Abstract
Spatio-temporal count data relating to a set of non-overlapping areal units are prevalent in many fields, including epidemiology and social science. The spatial autocorrelation inherent in these data is typically modelled by a set of random effects that are assigned a conditional autoregressive prior distribution, which is a special case of a Gaussian Markov random field. The autocorrelation structure implied by this model depends on a binary neighbourhood matrix, where two random effects are assumed to be partially autocorrelated if their areal units share a common border, and are conditionally independent otherwise. This paper proposes a novel graph-based optimisation algorithm for estimating either a static or a temporally varying neighbourhood matrix for the data that better represents its spatial correlation structure, by viewing the areal units as the vertices of a graph and the neighbour relations as the set of edges. The improved estimation performance of our methodology compared to the commonly used border sharing rule is evidenced by simulation, before the method is applied to a new respiratory disease surveillance study in Scotland between 2011 and 2017.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Spatio-temporal count data relating to K non-overlapping areal units for N consecutive time periods are prevalent in many fields, including epidemiology (Stoner et al. 2019) and social science (Bradley et al. 2016). The spatial and temporal correlations in these data are typically modelled by sets of random effects, that are assigned conditional autoregressive (CAR, Besag et al. 1991) and autoregressive prior distributions respectively. A large volume of research has developed models to estimate a range of spatio-temporal dynamics in such data, including spatially correlated linear time trends (Bernardinelli et al. 1995), a spatially correlated multivariate temporal autoregressive process (Rushworth et al. 2014), time period specific spatially correlated surfaces (Waller et al. 1997), and a decomposition of the data into spatial and temporal main effects and a spatio-temporal interaction (Knorr-Held 2000). Each of these approaches makes different assumptions about the spatio-temporal structure in the data, with for example Knorr-Held (2000) modelling the data as a convolution of three processes, a single spatial surface common to all time periods, a single temporal trend common to all areal units, and a set of interactions that allow for deviations from these common trends. In contrast, Waller et al. (1997) model the data as a convolution of two processes, which are a single temporal trend common to all areal units, and a separate (uncorrelated) spatial process for each time period.
Regardless of the spatio-temporal structure assumed, the random effects used to capture spatial correlation in areal unit data are typically assigned CAR-type prior distributions, which form part of the overall Bayesian hierarchical model. The spatial correlation structure implied by these CAR models depends on a \(K\times K\) geographical neighbourhood or adjacency matrix \({\mathbf {W}}\), which specifies which pairs of areal units are close together in space. A binary specification is typically adopted, where \(w_{kj}=1\) if areal units (k, j) share a common border (are spatially close), \(w_{kj}=0\) otherwise, and \(w_{kk}=0~\forall ~k\). In terms of spatial correlation, CAR models assume data in neighbouring areal units (k, j) (those with \(w_{kj}=1\)) are partially autocorrelated, while those relating to non-neighbouring areal units (k, j) (those with \(w_{kj}=0\)) are conditionally independent. Thus while the spatial autocorrelation structure implied by these CAR models depends on \({\mathbf {W}}\), the appropriateness of \({\mathbf {W}}\) for the data at hand or the sensitivity of the results to changing its specification are rarely assessed in the modelling. This is in sharp contrast to geostatistics for point level data, where variogram analysis is routinely used to identify an appropriate spatial autocorrelation structure for the data, such as assessing the validity of isotropy.
Furthermore, specifying \({\mathbf {W}}\) based on the simple border sharing rule is unlikely to provide an appropriate autocorrelation structure for the data under study, because spatial autocorrelation is unlikely to be present universally throughout the study region. Instead, there are likely to be pairs of neighbouring areal units that exhibit large differences between their data values, which can be driven by complex environmental and/or social process (Mitchell and Lee 2014). Examples that illustrate this phenomenon include the fields of spatial clustering (Knorr-Held and Raßer 2000) and boundary analysis (Lu and Carlin 2005), where identifying the locations of these differences (step-changes) is of primary interest. These two fields differ in the unit of primary interest, because in spatial clustering groups of areas are identified as being similar or different, while in boundary analysis it is the differences between each pair of geographically adjacent data values that is estimated to be either large (a boundary) or small (no boundary).
Numerous approaches have been proposed for identifying spatial step-changes in areal unit count data, including specifying piecewise constant mean models for clustering (e.g. Knorr-Held and Raßer 2000), and treating each \(w_{kj}\) element that corresponds to a pair of neighbouring areal units as a binary random quantity for boundary analysis. The latter allows one to estimate the spatial partial autocorrelation structure in the data, and Ma et al. (2010) model each \(w_{kj}\) as a Bernoulli random variable. However this suffers from parameter identifiability problems, because there are many more elements in \({\mathbf {W}}\) to estimate than there are areal units (data points). A solution to this proposed by Lee and Mitchell (2012) modelled the elements in \({\mathbf {W}}\) with a simple log-linear parametric regression model, where the covariates in the model are measures of the dissimilarity between geographically adjacent areas. This approach was extended to the spatio-temporal domain for normal, probit and tobit data models by Berchuck et al. (2019), by novelly including temporally varying regression parameters and thus allowing \({\mathbf {W}}\) to change over time. However, this class of regression models for \({\mathbf {W}}\) may be restricted in its estimation of \({\mathbf {W}}\) by the parametric nature of the model and the availability of covariates quantifying the dissimilarities between adjacent areal units.
Therefore this paper proposes a novel graph-based optimisation algorithm for estimating an appropriate neighbourhood matrix for the data, which overcomes the two parameterisation issues highlighted above. Our approach either allows the estimated neighbourhood matrix to be static over time which we denote by \({\mathbf {W}}_{E}\), or evolve dynamically over time t which we denote by \({\mathbf {W}}_{E_t}\). In either case its estimation is based on an initial graph G, where the K areal units comprise the vertex-set V(G), and the edge-set E(G) is defined by \({\mathbf {W}}\) via \(E(G)=\{(k,j)|w_{kj}=1\}\) (so \({\mathbf {W}}\) is the adjacency matrix of G). The algorithm estimates whether each edge in the graph should be removed or not, with the mild restriction that every vertex must retain at least one incident edge. Our novel estimation algorithm thus has two stages, the first of which estimates \({\mathbf {W}}_{E}\) or \({\mathbf {W}}_{E_t}\) from the data after covariate effects have been accounted for, which is akin to using variogram analysis on detrended geostatistical data to estimate an appropriate correlation structure. The second stage of our estimation algorithm fits a Poisson log-linear model with spatio-temporally correlated random effects to the count data based on \({\mathbf {W}}_{E}\) or \({\mathbf {W}}_{E_t}\), with inference in a Bayesian paradigm using integrated nested Laplace Approximations (INLA, Rue et al. 2009).
The general Poisson log-linear count data model we use is presented in Sect. 2, while our novel graph-based optimisation algorithm is described in Sect. 3. Section 4 presents a simulation study showing that using (\({\mathbf {W}}_{E}\), \({\mathbf {W}}_{E_t}\)) outperforms using \({\mathbf {W}}\) in terms of the estimation of key quantities of interest as long as there are at least \(N=3\) time periods of data. In Sect. 5 our approach is applied to a new respiratory disease surveillance study in Scotland, which in addition to showing better model fit and more precise inference compared to using \({\mathbf {W}}\), allows additional inferences to be made on the locations of boundaries in data. Finally, Sect. 6 concludes the paper.
2 Spatio-temporal count data modelling
The outcome variable \(Y_{kt}\) is a spatio-temporally aggregated count of the number of events that occur in areal unit \(k=1,\ldots ,K\) during time period \(t=1,\ldots ,N\), and is accompanied by a vector of p covariates \({\mathbf {x}}_{kt}\) and an expected count \(e_{kt}\). The latter allows for the fact that the areal units have different population sizes and age-sex demographics, which thus affects the observed count. A general Bayesian hierarchical model for these data is given by
where throughout this paper \(\text{ N }(a,b)\) denotes a normal distribution with mean a and variance b. Here \(\theta _{kt}\) denotes the risk or rate of the outcome variable \(Y_{kt}\) relative to the expected count \(e_{kt}\), and the spatio-temporal variation in this risk (rate) is modelled by covariates \(\{{\mathbf {x}}_{kt}\}\) and random effects \(\{\psi _{kt} = \phi _{kt} + \delta _{t}\}\). The covariate regression parameters \(\varvec{\beta }=(\beta _1,\ldots , \beta _p)\) are assumed to be a-priori independent of each other, and each \(\beta _j\) is assigned an independent weakly informative zero-mean Gaussian prior distribution with a large variance, to ensure the data play the dominant role in estimating its value.
