Abstract
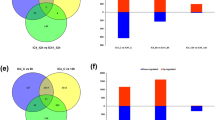
Broomcorn millet (Panicum miliaceum L.) is an early domesticated C4 cereal crop with high water utilization ability, nutrient content, favored agronomic traits, and tolerance to a broad spectrum of abiotic stresses. Hyperosmotic stress is an inevitable consequence of strong drought on plant, which causes severe dehydration and adversely affects the growth and yield of plants. In this study, we used PEG-6000 to simulate drought conditions and identified differentially expressed gene in broomcorn millet root and shoot under different PEG-6000 concentrations. A total of 11,452 and 14,952 significantly different expression genes (DEGs) were detected separately in root and shoot under at least one PEG treatment. Of these, 2991 and 2051 DEGs were commonly detected at all different PEG levels in root and shoot, respectively, and 535 DEGs were shared in both root and shoot at all different PEG levels, including six genes differently regulated in root and shoot. Enrichment analysis categorized these DEGs into 85 and 56 GO terms, or 17 and 6 significant KEGG pathways in root and shoot, respectively. And 39 hub genes especially longmi029229 associated with hyperosmotic stress were detected by weighted gene co-expression network analysis. In addition, 29 TCP transcription factors were identified and characterized in broomcorn millet, which were divided into three clades. Phylogenetic analysis indicated that the TCPs of monocot broomcorn millet and rice were closer than the TCPs of monocot broomcorn millet and dicot Arabidopsis. Analysis of the expression pattern of PmTCPs to drought stress indicated that 11 and 20 TCP genes showed significant change under PEG-6000 treatment in root and shoot separately. These DEGs and PmTCP genes with differential expression patterns in shoot and root deserve further study for uncovering the mechanism of drought resistance in broomcorn millet.








Similar content being viewed by others
References
Ali MA, Jabran K, Awan SI, Abbas A, Ehsanullah ZM et al (2011) Morpho-physiological diversity and its implications for improving drought tolerance in grain sorghum at different growth stages. Aust J Crop Sci 5:308–317
Anjum SA, Ashraf U, Zohaib A, Tanveer M, Naeem M, Ali I et al (2017) Growth and developmental responses of crop plants under drought stress: a review. Zemdirbyste 104:267–276. https://doi.org/10.13080/z-a.2017.104.034
Barth O, Vogt S, Uhlemann R, Zschiesche W, Humbeck K (2009) Stress induced and nuclear localized HIPP26 from Arabidopsis thaliana interacts via its heavy metal associated domain with the drought stress related zinc finger transcription factor ATHB29. Plant Mol Biol 69:213–226. https://doi.org/10.1007/s11103-008-9419-0
Barth O, Zschiesche W, Siersleben S, Humbeck K (2004) Isolation of a novel barley cDNA encoding a nuclear protein involved in stress response and leaf senescence. Physiol Plantarum 121:282–293. https://doi.org/10.1111/j.0031-9317.2004.00325.x
Bhaskara GB, Nguyen TT, Verslues PE (2018) Unique drought resistance functions of the highly ABA-induced clade A protein phosphatase 2Cs. Plant Physiol 178:1423–1423. https://doi.org/10.1104/pp.18.01168
Bohnert HJ, Jensen RG (1996) Strategies for engineering water-stress tolerance in plants. Trends Biotechnol 14:89–97. https://doi.org/10.1016/0167-7799(96)80929-2
Bray NL, Pimentel H, Melsted P, Pachter L (2016) Near-optimal probabilistic RNA-seq quantification. Nat Biotechnol 34:525–527. https://doi.org/10.1038/nbt.3519
Brini F, Masmoudi K (2012) Ion transporters and abiotic stress tolerance in plants. ISRN Mol Biol 2012:927436. https://doi.org/10.5402/2012/927436
Bui H, Greenhalgh R, Ruckert A, Gill GS, Lee S, Ramirez RA et al (2018) Generalist and specialist mite herbivores induce similar defense responses in maize and barley but differ in susceptibility to benzoxazinoids. Front Plant Sci 9:1222. https://doi.org/10.3389/fpls.2018.01222
