Abstract
Recently-proposed particle MCMC methods provide a flexible way of performing Bayesian inference for parameters governing stochastic kinetic models defined as Markov (jump) processes (MJPs). Each iteration of the scheme requires an estimate of the marginal likelihood calculated from the output of a sequential Monte Carlo scheme (also known as a particle filter). Consequently, the method can be extremely computationally intensive. We therefore aim to avoid most instances of the expensive likelihood calculation through use of a fast approximation. We consider two approximations: the chemical Langevin equation diffusion approximation (CLE) and the linear noise approximation (LNA). Either an estimate of the marginal likelihood under the CLE, or the tractable marginal likelihood under the LNA can be used to calculate a first step acceptance probability. Only if a proposal is accepted under the approximation do we then run a sequential Monte Carlo scheme to compute an estimate of the marginal likelihood under the true MJP and construct a second stage acceptance probability that permits exact (simulation based) inference for the MJP. We therefore avoid expensive calculations for proposals that are likely to be rejected. We illustrate the method by considering inference for parameters governing a Lotka–Volterra system, a model of gene expression and a simple epidemic process.






Similar content being viewed by others
References
Andrieu, C., Doucet, A., Holenstein, R.: Particle Markov chain Monte Carlo for efficient numerical simulation. In: L’Ecuyer, P., Owen, A.B. (eds.) Monte Carlo and Quasi-Monte Carlo methods 2008, pp. 45–60. Spinger, Berlin, Heidelberg (2009)
Andrieu, C., Doucet, A., Holenstein, R.: Particle Markov chain Monte Carlo methods (with discussion). J. R. Stat. Soc. B 72(3), 1–269 (2010)
Andrieu, C., Roberts, G.O.: The pseudo-marginal approach for efficient computation. Ann. Stat. 37, 697–725 (2009)
Bailey, N.T.J.: The Mathematical Theory of Infectious Diseases and Its Applications, 2nd edn. Hafner Press [Macmillan Publishing Co., Inc.], New York (1975)
Ball, F., Neal, P.: Network epidemic models with two levels of mixing. Math. Biosci. 212(1), 69–87 (2008)
Beaumont, M.A.: Estimation of population growth or decline in genetically monitored populations. Genetics 164, 1139–1160 (2003)
Boys, R.J., Giles, P.R.: Bayesian inference for stochastic epidemic models with time-inhomogeneous removal rates. J. Math. Biol. 55, 223–247 (2007)
Boys, R.J., Wilkinson, D.J., Kirkwood, T.B.L.: Bayesian inference for a discretely observed stochastic-kinetic model. Stat. Comput. 18, 125–135 (2008)
Christen, J.A., Fox, C.: Markov chain Monte Carlo using an approximation. J. Comput. Graph. Stat. 14, 795–810 (2005)
Del Moral, P.: Feynman-Kac Formulae: Genealogical and Interacting Particle Systems with Applications. Springer, New York (2004)
Doucet, A., Pitt, M. K., Kohn, R.: Efficient implementation of Markov chain Monte Carlo when using an unbiased likelihood estimator (2013). Available from http://arxiv.org/pdf/1210.1871.pdf
Elf, J., Ehrenberg, M.: Fast evolution of fluctuations in biochemical networks with the linear noise approximation. Genome Res. 13(11), 2475–2484 (2003)
Elowitz, M.B., Levine, A.J., Siggia, E.D., Swain, P.S.: Stochastic gene expression in a single cell. Science 297(5584), 1183–1186 (2002)
Fearnhead, P., Giagos, V., Sherlock, C.: Inference for reaction networks using the Linear Noise Approximation (to appear in Biometrics) (2014)
Fearnhead, P., Meligkotsidou, L.: Exact filtering for partially observed continuous time models. J. R. Stat. Soc. B. 66(3), 771–789 (2004)
Ferm, L., Lötstedt, P., Hellander, A.: A hierarchy of approximations of the master equation scaled by a size parameter. J. Sci. Comput. 34(2), 127–151 (2008)
Gillespie, D.T.: Exact stochastic simulation of coupled chemical reactions. J. Phys. Chem. 81, 2340–2361 (1977)