An appropriate random effects structure depends on both the residual spatio-temporal structure in the data and the goal of the analysis. Here we utilise the general spatio-temporal structure proposed by Waller et al. (1997), which models the data with an overall temporal trend \(\varvec{\delta }=(\delta _1,\ldots ,\delta _N)\) common to all areal units, and a separate spatial surface \(\varvec{\phi }_t=(\phi _{1t},\ldots ,\phi _{Kt})\) that is independent for each time period t. We adopt this structure over the alternatives described in the introduction because it does not enforce any temporal smoothing constraints on the residual spatial surfaces as Bernardinelli et al. (1995) and Rushworth et al. (2014) do, thus allowing them to change over time. We model the temporal trend by the first order autoregressive process:
The prior distributions and their parameterisations (e.g. via the precision \(\sigma ^{-2}\)) are chosen to be weakly informative, and are the default specifications suggested by the INLA software (Rue et al. 2009) that we use for inference. We model the residual spatial trend for time period t using the conditional autoregressive prior proposed by Leroux et al. (2000) which is given by
where \(\varvec{\phi }_{-kt}=(\phi _{1t},\ldots ,\phi _{k-1,t}, \phi _{k+1,t},\ldots , \phi _{Kt})\). Spatial autocorrelation is induced into these random effects by the neighbourhood matrix \({\mathbf {W}}\), and we adopt the commonly used binary border sharing definition described in the introduction. The level of spatial dependence at time t is controlled globally by \(\rho _t\), with \(\rho _t=0\) corresponding to spatial independence (as \(\phi _{kt}\) in (3) no longer depends on its neighbours), while if \(\rho _t=1\) then (3) becomes the intrinsic CAR model for strong spatial autocorrelation proposed by Besag et al. (1991). A weakly-informative normal prior on the logit scale is specified for the spatial dependence parameter \(\rho _t\), while a weakly informative log-gamma prior is specified for the log of the spatial precision \(\tau _t^{-2}\), again following the defaults suggested by the INLA software. The partial spatial autocorrelation structure implied by this model is given by
where \({\mathcal {C}}_{kj,t}=\text{ Corr }(\phi _{kt}, \phi _{jt} |\varvec{\phi }_{-kjt})\) and \(\varvec{\phi }_{-kjt} = \varvec{\phi }_{t}\setminus \{\phi _{kt}, \phi _{jt}\}\). Thus \({\mathbf {W}}\) controls the partial spatial autocorrelation structure in \(\varvec{\phi }_t\), because if \(w_{kj}=1\) then \((\phi _{kt}, \phi _{jt})\) are partially autocorrelated with the strength of that autocorrelation controlled globally for all pairs of neighbouring areas by \(\rho _t\), whereas if \(w_{kj}=0\) then \((\phi _{kt}, \phi _{jt})\) are conditionally independent. Thus while \({\mathbf {W}}\) is crucial to the model because it determines the spatial autocorrelation structure in the data, its appropriateness for the data or the sensitivity of the results to changing its specification are rarely assessed. Furthermore, specifying \({\mathbf {W}}\) via border sharing implies that all pairs of geographically adjacent areal units will have correlated random effects, which precludes the identification of boundaries in the spatial surface. Therefore in the next section we propose a novel graph-based optimisation algorithm for estimating a more appropriate neighbourhood matrix for the data that leads to improved inference.
3 Methodology
We propose a novel two-stage approach for jointly estimating the parameters from (1)–(3) denoted by \(\varvec{\varTheta }=(\varvec{\beta },\varvec{\delta }, \sigma ^2, \alpha , \varvec{\phi }_1,\ldots ,\varvec{\phi }_N, \rho _1,\ldots ,\rho _N, \tau _1^{2},\ldots ,\tau _N^{2})\) and an appropriate neighbourhood matrix (matrices) for the data, which extends the current approach of using \({\mathbf {W}}\) constructed from the border sharing rule. We propose approaches for estimating both static (\({\mathbf {W}}_E\)) and time-varying (\({\mathbf {W}}_{E_{t}}\)) neighbourhood matrices, where for the former \({\mathbf {W}}_E\) is used in (3) for all time periods t while for the latter a separate \({\mathbf {W}}_{E_{t}}\) is used in (3) for each time period t. In stage 1 we estimate (\({\mathbf {W}}_{E}\), \({\mathbf {W}}_{E_{t}}\)) using a graph-based optimisation algorithm, and in stage 2 we estimate the posterior distribution
\(f(\varvec{\varTheta } |{\mathbf {W}}_{E}, {\mathbf {Y}})\) or \(f(\varvec{\varTheta } |{\mathbf {W}}_{E_1},\ldots ,{\mathbf {W}}_{E_N}, {\mathbf {Y}})\) conditional on the estimated neighbourhood matrices. Our methodology thus brings areal unit modelling into line with standard practice in geostatistical modelling, which is to first estimate a trend model and then identify an appropriate autocorrelation structure via residual analysis.
3.1 Stage 1: Estimating \({\mathbf {W}}_{E}\) or \(({\mathbf {W}}_{E_1},\ldots ,{\mathbf {W}}_{E_N})\)
3.1.1 Estimating the residual spatial structure
The random effects \(\varvec{\phi }_{t}\) model the residual variation in the data at time t after the effects of the covariates have been removed, so the first step to estimating \({\mathbf {W}}_{E}\) or \(({\mathbf {W}}_{E_1},\ldots ,{\mathbf {W}}_{E_N})\) is to estimate this residual structure in the data. The count data model (1) has expectation \({\mathbb {E}}[Y_{kt}]=e_{kt}\exp ({\mathbf {x}}_{kt}^{\top }\varvec{\beta } + \phi _{kt} + \delta _t)\), which can be re-arranged to give
The latter approximation replaces the unknown \({\mathbb {E}}[Y_{kt}]\) with the observed data \(Y_{kt}\), and the temporal random effects \(\{\delta _t\}\) are removed as they are constant over space and hence do not impact on the estimation of the spatial correlation structure. The regression parameters \(\varvec{\beta }\) are estimated in this initial stage from a simpler model with no random effects (i.e. \(\{\phi _{kt}, \delta _t\}\) are removed from (1)) using maximum likelihood estimation, and are denoted by \(\hat{\varvec{\beta }}\). We consider two cases for how to use these residuals \(\{\phi _{kt}\}\) in our graph-based optimisation algorithm.
Case A: Static \({\mathbf {W}}_{E}\)
If the residual spatial surfaces are similar over time, then estimating a single \({\mathbf {W}}_{E}\) common to all time periods is appropriate. In this case we estimate a single residual spatial surface by averaging the residuals over the N time periods, that is
and use this single residual spatial surface in our graph-based optimisation algorithm.
Case B: Temporally varying \(({\mathbf {W}}_{E_1},\ldots ,{\mathbf {W}}_{E_N})\) If the residual spatial surface evolves significantly over time, then an appropriate neighbourhood structure will also evolve over time. The simplest approach is to apply the graph-based optimisation algorithm to the residuals \((\hat{\phi }_{1t},\ldots , \hat{\phi }_{Kt})\) from (5) separately for each time period t, yielding a separate matrix \({\mathbf {W}}_{E_t}\) for each time period. However, as we show in the simulation study (Sect. 4.2) multiple realisations of the residual spatial surface are required to estimate \({\mathbf {W}}_{E_t}\) well, which makes this simple approach inappropriate. Therefore instead we estimate the residual spatial surface for time t using a \(2q+1\) time period moving average of the residuals from (5), that is
with appropriate adjustments for the end time periods. For example, if \(q=1\) then for \(t=1\), \({\tilde{\phi }}_{k}=(1/3)\sum _{r=1}^{3}\hat{\phi }_{kr}\) and for \(t=N\), \({\tilde{\phi }}_{k}=(1/3)\sum _{r=N-2}^{N}\hat{\phi }_{kr}\). Thus in the static case A we estimate \({\mathbf {W}}_E\) from a set of spatial residuals \(\tilde{\varvec{\phi }}=({\tilde{\phi }}_1,\ldots , {\tilde{\phi }}_K)\) computed from all the data, while in the time-varying case B we estimate \({\mathbf {W}}_{E_t}\) separately for each time period t using a set of spatial residuals \(\tilde{\varvec{\phi }}=({\tilde{\phi }}_{1},\ldots , {\tilde{\phi }}_{K})\) that is computed separately for each year t.