Chini A, Fonseca S, Fernandez G, Adie B, Chico JM, Lorenzo O et al (2007) The JAZ family of repressors is the missing link in jasmonate signalling. Nature 448:666-+. https://doi.org/10.1038/nature06006
Cubas P, Lauter N, Doebley J, Coen E (1999) The TCP domain: a motif found in proteins regulating plant growth and development. Plant J 18:215–222. https://doi.org/10.1046/j.1365-313X.1999.00444.x
Dai M, Huang SZ, Huang Q, Leng GY, Guo Y, Wang L et al (2020) Assessing agricultural drought risk and its dynamic evolution characteristics Agr Water Manage 231. https://doi.org/10.1016/j.agwat.2020.106003
Das S, Khound R, Santra M, Santra DK (2019) Beyond bird feed: proso millet for human health and environment Agriculture-Basel 9. https://doi.org/10.3390/agriculture9030064
de Zelicourt A, Colcombet J, Hirt H (2016) The role of MAPK modules and ABA during abiotic stress signaling. Trends Plant Sci 21:677–685. https://doi.org/10.1016/j.tplants.2016.04.004
Ding SC, Cai ZZ, Du HW, Wang HW (2019) Genome-wide analysis of TCP family genes in Zea mays L. identified a role for ZmTCP42 in drought tolerance. Int J Mol Sci 20. https://doi.org/10.3390/ijms20112762
Ding XP, Li XK, Xiong LZ (2013) Insight into differential responses of upland and paddy rice to drought stress by comparative expression profiling analysis. Int J Mol Sci 14:5214–5238. https://doi.org/10.3390/ijms14035214
Ding Y, Lapko H, Ndamukong I, Xia Y, Al-Abdallat A, Lalithambika S et al (2009) The Arabidopsis chromatin modifier ATX1, the myotubularin-like AtMTM and the response to drought. Plant Signal Behav 4:1049–1058. https://doi.org/10.4161/psb.4.11.10103
Fang P, Yao QL, Chen FB (2011) Morpho-physiological characteristics of maize (Zea mays L.) landraces under water stress. Philipp Agric Sci 94:323–328
Feng ZJ, Xu SC, Liu N, Zhang GW, Hu QZ, Gong YM (2018) Soybean TCP transcription factors: evolution, classification, protein interaction and stress and hormone responsiveness. Plant Physiol Biochem 127:129–142. https://doi.org/10.1016/j.plaphy.2018.03.020
Hejnak V, Tatar O, Atasoy GD, Martinkova J, Celen AE, Hnilicka F et al (2015) Growth and photosynthesis of upland and pima cotton: response to drought and heat stress. Plant Soil Environ 61:507–514. https://doi.org/10.17221/512/2015-Pse
Horn S, Pabon-Mora N, Theuss VS, Busch A, Zachgo S (2015) Analysis of the CYC/TB1 class of TCP transcription factors in basal angiosperms and magnoliids. Plant J 81:559–571. https://doi.org/10.1111/tpj.12750
Hunt HV, Badakshi F, Romanova O, Howe CJ, Jones MK, Heslop-Harrison JSP (2014) Reticulate evolution in Panicum (Poaceae): the origin of tetraploid broomcorn millet. P Miliaceum J Exp Bot 65:3165–3175. https://doi.org/10.1093/jxb/eru161
Jaleel CA, Manivannan P, Wahid A, Farooq M, Al-Juburi HJ, Somasundaram R et al (2009) Drought stress in plants: a review on morphological characteristics and pigments composition. Int J Agric Biol 11:100–105
Janiak A, Kwasniewski M, Sowa M, Kuczynska A, Mikolajczak K, Ogrodowicz P et al (2019) Insights into barley root transcriptome under mild drought stress with an emphasis on gene expression regulatory mechanisms Int J Mol Sci 20. https://doi.org/10.3390/ijms20246139
Kole C, Muthamilarasan M, Henry R, Edwards D, Sharma R, Abberton M et al (2015) Application of genomics-assisted breeding for generation of climate resilient crops: progress and prospects Front Plant Sci 6. https://doi.org/10.3389/fpls.2015.00563
Langfelder P, Horvath S (2008) WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9:559
Lei N, Yu X, Li S, Zeng C, Zou L, Liao W et al (2017) Phylogeny and expression pattern analysis of TCP transcription factors in cassava seedlings exposed to cold and/or drought stress. Sci Rep 7:10016. https://doi.org/10.1038/s41598-017-09398-5
Li B, Liu Y, Cui XY, Fu JD, Zhou YB, Zheng WJ et al (2019) Genome-wide characterization and expression analysis of soybean TGA transcription factors identified a novel TGA gene involved in drought and salt tolerance Front Plant Sci 10. https://doi.org/10.3389/fpls.2019.00549
