant baseline coefficients is a model with predictor
where \(tr_0(\cdot )\) is a second tree partitioning the observations into subsets \(N_{0 m_0}, m_0 = 1,\ldots ,M_0\), with regard to the time t, and assigning a parameter \(\gamma _{0 m_0}\) to each subset \(N_{0 m_0}\), and \(tr(\cdot )\) is a tree structure defined as in (8). More formally, we have that
which represents a piecewise constant function over time. Both trees \(tr_0(\cdot )\) and \(tr(\cdot )\) are constructed using a similar stepwise procedure as described in Sect. 3.1. Assuming now that a split in t at \(c_t\) is selected in the first step yields the predictor
Then, a split in explanatory variable \(x_{j}\) at split point \(c_j\), in the second step, results in a predictor of the form
where the node \(I(x_{ij} > c_j)\) serves as reference node. In each iteration of the algorithm (see Sect. 3.4 for details), a split in either t or in one of the explanatory variables is performed, expanding \(tr_0(\cdot )\) or \(tr(\cdot )\), respectively. As a result, the predictor comprises piecewise constant effects over time and tree-structured effects of the explanatory variables. Note that the algorithm treats t as an ordinal variable. Thus, the total number of possible splits in \(tr_0(\cdot )\) is \(k-2\). When treating t as nominal variable instead, certain forms of the effects of time (e.g. U-shapes) might be detected more easily, particularly in small samples. Yet, this approach would be much more computationally expensive, as the number of possible splits grows exponentially with the number of time points. More specifically, \(2^{(k-1)-1} -1\) possible binary partitions would have to be considered for the first split in \(tr_0(\cdot )\). The design of the augmented data matrices for fitting the model with predictor (15) is illustrated in Online Resource Supplement 1, see Equations (S3) and (S4).
3.3 Modeling the effects of t and \(\varvec{x}\) by one single tree
The proposed models (7) and (12) are an attractive choice, as the effects can be captured in a very flexible way and the representation as a tree facilitates their interpretation. At the same time the common additive structure of parametric models (separating the effects of \(\varvec{x}\) and t) is kept.
If one suspects that interactions between time and explanatory variables are present, the use of a discrete hazard model with predictor
may be more appropriate. Here, the function \(tr(\cdot ,\cdot )\) partitions the observations based on the time t and the values of the explanatory variables \(\varvec{x_i}\), that is,
The construction of the tree \(tr(\cdot ,\cdot )\) is performed in the same manner as described in Sect. 3.1, but now in each step the time t (treated as an ordinal variable) and the explanatory variables are treated together. Essentially, this modeling approach is equivalent to the survival tree proposed by Schmid et al. (2016), as in both cases a single tree is built, where the time t and the explanatory variables are simultaneously considered as candidates for splitting. Schmid et al. (2016) proposed to use the Gini impurity measure for the selection of split points, whereas our approach selects the splits based on likelihood ratio (LR) test statistics (see Sect. 3.4), which is equivalent to the entropy criterion (Breiman et al. 1984). If the same splitting criterion is chosen, the model with predictor (16) and the survival tree yield the same results.
Model (16) is highly flexible, as an interaction between \(\varvec{x}\) and t implies the presence of time-varying effects on the hazards. This, however, comes at the price that the tree structure is harder to interpret, because each terminal node corresponds to a subset defined by the explanatory variables and to a time interval. An example of the augmented data matrices for fitting Model (16) is given in Equations (S5) and (S6) in Online Resource Supplement 1.
3.4 Fitting procedure
In each step of the tree-building algorithm, the best split among all candidate variables (that is, one component of \(\varvec{x}\) or t) and among all possible split points is selected. In order to do so, the two parameters corresponding to the two subsets resulting from the new split, \(\gamma _q^{[\ell ]}\) and \(\gamma _{q+1}^{[\ell ]}\) (where q is the current number of terminal nodes and \(\ell =q-1\) is the current number of splits), are tested for equivalence. More specifically, one examines all the null hypotheses \(H_0\): \(\gamma _{q}^{[\ell ]}=\gamma _{q+1}^{[\ell ]}\) against the alternatives \(H_1\): \(\gamma _q^{[\ell ]} \ne \gamma _{q+1}^{[\ell ]}\) by means of likelihood ratio (LR) tests. For a model with predictor (7), the splitting variable \(x_{q}\) and split point \(c_q\) related to the largest LR test statistic are selected. For models with predictor (12) and (16), the split is also selected based on the largest LR test statistic but can either be in t or in one of the explanatory variables (in the second or the same tree structure). Note that in each step of the algorithm all observations (of the augmented data matrix) are used to derive estimates of the model coefficients. That means, all parameters are refitted in each iteration and no previously estimated parameters are kept. Consequently, in case of the models with two components (introduced in Sects. 3.1 and 3.2), the parameter estimates of either of the two components are adjusted for the change through a split in the other. For the model with only one tree component described in Sect. 3.3, an additional split, however, does not change the parameter estimates in the remaining nodes (i.e. there would be no necessity to consider all the observations). Hence, the mechanism is equivalent to the fitting of a common tree, where only the subset of all observations in one node are used to guide the next split.
Three approaches for determining the size of the tree(s) are considered:
(i) The first alternative, which is based on the algorithm proposed by Berger et al. (2019), applies permutation tests. Let \(x_{q}\) be the variable selected for splitting. Then, the p-value obtained from the distribution of the maximally selected test statistic \(T_q = \max _{c_q} T_{q,c_q}\) provides a measure for the dependence between the time-to-event outcome and variable \(x_{q}\) while accounting for the number of possible split points (Hothorn and Lausen 2003). Therefore, one explicitly accounts for the involved multiple testing problem with regard to the number of split points for \(x_q\). The construction of the tree(s) is terminated when the null hypothesis of independence between the time-to-event outcome and the selected explanatory variable (or the time t) cannot be rejected (based on a prespecified error level \(\alpha \)). To determine the asymptotic distribution of \(T_q\) under the null hypothesis and to derive a test decision, a permutation test is performed. That means one permutes the values of the splitting variable \(x_{q}\) (or t) in the original augmented data matrix and computes the value of the maximally selected LR test statistic based on the permuted data. When this procedure is done repeatedly, an approximation of the distribution of \(T_q\) under the null hypothesis can be derived. Each permutation test is performed with local type I error level \(\alpha _l = \alpha /s\), where s is the number of candidate variables for splitting in the selected node. Hence, for each tree the probability of falsely identifying at least one variable as splitting variable is bounded by \(\alpha \) (i.e. on the tree level the family-wise error rate is controlled by \(\alpha \)). To determine the corresponding p-values with sufficient accuracy, the number of permutations should increase with the number of candidate splitting variables.