3.1.2 Deriving an objective function to optimise
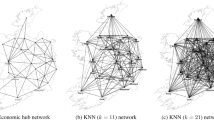
The CAR model (3) represents a graph G whose vertex-set V(G) is the set of K areal units, and whose edge-set is \(E(G)=\{(k,j)|w_{kj}=1\}\), a subset of un-ordered pairs of elements of V(G). In graph theoretic terms G is the simple graph with adjacency matrix \({\mathbf {W}}=(w_{kj})\) defined by the border sharing rule, and the graph for the Scotland respiratory disease study presented in Sect. 5 is shown in panel (a) of Fig. 1. Given \(\tilde{\varvec{\phi }}\) we estimate \({\mathbf {W}}_{E}\) or \({\mathbf {W}}_{E_t}\) by searching for a suitable subgraph of G that maximises the value of an objective function \(J(\tilde{\varvec{\phi }})\), and the sub-graph corresponding to \({\mathbf {W}}_{E}\) that was estimated for the Scotland study is presented in panel (b) of Fig. 1. This estimated graph has 47% fewer edges compared with the border sharing graph, and 90% of the vertices have had at least one edge removed.
We base the objective function on the natural log of the product of full conditional distributions \(f({\tilde{\phi }}_{k}|\tilde{\varvec{\phi }}_{-k})\) from (3) over all spatial units \(k=1,\ldots ,K\). We additionally enforce the restriction that \(\rho _t=1\), because from (4) this ensures strong spatial autocorrelation globally that can be altered locally by removing edges when estimating \({\mathbf {W}}_E\) or \({\mathbf {W}}_{E_t}\). These removed edges correspond to boundaries in the random effects surface, because if \(w_{E_{kj}}=0\) the corresponding random effects are conditionally independent. Thus dropping the subscript t for notational simplicity as we are dealing with a purely spatial objective function and fixing \(\rho =1\) in (3) as described above, we obtain the following objective function
We base the objective function on a product of full conditional distributions rather than the joint distribution for \(\tilde{\varvec{\phi }}\), because when \(\rho =1\) the latter is not a proper distribution as its precision matrix is singular. One could use the joint probability density function up to a proportionality constant but this leads to all edges being removed by the algorithm, and details are given in Sect. 2 of the supplementary information. As (8) depends on the unknown variance parameter \(\tau ^2\), we profile it out by maximising \(J(\tilde{\varvec{\phi }})\) with respect to \(\tau ^2\) which gives \(\hat{\tau }^2= \sum _{k=1}^{K}\left( \sum _{j=1}^{K}w_{kj}\right) \left( {\tilde{\phi }}_k - \frac{\sum _{r=1}^{K}w_{kr}{\tilde{\phi }}_{r}}{\sum _{r=1}^{K}w_{kr}}\right) ^{2} \Bigg / K\). This estimator is then plugged into (8) to yield the final objective function
3.1.3 Graph-based optimisation
Let H be generic notation for any graph, then we use the following graph theoretic terminology in this section: (i) we write uv for the edge \(\{u,v\}\) with endpoints u and v; (ii) an edge \(e \in E(H)\) is said to be incident with a vertex \(v \in V(H)\) if v is an endpoint of e; (iii) the number of edges in H incident with any single vertex v, written \(d_H(v)\), is called the degree of v in H; (iv) we write \(N_H(v)\) for the set \(\{u \in V(H) \setminus \{v\}: uv \in E(H)\}\) of neighbours of v in H; (v) a graph \(H'\) is a subgraph of H if \(V(H') \subseteq V(H)\) and \(E(H') \subseteq E(H)\); and (vi) if \(H'\) is a subgraph of H and these two graphs also have the same vertex set, we say that \(H'\) is a spanning subgraph of H.
The graph G based on \({\mathbf {W}}\) has vertex-set V(G) and edge-set E(G), and we assume that edges \(e \in E(G)\) can be removed from the graph but that new edges cannot be added in. This means that one can estimate \(w_{E_{kj}}=\{0,1\}\) if \(w_{kj}=1\), but if \(w_{kj}=0\) then \(w_{E_{kj}}\) remains fixed at zero. Additionally, we assume that each area (vertex) must retain at least one edge in the graph, which corresponds to the constraint \(\sum _{j=1}^{K}w_{E_{kj}}>0\) for all k. This ensures that we do not divide by 0 in (9). Let \(f(H,\tilde{\varvec{\phi }})\) denote the value of \(J(\tilde{\varvec{\phi }})\) corresponding to \({\mathbf {W}}_H\), the adjacency matrix corresponding to the sub-graph H of G. Then the goal of our optimisation problem can be phrased as finding a spanning subgraph \({\tilde{G}}\) of G, with minimum degree at least one, which maximises \(f({\tilde{G}}, \tilde{\varvec{\phi }})\).
This graph optimisation problem is known to be NP-hard (Enright et al. 2021), and so is extremely unlikely to admit an exact algorithm which will terminate in polynomial time on all possible inputs. Moreover, this intractability result holds even if we assume that the input graph G is planar; our input graph is necessarily planar because it is derived from the adjacencies of non-overlapping regions in the plane. In this work we therefore adopt a heuristic local search approach, which we emphasise is not guaranteed to find the global optimal solution. We leave a more in-depth study of the existence or otherwise of algorithms with provable performance guarantees for future work.
A brute force optimisation strategy would consider all possible subsets of edges to delete (which is exponential in the number of edges in the original graph), and choose the one which maximises the objective function. However such a running-time is already infeasible in our relatively small simulation study example which has 671 edges. To avoid this, we instead obtain an improved matrix \({\mathbf {W}}_{E}\) by carrying out a sequence of local optimisation operations, which is much faster.
While many heuristic graph searching algorithms exist, we were unable to identify any existing approaches which can be applied directly to this optimisation problem. The objective function (9) considered here has the unusual feature that it contains both a log-of-sums over vertices and a sum-of-logs: the way in which these interact makes it trivial to find examples where an optimal subgraph may no longer be optimal if a disconnected and isolated edge is added elsewhere in the graph. This subtlety rules out any exact or heuristic local search method. Additionally, the nature of the objective function rules out any heuristic that relies on a matrix representation of the objective function as well as other common techniques from operational research.
The starting point for our bespoke heuristic local optimisation is the following fairly standard approach. We consider the vertices of the graph in some fixed order, and attempt to optimise the set of edges incident with each vertex in turn. Since the effect of deleting the edge uv depends on the set of edges incident at both u and v, we have to decide whether or not to retain each edge incident with v without knowing precisely what effect this will have (as the neighbourhood of any neighbour u of v may not yet be fixed). We therefore decide whether or not to delete an edge by considering the difference between the contribution to the objective function from u (respectively v) from the best possible set of incident edges at u (respectively v) that does include the edge uv, and the best possible set that does not include this edge.
In order to apply this strategy, we need to express the objective function as a sum of contributions associated with each vertex of the graph, so that we can assess the impact of making local changes associated with an individual vertex; the main novelty of our approach lies in this derivation of a suitable bound on the contribution from each vertex that can be computed locally. As a first step, we reformulate (9) in more graph theoretic notation. To do this, we set \(V = V(G)\) (observing that we use the same vertex set throughout), and note that \(|V| = K\). For the vertex v corresponding to region k in the matrix, we set \({\tilde{\phi }}_v = {\tilde{\phi }}_k\). This gives
To simplify notation, we will write \({{\,\mathrm{ND}\,}}_H(v,\tilde{{\phi }})\) for the neighbourhood discrepancy defined as
It is now clear that, to maximise the right-hand side of (10), on the one-hand we would like to retain as many edges as possible to maximise the first term, but on the other hand we minimise the second term by deleting edges to decrease the neighbourhood discrepancy at each vertex. We can now associate with a given vertex v the following contribution, \({{\,\mathrm{cont}\,}}(v,H,\tilde{\varvec{\phi }})\), to the right-hand side of (10):
We then have that \(f(H,\tilde{\varvec{\phi }}) \propto \sum _{v \in V} {{\,\mathrm{cont}\,}}(v,H,\tilde{\varvec{\phi }})\). The remaining barrier to using this expression to carry out locally optimal modifications is that the value of \(\sum _{w \in V \setminus \{v\}} d_H(w) {{\,\mathrm{ND}\,}}_H(w,\tilde{{\phi }})\) depends on the entire graph, not just the edges incident with v, so we cannot compute the value of \({{\,\mathrm{cont}\,}}(v,H,\tilde{{\phi }})\) knowing only the neighbours of v in H. To deal with this, we define the adjusted contribution of v in H, with respect to a second graph \(H'\):
Observe that, if H is a spanning subgraph of \(H'\), we have \(\sum _{v \in V} \ln (d_H(v)) \le \sum _{v \in V} \ln (d_{H'}(v))\) and so, if \(f(H,\tilde{{\phi }}) > f(H',\tilde{{\phi }})\), we must have
This tells us that, if \({{\,\mathrm{adjcont}\,}}_H(v,H \setminus \{e\},\tilde{{\phi }})\) is strictly greater than \({{\,\mathrm{adjcont}\,}}_H(v,H,\tilde{{\phi }})\), then the contribution at v is still increased by deleting e even when deletions are also carried out elsewhere in the graph to decrease the weighted sum of neighbourhood discrepancies.