Ling L, Zhang WR, An YM, Du BH, Wang D, Guo CH (2020) Genome-wide analysis of the TCP transcription factor genes in five legume genomes and their response to salt and drought stresses. Funct Integr Genomic 20:537–550. https://doi.org/10.1007/s10142-020-00733-0
Liu HL, Gao YM, Wu M, Shi YN, Wang H, Wu L et al (2020) TCP10, a TCP transcription factor in moso bamboo (Phyllostachys edulis), confers drought tolerance to transgenic plants Environ Exp Bot 172. https://doi.org/10.1016/j.envexpbot.2020.104002
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2 -△△CT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Lu HY, Zhang JP, Liu KB, Wu NQ, Li YM, Zhou KS et al (2009) Earliest domestication of common millet (Panicum miliaceum) in East Asia extended to 10,000 years ago. P Natl Acad Sci USA 106:7367–7372. https://doi.org/10.1073/pnas.0900158106
Mahmood T, Khalid S, Abdullah M, Ahmed Z, Shah MKN, Ghafoor A et al (2020) Insights into drought stress signaling in plants and the molecular genetic basis of cotton drought tolerance Cells-Basel 9. https://doi.org/10.3390/cells9010105
Martin-Trillo M, Cubas P (2010) TCP genes: a family snapshot ten years later. Trends Plant Sci 15:31–39. https://doi.org/10.1016/j.tplants.2009.11.003
Mortazavi A, Williams BA, Mccue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-seq. Nat Methods 5:621–628. https://doi.org/10.1038/nmeth.1226
Mukhopadhyay P, Tyagi AK (2015) OsTCP19 influences developmental and abiotic stress signaling by modulating ABI4-mediated pathways. Sci Rep 5:9998. https://doi.org/10.1038/srep12381
Ndamukong I, Jones DR, Lapko H, Divecha N, Avramova Z (2010) Phosphatidylinositol 5-phosphate links dehydration stress to the activity of Arabidopsis trithorax-like factor ATX1 PLoS One 5. https://doi.org/10.1371/journal.pone.0013396
Pandey V, Shukla A (2015) Acclimation and tolerance strategies of rice under drought stress. Rice Sci 22:147–161. https://doi.org/10.1016/j.rsci.2015.04.001
Qu YY, Mu P, Zhang HL, Chen CY, Gao YM, Tian YX et al (2008) Mapping QTLs of root morphological traits at different growth stages in rice. Genetica 133:187–200. https://doi.org/10.1007/s10709-007-9199-5
Ravelombola W, Shi AN, Qin J, Weng YJ, Bhattarai G, Zia B et al (2018) Investigation on various aboveground traits to identify drought tolerance in cowpea seedlings. HortScience 53:1757–1765. https://doi.org/10.21273/Hortsci13278-18
Riemann M, Dhakarey R, Hazman M, Miro B, Kohli A, Nick P (2015) Exploring jasmonates in the hormonal network of drought and salinity responses Front Plant Sci 6. https://doi.org/10.3389/fpls.2015.01077
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140. https://doi.org/10.1093/bioinformatics/btp616
Shan ZY, Jiang YM, Li HQ, Guo JJ, Dong M, Zhang JA et al (2020) Genome-wide analysis of the NAC transcription factor family in broomcorn millet (Panicum miliaceum L.) and expression analysis under drought stress. BMC Genomics 21. https://doi.org/10.1186/s12864-020-6479-2
Sharma R, Kapoor M, Tyagi AK, Kapoor S (2010) Comparative transcript profiling of TCP family genes provide insight into gene functions and diversification in rice and Arabidopsis. J Plant Mol Biol Biotechnol 1:24–38
Shi JP, Ma XX, Zhang JH, Zhou YS, Liu MX, Huang LL et al (2019) Chromosome conformation capture resolved near complete genome assembly of broomcorn millet Nat Commun 10. https://doi.org/10.1038/s41467-018-07876-6
Shinozaki K, Yamaguchi-Shinozaki K (2007) Gene networks involved in drought stress response and tolerance. J Exp Bot 58:221–227. https://doi.org/10.1093/jxb/erl164
Tanaka Y, Fujii K, Shiraiwa T (2010) Variability of leaf morphology and stomatal conductance in soybean [Glycine max (L.) Merr.] cultivars. Crop Sci 50:2525–2532. https://doi.org/10.2135/cropsci2010.02.0058
Tian XM, Zhang L, Feng SS, Zhao ZY, Wang XP, Gao H (2019) Transcriptome analysis of apple leaves in response to powdery mildew (Podosphaera leucotricha) infection Int J Mol Sci 20. https://doi.org/10.3390/ijms20092326
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The Clustal_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882. https://doi.org/10.1093/nar/25.24.4876