(ii) A second alternative is a minimal node size criterion, which is widely used in tree-building algorithms (Probst et al. 2019). With the minimal node size criterion, the minimal number of observations in the nodes is considered as the main tuning parameter for tree construction. If the number of observations in a current node falls below a prespecified threshold, the node is flagged as terminal node and is no longer available for further splits. The construction of the tree(s) is terminated when all current nodes are flagged as terminal nodes. To find the optimal minimal node size we propose to use the predictive log-likelihood of the model (evaluated on a separate validation sample or by cross-validation). It appears to be a natural choice, because split selection is also based on the likelihood. Note that, for the model with predictor (12), the optimal minimal node size must be determined for both trees, which requires optimization on a two-dimensional grid.
(iii) The third alternative is a post-pruning strategy, where a large tree is grown first and is then pruned to an adequate size to avoid overfitting. More specifically, after building the large tree, the sequence of nested subtrees is evaluated with regard to its predictive performance, for example, by using a validation sample or a k-fold cross validation scheme. Finally, the best-performing subtree is selected. As with the minimal node size criterion, we propose to consider the predictive log-likelihood as evaluation criterion. Note that, also for model (12) with two tree components, this strategy requires a one-dimensional optimization only.
To prevent the algorithm from building extremely small nodes (with only a few observations), an additional minimal bucket size constraint may be applied. With the minimal bucket size constraint, the minimum number of observations required in any terminal node is limited downward.
To summarize, the following steps are performed during the fitting procedure, if the permutation test is applied:
-
1.
Initial model: Fit the model without any explanatory variables, yielding a single estimate of the intercept \({\hat{\gamma }}_0\) (or a P-spline function modeling the baseline effects).
-
2.
Tree building:
-
(a)
Fit all candidate models with one additional split regarding one of the explanatory variables (or t in case of a model with predictor (12) or (16)), that fulfill the minimal bucket size constraint, in one of the already built nodes. If none of the additional splits meets the minimal bucket size constraint, terminate the algorithm.
-
(b)
Select the best model based on the maximal LR test statistic.
-
(c)
Carry out the permutation test for the variable associated with the selected split with error level \(\alpha _l\). If significant, fit the selected model and continue with step (2a). Otherwise, terminate the algorithm.
-
(a)
If the minimal node size criterion is used, the algorithm can be summarized as follows:
-
1.
Initial model: Fit the model without any explanatory variables.
-
2.
Tree building:
-
(a)
Fit all candidate models with one additional split regarding one of the explanatory variables (or t), that fulfill the minimal bucket size constraint, in one of the already built nodes. If none of the additional splits meets the minimal bucket size constraint, terminate the algorithm.
-
(b)
Select the best model based on the maximal LR test statistic considering the already built nodes that fulfill the minimal node size criterion, only. If none of the already built nodes meets the minimal node size criterion, terminate the algorithm.
-
(c)
Fit the selected model and continue with step (2a).
-
(a)
Using the post-pruning method, the fitting procedure is as follows:
-
1.
Initial model: Fit the model without any explanatory variables.
-
2.
Tree building:
-
(a)
Fit all candidate models with one additional split regarding one of the explanatory variables (or t), that fulfill the minimal bucket size constraint, in one of the already built nodes. If none of the additional splits meets the minimal bucket size constraint, continue with step (3).
-
(b)
Select the best model based on the maximal LR test statistic.
-
(c)
Fit the selected model and continue with step (2a).
-
(a)
-
3.
Pruning: Select the best model with the minimal predictive log-likelihood from the sequence of models built in steps (1) and (2). Terminate the algorithm.
In R, the augmented data matrix for fitting discrete time-to-event models can be obtained by applying the function dataLong() of the package discSurv (Welchowski et al. 2022). Technically, the proposed algorithm can be embedded into the framework of tree-structured varying coefficients models (TSVC; Berger et al. 2019). The models can therefore be fitted by the eponymous R add-on package TSVC (Berger 2021), where the explanatory variables (and the time t) serve as effect modifiers, modifying the effect of a constant auxiliary variable. Online Resource Supplement 2 contains a collection of code examples demonstrating how the proposed models can be fitted using TSVC.
4 Simulation study
To assess the performance of the proposed tree-structured models and to illustrate their properties, we considered different simulation scenarios. In all scenarios, the aims were (i) to evaluate how the performance of the approaches is affected by the degree of censoring and the number of noise variables, (ii) to compare the three approaches for determining the size of the tree(s), and (iii) to compare the proposed approaches to alternative methods, in particular to penalized maximum likelihood (ML) estimation and Bayesian additive regression trees. In addition, the computation times of the models were considered. The scenarios were based on a true model with predictor (12) composed of a piecewise constant baseline function and tree-structured effects of the explanatory variables (scenario 1), a true model with predictor (4) composed of a smooth baseline function and linear effects of the explanatory variables (scenario 2), and a true model with predictor (16) including interaction effects between time and explanatory variables (scenario 3). More details on the data-generating models will be given in the following subsections.
In each scenario, the following models were fitted to the simulated data:
-
(i)
the tree-structured model with smooth baseline effects as described in Sect. 3.1, referred to as SB,
-
(ii)
the tree-structured model with piecewise constant baseline coefficients as described in Sect. 3.2, referred to as PCB,
-
(iii)
the tree-structured model including t and \(\varvec{x}\) in a single tree as described in Sect. 3.3, referred to as ST,
-
(iv)
a parametric model (4) with a LASSO penalty on each of the regression coefficients and a group LASSO penalty on the baseline coefficients, referred to as LASSO,
-
(v)
a nonparametric model using Bayesian additive regression trees with logistic link function, referred to as BART,
-
(vi)
a fully specified parametric model (4) including dummy-coded baseline coefficients and all explanatory variables (Full),
-
(vii)
a model without any explanatory variables and a constant baseline coefficient only (Null), and
-
(viii)
the true data-generating model (Perfect).
For the SB, the PCB and the ST model, permutation tests (PT), the minimal node size criterion (MNS) as well as the post-pruning method (PR) were applied to determine the optimal sizes of the trees. The baseline coefficients of the SB model were estimated by a P-spline comprising five cubic B-spline basis functions and using a second order difference penalty. The optimal penalty parameter \(\varepsilon \) was determined by generalized cross-validation (see Wood 2017).
The LASSO model contains a penalty term for each explanatory variable and for the time t. The group LASSO penalty on the baseline coefficients guarantees that either all or none of the corresponding estimates are equal to zero. The degree of regularization is controlled by a joint penalty parameter \(\varepsilon \). See Yuan and Lin (2006) and Meier et al. (2008) for details.
The BART model, as introduced for survival analysis by Sparapani et al. (2016), is built of a sum of trees modeling the effects of time and the explanatory variables jointly (analogously to the ST model) and is fitted within a Bayesian framework. To achieve maximum comparability, we used a logistic link function (instead of the default probit link) and set the number of trees to one. Following Sparapani et al. (2016), the posterior distributions were estimated based on 2000 draws using a Markov chain Monte Carlo (MCMC) algorithm with 100 burn-in draws, a thinning factor of 10, and the default priors (see Sparapani et al. 2021).