These observations motivate our iterative approach. At the first step we consider the first vertex v and use the original graph G to identify a set of edges incident with v to delete (by considering the best possible adjusted contribution with respect to G that can be achieved at both endpoints of the edges in question). We then delete these edges to obtain a new graph \(G'\) and continue with the next vertex, this time considering the adjusted contribution with respect to \(G'\). We continue in this way, returning to the start of the vertex list when we reach the end, until we complete a pass through all remaining feasible vertices (that is, those which still have more than one neighbour in the modified graph) without identifying any deletions that increase the objective function.
The algorithm is summarised in pseudocode as Algorithm 1 in the appendix. We note that the running-time depends exponentially on the maximum degree (as we consider all possible subsets of neighbours to retain at each vertex in order to identify the “best” possible neighbourhood), but only polynomially on the number of edges. It is not unreasonable to expect that the maximum degree will in practice be small compared with the total number of vertices or edges: it is unlikely that any one areal unit will border a very large number of other units (in our simulation study example the maximum degree is 22). Software to implement the optimisation are available in the R spatio-temporal modelling package CARBayesST (Lee et al. 2018). We discuss the performance of this algorithm (and several variations) on our example inputs in the appendix.
3.2 Stage 2: Estimating \(\varvec{\varTheta }\) given \({\mathbf {W}}_{E}\) or \(({\mathbf {W}}_{E_1},\ldots ,{\mathbf {W}}_{E_N})\)
We fit model (1)–(3) with \({\mathbf {W}}_{E}\) or \(({\mathbf {W}}_{E_1},\ldots ,{\mathbf {W}}_{E_N})\) replacing \({\mathbf {W}}\) in a Bayesian setting using integrated nested Laplace approximations (INLA, Rue et al. 2009). We use INLA due to its computational speed in fitting the models, but we could have used Markov chain Monte Carlo (MCMC) simulation methods, for example using the CARBayesST package.
4 Simulation study
This section presents a simulation study that compares the performance of model (1)–(3) when it is used in conjunction with neighbourhood matrices specified by: (i) the border sharing rule (denoted \({\mathbf {W}}\)); (ii) graph-based optimisation and forced to be static over time (denoted \({\mathbf {W}}_E\)); and (iii) graph-based optimisation and allowed to vary over time (denoted \({\mathbf {W}}_{E_t}\)). For the latter we use a 3-year moving average to estimate \({\tilde{\phi }}_{kt}\), (that is \(q=1\)) as this allows the most variation in \({\mathbf {W}}_{E_t}\) over time. Additionally, we also fit model (1) with the adjustment that the spatial random effects for each time period are modelled by the CAR prior proposed by Besag et al. (1991), because it is the most commonly used CAR model in the literature. This model is denoted by BYM in what follows, and is described in Sect. 3 of the supplementary information.
4.1 Data generation
The study region is the \(K=257\) Intermediate Zones (IZ) that make up the Greater Glasgow and Clyde Health Board in Scotland, which is the largest health board (and contains the largest city) in our mainland Scotland case study presented in Sect. 5. We choose this region as the template for this study due to the large number of simulated data sets and models we run, which would be computationally prohibitive to do for all of mainland Scotland.
Count data are generated for this region from model (1), and we consider scenarios with differing numbers of time periods (denoted by N) to see how this affects the performance of our methodology. We also examine how the size of the counts \(\{Y_{kt}\}\) affects estimation performance, by considering scenarios where the expected counts \(\{e_{kt}\}\) are drawn uniformly within the ranges: (i) [10, 30] (rare events); and (ii) [150, 250] (common events). Finally, we also vary the sizes of the boundaries we generate in the residual surface \(\varvec{\phi }_t\).
Each simulated data set includes an independent (\({\mathbf {x}}_1\)) and a spatially autocorrelated (\({\mathbf {x}}_2\)) covariate, and the corresponding regression parameters are fixed at \(\beta _1=\beta _2=0.05\). Both covariates are generated from zero-mean multivariate normal distributions with a standard deviation of 0.5 separately for each time period, with the independent covariate \({\mathbf {x}}_1\) having the identity correlation matrix. The correlation matrix for \({\mathbf {x}}_2\) is defined by the spatial exponential correlation matrix \(\varvec{\varSigma }=\exp (-\xi {\mathbf {D}})\), where \({\mathbf {D}}\) is a \(K\times K\) distance matrix between the centroids of the K IZs. The spatial range parameter \(\xi \) was chosen to ensure the covariate was visually spatially smooth, which was achieved by fixing \(\xi \) so that the mean correlation across all pairs of IZs was 0.25.
Temporal autocorrelation is induced into each simulated data set by a first order autoregressive process, with AR(1) coefficient \(\alpha =0.8\). Similarly, spatial autocorrelation is induced via a multivariate normal distribution with a spatial exponential correlation matrix \(\varvec{\varSigma }=\exp (-\xi {\mathbf {D}})\), where \(\xi \) was chosen so that the mean pairwise correlation across all IZs was 0.15. We consider scenarios in this study where the locations of the boundaries are either static or vary over time, and the generation of the random effects are described in both cases below.
Case A: Static boundaries
When the boundaries are static the residual spatial surfaces \(\varvec{\phi }_t\) are similar for each time period, and are generated as \(\varvec{\phi }_t=\varvec{\phi } + \varvec{\phi }^{*}_t\). This has a common spatial surface \(\varvec{\phi }\) for all time periods and time period specific deviations \(\varvec{\phi }^{*}_t\) with a lower variance, the latter ensuring the random effects surfaces are similar but not identical over time. Boundaries are induced into the spatial surface through the mean of \(\varvec{\phi }\), which is denoted by \(\varvec{\mu }\). This mean is a piecewise constant surface with 3 distinct values \((-\lambda , 0, \lambda )\), where \(\lambda \) determines the size of the boundaries and larger values of \(\lambda \) result in larger boundaries. Thus two neighbouring areas (k, j) that have the same mean (i.e. \(\mu _k=\mu _j\)) will have no boundary between them, while if \(\mu _k\ne \mu _j\) then their simulated random effects \((\phi _k,\phi _j)\) will be very different corresponding to a boundary between these areas. The values of \((-\lambda , 0, \lambda )\) are assigned to the mean vector \(\varvec{\mu }\) to match the spatial structure of the case study data as closely as possible, with for example IZs that exhibit comparatively high rates \(\{\theta _{kt}\}\) being assigned a mean value of \(\lambda \). The template used in the simulation study for generating spatially correlated data with boundaries, as well as example realisations of the random effects surfaces generated are presented in Sect. 4 of the supplementary information.
Case B: Time-varying boundaries
Spatio-temporal random effects \(\{\phi _{kt}\}\) with temporally varying boundaries are generated in a similar way to the static boundary case described above. Specifically, we generate \(\varvec{\phi }_t=\varvec{\phi } + \varvec{\phi }^{*}_t\), where the temporally constant component \(\varvec{\phi }\) is generated from a mean-zero multivariate normal distribution with a spatially correlated covariance matrix as described above. The temporally varying component \(\varvec{\phi }^{*}_t\) is generated via the same mechanism but with a mean \(\varvec{\mu }_{t}\), which again is a piecewise constant surface with only 3 distinct values \((-\lambda , 0, \lambda )\). This time the allocation of these three values to each component of the mean vector varies over time, resulting in boundaries that also vary over time. This variation in mean \(\varvec{\mu }_t\) is controlled so that either 5%, 10% or 15% of the boundaries in the random effects surfaces change from one time period to the next.
4.2 Results: Spatial data (\(N=1\))
We first examine how well our graph-based optimisation algorithm works for purely spatial data with only \(N=1\) time period, by generating data under 4 different scenarios that include all pairwise combinations of: (i) \(e_{kt}\in [10, 30]\) (rare events) and \(e_{kt}\in [150, 250]\) (common events); and (ii) \(\lambda =0.25\) (small boundaries) and \(\lambda =0.5\) (large boundaries). One hundred data sets are generated under each of these 4 scenarios, and the results are displayed in Sect. 5 of the supplementary information. These results relate to the accuracy with which the risks (rates) \(\{\theta _{kt}\}\) and the covariate effects \((\beta _1, \beta _2)\) can be estimated using each specification of the neighbourhood matrix, as well as the accuracy of the boundary detection when using \({\mathbf {W}}_E\). Overall, using \({\mathbf {W}}_{E}\) leads to slightly worse results than using \({\mathbf {W}}\), with slightly larger RMSEs and reduced coverage percentages. This general poor performance of \({\mathbf {W}}_{E}\) occurs because when \(N=1\) the estimate \(\tilde{\varvec{\phi }}\) used in \(J(\tilde{\varvec{\phi }})\) is only based on one realisation of the residual spatial surface, which is contaminated with noise and hence not a good estimate of the true residual spatial structure in the data.