Gillespie, D.T.: A rigorous derivation of the chemical master equation. Physica A 188, 404–425 (1992)
Gillespie, D.T.: The chemical Langevin equation. J. Chem. Phys. 113(1), 297–306 (2000)
Golightly, A., Wilkinson, D.J.: Bayesian inference for stochastic kinetic models using a diffusion approximation. Biometrics 61(3), 781–788 (2005)
Golightly, A., Wilkinson, D.J.: Bayesian inference for nonlinear multivariate diffusion models observed with error. Comput. Stat. Data Anal. 52, 1674–1693 (2008)
Golightly, A., Wilkinson, D.J.: Bayesian parameter inference for stochastic biochemical network models using particle Markov chain Monte Carlo. Interface Focus 1(6), 807–820 (2011)
Gordon, N.J., Salmond, D.J., Smith, A.F.M.: Novel approach to nonlinear/non-Gaussian Bayesian state estimation. IEE. Proc-F. 140, 107–113 (1993)
Jewell, C.P., Keeling, M.J., Roberts, G.O.: Predicting undetected infections during the 2007 foot-and-mouth disease outbreak. J. R. Soc. Interface 6, 1145–1151 (2009)
Kitano, H.: Computational systems biology. Nature 420(6912), 206–210 (2002)
Komorowski, M., Finkenstadt, B., Harper, C., Rand, D.: Bayesian inference of biochemical kinetic parameters using the linear noise approximation. BMC Bioinformatics 10(1), 343 (2009)
Kurtz, T.G.: Solutions of ordinary differential equations as limits of pure jump markov processes. J. Appl. Probab. 7, 49–58 (1970)
Lee, A., Yau, C., Giles, M.B., Doucet, A.: On the utility of graphics cards to perform massively parallel simulation with advanced Monte Carlo methods. J. Comput. Graph. Stat. 19(4), 769–789 (2010)
Liu, J.S.: Monte Carlo Strategies in Scientific Computing. Springer, New York (2001)
O’Neill, P.D., Roberts, G.O.: Bayesian inference for partially observed stochastic epidemics. J. R. Stat. Soc. A. 162, 121–129 (1999)
Petzold, L.: Automatic selection of methods for solving stiff and non-stiff systems of ordinary differential equations. SIAM J. Sci. Stat. Comput. 4(1), 136–148 (1983)
Pitt, M.K., dos Santos Silva, R.: On some properties of Markov chain Monte Carlo simulation methods based on the particle filter. J. Econom 171(2), 134–151 (2012)
Purutcuoglu, V., Wit, E.: Bayesian inference of the kinetic parameters of a realistic MAPK/ERK pathway. BMC Syst. Biol. 1, P19 (2007)
Sherlock, C., Thiery, A., Roberts, G.O., Rosenthal, J.S.: On the effciency of pseudo-marginal random walk Metropolis algorithms (2013). Available from http://arxiv.org/abs/1309.7209
Smith, M.E.: Estimating nonlinear economic models using surrogate transitions (2011). Available from https://files.nyu.edu/mes473/public/Smith_Surrogate.pdf
Stathopoulos, V., Girolami, M.: Markov chain Monte Carlo inference for Markov jump processes via the linear noise approximation. Philos. Trans. R. Soc. A 371, 20110549 (2013)
Swain, P.S., Elowitz, M.B., Siggia, E.D.: Intrinsic and extrinsic contributions to stochasticity in gene expression. PNAS 99(20), 12795–12800 (2002)
van Kampen, N.G.: Stochastic Processes in Physics and Chemistry. North-Holland, Amsterdam (2001)
Wilkinson, D.J.: Stochastic modelling for quantitative description of heterogeneous biological systems. Nat. Rev. Genet. 10, 122–133 (2009)
Wilkinson, D.J.: Stochastic Modelling for Systems Biology, 2nd edn. Chapman and Hall/CRC Press, London (2012)
Acknowledgments
The authors would like to thank the editor and two anonymous referees for their suggestions for improving this paper.
Author information
Authors and Affiliations
Corresponding author
Appendix
Appendix
Recall that \(\mathbf {x}=\{x_{t}\,|\, 1\le t \le T\}\) denotes values of the latent MJP and \(\mathbf {y}=\{y_{t}\,|\, t=1,2,\ldots ,T\}\) denotes the collection of (noisy) observations on the MJP at discrete times. In addition, we define \(\mathbf {x}_{t}=\{x_{s}\,|\, t-1<s\le t\}\) and \(\mathbf {y}_{t}=\{y_{s}\,|\, s=1,2,\ldots , t\}\).
1.1 PMMH scheme
The PMMH scheme has the following algorithmic form.
-
1.
