Abstract
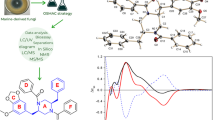
In this study, the activation of pyruvate kinase enzyme in vitro via different urea substituents in the para position as functional groups of 1,4-dihydropyridine derivatives synthesized by Hantzsch reaction method was investigated. Elemental analysis, 1H-NMR, 13C-NMR and FT-IR spectroscopy were used to identify the ureido phenyl substituted 1,4-dihydropyridine derivatives. Virtual screening based on molecular docking supported the results of possible in vitro pyruvate kinase (PK) activators among the synthesized substances. The results showed that all compounds successfully activated PK. The strongest activator effect was shown by ethyl-4-(4-(4-(3-(3-methoxyphenyl)thioureido)phenyl)-2,7,7-trimethyl-5-oxo-1,4,5,6,7,7,8-hexahydroquinolin-3 (7) with an AC50 value of 87.70 µM. In molecular docking studies, full compatibility (− 3016.93 FF), binding affinities (ΔG = − 8.58 kcal/mol), LUMO–HOMO energy gap (ΔE = 7.85 eV) in Density functional theory (DFT) studies and drug similarity score of the compounds were found to be 0.69. These results shed light on the therapeutic potential of the produced compounds to treat PK-related diseases.


Similar content being viewed by others
Data availability
All data are available in the article and its supplementary material.
References
V. Gupta, R.N.K. Bamezai, Protein Sci. 505, 19 (2010). https://doi.org/10.1002/pro.505
N. Wong, J. De Melo, D. Tang, Int. J. Cell Biol. 2013, 242513 (2013). https://doi.org/10.1155/2013/242513
A. Wolf, S. Agnihotri, A. Guha, Oncotarget 190, 1 (2010). https://doi.org/10.18632/oncotarget.190,
S. Strumilo, A. Tylicki, J. Evol. Biochem. Physiol. 1132, 51 (2015). https://doi.org/10.1134/S0022093015020040
T.L. Dayton, T. Jacks, M.G. Vander Heiden, EMBO Rep. 17, 12 (2016). https://doi.org/10.15252/embr.201643300
W.J. Israelsen, M.G. Vander Heiden, Semin. Cell Dev. Biol. 8, 43 (2015). https://doi.org/10.1016/j.semcdb.2015.08.004
Bücher T, Pfleiderer G. Methods in Enzymology, (Academic Press, 1955), pp. 1 DOI: https://doi.org/10.1016/0076-6879(55)01071-9
T.M. Larsen, L.T. Laughlin, H.M. Holden, I. Rayment, G.H. Reed, Biochemistry 33, 20 (1994). https://doi.org/10.1021/bi00186a033
R.P. Tomás, I. Pérez-Guillén, Cancers Basel 12, 11 (2020). https://doi.org/10.3390/cancers12113244
E.M.P. McDermott, A.M. Curtis, G. Goel, Cell Metab. 21, 1 (2015). https://doi.org/10.1016/j.cmet.2014.12.005
S. Patel, C. Globisch, P. Pulugu, P. Kumar, A. Jain, A. Shard, Eur. J. Pharm. Sci. 2012, 170 (2022). https://doi.org/10.1016/j.ejps.2021.106112e
D. Anastasiou, Y. Yu, W.J. Israelsen et al., Nat. Chem. Biol. 57, 8 (2012). https://doi.org/10.1038/nchembio.1060
H. Lineweaver, D. Burk, J. Am. Chem. 56, 3 (1934). https://doi.org/10.1021/ja01318a036
I. Harris, S. McCracken, T.W. Mak, Cell Res. 203, 22 (2012). https://doi.org/10.1038/cr.2011.203
Q. Sun, X. Chen, J. Ma, Proc. Natl. Acad. Sci. 108, 10 (2011). https://doi.org/10.1073/pnas.1014769108
D. Anastasiou, G. Poulogiannis, J.M. Asara et al., Science 334, 6060 (2011). https://doi.org/10.1126/science.1211485