4.3 Results: Spatio-temporal data (\(N>1\))
The main results for the simpler static boundaries (Case A) are presented below, while those relating to time-varying boundaries (Case B) are presented in Sect. 7 of the supplementary information. For Case A one hundred data sets are generated under each of 12 scenarios, which include all possible combinations of: (i) \(N=3,5,7\); (ii) \(e_{kt}\in [10, 30], [150, 250]\); and (iii) \(\lambda =0.25,0.5\). The accuracy of the risk (rate) estimates \(\{\hat{\theta }_{kt}\}\) from all 4 models compared in this study are summarised in Table 1, while the accuracy of the boundary identification from using \(({\mathbf {W}}_{E}, {\mathbf {W}}_{E_t})\) is summarised in Table 2. In contrast, to preserve space the accuracy of the covariate effect estimates \(\hat{\varvec{\beta }}\) are presented in Sect. 6 of the supplementary information.
Table 1 displays the root mean square errors (RMSE) of the risk (rate) estimates \(\{\hat{\theta }_{kt}\}\), as well as the coverage percentages and average widths of the associated 95% credible intervals. The table shows clear evidence of improved risk (rate) estimation when using \({\mathbf {W}}_E\) (and \({\mathbf {W}}_{E_{t}}\)) compared with using \({\mathbf {W}}\), which is consistent over all sizes of count data (controlled by \(e_{kt}\)), boundary sizes (controlled by \(\lambda \)) and numbers of time points (N) considered in this study. This improved performance is evidenced by lower RMSE values and narrower 95% credible intervals, the latter being achieved while retaining coverage percentages around 95%. The RMSE when using \({\mathbf {W}}_E\) compared with using \({\mathbf {W}}\) reduces by between 6.6% and 24.0%, while the credible intervals are narrower by between 9.5% and 19.3%. The reduced widths of the 95% credible intervals when using \({\mathbf {W}}_{E}\) is because this matrix does not enforce correlation between neighbouring areas that exhibit a boundary between them. Thus the spatial variance \(\tau ^2_t\) is not inflated to account for the spatial smoothing that is enforced between those areal units with very different data values.
The commonly used BYM model (based on \({\mathbf {W}}\)) performs almost identically to the Leroux CAR model based on \({\mathbf {W}}\), which corroborates an existing comparison of these models by Lee (2011). Furthermore, using \({\mathbf {W}}_E\) rather than \({\mathbf {W}}_{E_{t}}\) leads to better estimation performance, which is not surprising given the boundaries do not change over time. The model using \({\mathbf {W}}_E\) provides better RMSE values as the number of time periods increases, which occurs because \(\tilde{\varvec{\phi }}\) is estimated using more replications of the spatial surface which in turn leads to better estimation of \({\mathbf {W}}_E\). The estimation of \(\{\theta _{kt}\}\) also improves as \(e_{kt}\) increases, which again is due to an increased amount of data (more events) upon which to base inference.
Table 2 shows the sensitivity and specificity of the boundary identification from using \(({\mathbf {W}}_{E}, {\mathbf {W}}_{E_t})\), where the sensitivity is the percentage of the true boundaries identified as such, while the specificity is the percentage of the non-boundaries correctly identified as such. The results show that the sensitivity ranges between 67.1% and 97.2%, and is substantially higher if the count data are not rare (i.e. \(e_{kt}\in [150,250]\)) and the boundaries are larger as expected. In contrast the specificity ranges between 61.7% and 74.8%, and again increases for less rare data with larger boundaries. The sensitivity is also higher than the specificity in all cases, suggesting that \(({\mathbf {W}}_{E}, {\mathbf {W}}_{E_t})\) tend to identify too many rather than too few boundaries on average. However, this is likely to be an artifact of the data generating process, because the boundaries regarded as ‘true’ are only those defined by the spatially varying mean of the random effects \(\varvec{\mu }\). In contrast, the independent random variation induced into the count data by generating \(\{Y_{kt}\}\) from the Poisson model may induce extra boundaries not classified as ‘true’ in the above table, leading to the lower specificity.
5 Respiratory disease in Scotland
The methodology proposed in this paper was motivated by a new study investigating the spatio-temporal dynamics of respiratory disease risk in mainland, Scotland, which exhibits some of the poorest health in western Europe (Walsh et al. 2016). We focus on respiratory disease because it is one of the leading causes of death in Scotland, and our aim is to answer the following key public health questions.
-
1.
What effects do socio-economic deprivation and air pollution concentrations have on respiratory disease risk?
-
2.
Which areas exhibit substantially elevated disease risks and to what extent do these high risk areas change over time?
-
3.
Where are the boundaries in the risk surface that separate geographically adjacent areas that have very different risks?
In answering the second and third questions we focus attention on the Greater Glasgow and Clyde health board region, because Glasgow exhibits the highest disease risks in Scotland (see Fig. 2), and also because its small geographical scale means the risk maps and boundaries are much easier to see at the small area scale compared to the equivalent Scotland-wide maps.
5.1 Data description
Data were collected summarising the yearly numbers of respiratory hospitalisations (ICD-10 codes J00 - J99) between 2011 and 2017 in each of the \(K=1,252\) Intermediate Zones (IZ) that make up mainland Scotland. These yearly disease counts \(\{Y_{kt}\}\) are accompanied by expected counts \(\{e_{kt}\}\) computed using indirect standardisation, which allow for the different population demographics in each IZ. The commonly used exploratory estimate of disease risk \(\theta _{kt}\) is the standardised morbidity ratio (SMR) computed as SMR\(_{kt}=Y_{kt} / e_{kt}\), and SMRs that are respectively greater/less than one indicate IZs that exhibit respectively higher/lower risks than the Scottish average over the study duration.
The temporal (a) and spatial (b) trends in the SMR are displayed in Fig. 2, where in panel (a) jittering has been added to the Year direction to improve the visibility of the points, and a trend line has been estimated using LOESS smoothing. Panel (a) shows a small increasing trend in the SMR over time, with average SMRs of 0.91 (a 9% decreased risk) in 2011 and 1.10 (a 10% increased risk) in 2017 compared to the Scottish average. Panel (b) displays the spatial pattern in the overall SMR across the 7-year period (i.e. SMR\(_k=\sum _{t=1}^{7}Y_{kt} / \sum _{t=1}^{7}e_{kt}\)), which shows that the high risk areas are mostly situated in Glasgow and south west Scotland, with Dundee in the east also showing elevated risks.
We obtained covariate data to explain the spatio-temporal pattern in disease risk, the most important of which is the Scottish Index of Multiple Deprivation (SIMD, http://www.gov.scot/Topics/Statistics/SIMD). Deprivation (poverty) is a key driving factor in population level ill health (NHS Health Scotland 2016), and is commonly used as a proxy measure for smoking as real smoking data are unavailable. The SIMD is not computed each year using the same methodology, so here we use the index for 2016 as a purely spatial covariate. The SIMD is a composite index comprising indicators relating to access to services, crime, education, employment, health, housing, and income, and we consider each of these as possible covariates except for health as our outcome variable is health related. Finally, the income, employment, and education domains are all collinear, having pairwise correlations between 0.87 and 0.98.
We also obtained data on fine particulate matter air pollution, called PM\(_{2.5}\), because existing studies have associated it with respiratory ill health in Scotland (Lee et al. 2019). The data come from the Pollution Climate Mapping (PCM) model (https://uk-air.defra.gov.uk/data/pcm-data), because measured data are not available at our small area IZ scale. The model produces annual average concentrations on a 1\(km^2\) grid, which are spatially realigned to the IZ scale by averaging.
5.2 Modelling
We first built a mean-model for the data using just the covariates (i.e. model (1) with no random effects), which allows us to estimate the residual spatial structure via (5). Initially, we included the three collinear SIMD indicators (education, employment and income) in separate models with the remaining covariates, and the model with income had the lowest AIC and was thus retained with the education and employment variables not being considered further. The remaining covariates (crime, housing, access and PM\(_{2.5}\)) exhibited significant effects at the 5% level from this simple model and were hence retained. The residuals from this model display substantial overdispersion with respect to the Poisson assumption (\(\text{ Var }(Y_{kt})=4.18\times {\mathbb {E}}[Y_{kt}]\)), as well as significant spatial autocorrelation in all 7 years based on a Moran’s I permutation test.