In all simulation scenarios, the time-to-event outcome was related to one binary explanatory variable \(x_1 \sim B(1, 0.5)\) and two continuous explanatory variables \(x_2,x_3 \sim N(0,1)\). We considered a low-dimensional setting with five additional binary noise variables (\(\sim B(1,0.5)\)) and two additional standard normally distributed variables, as well as a high-dimensional setting with 90 additional binary noise variables and seven additional standard normally distributed noise variables. In each scenario we performed 100 replications and simulated a training sample, a validation sample and a test sample of size \(n=500\), respectively. The number of discrete time points was set to \(k=11\). The validation sample was used to determine the optimal minimal node sizes and best performing subtrees (post-pruning) for the SB, the PCB and the ST model and the optimal penalty parameter \(\varepsilon \) for the LASSO model by maximization of the predictive log-likelihood. The permutation tests were based on 1000 permutations with error level \(\alpha = 0.05\). The censoring times \(C_i\) were sampled independently of the event times \(T_i\) using the probability mass function \(P(C_i=t) = b^{k+1-t}/ \sum _{j=1}^{k}b^j,\ t = 1,\ldots ,k\). This resulted in censoring rates of approximately \(30\%\) (\(b=0.7\)), \(50\%\) (\(b = 1\)) and \(70\%\) (\(b =1.3\)).
To assess the performance of the different approaches, the predictive log-likelihood was calculated on the test samples. In addition, we considered prediction error (PE) curves as a time-dependent measure of prediction error. In case of BART, we used the mean (predicted) hazards over the MCMC draws to calculate the predictive log-likelihood values and the PE curves. The PE at time point t is given by
where \({\hat{S}}_i (t) = \prod _{s=1}^{t}(1-{\hat{\lambda }} (s\vert \varvec{x}_i))\) denotes the estimated survival function, \({\tilde{S}}_i(t) = I(t < {\tilde{T}}_i)\) denotes the observed survival function of individual i at time t, and \(w_i(t)\) are inverse-probability-of-censoring weights, see van der Laan and Robins (2003). We also computed the integrated PE given by
The marginal probabilities \(P(T =t)\) were estimated using a logistic discrete hazard model with dummy variables for each time point. For more details on PE curves, see Tutz and Schmid (2016).
Furthermore, true positive rates (TPR) and false positive rates (FPR) for the explanatory variables were considered. The true positive rate specifies the proportion of explanatory variables that were correctly identified to have an effect on the hazard (that is, correctly selected for splitting). It is given by
where \(\vartheta _j = 1\) if \(x_j\) has an effect on the hazard and \(\vartheta _j = 0\) otherwise. The false positive rate describes the proportion of all noise variables that were falsely identified to have an effect on the hazard and is given by
In case of BART, TPR and FPR were determined by averaging over the MCMC draws.
For the settings with tree-structured effects of time, we also determined a true positive rate for the thresholds in t, which is given by
where \(\delta _t=1\) if there is a split at t and \(\delta _t = 0\) otherwise. Analogously, the false positive rate for the thresholds in t is given by
Results of the simulation study: Predictive performance (scenario 1). The figure shows the predicted log-likelihood obtain from fitting the different models for the low-dimensional setting (left panel) and the high-dimensional setting (right panel) with low censoring (30%). For the results with medium and high censoring, see Figure S3 and S4 of Online Resource Supplement 3
Results of the simulation study: Prediction error curves (scenario 1). The figures show the prediction error curves (averaged over 100 replications) of the proposed tree-structured model with the lowest integrated prediction error (SB PR) and the competing models for the low-dimensional setting (left panel) and the high-dimensional setting (right panel) with low censoring (30%). For the results of the other proposed tree-structured models, see Figures S9 and S10 of Online Resource Supplement 3. Values of the integrated PE for all settings (with low, medium and high censoring) are given in Table S3 of Online Resource Supplement 3
4.1 Model with piecewise constant baseline effects and tree-structured effects of the explanatory variables
In the first scenario, the predictor of the true data-generating model had the form
with \(\gamma _1 = -2.5\), \(\gamma _2 = -1.5\), \(\gamma _3 = -0.5 \), \(\gamma _4 = 0.5\), and baseline function
Figure S1 in Online Resource Supplement 3 shows the effects of the explanatory variables represented as a tree structure.
From the results in Table 1 it is seen that all the proposed models were very efficient in detecting the informative variables. The TPR (upper panel) were higher than 0.6 throughout all settings, even for high-dimensional data with strong censoring. Compared to the tree-structured models, LASSO yielded much higher FPR, indicating that the selected models were far too large, while BART yielded much lower TPR. This also resulted in a considerably worse predictive performance of LASSO and BART compared to the proposed models (see Fig. 1, which show the results for the settings with low censoring). The corresponding results for medium and high censoring are given in Online Resource Supplement 3.
It is also seen that the SB and PCB model performed similarly well, whereas the ST model performed worst among the proposed models. This indicates that the true underlying piecewise constant baseline function could be approximated sufficiently by the smooth function of the SB model. Modeling the effects of time and the explanatory variables in one tree, however, did not fit the data very well.
Furthermore, models where permutation tests were applied as stopping criterion mostly showed lower FPR than models using the minimal node size criterion or the post-pruning method (see lower panel of Table 1). This is because the permutation test procedure is intended to control the family-wise error rate \(\alpha \) (see Sect. 3.4). Regarding the TPR, the permutation test procedure showed slightly better performance in the settings with low censoring, whereas applying the minimal node size criterion was superior in the settings with high censoring. The post-pruning method performed particularly well in the high-dimensional settings. Of note, in the present setting the true underlying model was determined by a tree with similarly sized terminal nodes. In a more unbalanced setting, the minimal node size criterion tended to be inferior to the permutation test and the post-pruning method (see, for example, Figure S8 of scenario 3 in Online Resource Supplement 3).
Figure 2 shows the PE curves for the settings with low censoring. The figure depicts the results obtained from the proposed tree-structured model with the lowest integrated prediction error (SB PR) and the competing models. It is seen that the differences in PE were larger for later time points, with the proposed SB PR model resulting in the lowest PE values among the competitors across all time points. This result appears to be unaffected by the dimensionality of the data. For the PE curves of the other proposed-tree-structured models, we refer to Figures S9 and S10 of Online Resource Supplement 3.
The selection rates obtained for the thresholds in t given in Table 7 of Online Resource Supplement 3 demonstrate that the PCB model with minimal node size criterion carried out considerably more splits in time yielding higher TPR and FPR. In case of the ST model, most splits were performed when the post-pruning method was applied.