Initialisation, \(i=0\),
-
(a)
set \(c^{(0)}\) arbitrarily and
-
(b)
run an SMC scheme targeting \(p(\mathbf {x}|\mathbf {y},c^{(0)})\), and let \(\widehat{p}(\mathbf {y}|c^{(0)})\) denote the marginal likelihood estimate
-
(a)
-
2.
For iteration \(i\ge 1\),
-
(a)
sample \(c^{*}\sim q(\cdot | c^{(i-1)})\),
-
(b)
run an SMC scheme targeting \(p(\mathbf {x}|\mathbf {y},c^{*})\), and let \(\widehat{p}(\mathbf {y}|c^{*})\) denote the marginal likelihood estimate,
-
(c)
with probability min\(\{1,A\}\) where
$$\begin{aligned} A=\frac{\widehat{p}(\mathbf {y}|c^{*}) p(c^{*})}{\widehat{p}(\mathbf {y}|c^{(i-1)}) p(c^{(i-1)})} \times \frac{q(c^{(i-1)} | c^{*})}{q(c^{*} | c^{(i-1)})} \end{aligned}$$accept a move to \(c^{*}\) otherwise store the current values
-
(a)
Note that the PMMH scheme can be used to sample the joint posterior \(p(c,\mathbf {x}|\mathbf {y})\). Essentially, a proposal mechanism of the form \(q(c^{*}|c)\widehat{p}(\mathbf {x}^{*}|\mathbf {y},c^{*})\), where \(\widehat{p}(\mathbf {x}^{*}|\mathbf {y},c^{*})\) is an SMC approximation of \(p(\mathbf {x}^{*}|\mathbf {y},c^{*})\), is used. The resulting MH acceptance ratio is as above. Full details of the PMMH scheme including a proof establishing that the method leaves the target \(p(c,\mathbf {x}|\mathbf {y})\) invariant can be found in Andrieu et al. (2010).
1.2 SMC scheme
A sequential Monte Carlo estimate of the marginal likelihood \(p(\mathbf {y}|c)\) under the MJP can be constructed using (for example) the bootstrap filter of Gordon et al. (1993). Algorithmically, we perform the following sequence of steps.
-
1.
Initialisation.
-
(a)
Generate a sample of size \(N\), \(\{x_{1}^{1},\ldots ,x_{1}^{N}\}\) from the initial density \(p(x_{1})\).
-
(b)
Assign each \(x_{1}^{i}\) a (normalised) weight given by
$$\begin{aligned} w_{1}^{i}=\frac{w_{1}^{*i}}{\sum _{i=1}^{N}w_{1}^{*i}}, \quad \text {where}\quad w_{1}^{*i}=p(y_{1}|x_{1}^{i},c)\,. \end{aligned}$$ -
(c)
Construct and store the currently available estimate of marginal likelihood,
$$\begin{aligned} \widehat{p}({y}_{1}|c) = \frac{1}{N}\sum _{i=1}^{N} w_{1}^{*i}\,. \end{aligned}$$ -
(d)
Resample \(N\) times with replacement from \(\{x_{1}^{1},\ldots ,x_{1}^{N}\}\) with probabilities given by \(\{w_{1}^{1},\ldots ,w_{1}^{N}\}\).
-
(a)
-
2.
For times \(t=1,2,\ldots ,T-1\),
-
(a)
For \(i=1,\ldots ,N\): draw \(\mathbf {X}_{t+1}^{i}\sim p\big (\mathbf {x}_{t+1}|{x}_{t}^{i},c\big )\) using the Gillespie algorithm.
-
(b)
Assign each \(\mathbf {x}_{t+1}^{i}\) a (normalised) weight given by
$$\begin{aligned} w_{t+1}^{i}=\frac{w_{t+1}^{*i}}{\sum _{i=1}^{N}w_{t+1}^{*i}}, \quad \text {where}\quad w_{t+1}^{*i}=p(y_{t+1}|x_{t+1}^{i},c)\, . \end{aligned}$$ -
(c)
Construct and store the currently available estimate of marginal likelihood,
$$\begin{aligned} \widehat{p}(\mathbf {y}_{t+1}|c)&= \widehat{p}(\mathbf {y}_{t}|c)\widehat{p}(y_{t+1}|\mathbf {y}_{t},c)\\&=\widehat{p}(\mathbf {y}_{t}|c)\frac{1}{N}\sum _{i=1}^{N} w_{t+1}^{*i}\,. \end{aligned}$$ -
(d)
Resample \(N\) times with replacement from \(\{\mathbf {x}_{t+1}^{1},\ldots ,\mathbf {x}_{t+1}^{N}\}\) with probabilities given by \(\{w_{t+1}^{1},\ldots ,w_{t+1}^{N}\}\).