Based on these covariates we compute the spatial residuals from (5), and hence estimate static (\({\mathbf {W}}_{E}\)) and time-varying (\({\mathbf {W}}_{E_t}\)) neighbourhood matrices for the data. For the latter we use a 3-year moving average to estimate \({\tilde{\phi }}_{kt}\), that is \(q=1\), as this allows the most variation in \({\mathbf {W}}_{E_t}\) over time. The graph based on \({\mathbf {W}}\) contains 3,281 edges compared to 1,735 for the graph based on \({\mathbf {W}}_{E}\), and both are displayed in Fig. 1. The latter contains 1 large sub-graph containing 1,189 IZs and 19 small disconnected sub-graphs each containing between 2 and 10 IZs. The temporally varying neighbourhood structures \(({\mathbf {W}}_{E_2},\ldots ,{\mathbf {W}}_{E_{6}})\) (note, \({\mathbf {W}}_{E_1}={\mathbf {W}}_{E_2}\) and \({\mathbf {W}}_{E_{6}}={\mathbf {W}}_{E_7}\)) show evidence of some evolution over time, with 20% of the edges in the graph being identified as boundaries in all 5 neighbourhod matrices, while 75% of the edges are identified as boundaries in multiple years.
Four different specifications of model (1) are now fitted to the data, the first two of which use the border sharing \({\mathbf {W}}\) in conjunction with random effects \(\{\phi _{kt}\}\) modelled by either (3) or the BYM CAR prior described in Sect. 3 of the supplementary information. We compare these approaches to using model (3) in conjunction with the estimated neighbourhood matrices that are either constant (\({\mathbf {W}}_{E}\)) or varying (\([{\mathbf {W}}_{E_1},\ldots ,{\mathbf {W}}_{E_N}]\)) over time. Additionally, to assess the sensitivity of our results we re-fit the three models with the Leroux CAR prior (3) with the spatial dependence parameter \(\rho \) fixed at 1, because this restriction was made when creating the objective function \(J(\tilde{\varvec{\phi }})\) in Sect. 3. The results of this sensitivity analysis are displayed in Sect. 8 of the supplementary information, and show little change to the results presented below.
5.3 Results: Overall model fit
A summary of the overall fit of each model to the data via the deviance information criterion (DIC) and the effective number of independent parameters (p.d) is presented in Table 3, together with other key model parameters. The table shows that using either the static or the time-varying estimated neighbourhood matrices results in better model fit compared with using the border sharing specification, with a maximum difference in DIC of 2,172 which corresponds to a 3.2% reduction. Using a time-varying estimated neighbourhood matrix fits these data better than a static matrix as measured by the DIC, while the BYM model (using \({\mathbf {W}}\)) provides a slightly improved fit compared to the Leroux model (using \({\mathbf {W}}\)). The improvement in overall model fit from estimating the neighbourhood matrix is not caused by an improved fit at only a small number of data points, for example where edges have been removed, but instead results from an improved fit at the majority of the data points. For example, using \({\mathbf {W}}_{E_t}\) rather than \({\mathbf {W}}\) leads to smaller contributions to the DIC at 80% of the \(K\times N\) data points when using the Leroux CAR prior.
Using the estimated neighbourhood matrices \({\mathbf {W}}_{E}\) or \({\mathbf {W}}_{E_t}\) also provides better predictive performance compared with using \({\mathbf {W}}\), which is summarised by the log marginal predictive likelihood (LMPL) in Table 3 that one wishes to maximise. The LMPL is often used in disease surveillance applications and is calculated as \(\hbox {LMPL}=\sum _{kt}\ln [f(Y_{kt}|{\mathbf {Y}}_{-kt})]\), where \({\mathbf {Y}}_{-kt}\) denotes all observations except for \(Y_{kt}\), and further details are given by Corberán-Vallet and Lawson (2011).
Estimating the neighbourhood structure also provides a more parsimonious description of the data, as the models using \({\mathbf {W}}_{E}\) or \({\mathbf {W}}_{E_t}\) have fewer effective independent parameters as measured by p.d. This increase in parsimony is due to an increase in the estimated precisions \((\hat{\tau }_1^{-2},\ldots ,\hat{\tau }_N^{-2})\) when using \({\mathbf {W}}_E\) or \({\mathbf {W}}_{E_t}\), which are displayed for the three models using the Leroux prior (3) in Table 3. Note, the BYM model does not have a single comparable precision parameter. These increased precisions occur because unlike \({\mathbf {W}}\), \(({\mathbf {W}}_{E}, {\mathbf {W}}_{E_t})\) do not include edges between pairs of geographically adjacent IZs that exhibit large differences in their residuals, which reduces the amount of variation between \(\phi _{kt}\) and its spatially weighted mean from (3). This also increases the amount of spatial dependence in each spatial surface, which can be seen by the increases in \((\hat{\rho }_1,\ldots ,\hat{\rho }_N)\) when using \({\mathbf {W}}_{E}\) or \({\mathbf {W}}_{E_t}\).
5.4 Results: Covariate effects
Estimated relative risks (posterior medians) and 95% credible intervals for the covariates are displayed in Table 3, where each relative risk relates to the realistic increase in each covariate (i.e. close to the standard deviation of that covariate) given in column 1 of the table. The table shows that particulate air pollution (PM\(_{2.5}\)) is significantly associated with respiratory hospitalisations, with a 1\(\mu gm^{-3}\) increase in concentrations estimated to increase hospitalisations by between 4% and 5% depending on the model. The impact of income deprivation (poverty) is also clear and consistent, with increases in income deprivation being associated with between a 27% and a 30% increased risk.
In contrast the effects of housing deprivation after controlling for income deprivation are mixed, with the models based on \({\mathbf {W}}\) estimating a significant effect (the 95% credible interval does not include the null relative risk of 1), while if \({\mathbf {W}}_{E}\) or \({\mathbf {W}}_{E_t}\) are used the estimated risk is greatly attenuated close to one and is not significant. The results from the simulation study suggest that using \({\mathbf {W}}_{E}\) or \({\mathbf {W}}_{E_t}\) results in better covariate effect estimation compared with using \({\mathbf {W}}\) in almost all the scenarios considered, so it seems likely that housing deprivation has no effect after controlling for income deprivation. Finally, the widths of the 95% credible intervals are either the same or narrower for the model using \({\mathbf {W}}_{E}\) (and \({\mathbf {W}}_{E_{t}}\)) compared with the models using \({\mathbf {W}}\), which is most prominent for the PM\(_{2.5}\) effect with a 10–15% reduction in width.
5.5 Results: Disease surveillance
Our second aim is to use the models for disease surveillance, which requires us to estimate the spatio-temporal patterns in disease risk and identify areas with elevated risks of disease. The posterior mean risk estimates from the 4 models are broadly similar in most cases, with for example the estimates from the models using \({\mathbf {W}}\) and \({\mathbf {W}}_{E_t}\) differing on average by around 3% on the risk scale. The main differences in risk estimation relate to the uncertainty quantification, as the 95% credible intervals are 7–9% wider on average when using \({\mathbf {W}}\) compared with using \({\mathbf {W}}_E\) or \({\mathbf {W}}_{E_{t}}\). Thus estimating the neighbourhood structure provides more precise inference on disease risk, which the simulation study suggests does not come at the expense of poorer coverage.
The spatial patterns in disease risk show some relatively small evolution over the 7-year period, with the correlation (based on the model with \({\mathbf {W}}_{E_t}\)) between any two year’s risk surfaces ranging between 0.86 and 0.94, while the corresponding mean absolute differences (after accounting for the temporal trend) range between 0.096 and 0.16 on the risk scale. The temporal dynamics in disease risk are summarised in Sect. 9 of the supplementary information, while the spatial dynamics are illustrated here by posterior exceedance probabilities (PEP).
PEP are commonly used to identify areas that exhibit elevated risks of disease (see for example Kavanagh et al. (2012)), which allow public health professionals to target an intervention where it is most needed. The PEP is computed as \(\pi _{kt} = {\mathbb {P}}(\theta _{kt}> C|{\mathbf {Y}})\), the posterior probability that the risk \(\theta _{kt}\) exceed a certain threshold risk level C. The specification of C is somewhat arbitrary and chosen following discussions with public health experts, and here we choose \(C=1.5\) which represents a 50% elevated risk compared to the Scottish average.