Results of the simulation study: Predictive performance (scenario 2). The figure shows the predicted log-likelihood obtained from fitting the different models for the low-dimensional setting (left panel) and the high-dimensional setting (right panel) with low censoring (30%). For the results with medium and high censoring, see Figure S5 and S6 of Online Resource Supplement 3
Results of the simulation study: Prediction error curves (scenario 2). The figures show the prediction error curves (averaged over 100 replications) of the proposed tree-structured model with the lowest integrated prediction error (SB MNS) and the competing models for the low-dimensional setting (left panel) and the high-dimensional setting (right panel) with low censoring (30%). For the results of the other proposed tree-structured models, see Figures S11 and S12 of Online Resource Supplement 3. Values of the integrated PE for all settings (with low, medium and high censoring) are given in Table S4 of Online Resource Supplement 3
4.2 Model with smooth baseline effects and linear effects of the explanatory variables
The second scenario was based on a predictor of the form
with \(\gamma _1 = -0.5\), \(\gamma _2 = -0.25\), \(\gamma _3 = 0.25\) and a sigmoid baseline function
Table 2 and Fig. 3 show that the proposed models yielded lower TPR than in scenario 1 throughout all settings and were outperformed by the LASSO in terms of TPR and predictive ability. The TPR particularly deteriorated in the settings with high censoring. The BART model appeared to be less affected by an increasing censoring rate, but showed much lower TPR in the high-dimensional settings.
Yet, the proposed models showed substantially better predictive performance compared to the Null model in the low- and high-dimensional setting as well as compared to the Full model in the high-dimensional setting, which reflects the added value of the variable selection procedure.
Again, the ST models achieved the lowest predictive log-likelihood values among the tree-structured models (seventh, eighth and ninth boxplot of Fig. 3), which demonstrates that fitting one single tree comprising the effects of time and the explanatory variables is not adequate if the underlying model has an additive structure.
The models applying the minimal node size criterion performed slightly better than models using permutation tests and the post-pruning method. The rather weak performance of the permutation test indicates that the test procedure is too conservative to detect numerous informative variables in settings with moderately sized linear effects.
Figure 4 shows that the PE for the settings with low censoring increased until \(t=7\) for all of the models. The LASSO performed best across all time points in both the low- and high-dimensional setting. While the SB MNS model was outperformed by the LASSO, it was superior to the Null and BART model in the low-dimensional setting as well as to the Full model in the high-dimensional setting. For the PE curves of the other proposed-tree-structured models, we refer to Figures S11 and S12 of Online Resource Supplement 3.
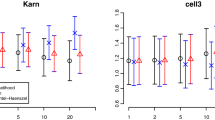
Results of the simulation study: Predictive performance (scenario 3). The figure shows the predicted log-likelihood obtained from fitting the different models for the low-dimensional setting (left panel) and the high-dimensional setting (right panel) with low censoring (30%). For the results with medium and high censoring, see Figure S7 and S8 of Online Resource Supplement 3
Results of the simulation study: Prediction error curves (scenario 3). The figures show the prediction error curves (averaged over 100 replications) of the proposed tree-structured model with the lowest integrated prediction error (ST PR) and the competing models for the low-dimensional setting (left panel) and the high-dimensional setting (right panel) with low censoring (30%). For the results of the other proposed tree-structured models, see Figures S13 and S14 of Online Resource Supplement 3. Values of the integrated PE for all settings (with low, medium and high censoring) are given in Table S5 of Online Resource Supplement 3
4.3 Tree-structured model with interaction effects between time and explanatory variables
The data of the third scenario was generated based on a predictor of the form
with \(\gamma _0 = -0.5\), \(\gamma _1 = -4\), \(\gamma _2 = -3\), \(\gamma _3=-2\), \(\gamma _4=-1\), \(\gamma _5=0\). Figure S2 in Online Resource Supplement 3 illustrates the predictor function represented as a tree structure.
From Table 3 and Fig. 5 it is seen that the performance of the different models deviated from each other more strongly than in the previous scenarios. While the LASSO yielded higher TPR than the tree-structured models, the proposed models resulted in much lower FPR than LASSO and BART and exhibited good predictive performance. In particular, they performed substantially better than the Null model as well as the Full model in the high-dimensional settings).
Among the proposed models, the ST model (seventh, eighth and ninth boxplots in Fig. 5), whose structure corresponds to the form of the true underlying predictor function, was clearly superior to the SB and PCB models. Further, the difference in predictive performance between the SB and PCB models shows that the abrupt change in the hazard at time point \(t=4\) could be captured less adequately by the smooth function of the SB model. According to the TPR in Table S2 in Online Resource Supplement 3, a split in \(t=4\) was carried out by the PCB and ST models in each replication (as all values were equal to one).
In addition, Table 3 shows that the ST model resulted in much lower TPR when applying the minimal node size criterion compared to the permutation tests or the post-pruning procedure in the settings with medium and high censoring. This may be explained by the unbalanced size of the terminal nodes of the tree according to the true underlying model. This also results in a decreased predictive performance (see Figures S7 and S8 in Supplement 3 of the Online Resource).
The PE curves shown in Fig. 6 were very close at early time points, but for later time points (\(t>4\)) in particular the BART model performed worse than the other models, which may be caused by the low number of observations at subsequent time points (leading to a stronger influence of the prior distributions). The proposed ST PR model showed the lowest PE among the competitors across all time points for low- and high-dimensional data. For the PE curves of the other proposed-tree-structured models, we refer to Figures S13 and S14 of Online Resource Supplement 3.
4.4 Run-time
In the last part of the simulation study we investigated the computing times of the proposed tree-structured models and compared them to the alternative approaches LASSO and BART. To do so, we evaluated the run-times in simulation scenario 2 with low censoring. This setting was chosen because increasing the degree of censoring results in a lower number of rows in the augmented data matrix \(({\tilde{n}})\) and therefore generally reduces run-times. Table 4 shows that the run-times differ strongly depending on the structure of the model (SB, PCB or ST) and the stopping criterion (PT, MNS or PR). Run-times were consistently longer compared to the LASSO model, which on average took only one minute or less in both settings, and also longer than the BART model in some cases. Among the proposed tree-structured approaches the post-pruning method was least expensive, in particular for the ST and PCB model. A higher number of noise variables (high-dimensional setting) increased the run-times when using the minimal node size criterion or the post-pruning method. In contrast, the run-times decreased with a larger number of noise variables if permutation tests were applied. This is because the effort of the permutation test approach directly depends on the actual size of the fitted trees, which was considerably smaller in the high-dimensional setting (resulting in lower TPR, cf. Table 2). Moreover, the long run-times of the SB model demonstrate the computational demand of refitting the smooth baseline function \(\gamma _0(t)\) in each iteration, and the low performance of the PCB MNS model resulted from the optimization of the minimal node size on a two-dimensional grid.
5 Application
To illustrate the use of the proposed models, two real-world examples were considered. In both applications, we compared the different approaches also considered in the simulation study and selected the best-performing approach using a cross-validation procedure. The corresponding results are presented in the following.
5.1 Patients with acute odontogenic infection
We considered data of a five-year retrospective study investigating hospitalized patients with abscess of odontogenic origin conducted between 2012 and 2017 by the Department of Oral and Cranio-Maxillo and Facial Plastic Surgery at the University Hospital Bonn. Patients with an acute odontogenic infection suffer from pain, swelling, erythema and hyperthermia. If not treated at an early stage, such infections may spread into deep neck spaces and lead to perilous complications by menacing anatomical structures, such as major blood vessels, the upper airway and the mediastinum (Biasotto et al. 2004). The primary objective of the study was to identify risk factors that are associated with a prolonged length of stay (LOS) in the treatment of severe odontogenic infections. As the LOS was recorded in 24-h intervals (that is, time was measured in days \(t=1,\ldots ,18\)), the use of a discrete hazard model is appropriate.