-
(a)
1.3 Marginal likelihood under the linear noise approximation
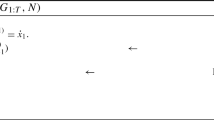
Assume an observation regime of the form
where \(G\) is a constant matrix of dimension \(u\times p\) and \(\varepsilon _{t}\) is a length-\(p\) Gaussian random vector.
Now suppose that \(X_{1}\sim N(a,C)\) a priori. The marginal likelihood under the LNA, \(p_{a}(\mathbf {y}|c)\) can be obtained as follows.
-
1.
Initialisation. Compute
$$\begin{aligned} p_{a}(y_{1}|c)=\phi \left( y_{1}\,;\, G'a\,,\,G'CG+\varSigma \right) \end{aligned}$$where \(\phi (\cdot \,;\,a\,,\,C)\) denotes the Gaussian density with mean vector \(a\) and variance matrix \(C\). The posterior at time \(t=1\) is therefore \(X_{1}|y_{1}\sim N(a_{1},C_{1})\) where
$$\begin{aligned} a_{1}&= a+CG\left( G'CG+\varSigma \right) ^{-1}\left( y_{1}-G'a\right) \\ C_{1}&= C-CG\left( G'CG+\varSigma \right) ^{-1}G'C\,. \end{aligned}$$ -
2.
For times \(t=1,2,\ldots ,T-1\),
-
(a)
Prior at \(t+1\). Initialise the LNA with \(z_{t}=a_{t}\), \(m_{t}=0\) and \(V_{t}=C_{t}\). Note that this implies \(m_{s}=0\) for all \(s>t\). Therefore, integrate the ODEs (6) and (10) forward to \(t+1\) to obtain \(z_{t+1}\) and \(V_{t+1}\). Hence
$$\begin{aligned} X_{t+1}|\mathbf {y}_{t}\sim N(z_{t+1},V_{t+1})\,. \end{aligned}$$ -
(b)
One step forecast. Using the observation equation, we have that
$$\begin{aligned} Y_{t+1}|\mathbf {y}_{t}\sim N\left( G'z_{t+1},G'V_{t+1}G+\varSigma \right) . \end{aligned}$$Compute
$$\begin{aligned} p_{a}(\mathbf {y}_{t+1}|c)&=p_{a}(\mathbf {y}_{t}|c)p_{a}(y_{t+1}|\mathbf {y}_{t},c)\\&=p_{a}(\mathbf {y}_{t}|c)\,\phi \left( y_{t+1}\,;\, G'z_{t+1}\,,\,G'V_{t+1}G+\varSigma \right) . \end{aligned}$$ -
(c)
Posterior at \(t+1\). Combining the distributions in (a) and (b) gives the joint distribution of \(X_{t+1}\) and \(Y_{t+1}\) (conditional on \(\mathbf {y}_{t}\) and \(c\)) as
$$\begin{aligned} \left( \begin{array}{c} X_{t+1} \\ Y_{t+1} \end{array}\right) \sim N\left\{ \left( \begin{array}{c} z_{t+1} \\ G'z_{t+1} \end{array} \right) \,,\, \left( \begin{array}{cc} V_{t+1} &{} V_{t+1}G \\ G'V_{t+1} &{} G'V_{t+1}G+\varSigma \end{array} \right) \right\} \end{aligned}$$and therefore \(X_{t+1}|\mathbf {y}_{t+1}\sim N(a_{t+1},C_{t+1})\) where
$$\begin{aligned} a_{t+1}&= z_{t+1}+V_{t+1}G\left( G'V_{t+1}G+\varSigma \right) ^{-1}\left( y_{t+1}-G'z_{t+1}\right) \\ C_{t+1}&= V_{t+1}-V_{t+1}G\left( G'V_{t+1}G+\varSigma \right) ^{-1}G'V_{t+1}\,. \end{aligned}$$
-
(a)
Rights and permissions
About this article
Cite this article
Golightly, A., Henderson, D.A. & Sherlock, C. Delayed acceptance particle MCMC for exact inference in stochastic kinetic models. Stat Comput 25, 1039–1055 (2015). https://doi.org/10.1007/s11222-014-9469-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11222-014-9469-x