Wang J, Zhang S, Fu Y, He T, Wang X (2020) Analysis of dynamic global transcriptional atlas reveals common regulatory networks of hormones and photosynthesis across Nicotiana varieties in response to long-term drought Front Plant Sci 11. https://doi.org/10.3389/fpls.2020.00672
Wang YN, Liu C, Li KX, Sun FF, HZ Hu X Li et al (2007) Arabidopsis EIN2 modulates stress response through abscisic acid response pathway Plant Mol Biol 64:633-644. https://doi.org/10.1007/s11103-007-9182-7
Wu QS, Xia RX, Zou YN (2008) Improved soil structure and citrus growth after inoculation with three arbuscular mycorrhizal fungi under drought stress. Eur J Soil Biol 44:122–128. https://doi.org/10.1016/j.ejsobi.2007.10.001
Yue H, Wang M, Liu SY, Du XH, Song WN, Nie XJ (2016) Transcriptome-wide identification and expression profiles of the WRKY transcription factor family in broomcorn millet (Panicum miliaceum L.). BMC Genomics 17. https://doi.org/10.1186/s12864-016-2677-3
Zhang J, Li DD, Zou D, Luo F, Wang XL, Zheng Y et al (2013) A cotton gene encoding a plasma membrane aquaporin is involved in seedling development and in response to drought stress. Acta Bioch Bioph Sin 45:104–114. https://doi.org/10.1093/abbs/gms096
Zhang YY, Gao XL, Li J, Gong XW, Yang P, Gao JF et al (2019) Comparative analysis of proso millet (Panicum miliaceum L.) leaf transcriptomes for insight into drought tolerance mechanisms. Bmc Plant Biol 19. https://doi.org/10.1186/s12870-019-2001-x
Zhang ZL, Zhang SP, Zhang Y, Wang X, Li D, Li QL et al (2011) Arabidopsis floral initiator SKB1 confers high salt tolerance by regulating transcription and pre-mRNA splicing through altering histone H4R3 and small nuclear ribonucleoprotein LSM4 methylation. Plant Cell 23:396–411. https://doi.org/10.1105/tpc.110.081356
Zhou M, Li DY, Li ZG, Hu Q, Yang CH, Zhu LH et al (2013) Constitutive expression of a miR319 gene alters plant development and enhances salt and drought tolerance in transgenic creeping bentgrass. Plant Physiol 161:1375–1391. https://doi.org/10.1104/pp.112.208702
Zhu JK (2016) Abiotic stress signaling and responses in plants. Cell 167:313–324. https://doi.org/10.1016/j.cell.2016.08.029
Zou CS, Li LT, Miki D, Li DL, Tang QM, Xiao LH et al (2019) The genome of broomcorn millet Nat Commun 10. https://doi.org/10.1038/s41467-019-08409-5
Zou YN, Wang P, Liu CY, Ni QD, Zhang DJ, Wu QS (2017) Mycorrhizal trifoliate orange has greater root adaptation of morphology and phytohormones in response to drought stress Sci Rep 7. https://doi.org/10.1038/srep41134
Funding
This research was financially supported by the Second Tibetan Plateau Scientific Expedition and Research Program (STEP) (2019QZKK0303), the China Agriculture Research System (CARS-06–14.5-A8), Special Project of Agricultural Science and Technology Innovation of GAAS (2021GAAS02), and Science & Technology Supporting Project for Young Talents in Gansu Province (2018).
Author information
Authors and Affiliations
Contributions
T.L. and W.W. carried out the experiments and data analysis and wrote the first draft. J.H., K.D., R.R., and L.Z. participated in the phenotypic measurements. X.W. contributed to manuscript preparation. Y.H. and M.W. finished the RNA extraction, cDNA synthesis, and qRT-PCR analysis. P.Y. contributed to primer design. Z.Z. and T.Y. contributed to experimental design. All the authors have read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key Message
Broomcorn millet (Panicum miliaceum L.) is a water-saving and drought-tolerant crop. DEGs, hub genes, and TCP transcription factors identified in shoot and root offer insights into dissecting drought tolerance mechanism of broomcorn millet.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, T., Wang, W., He, J. et al. Transcriptome Profiling and TCP Family Analysis of Broomcorn Millet (Panicum miliaceum L.) Seedlings Under Hyperosmotic Stress. Plant Mol Biol Rep 41, 277–291 (2023). https://doi.org/10.1007/s11105-022-01365-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-022-01365-3