Analysis of the odontogenic infection data. The figure shows the results obtained from fitting the SB model with minimal node size criterion to the odontogenic infection data. On the left, the estimated smooth baseline function is presented. The graph on the right shows the estimated tree obtained from fitting the SB model with minimal node size criterion. The estimated coefficients \(\gamma _m\) are given in each leaf of the tree, where the right node on the first tree level serves as reference node
Here data from 303 patients that underwent surgical treatment in terms of incision and drainage of the abscess were considered. Intravenous antibiotics were administered during the operation and for the length of inpatient treatment. Further details on the study can be found in Heim et al. (2019). The characteristics of the patients considered for modeling were: age in years, gender (0: female, 1: male), an indicator of whether the infection spread into other facial spaces (0: no, 1: yes), the location of the infection focus (0: mandible, 1: maxilla), the administered antibiotics (0: ampicillin, 1: clindamycin), the presence of diabetes mellitus type 2 (0: no, 1: yes) and an indicator of whether the infection was already removed at admission (0: no, 1: yes). Basic statistics of the LOS and the patients characteristics are summarized in Table S6 in Online Resource Supplement 4.
The logistic discrete hazard model with predictor (4) including linear effects of the explanatory variables that was recently applied for statistical analysis by Heim et al. (2019) indicated that age and spreading of the infection focus into facial spaces significantly affected the LOS (with error level \(\alpha =0.05\)), while all the other variables showed no evidence for an effect.
In the first step of our analysis, we compared the tree-structured models (SB, PCB, ST, and BART), the LASSO model, the Full and the Null model with regard to their predictive performance. More specifically, we calculated the predictive log-likelihood values using ten-fold cross-validation. In case of the proposed tree-structured models, permutation tests, the minimal node size criterion, and the post-pruning method were used to limit the size of the trees. Additionally, a minimal bucket size constraint of \(mb = \lfloor 0.1 {\tilde{n}} \rfloor \), where \({\tilde{n}}\) is the number of observations (i.e. rows) in the augmented data matrix, was set for all tree-structured models. The optimal minimal node size, the optimal subtree, and the optimal LASSO penalty parameter were respectively determined by means of leave-one-out cross-validation on each of the ten training samples. Accordingly, predictive log-likelihood values were calculated for each of the ten test samples. Afterwards, the minimal node size, the subtree, and the penalty parameter value with the largest average predictive likelihood were selected.
Table 5 shows that the SB model applying the post-pruning method achieved the best performance, that is, the highest cross-validated log-likelihood value. In the second step of the analysis, we therefore fitted the SB PR model to the complete data. To determine the optimal subtree, we again performed leave-one-out cross-validation and selected the model with the highest predictive log-likelihood. The smooth baseline function of the SB model was fitted by five cubic P-splines with a second order difference penalty.
Figure 7a shows the estimated smooth baseline function \(\gamma _0(t)\) obtained from fitting the SB PR model. The figure illustrates that it was highly unlikely for patients to be discharged in the first few days after surgery. The probability of being discharged strongly increased until day five, but subsequently remained constant until day 18. Figure 7b shows the estimated tree-structured effects of the explanatory variables on the LOS. According to the first split, patients with spreading into facial spaces (Spreading=1) had a lower probability of being discharged. In the group of patients without spreading into facial spaces (Spreading=0), age was identified as an additional risk factor affecting the LOS. Patients who were older than 68 years were less likely being discharged than patients who were 68 years of age or younger. For patients 68 years of age or younger without spreading the continuation ratio was increased by the factor \(\exp (1.044)=2.841\) compared to patients with spreading. Figure 8 depicts the corresponding estimated survival functions for the three groups of patients determined by the tree.
Overall, our results strongly coincide with the findings by Heim et al. (2019). Based on the predictive log-likelihood, however, the tree-structured models SB and PCB surpassed the BART as well as the unrestricted and penalized models with linear effects (Full, Null and LASSO). This indicates that the tree-structured model (detecting an interaction between Spreading and Age) describes the association between the LOS and the explanatory variables best.
5.2 Patients with lymphatic filariasis
As a second example, we considered data from a randomized controlled trial in patients with lymphatic filariasis that was carried out in the western region of Ghana. Lymphatic filariasis is a filarial worm disease transmitted by mosquitoes that affects approximately 120 million persons worldwide and can lead to the development of severe lymphedema (LE) or hydroceles. The main objective of the study was to investigate the effect of antibiotic doxycycline in LE patients. Doxycycline has been shown previously to ameliorate LE severity in a subgroup of the population (Debrah et al. 2006). While the primary outcome of the study was change in LE stages, a key secondary outcome was time to occurrence of acute attacks, which are caused by secondary infections through skin lesions. Patients were examined 3 (\(t=1\)), 12 (\(t=2\)) and 24 (\(t=3\)) months after treatment onset, i.e., time was measured on a discrete scale with unequally spaced time intervals.
A sample of 118 patients was analyzed. The empirical distribution of the event times was \(23.73\%\), \(31.36\%\) and \(44.92\%\) for \({\tilde{T}}=1, 2, 3\), respectively. The censoring rate was \(34.75\%\) (patients, who did not suffer from an attack during follow-up). For details on how the trial was conducted, see Mand et al. (2012). The characteristics of the patients that were included in the analysis are: age in years, weight in kilograms, microfiliariae count, gender (0: female, 1: male), infection status (0: negative, 1: positive), administered drug (0: placebo, 1: 6-week course of amoxicillin at 1000 mg/d, 2: 6-week course of doxycycline at 200 mg/d), lymphedema stage before treatment and hygiene status before treatment (0: poor, 1: moderate, 2: good). Based on the inclusion criteria of the study, only patients with LE stages 1–5 according to the scheme by Dreyer et al. (2002) were considered. Two patients were excluded from the analysis because of missing values in the explanatory variables. Summary statistics on the characteristics of the patients are given in Table 12 in Supplement S7 of the Online Resource.
Based on a log-rank test, Mand et al. (2012) found a significant difference between the doxycycline and the placebo group with regard to the occurrence of acute attacks (with error level \(\alpha = 0.05\)). However, there was no evidence for a difference between the doxycycline and the amoxicillin group. More recently, the study data was analyzed with the survival tree method by Schmid et al. (2016). In their analysis, splits in hygiene status, LE stage and drug were performed, indicating that these characteristics affect the risk for an acute attack.
Our analysis of the data was performed analogously to the analysis of the odontogenic infection data (using ten-fold cross-validation and the minimal bucket size constraint \(mb = \lfloor 0.1 {\tilde{n}} \rfloor \)). Note that the three patients with LE stage 1 and 4 were excluded from the cross-validation step. Because of the small number of time intervals, fitting a smooth baseline function was not reasonable and the SB model was omitted.