The PEP for the Greater Glasgow and Clyde health board for 2015 is displayed in Fig. 3, with the maps in panels (a) and (b) relating to the Leroux model based on \({\mathbf {W}}\) and \({\mathbf {W}}_{E_t}\) respectively. The largest and most well known high risk cluster is the east end of Glasgow, which is caused in part by a cycle of multi-generational poverty (NHS Health Scotland 2016). In contrast, the single IZ in the north east of the health board near Kirkintilloch exhibits an elevated risk unlike its geographical neighbours, and would warrant further investigation by the health board into why it exhibits a very high PEP.
The maps also show that most areas have a PEP that is very close to either 0 (58% have a PEP less than 0.05) or 1 (18% have a PEP greater than 0.95), as the posterior uncertainty in the risk estimates is relatively small. Therefore the biggest differences between the PEP from the two models comes when the models are uncertain as to whether the risk exceeds 1.5, which can be seen for a small number of the IZs in the figure. For example, for those areas whose PEP is between (0.25, 0.75) the mean absolute difference in the PEP is 0.11, with a maximum difference of 0.36 on the probability scale.
5.6 Results: Boundary identification
Our final motivating question concerns the identification of boundaries in the spatial risk surface, which are locations where geographically adjacent areal units have very different disease risks. Boundary identification for areal unit data was first introduced by Womble (1951) and then used in a disease risk context by Lu and Carlin (2005). The identification of boundaries is important for social epidemiologists because their locations are likely to represent the demarcation between different neighborhoods as well as reflecting ‘underlying biological, physical, and/or social processes’ (Jacquez et al. 2000).
Our estimation of the neighbourhood structure via (\({\mathbf {W}}_E\), \({\mathbf {W}}_{E_{t}}\)) automatically identifies these boundaries, which is additional inference gained from our approach compared with using \({\mathbf {W}}\). Boundaries occur where two geographically adjacent areal units (k, j) (i.e. where \(w_{kj}=1\)) have \(w_{E_{kj}}=0\) or \(w_{E_{t_{kj}}}=0\), because the edge in the graph that induces correlation between their random effects \((\phi _{kt}, \phi _{jt})\) (see (4)) has been removed making them conditionally independent. As these boundaries are identified in the random effects surface, their interpretation depends on whether one includes covariates in the model. If no covariates are included then the boundaries relate directly to the risk surface because the random effects and risks have the same spatial structure as \(\theta _{kt}=\exp (\beta _0\,+\,\phi _{kt}\,+\,\delta _t)\). In contrast, if covariates are included in the model then the boundaries relate to the residual (unexplained) component of the risk surface after covariate adjustment, i.e. \(\exp (\phi _{kt})\) from \(\theta _{kt}=\exp ({\mathbf {x}}_{kt}^{\top }\varvec{\beta }\,+\,\delta _t)\exp (\phi _{kt})\).
Therefore we fit the model based on \({\mathbf {W}}_{E_{t}}\) (the best fitting model) separately with and without covariates, and present the resulting boundaries in both the risk and residual risk surfaces in Fig. 4. The figure presents maps of the average (over time) risks (panel a) and residual risks (panel b) in the Glasgow region, and we focus on this region for the reasons outlined above. The simulation study showed that while the sensitivity of boundary identification was very high when the disease counts are not rare (the case here as the mean of \(\{Y_{kt}\}\) is 75), the specificity is likely to be between 70% and 75%. Therefore to identify the clearest boundaries and avoid presenting false positives, we only present the boundaries that are consistently identified over multiple time periods. In our 7-year study period we estimate 5 different neighbourhood structures because \({\mathbf {W}}_{E_{1}}={\mathbf {W}}_{E_{2}}\) and \({\mathbf {W}}_{E_{6}}={\mathbf {W}}_{E_{6}}\), and the yellow and orange dots in the figure denote boundaries that were identified for all 5 (yellow) and 4 out of the 5 (orange) years. Note, the river Clyde running north-west divides the Glasgow region into 2 sub-regions in terms of \({\mathbf {W}}\) and its corresponding graph, and hence boundaries cannot be identified across the river.
The figure shows that neither the risk surface nor the residual risk surface are wholly spatially smooth, which explains why the models based on (\({\mathbf {W}}_E\), \({\mathbf {W}}_{E_{t}}\)) fit the data better and are hence more appropriate than those based on \({\mathbf {W}}\). The boundaries identified mainly correspond to locations where there visually appear to be step changes in these surfaces, suggesting that they do appear to represent the boundaries between different neighbourhoods. An in-depth exploration of the risk boundaries in panel a highlights that a number of them correspond to physical barriers such as rivers, railway lines, motorways, etc, which are difficult to cross and thus make it hard for the two communities on either side to mix. Examples of these physical barrier boundaries are illustrated in Sect. 10 of the supplementary information, and provide insight that these physical barriers may help the formation of neighbourhoods in urban environments.
6 Discussion
This paper has presented a novel graph-based optimisation algorithm for estimating the neighbourhood matrix when modelling spatio-temporal areal unit count data, and has provided software to allow others to utilise our methods. Our approach estimates an appropriate spatial autocorrelation structure for the data at hand, rather than naively using the simple border sharing rule. We have proposed related approaches for assuming the neighbourhood matrix is either static or evolves dynamically over time, and the suitability of each will depend on the consistency of the spatial variation in the data over time. Our approach estimates the residual spatial autocorrelation structure in the data using the residuals from a covariate only model, which is akin to applying variogram analysis to detrended geostatistical data to identify an appropriate spatial correlation model. Thus we recommend that, as in geostatistics, standard practice in spatio-temporal areal unit modelling should involve estimating both the mean model and the residual spatial dependence structure. This contrasts with current practice that assumes the latter is well represented by the border sharing rule, with little assessment of its suitability to the data being modelled (e.g. Quick et al. 2017; Lee et al. 2019).
The simulation study showed strong evidence that estimating the neighbourhood structure delivers improved estimation of risks (rates) and covariate effects for spatio-temporal data that contain small or large boundaries, as long as the number of time periods is at least \(N=3\). Additionally, the uncertainty intervals are narrower compared to using a border sharing neighbourhood structure, whilst retaining appropriate coverage percentages. These improvements were consistently observed for boundaries that did and did not change over time, and the sensitivity and specificity for boundary identification were between 85–97% (sensitivity) and between 70–75% (specificity) as long as the count data were not rare. However, as previously discussed this specificity result from the simulation study is likely to be artificially low, because it ignores the independent random variation induced into the count data by the Poisson data model that is likely to cause additional unintended boundaries.
In contrast, our approach does not work well for purely spatial data (\(N=1\)), because the residual spatial surface \(\tilde{\varvec{\phi }}\) is not well estimated using only 1 time period of data due to the presence of random noise in \(\{Y_{kt}\}\). However, as N increases this random noise is reduced by averaging the residuals over time, leading to the improved performance described above. Thus to apply this approach to purely spatial data we suggest estimating \(\tilde{\varvec{\phi }}\) from multiple sets of external data that have a similar residual spatial structure to the study data. Possible candidates in this regard are the same data but for earlier time periods, or data with a related response variable such as a different disease with a similar etiology. This last point illustrates the fact that while the estimated neighbourhood matrix \({\mathbf {W}}_E\) is specific to each response variable and covariate combination, the estimated matrices for two variables with similar spatial surfaces should themselves be similar. Thus if one is studying the risks from multiple chronic diseases which often have similar spatial patterns (see Jack et al. 2019 for an example), then their estimated neighbourhood matrices are also likely to be similar.
Our motivating Scotland study has illustrated the importance of obtaining improved estimation and uncertainty quantification of the drivers and spatio-temporal patterns in disease risk, because it leads to more accurate inferences with less uncertainty. This is particularly relevant to disease surveillance metrics such as PEPs, because their construction depends on the full posterior distribution of risk. Our motivating study also illustrates the additional inference that is gained from our approach, namely the identification of boundaries in the spatial data that separate areas that are geographically adjacent but have very different data values. We found that numerous boundaries exist in the Greater Glasgow region that separate different communities, and that one possible driver of these boundaries is the presence of physical barriers such as railway lines that prevent population mixing.
There is a wealth of exciting research directions for extending this work, the most obvious of which is to extend the class of data and models that our graph-based optimisation approach can be used with. These include extending the methods away from count data to deal with Gaussian and binomial type responses, considering multivariate rather than spatio-temporal data structures, and using different spatio-temporal random effects structures to that considered here. Additionally, our motivating study has shown that similar levels of disease risk are more commonly observed between areas with similar levels of socio-economic deprivation rather than those that happen to be geographically close. This suggests that one might want to additionally allow for autocorrelation between areas with similar levels of socio-economic deprivation, perhaps via the introduction of a second neighbourhood matrix based on socio-economic rather than physical adjacency. This would result in the data having an autocorrelation structure based on a multilayer graph, and our optimisation approach would need to be extended to allow for this multilayer scenario.