C.S. Deane, B.E. Phillips, C.R.G. Willis et al., Gero Sci. (2022). https://doi.org/10.1007/s11357-022-00658-5
M.J. Kim, I.S. Sinam, Z. Siddique, J.H. Jeon, I.K. Lee, Diabetes Metab. J. 47, 2 (2023). https://doi.org/10.4093/dmj.2022.0305
T. Demirci, B. Çelik, Y. Yildiz, S. Eriş, M. Arslan, F. Sen, B. Kilbas, RSC Adv. 6, 80 (2016). https://doi.org/10.1039/C6RA13142E
M. Mamaghani, M. Sheykhan, M. Sadeghpour, F. Tavakoli, Monatshefte Chem. 13, 149 (2018). https://doi.org/10.1007/s00706-018-2166-2
H. Ahankar, A. Ramazani, S.W. Joo, Res. Chem. Intermed. 21, 42 (2016). https://doi.org/10.1007/s11164-015-2163-6
M. Patil, S. Karhale, A. Kudale, A. Kumbhar, S. More, V. Helavi, Curr. Sci. 116, 6 (2019). https://doi.org/10.18520/cs/v116/i6/936-942
S. Besoluk, M. Kucukislamoglu, M. Nebioglu, M. Zengin, M. Arslan, J. Iran. Chem. Soc. 17, 5 (2008). https://doi.org/10.1007/BF03245816
G.M. Ziarani, P. Gholamzadeh, A. Badiei, S. Asadi, A.A. Soorki, J. Chil. Chem. Soc. 60, 2 (2015). https://doi.org/10.4067/S0717-97072015000200024
D.M. Stout, A.I. Meyers, Chem. Rev. 82, 222 (1982). https://doi.org/10.1021/cr00048a004
K. Sharma, S. Chromone, Chem. Sel. 7, e202200540 (2022). https://doi.org/10.1002/slct.202200540
A.O. Bryzgalov, M.P. Dolgikh, I.V. Sorokina, T.G. Tolstikova, V.F. Sedova, O.P. Shkurko, Bioorg. Med. Chem. Lett. 16, 1418 (2006). https://doi.org/10.1016/j.bmcl.2005.11.043
S. Viveka, L.N. Madhu, G.K. Nagaraja, Monatshefte Chem. 18, 146 (2015). https://doi.org/10.1007/s00706-015-1428-5
A. Trivedi, D. Dodiya, B. Dholariya, V. Kataria, V. Bhuva, V. Shah, Chem. Biol. Drug Des. 78, 875 (2011). https://doi.org/10.1111/j.1747-0285.2011.01233.x
M. Khoshneviszadeh, N. Edraki, K. Javidnia, A. Alborzi, B. Pourabbas, J. Mardaneh, R. Miri, Bioorganic. Med. Chem. 14, 14574 (2009). https://doi.org/10.1016/j.bmc.2008.12.070
S. Ulloora, R. Shabaraya, R. Ranganathan, A.V. Adhikari, Eur. J. Med. Chem. 19, 70 (2013). https://doi.org/10.1016/j.ejmech.2013.10.010
R.H. Tale, A.H. Rodge, G.D. Hatnapure, A.P. Keche, K.M. Patil, R.P. Pawar, Med. Chem. Res. 53, 1450 (2013). https://doi.org/10.1007/s00044-012-0109-8
P.J. Marín, A.G.L. Pardo, C.P. Rossignoli, M.G. Durruthy, E.O. Rodríguez, Y.V. Reyes, R.F. Acosta, S.A. Uyemura, L.C. Alberici, Toxicol. Vitr. 17, 42 (2017). https://doi.org/10.1016/j.tiv.2017.03.011
Z.C. Nava, S.S. Estrada, Á.J. Guerrero, Bioorganic Med. Chem. 18, 6398 (2010). https://doi.org/10.1016/j.bmc.2010.07.008
P.A. Datar, P.B. Auti, J. Comput. Methods Mol. Des. 2, 85 (2012). https://doi.org/10.13140/RG.2.2.25106.76487
X. Huang, J. Su, A.U. Rao et al., Bioorganic Med. Chem. Lett. 22, 854 (2012). https://doi.org/10.1016/j.bmcl.2011.12.041
M. Alvala, S. Bhatnagar, A. Ravi, V.U. Jeankumar, T.H. Manjashetty, P. Yogeeswari, D. Sriram, Bioorganic Med. Chem. Lett. 22, 3256 (2012). https://doi.org/10.1016/j.bmcl.2012.03.030
M. Teleb, F.X. Zhang, J. Huang, V.M. Gadotti, A.M. Farghaly, O.M. AboulWafa, G.W. Zamponi, H. Fahmy, Bioorganic. Med. Chem. (2017). https://doi.org/10.1016/j.bmc.2017.02.015
D.J. Triggle, Cell Mol. Neurobiol. 23, 23 (2003). https://doi.org/10.1023/a:1023632419813