The results of ten-fold cross-validation are shown in Table 6. The highest predictive log-likelihood value was achieved by the ST model applying the minimal node size criterion, which is (apart from the criterion for split selection) equivalent to the survival tree by Schmid et al. (2016). We therefore fitted the ST MNS model to the complete data (after determining the optimal minimal node size using again leave-one-out cross-validation).
Figure 9 shows the estimated tree structure, which concurs with the findings by Schmid et al. (2016), cf. Figure 7 therein. The first split was performed in hygiene showing the importance of good hygiene for lowering the risk for an acute attack. The subsequent splits reveal an interaction between time, lymphedema stage and drug for patients with poor to moderate hygiene. That is, within the first three months of the study, a higher lymphedema stage was shown to be a relevant risk factor, whereas after three months patients treated with doxycycline were shown to be at lower risk for an acute attack than patients in the placebo or amoxicillin group. The continuation ratio for patients with poor to moderate hygiene was increased by the factor \(\exp (1.163)=3.199\) after three months for patients treated with placebo or amoxicillin compared to patients treated with doxycycline. This finding confirms the results of the analysis by Mand et al. (2012) who found that the effects of doxycycline for the risk of an acute attack develop over the course of the second and third follow-up interval.
The predictive log-likelihood values for the considered models hardly deviate from each other (apart from the PCB PR model) indicating a lack of evidence for the superiority of one or several models (Burnham and Anderson 2002). However, the estimated ST model coincides with the survival tree established by Schmid et al. (2016), confirming their findings and supporting the validity of our approach.
6 Summary and discussion
The results of the simulation study and the applications to real-world data indicate that the proposed tree-structured models are promising tools for modeling discrete event times. The tree-structured models, unlike common parametric approaches, are able to capture non-linear effects and interactions between the explanatory variables, as illustrated by the results in Figs. 7 and 9. Even though the linear modeling approach surpassed the tree-structured approaches in the simulations, where the true effects were linear (see Sect. 4.2), the proposed models were shown to be highly effective in identifying the informative variables, particularly in high-dimensional settings. The ST model has a more flexible form than the SB and PCB model, as splitting in t allows to account for time-varying effects of the explanatory variables on the hazard. Yet, this may come at the price of interpretability of the tree structure and corresponding effects.
Our approach differs from traditional recursive partitioning algorithms: (i) Fitting of our tree-structured models is done within the framework of parametric discrete hazard models, which allows to apply software for binary regression modeling. Split selection and pruning of the built tree(s) are therefore naturally based on the likelihood. (ii) The SB and PCB model exhibit the common additive form of parametric discrete hazard models, which facilitates the interpretation of effects in terms of continuation ratios. (iii) The predictor function can be specified in a very flexible way including the survival tree by Schmid et al. (2016) as special case. (iv) The framework is easily generalizable, for example, to an additive discrete hazard model of the form
where \(\varvec{z}\) is an additional set of explanatory variables with linear effects on the outcome. An exemplary code how to fit such a model using TSVC is given in Online Resource Supplement 2.
One of the most important parameters for tree building is the number of splits that determines the size of the tree. Hence, before fitting one of our proposed models, one needs to consider which of the three criteria (PT, MNS or PR) to apply. In terms of predictive performance, the permutation test, the minimal node size criterion, and the post-pruning method yielded very similar results. The permutation test, which is the most conservative criterion (resulting in much lower FPR), appears slightly favorable in settings with low censoring, whereas the minimal node size criterion appears advantageous (showing higher TPR) for data with higher censoring rates, and the post-pruning methods appeared beneficial in high-dimensional settings. Both, performing permutation tests as well as determining the optimal minimal node size, may become computationally expensive, depending on the number of permutations or the resampling scheme used. In the latter case, for example, optimization on a two-dimensional grid (within the PCB model) using (leave-one-out) cross-validation requires high computing time (cf. Table 4). In terms of run-time, the post-pruning method proved to be superior, as the optimal subtree is selected from the sequence of nested subtrees that were already built on the training sample during iteration.
Alongside the logistic link function defined by the logistic distribution function (which we focused on in this article), the probit link function based on the standard normal distribution, and the complementary log-log (cloglog) link function defining the Gompertz model are popular choices. A comparison of parametric discrete hazard models with different link functions (considering information criteria) has been conducted by Hashimoto et al. (2011). All of the corresponding discrete hazard models postulate a proportionality property that affects the interpretation of parameters (Tutz and Schmid 2016, Chapter 3). In case of the logistic link function, the SB and PCB model yield proportionality with respect to the continuation ratios. Assume an SB model (7) with one split in \(x_j\) at split point \(c_j\), where \(\gamma _1\) is the coefficient of the left node and \(\gamma _2\) is the coefficient of the right node. Then the continuation ratio at time t is given by
Consequently, the term
is the factor by which the continuation ratio changes if \(x_j\) increases such that it exceeds \(c_j\). Since the effects of the explanatory variable \(x_j\) are independent of time t, if \(\gamma _2 >\gamma _1\), the continuation ratio of the second subgroup is higher at all time points. For the ST model such proportionality only holds in specific time intervals (see, for example, the tree in Fig. 9).
Regarding the choice of the link function one should note, that the logistic and the probit link function are based on density functions that are symmetric about the y-axis, whereas the Gompertz function, which is the basis of the cloglog model, is asymmetric in the sense that the right-hand asymptote of the function is approached much more gradually. For binary regression, Chen et al. (1999) recommended the use of an asymmetric link function in cases where the number of zeros and the number of ones in the data is highly unequally distributed. This is the case here, because the augmented data matrices used for tree building comprise a disproportionately high number of zero values. In addition, the Gompertz model is equivalent to the Cox proportional hazards model for continuous event times (with regard to the effects of the explanatory variables), if the original data were generated by the Cox model but only grouped event times are recorded (Tutz and Schmid 2016). Hence, the use of the cloglog link function in the scope of the proposed tree-structured hazard models might by worth investigating. For more details on link functions for binary regression models, see also Czado and Santner (1992) and Prasetyo et al. (2019).
Although only time-constant values of the explanatory variables were considered in the simulation study and the presented applications, it is also possible to deal with time-varying information. Instead of repeating the vector of explanatory variable row-wise, the vector of values measured at each time point could be entered in the corresponding rows of the augmented data matrix. The use of time-varying information, however breaks the relation between the survival function and the hazard function (2), because an observed value at t indicates that the individual must have survived up to t and thus \(S(t\vert \ \varvec{x}_{it})=1\).
In future research the computation of standard errors for the parameters \({\hat{\gamma }}\) or for the hazards \({\hat{\lambda }}(t\vert \ \varvec{x}_i)\) directly, for example by bootstrap procedures, needs to be investigated. Moreover, the proposed class of models can be extended to an ensemble method, as survival forests for continuous (Ishwaran et al. 2008; Moradian et al. 2017; Wang et al. 2018) and discrete-time data (Bou-Hamad et al. 2011b; Schmid et al. 2020; Moradian et al. 2021), and adapted to competing risk data, similar to the approach by Berger et al. (2019).