Finally, there is scope to improve the performance of the graph-based optimisation algorithm used to estimate \({\mathbf {W}}_{E}\), as the current implementation makes use of a local search method that is not guaranteed to find the best possible matrix \({\mathbf {W}}_{E}\) with respect to the objective function. The fact that the optimisation problem is NP-hard in general means that we are very unlikely to find an algorithm that is guaranteed to perform the optimsation exactly within a reasonable length of time for all possible inputs. Nevertheless, it may be possible to obtain an efficient approximation algorithm that achieves a guaranteed performance ratio (for example, computing a matrix for which the objective function is at most \(5\%\) worse than the best possible), or parameterised algorithms which have exponential running-time in the worst case but are guaranteed to perform much faster on inputs with specific structural properties. Further work is needed to establish the feasibility or otherwise of both approaches.
References
Berchuck, S., Mwanza, J., Warren, J.: Diagnosing glaucoma progression with visual field data using a spatiotemporal boundary detection method. J. Am. Stat. Assoc. 114, 1063–1074 (2019)
Bernardinelli, L., Clayton, D., Pascutto, C., Montomoli, C., Ghislandi, M., Songini, M.: Bayesian analysis of space-time variation in disease risk. Stat. Med. 14, 2433–2443 (1995)
Besag, J., York, J., Mollié, A.: Bayesian image restoration with two applications in spatial statistics. Annals of the Institute of Statistics and Mathematics 43, 1–59 (1991)
Bradley, J., Wikle, C., Holan, S.: Bayesian spatial change of support for count-valued survey data with application to the American community survey. J. Am. Stat. Assoc. 111, 472–487 (2016)
Corberán-Vallet, A., Lawson, A.: Conditional predictive inference for online surveillance of spatial disease incidence. Stat. Med. 30, 3095–3116 (2011)
Enright, J., Lee, D., Meeks, K., Pettersson, W., Sylvester, J.: (2021) The Complexity of Finding Optimal Subgraphs to Represent Spatial Correlation (2021). arXiv:2010.10314
Fisher, R.A., Yates, F.: Statistical Tables for Biological, Agricultural and Medical Research, 3rd edn. Oliver & Boyd, London (1948)
Jack, E., Lee, D., Dean, N.: Estimating the changing nature of Scotland’s health inequalities by using a multivariate spatiotemporal model. J. R. Stat. Soc. Ser. A 182, 1061–1080 (2019)
Jacquez, G., Maruca, S., Fortin, M.: From fields to objects: a review of geographic boundary analysis. J. Geogr. Syst. 2, 221–241 (2000)
Kavanagh, K., Robertson, C., Murdoch, H., Crooks, G., McMenamin, J.: Syndromic surveillance of influenza-like illness in scotland during the influenza a h1n1v pandemic and beyond. J. R. Stat. Soc.: Ser. A 175, 939–958 (2012)
Knorr-Held, L.: Bayesian modelling of inseparable space-time variation in disease risk. Stat. Med. 19, 2555–2567 (2000)
Knorr-Held, L., Raßer, G.: Bayesian Detection of Clusters and Discontinuities in Disease Maps. Biometrics 56, 13–21 (2000)
Lee, D.: A comparison of conditional autoregressive models used in Bayesian disease mapping. Spatial and Spatio-temporal Epidemiology 2, 79–89 (2011)
Lee, D., Mitchell, R.: Boundary detection in disease mapping studies. Biostatistics 13, 415–426 (2012)
Lee, D., Rushworth, A., Napier, G.: Spatio-temporal areal unit modeling in R with conditional autoregressive priors using the CARBayesST package. J. Stat. Softw. 84(9), 1–39 (2018)
Lee, D., Robertson, C., Ramsay, C., Gillespie, C., Napier, G.: Estimating the health impact of air pollution in Scotland, and the resulting benefits of reducing concentrations in city centres. Spat. Spatio-temporal Epidemiol. 29, 85–96 (2019)
Leroux, B., Lei, X., Breslow, N.: Estimation of disease rates in small areas: a new mixed model for spatial dependence. In: Halloran, M., Berry, D. (Eds.) Statistical Models in Epidemiology, the Environment and Clinical Trials, pp. 135–178. Springer, New York (2000)
Lu, H., Carlin, B.: Bayesian areal wombling for geographical boundary analysis. Geogr. Anal. 37, 265–285 (2005)
Ma, H., Carlin, B., Banerjee, S.: Hierarchical and joint site-edge methods for Medicare hospice service region boundary analysis. Biometrics 66, 355–364 (2010)
Mitchell, R., Lee, D.: Is there really a ‘wrong side of the tracks’ in urban areas and does it matter for spatial analysis? Ann. Assoc. Am. Geogr. 104, 432–443 (2014)
NHS Health Scotland (2016) Health inequalities–what are they and how do we reduce them?http://www.healthscotland.scot/media/1086/health-inequalities-what-are-they-how-do-we-reduce-them-mar16.pdf
Quick, H., Waller, L., Casper, M.: Multivariate spatiotemporal modeling of age-specific stroke mortality. Ann. Appl. Stat. 11, 2165–2177 (2017)
Rue, H., Martino, S., Chopin, N.: Approximate Bayesian inference for latent Gaussian models using integrated nested laplace approximations (with discussion). J. R. Stat. Soc. Ser. B 71, 1 (2009)
Rushworth, A., Lee, D., Mitchell, R.: A spatio-temporal model for estimating the long-term effects of air pollution on respiratory hospital admissions in Greater London. Spat. Spatio-temporal Epidemiol. 10, 29–38 (2014)
Stoner, O., Economou, T., da Silva, G.: A hierarchical framework for correcting under-reporting in count data. J. Am. Stat. Assoc. 114(528), 1481–1492 (2019)
Waller, L., Carlin, B., Xia, H., Gelfand, E.: Hierarchical spatio-temporal mapping of disease rates. J. Am. Stat. Assoc. 92(438), 607–617 (1997)
Walsh, D., McCartney, G., Collins, C., Taulbut, M., Batty, D.: History, Politics and Vulnerability: Explaining Excess Mortality in Scotland and Glasgow. Glasgow Centre for Population Health (2016)
Womble, W.: Differential systematics. Science 114, 315–322 (1951)
Acknowledgements
The authors are grateful to Jessica Enright and John Sylvester for providing helpful comments on an initial draft of this manuscript, and to two anonymous reviewers whose comments improved the motivation for and content of this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
None
Availability of data
The respiratory hospitalisation data were provided confidentially by Public Health Scotland (PHS), and others would need to apply to them to access the data.
Code availability
Software to implement the graph theoretic algorithm is available in the CARBayesST package on CRAN.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
All authors gratefully acknowledge funding from the Engineering and Physical Sciences Research Council (ESPRC) grant number EP/T004878/1 for this work, while the work of the second author was also funded by a Royal Society of Edinburgh Personal Research Fellowship (funded by the Scottish Government).
Supplementary Information
Below is the link to the electronic supplementary material.
Appendix
Appendix
Algorithm 1 summarises our graph-based optimisation algorithm. We note here a number of interesting features we discovered while developing and using Algorithm 1.

Algorithm 1 iterates through the vertices in some order, which represents a starting point for the heuristic. For our simulation data on 257 vertices, we randomly sampled 1300 permutations using the Fisher-Yates shuffle (Fisher and Yates 1948). Over these 1300 permutations, we noted no substantial change to the value of the objective function. Whilst 1300 is still a minute proportion of the 257! possible permutations, this does suggest that the heuristic is relatively stable to changes in vertex ordering.
There is also the obvious potential that the algorithm may get stuck within local optima. We tested a simple simulated annealing model, but like the permutation tests, results showed that simulated annealing never led to any noticeable improvement of the value of the resulting graph, and occasionally lead to a worse result as a “good” edge was randomly removed. This removal of “good” edges also suggested that adding previously-removed edges back to our graph might be useful, if doing so would improve the value of our objective function. However, there was no clear improvement in the value achieved by the objective function when the heuristic was able to add edges back into graph.
We also investigated an approach where, for each vertex v, we first removed all edges to neighbouring vertices except for the the edge to the vertex whose weight is closest to v, and then used our heuristic to add some of the edges back. Again, however, we report that we noted no significant change in the value achieved by the objective function when compared to our other techniques.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, D., Meeks, K. & Pettersson, W. Improved inference for areal unit count data using graph-based optimisation. Stat Comput 31, 51 (2021). https://doi.org/10.1007/s11222-021-10025-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11222-021-10025-7