N. Pedemonte, D. Boido, O. Moran, M. Giampieri, M. Mazzei, L.J.V. Galietta, Mol. Pharmacol. 72, 191 (2007). https://doi.org/10.1124/mol.107.034702
H.R. Christofk, H.M.G. Vander, N. Wu, J.M. Asara, L.C. Cantley, Nature 96, 452 (2008). https://doi.org/10.1038/nature06667
A.J. Lakhter, T. Hato, B.M. Shankar, S.R. Naidu, S. Paul, S. Mahanta, PLoS ONE 13, e0191419 (2018). https://doi.org/10.1371/journal.pone.0191419
A. Turberville, H. Semple, G. Davies, D. Ivanov, G.A. Holdgate, SLAS Discov. 27, 418 (2022). https://doi.org/10.1016/j.slasd.2022.09.001
S. Adem, A. Aslan, I. Ahmed, K. Krohn, C. Guler, V. Comaklı, R. Demirdag, M. Kuzu, Arch. Pharm. 349, 230 (2016). https://doi.org/10.1002/ardp.201500357
A. Shrestha, M. Chi, K. Wagner, A. Malik, J. Korpik, A. Drake, K. Fulzele, S. Guichard, P. Malik, Blood Adv. 5, 9 (2021). https://doi.org/10.1182/bloodadvances.2020003604
I. Kaur, W. Jia, R.P. Kopreski, S. Selvarasah, M.R. Dokmeci, C. Pramanik, N.E. McGruer, G.P. Miller, J. Am. Chem. Soc. 130, 48 (2008). https://doi.org/10.1021/ja804515y
Ü. Çalışır, F. Çakır, Turk. J. Technol. Appl. Sci. 3, 11 (2022). https://doi.org/10.5281/zenodo.7236656
R.F.W. Bader, P.M. Beddall, P.E. Cade, J. Am. Chem. Soc. 93, 13 (1971). https://doi.org/10.1021/ja00742a001
R.J. Ouellette, J.D. Rawn. In Structure of Organic Compounds, ed. by R.J. Ouellette, J.D. Rawn. Principles of Organic Chemistry. (Elsevier, 2015), pp. 31 DOI: https://doi.org/10.1016/B978-0-12-802444-7.00001-X
Y. Mei, A.C. Simmonett, F.C. Pickard, R.A. DiStasio Jr., B.R. Brooks, Y. Shao, J. Phys. Chem. A 119, 22 (2015). https://doi.org/10.1021/acs.jpca.5b03159
A.K. Ghose, T. Herbertz, R.L. Hudkins, B.D. Dorsey, J.P. Mallamo, ACS Chem. Neurosci. 3, 12 (2012). https://doi.org/10.1021/cn200100h
S. Winiwarter, F. Ax, H. Lennernäs, A. Hallberg, C. Pettersson, A. Karlén, J. Mol. Graph. Model. 21, 4 (2003). https://doi.org/10.1016/s1093-3263(02)00163-8
E.M. Gad, M.S. Nafie, E.H. Eltamany, M.S.A.G. Hammad, A. Barakat, A.T.A. Boraei, Molecules 25, 11 (2020). https://doi.org/10.3390/molecules25112523
C.A. Lipinski, F. Lombardo, B.W. Dominy, P.J. Feeney, Adv. Drug Deliv. Rev. 46, 1 (2001). https://doi.org/10.1016/s0169-409x(00)00129-0
A.K. Ghose, V.N. Viswanadhan, J.J. Wendoloski, J. Comb. Chem. 1, 156 (1999). https://doi.org/10.1021/cc9800071
D.F. Veber, S.R. Johnson, H.Y. Cheng, B.R. Smith, K.W. Ward, K.D. Kopple, J. Med. Chem. 45, 12 (2002). https://doi.org/10.1021/jm020017n
M. Lobell, M. Hendrix, B. Hinzen, J. Keldenich, H. Meier, C. Schmeck, L.R. Schohe, T. Wunberg, A. Hillisch, Chem. Med. Chem. 1, 11 (2006). https://doi.org/10.1002/cmdc.200600168
A. Grosdidier, V. Zoete, O. Michielin, J. Comput. Chem. 32, 10 (2011). https://doi.org/10.1002/jcc.21797
A. Grosdidier, V. Zoete, O. Michielin, Nucleic Acids Res. 39, W270 (2011). https://doi.org/10.1093/nar/gkr366
E.F. Pettersen, T.D. Goddard, C.C. Huang, G.S. Couch, D.M. Greenblatt, E.C. Meng, T.E. Ferrin, Comput. Chem. 25, 1605 (2004). https://doi.org/10.1002/jcc.20084
N.S. Kaviyarasi, C.N. Prashantha, V.V.S. Suryanarayana. Int. J. Pharm. Pharm. Sci. 8:123 DOI: https://journals.innovareacademics.in/index.php/ijpps/article/view/10713
