Abstract
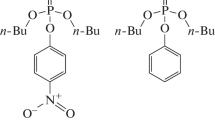
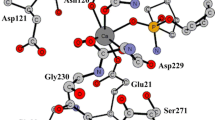
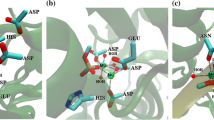
Full-atom molecular models of the phosphotriesterase dimer from the Pseudomonas diminuta bacterium with organophosphorus compounds bearing good and poor leaving groups, dibutyl p-nitrophenyl phosphate and dibutyl phenyl phosphate, respectively, were constructed. Molecular dynamic simulations with combined quantum mechanics/molecular mechanics (QM/MM) potentials depicted differences in the properties of intermediates of the hydrolysis reaction with pentacoordinated phosphorus for these substrates. In the case of a substrate with the good leaving group, the bond between the phosphorus and oxygen atoms of the leaving group is weaker than that between the phosphorus and oxygen atoms of the nucleophilic hydroxide anion, which leads to an almost barrierless formation of the reaction products from the intermediate. For the substrate with the poor leaving group, an opposite pattern is observed, which favors the return of the system from the intermediate to reactants. These conclusions were made on the basis of an analysis of the distributions of bond lengths along the trajectories, as well as from an analysis of the maps of the Laplacian of electron density in the reaction region.
Similar content being viewed by others
References
T. Reemtsma, J. B. Quintana, R. Rodil, M. Garcia-López, I. Rodriguez, TrAC Trends Anal. Chem., 2008, 27, 727–737; DOI: https://doi.org/10.1016/j.trac.2008.07.002.
J. Du, H. Li, S. Xu, Q. Zhou, M. Jin, J. Tang, Environ. Sci. Pollut. Res., 2019, 26, 22126–22136; DOI: https://doi.org/10.1007/s11356-019-05669-y.
P.-C. Tsai, N. Fox, A. N. Bigley, S. P. Harvey, D. P. Barondeau, F. M. Raushel, Biochemistry, 2012, 51, 6463–6475; DOI: https://doi.org/10.1021/bi300811t.
D. F. Xiang, A. N. Bigley, Z. Ren, H. Xue, K. G. Hull, D. Romo, F. M. Raushel, Biochemistry, 2015, 54, 7539–7549; DOI: https://doi.org/10.1021/acs.biochem.5b01144.
J. L. Vanhooke, M. M. Benning, F. M. Raushel, H. M. Holden, Biochemistry, 1996, 35, 6020–6025; DOI: https://doi.org/10.1021/bi960325l.
J. K. Grimsley, B. Calamini, J. R. Wild, A. D. Mesecar, Arch. Biochem. Biophys., 2005, 442, 169–179; DOI: https://doi.org/10.1016/j.abb.2005.08.012.
S. D. Aubert, Y. Li, F. M. Raushel, Biochemistry, 2004, 43, 5707–5715; DOI: https://doi.org/10.1021/bi0497805.
C. J. Jackson, J.-L. Foo, H.-K. Kim, P. D. Carr, J.-W. Liu, G. Salem, D. L. Ollis, J. Mol. Biol., 2008, 375, 1189–1196; DOI: https://doi.org/10.1016/j.jmb.2007.10.061.
J. Kim, P. C. Tsai, S. L. Chen, F. Himo, S. C. Almo, F. M. Raushel, Biochemistry, 2008, 47, 9497–9504; DOI: https://doi.org/10.1021/bi800971v.
J. M. Word, S. C. Lovell, J. S. Richardson, D. C. Richardson, J. Mol. Biol., 1999, 285, 1735–1747; DOI: https://doi.org/10.1006/jmbi.1998.2401.
W. Humphrey, A. Dalke, K. Schulten, J. Mol. Graph., 1996, 14, 33–38; DOI: https://doi.org/10.1016/0263-7855(96)00018-5.
J. C. Phillips, R. Braun, W. Wang, J. Gumbart, E. Tajkhorshid, E. Villa, C. Chipot, R. D. Skeel, L. Kalé, K. Schulten, J. Comput. Chem., 2005, 26, 1781–1802; DOI: https://doi.org/10.1002/jcc.20289.