References
Berger, M.: TSVC: tree-structured modelling of varying coefficients. R Package Vers. 1(2), 2 (2021)
Berger, M., Tutz, G., Schmid, M.: Tree-structured modelling of varying coefficients. Stat. Comput. 29(2), 217–229 (2019). https://doi.org/10.1007/s11222-018-9804-8
Berger, M., Schmid, M.: Semiparametric regression for discrete time-to-event data. Stat. Model. 18(3–4), 1–24 (2018). https://doi.org/10.1177/1471082X17748084
Berger, M., Welchowski, T., Schmitz-Valckenberg, S., Schmid, M.: A classification tree approach for the modeling of competing risks in discrete time. Adv. Data Anal. Classif. 13(4), 965–990 (2019). https://doi.org/10.1007/s11634-018-0345-y
Biasotto, M., Pellis, T., Cadenaro, M., Bevilacqua, L., Berlot, G., Lenarda, R.D.: Odontogenic infections and descending necrotising mediastinitis: case report and review of the literature. Int. Dent. J. 54(2), 97–102 (2004). https://doi.org/10.1111/j.1875-595x.2004.tb00262.x
Bou-Hamad, I., Larocque, D., Ben-Ameur, H.: A review of survival trees. Stat. Surv. 5, 44–71 (2011). https://doi.org/10.1214/09-SS047
Bou-Hamad, I., Larocque, D., Ben-Ameur, H.: Discrete-time survival trees and forests with time-varying covariates: application to bankruptcy data. Stat. Model. 11(5), 429–446 (2011). https://doi.org/10.1177/1471082X1001100503
Bou-Hamad, I., Larocque, D., Ben-Ameur, H., Mâsse, L.C., Vitaro, F., Tremblay, R.E.: Discrete-time survival trees. Can. J. Stat. 37(1), 17–32 (2009). https://doi.org/10.1002/cjs.10007
Breiman, L., Friedman, J.H., Olshen, R.A., Stone, J.C.: Classification and Regression Trees. Taylor and Francis, Moneterey, CA Wadsworth (1984)
Burnham, K.P., Anderson, D.R.: Model Selection and Multimodel Inference, 2nd edn. Springer, New York, NY (2002)
Carmelli, D., Zhang, H., Swan, G.E.: Obesity and 33-year follow-up for coronary heart disease and cancer mortality. Epidemiology 8(4), 378–383 (1997). https://doi.org/10.1097/00001648-199707000-00005
Chen, M.H., Dey, D.K., Shao, Q.M.: A new skewed link model for dichotomous qantal response data. J. Am. Stat. Assoc. 94(448), 1172–1186 (1999). https://doi.org/10.2307/2669933
Chipman, H.A., George, E.I., McCulloch, R.E.: BART: Bayesian additive regression trees. Ann. Appl. Stat. 4(1), 266–298 (2010). https://doi.org/10.1214/09-AOAS285
Cox, D.R.: Regression models and life tables. J. R. Stat. Soc. Ser. B (Stat. Methodol.) 34(2), 187–220 (1972). https://doi.org/10.1111/j.2517-6161.1972.tb00899.x
Czado, C., Santner, T.J.: The effect of link misspecification on binary regression inference. J. Stat. Plan. Inference 33(2), 213–231 (1992). https://doi.org/10.1016/0378-3758(92)90069-5
de Boor, C.: A Practical Guide to Splines. Springer, New York, NY (1978)
Debrah, A.Y., Mand, S., Narfo-Debrekyei, Y., Basta, L., Pfarr, K., Labri, J., Lawson, B., Taylor, M., Adjei, O., Hoerauf, A.: Doxycycline reduces plasma VEGF-C/sVEGFR-3 and improves pathology in lymphatic filariasis. PLoS Pathog. 9(2), e92 (2006). https://doi.org/10.1371/journal.ppat.0020092
Dreyer, G., Addiss, D., Dreyer, P., Noroes, J.: Basic lymphoedema management: treatment and prevention of problems associated with lymphatic filariasis. Hollis Publishing Company, Hollis, NH (2002)
Eilers, P.H.C., Marx, B.D.: Flexible Smoothing with B-splines and Penalties. Stat. Sci. 11(2), 89–121 (1996). https://doi.org/10.1214/ss/1038425655
Gordon, L., Olshen, R.A.: Tree-structured survival analysis. Cancer Treat. Rep. 69(10), 1065–1069 (1985)
Hashimoto, E.M., Ortega, E.M.M., Paula, G.A., Barreto, M.L.: Regression models for grouped survival data: estimation and sensitivity analysis. Comp. Stat. Data Anal. 55(2), 993–1007 (2011). https://doi.org/10.1016/j.csda.2010.08.004
Hastie, T., Tibshirani, R., Friedman, J.: The Elements of Statistical Learning, 2nd edn. Springer, New York, NY (2009)
Heim, N., Berger, M., Wiedemeyer, V., Reich, R., Martini, M.: A mathematical approach improves the predictability of length of hospitalization due to acute odontogenic infection. A retrospective invetigation of 303 patients. J. Cranio-Maxillofac. Surg. 47(2), 334–340 (2019). https://doi.org/10.3844/jmssp.2019.354.365
Hothorn, T., Hornik, K., Zeileis, A.: Unbiased recursive partitioning: a conditional inference framework. J. Comp. Graph. Stat. 15(3), 651–674 (2006). https://doi.org/10.1198/106186006X133933
Hothorn, T., Lausen, B.: On the exact distribution of maximally selected rank statistics. Comp. Stat. Data Anal. 43(2), 121–137 (2003). https://doi.org/10.1016/S0167-9473(02)00225-6
Ishwaran, H., Kogalur, U.B., Blackstone, E.H., Lauer, M.S.: Random survival forests. Ann. Appl. Stat. 2(3), 841–860 (2008). https://doi.org/10.1214/08-AOAS169
Kalbfleisch, J., Prentice, P.: The Statistical Analysis of Failure Time Data, 2nd edn. Wiley Inter-Science, New Jersey, NJ (2002)
Klein, J., Moeschberger, M.: Survival Analysis: Statistical Methods for Censored and Truncated Data. Springer, New York, NY (2003)
Kretowska, M.: Oblique survival trees in discrete event time analysis. IEEE J. Biomed. Health Inform. 24(1), 247–258 (2019). https://doi.org/10.1109/JBHI.2019.2908773
Kuss, O., Hoyer, A.: A proportional risk model for time-to-event analysis in randomized controlled trials. Stat. Methods Med. Res. 30(2), 411–424 (2021). https://doi.org/10.1177/0962280220953599
LeBlanc, M., Crowley, J.: Adaptive regression splines in the cox model. Biom. 55(1), 204–213 (2004). https://doi.org/10.1111/j.0006-341x.1999.00204.x
Mand, S., Debrah, A.Y., Klarmann-Schulz, U., Basta, L., Marfo-Debrekyei, Y., Kwarteng, A., Specht, S., Belda-Domene, A., Fimmers, R., Taylor, M., Adjei, O., Hoerauf, A.: Doxycycline improves filarial lymphedema independent of filarial infection: a randomized controlled trial. Clin. Infect. Dis. 55(5), 621–630 (2012). https://doi.org/10.1093/cid/cis486