R.O. Gould, A.M. Gray, P. Taylor, M.D. Walkinshaw, J. Am. Chem. Soc. 107, 5921 (1985). https://doi.org/10.1021/ja00307a016
V.D. Kharisma, S.L. Utami, W.C. Rizky, T.G.A. Dings, M.E. Ullah, V. Jakhmola, A.P. Nugraha, Dent. J. 56, 23 (2023). https://doi.org/10.20473/j.djmkg.v56.i1.p23-29
N.J. Zondlo, Aromatic–proline interactions: electronically tunable CH/π interactions. Acc. Chem. Res. 46, 1039 (2013). https://doi.org/10.1021/ar300087y
S.E. Scott, J.P. Fernandez, C.M. Hadad, A.A. MacKay, Environ. Sci. Technol. 56, 951 (2022). https://doi.org/10.1021/acs.est.1c06147
M. Kumari, U.K. Singh, P. Singh, R. Patel, Chem. Sel. 69, 1241 (2017). https://doi.org/10.1002/slct.201601477
A.K. Aranda-Rivera, A. Cruz-Gregorio, O.E. Aparicio-Trejo, A.J. Ortega-Lozano, J. Pedraza-Chaverri, Free Radic. Biol. Med. 56, 172 (2021). https://doi.org/10.1016/j.freeradbiomed.2021.05.034
U. Johnsen, A. Reinhardt, G. Landan, F.D.K. Tria, J.M. Turner, C. Davies, P. Schönheit, FEBS J. 78, 2471 (2019). https://doi.org/10.1111/febs.14837
M.O. Kaya, T. Demirci, O. Ozdemir, U. Calisir, F. Sonmez, M. Arslan, Med. Chem. Res. 23, 841 (2023). https://doi.org/10.1007/s00044-023-03029-7
N.M. O’Boyle, M. Banck, C.A. James, C. Morley, T. Vandermeersch, Hutchison GRJ Cheminform. 56, 33 (2011). https://doi.org/10.1186/1758-2946-3-33
Dassault Systèmes BIOVIA (2020) Discovery studio 2020 Client.
A. Daina, O. Michielin, V. Zoete, J. Chem. Inf. Model. 54, 3284 (2014). https://doi.org/10.1021/ci500467k
A. Daina, O. Michielin, V. Zoete, Sci. Rep. 12, 42717 (2017). https://doi.org/10.1038/srep42717
F. Neese, Softw. Updat. Comput. Mol. Sci. 12, e1606 (2022). https://doi.org/10.1002/wcms.1606
A.D. Becke, J. Chem. Phys. 98, 1372 (1993). https://doi.org/10.1063/1.464304
C. Lee, W. Yang, R.G. Parr, Phys. Rev. B 37, 785 (1988). https://doi.org/10.1103/PhysRevB.37.785
S.H. Vosko, L. Wilk, M. Nusair, Can. J. Phys. 58, 1200 (1980). https://doi.org/10.1139/p80-159
P.J. Stephens, F.J. Devlin, C.F. Chabalowski, M.J. Frisch, J. Phys. Chem. 98, 11623 (1994). https://doi.org/10.1021/j100096a001
S. Grimme, J. Antony, S. Ehrlich, H. Krieg, J. Chem. Phys. 63, 132 (2010). https://doi.org/10.1063/1.3382344
S. Grimme, S. Ehrlich, L. Goerigk, J. Comput. Chem. 32, 1456 (2011). https://doi.org/10.1002/jcc.21759
F. Weigend, Ahlrichs. Phys. Chem. Chem. Phys. 7, 3297 (2005). https://doi.org/10.1039/b508541a
Author information
Authors and Affiliations
Contributions
In this study, both M.O.K. and M.A. designed the study. M.O.K., T.D., O.Ö., Y.K. and Ü.Ç. and performed all experiments/analysis, enzymatic and computational studies. All authors evaluated the results and prepared the manuscript.
Corresponding author
Ethics declarations
Confict of interest
The authors declare that they have no confict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kaya, M.O., Demirci, T., Çalışır, Ü. et al. Synthesis, activatory effects, molecular docking and ADME studies as rabbit muscle pyruvate kinase activators of ureido phenyl substituted 1,4-dihydropyridine derivatives. Res Chem Intermed 50, 437–463 (2024). https://doi.org/10.1007/s11164-023-05149-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11164-023-05149-6