R. B. Best, X. Zhu, J. Shim, P. E. M. Lopes, J. Mittal, M. Feig, A. D. MacKerell, J. Chem. Theory Comput., 2012, 8, 3257–3273; DOI: https://doi.org/10.1021/ct300400x.
K. Vanommeslaeghe, E. Hatcher, C. Acharya, S. Kundu, S. Zhong, J. Shim, E. Darian, O. Guvench, P. Lopes, I. Vorobyov, A. D. Mackerell, J. Comput. Chem., 2009, 31, 671–690; DOI: https://doi.org/10.1002/jcc.21367.
W. L. Jorgensen, J. Chandrasekhar, J. D. Madura, R. W. Impey, M. L. Klein, J. Chem. Phys., 1983, 79, 926–935; DOI: https://doi.org/10.1063/1.445869.
S. Seritan, C. Bannwarth, B. S. Fales, E. G. Hohenstein, C. M. Isborn, S. I. L. Kokkila-Schumacher, X. Li, F. Liu, N. Luehr, J. W. Snyder, C. Song, A. V. Titov, I. S. Ufimtsev, L. Wang, T. J. Martínez, WIREs Comput. Mol. Sci., 2021, 11, e1494; DOI: https://doi.org/10.1002/wcms.1494.
M. C. R. Melo, R. C. Bernardi, T. Rudack, M. Scheurer, C. Riplinger, J. C. Phillips, J. D. C. Maia, G. B. Rocha, J. V. Ribeiro, J. E. Stone, F. Neese, K. Schulten, Z. Luthey-Schulten, Nat. Methods, 2018, 15, 351–354; DOI: https://doi.org/10.1038/nmeth.4638.
C. Adamo, V. Barone, J. Chem. Phys., 1999, 110, 6158–6170; DOI: https://doi.org/10.1063/1.478522.
S. Grimme, J. Antony, S. Ehrlich, H. Krieg, J. Chem. Phys., 2010, 132, 154104; DOI: https://doi.org/10.1063/1.3382344.
T. Lu, F. Chen, J. Comput. Chem., 2012, 33, 580–592; DOI: https://doi.org/10.1002/jcc.22885.
R. F. Bader, P. J. MacDougall, C. D. H. Lau, J. Am. Chem. Soc., 1984, 106, 1594–1605; DOI: https://doi.org/10.1021/ja00318a009.
V. G. Tsirelson, P. F. Zhou, T.-H. Tang, R. F. W. Bader, Acta Crystallogr., 1995, A51, 143–153; DOI: https://doi.org/10.1107/S0108767394009463.
A. V. Nemukhin, B. L. Grigorenko, S. V. Lushchekina, S. D. Varfolomeev, Russ. Chem. Bull., 2021, 70, 2084–2089; DOI: https://doi.org/10.1007/s11172-021-3319-8.
M. G. Khrenova, B. L. Grigorenko, A. V. Nemukhin, ACS Catal., 2021, 11, 8985–8998; DOI: https://doi.org/10.1021/acscatal.
M. G. Khrenova, A. M. Kulakova, A. V. Nemukhin, J. Chem. Inf. Model., 2021, 61, 1215–1225; DOI: https://doi.org/10.1021/acs.jcim.0c01308.
M. G. Khrenova, B. L. Grigorenko, A. V. Nemukhin, ACS Catal., 2021, 11, 8985–8998; DOI: https://doi.org/10.1021/acscatal.1c00582.
Author information
Authors and Affiliations
Corresponding author
Additional information
Based on materials of the XXXIII Symposium “Modern Chemical Physics” (September 24–October 4, 2021, Tuapse, Russia).
Published in Russian in Izvestiya Akademii Nauk. Seriya Khimicheskaya, No. 5, pp. 921–926, May, 2022.
The research is carried out using the equipment of the shared research facilities of HPC computing resources at Lomonosov Moscow State University.
This work was financially supported by the Russian Foundation for Basic Research (Project No. 21-33-70001).
No human or animal subjects were used in this research.
The authors declare no competing interests.
Rights and permissions
About this article
Cite this article
Kulakova, A.M., Mulashkina, T.I., Nemukhin, A.V. et al. Influence of the leaving group on the mechanism of hydrolysis of organophosphorus compounds by phosphotriesterase from bacterium Pseudomonas diminuta. Russ Chem Bull 71, 921–926 (2022). https://doi.org/10.1007/s11172-022-3491-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11172-022-3491-5