Meier, L., van de Geer, S., Bühlmann, P.: The Group Lasso for Logistic Regression. J. R. Stat. Soc. 70(1), 53–71 (2008). https://doi.org/10.1111/j.1467-9868.2007.00627.x
Moradian, H., Larocque, D., Bellavance, F.: L1 splitting rules in survival forests. Lifetime Data Anal. 23, 671–691 (2017). https://doi.org/10.1007/s10985-016-9372-1
Moradian, H., Yao, W., Larocque, D., Simonoff, J.S., Frydman, H.: Dynamic estimation with random forests for discrete-time survival data. Can. J. Stat. (published online) (2021). https://doi.org/10.1002/cjs.11639
Murdoch, W.J., Singh, C., Kumbier, K., Abbasi-Asl, R., Yu, B.: Definitions, methods, and applications in interpretable machine learning. Proc. Natl. Acad. Sci. 116(44), 22071–22080 (2019). https://doi.org/10.1073/pnas.1900654116
Prasetyo, R.B., Kuswanto, H., Iriawan, N., Sutijo, B., Ulama, S.: A comparison of some link functions for binomial regression models with application to school drop out rates in east java. AIP Conf. Proc. 2194, 020083 (2019)
Probst, P., Wright, M.N., Boulesteix, A.L.: Hyperparameters and tuning strategies for random forest. Wiley Interdisciip.: Rev. Data Min. Knowl. Discov. 9(3), 1301 (2019). https://doi.org/10.48550/arXiv.1804.03515
Puth, M.T., Tutz, G., Heim, N., Münster, E., Schmid, M., Berger, M.: Tree-based modeling of time-varying coefficients in discrete time-to-event models. Lifetime Data Anal. 26(3), 545–572 (2020). https://doi.org/10.1007/s10985-019-09489-7
Rancoita, P.M.V., Zaffalon, M., Zucca, E., Bertoni, F., De Campos, C.P.: Bayesian network data imputation with application to survival tree analysis. Comput. Stat. Data Anal. 93, 373–387 (2016). https://doi.org/10.1016/j.csda.2014.12.008
Schmid, M., Küchenhoff, H., Hoerauf, A., Tutz, G.: A survival tree method for the analysis of discrete event times in clinical and epidemiological studies. Stat. Med. 35(5), 734–1 (2016). https://doi.org/10.1002/sim.6729
Schmid, M., Welchowski, T., Wright, M.N., Berger, M.: Discrete-time survival forests with Hellinger distance. Data Min. Knowl. Discov. 34, 812–832 (2020). https://doi.org/10.1007/s10618-020-00682-z
Segal, M.R.: Extending the elements of tree-structured regression. Stat. Methods Med. Res. 4(3), 219–236 (1995). https://doi.org/10.1177/096228029500400304
Segal, M.R.: Features of tree-structured survival analysis. Epidemiology 8(4), 344–446 (1997)
Sleeper, L.A., Harrington, D.P.: Regression splines in the cox model with application to covariate effects in liver disease. J. Am. Stat. Soc. (1990). https://doi.org/10.1080/01621459.1990.10474965
Sparapani, R.A., Logan, B.R., McCulloch, R.E., Laud, P.W.: Nonparametric survival analysis using Bayesian Additive Regression Trees (BART). Stat. Med. 35(16), 2741–2753 (2016). https://doi.org/10.1002/sim.6893
Sparapani, R.A., Spanbauer, C., McCulloch, R.: Nonparametric machine learning and efficient computation with Bayesian additive regression trees: the BART R package. J. Stat. Software 97(1), 1–66 (2021). https://doi.org/10.18637/jss.v097.i01
Tiendrébéogo, S., Somé, B., Kouanda, S., Gbété, S.D.: Survival analysis of data in HIV infected persons receiving antiretroviral therapy using a model-based binary tree. J. Math. Stat. 15, 354–365 (2019)
Tutz, G., Schmid, M.: Modeling Discrete Time-to-Event-Data. Springer, New York, NY (2016)
van der Laan, M.J., Robins, J.M.: Unified Methods for Censored Longitudinal Data and Causality. Springer, New York (2003)
Wallace, M.L.: Time-dependent tree-structured survival analysis with unbiased variable selection through permutation tests. Stat. Med. 33(27), 4790–4804 (2014). https://doi.org/10.1002/sim.6261
Wang, H., Chen, X., Li, G.: Survival forests with R-squared splitting rules. J. Comp. Biol. 25(4), 388–395 (2018). https://doi.org/10.1089/cmb.2017.0107
Welchowski, T., Berger, M., Koehler, D., Schmid, M.: discSurv: Discrete Time Survival Analysis. R package version 2.0.0 (2022)
Willet, J.B., Singer, J.D.: Investigating onset, cessation, relapse, and recovery. J. Consult. Clin. Psychol. 61(6), 952–65 (1993). https://doi.org/10.1037/0022-006X.61.6.952
Wood, S.N.: Fast stable restricted maximum likelihood and marginal likelihood estimation of semi-parametric generalized linear models. J. R. Stat. Soc.: Ser. B (Stat. Methodol.) 73, 3–36 (2011). https://doi.org/10.1111/j.1467-9868.2010.00749.x
Wood, S.N.: Generalized Additve Models: An Introduction with R, 2nd edn. Chapman & Hall, Boca Raton, FL (2017)
Yuan, M., Lin, Y.: Model selection and estimation in regression with grouped variables. J. R. Stat. Soc.: Ser. B (Stat. Methodol.) 68(1), 49–67 (2006). https://doi.org/10.1111/j.1467-9868.2005.00532.x
Zhang, H., Singer, B.H.: Recursive Partitioning in the Health Sciences. Springer, New York, NY (1999)
Acknowledgements
Support by the German Research Foundation is gratefully acknowledged.
Funding
Open Access funding enabled and organized by Projekt DEAL. This work was supported by the German Research Foundation (DFG; Grant number BE 7543/1-1)
Author information
Authors and Affiliations
Contributions
Conceptualization: Moritz Berger; Methodology: Moritz Berger; Formal analysis and investigation: Nikolai Spuck; Writing—original draft preparation: Nikolai Spuck; Writing—review and editing: Nikolai Spuck, Matthias Schmid and Moritz Berger; Funding acquisition: Moritz Berger; Resources: Nils Heim, Ute Klarmann-Schulz and Achim Hörauf
Corresponding author
Ethics declarations
Conflict of interest
The authors have no competing interests to declare that are relevant to the content of this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Spuck, N., Schmid, M., Heim, N. et al. Flexible tree-structured regression models for discrete event times. Stat Comput 33, 20 (2023). https://doi.org/10.1007/s11222-022-10196-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11222-022-10196-x