Abstract
The j allele delays flowering and enhances yield of long juvenile (LJ) soybean under short day (SD) condition. However, the underlying mechanism of j in flowering pathway of soybean is not fully known. The objective of the study is to profile the transcriptome of Huaxia-3 (HX3), a typical long juvenile soybean variety with a loss of function allele (j) for the J gene. This helps to identify the genes implicated in delayed flowering in the long juvenile soybean variety and the pathways involved. Phenotypic analysis revealed HX3-delayed flowering and matured later than the transgenic line with the functional J gene under SD. RNA-Seq analysis was conducted to compare the transcriptome of HX3 relative to the one overexpressing J gene under SD and long day (LD). A total of 674,800,494 clean reads were generated, of which 626,517,161 (93%) were uniquely mapped to the soybean reference genome. A total of 31 and 2311 genes were differentially expressed in the HX3 under SD and LD conditions respectively. The circadian rhythm pathway was the most significantly enriched pathway in the HX3 under SD condition. The GmELF3a (Glyma.04G050200) and FLOWERING LOCUS T (FT) genes such as GmFT2a (Glyma.16G150700) and GmFT5a (Glyma.16G044100) were downregulated, whiles GmFT4 (Glyma.08G363100) was upregulated in the HX3 under SD. Under LD FT homologs, GmFT4 (Glyma.08G363100) was downregulated and GmFT1a (Glyma.18G299000) was upregulated. Our study suggests that these FT homologs may be involved in delayed flowering of LJ soybean under SD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Soybean [Glycine max (L.) Merr.] is a short-day plant domesticated about 5000 years ago in temperate region of China at approximately 35°N latitude (Hymowitz, 1970; Carter et al. 2004). Nowadays, soybean is one of the important field crops, providing dietary proteins, and edible oil worldwide, and the demand for soybean production in tropical regions is rising but the available high-yielding soybean varieties originally bred for middle and high latitudes perform poorly in tropical regions because of inadequate adaptation. When temperate elite soybean varieties are grown in the tropics, a majority of them bloom nearly 3 weeks after emergence leading to low yields (Destro et al. 2001). Short day (SD) (~ 12 h) leads to a significant reduction of the soybean vegetative growth. The short stature and a low leaf area index when the vegetative stage of development is stunted leads to reduced yields (Sinclair and Hinson 1992). Therefore, there is a need for sustained research and development aiming at adapting high yielding temperate varieties to tropical climates to increase global soybean production since the demand keeps rising.
The long juvenile (LJ) trait which delays flowering in soybean under SD conditions was discovered and has resulted in a major breakthrough in the expansion of soybean commercial production at latitudes below 22° (Hartwig and Kiihl 1979; Neumaier and James 1993; Spehar 1995). An assessment of the genetic control of the LJ trait in soybean by Destro et al. (2001) reported that up to five genes can control the LJ trait and that recessive alleles at two loci are necessary for the LJ trait. Studies by Yue et al. (2017) and Lu et al. (2017) concluded that J is the dominant functional allele of GmELF3a, an ortholog of ELF3 in Arabidopsis, which inhibits flowering, while j is the recessive, loss-of-function allele that causes delayed flowering and confers the LJ trait that is associated with the adaptation of soybean to tropical regions. In both studies, the J gene was confirmed by transgenic complementation of the GmELF3a gene in a LJ variety. A recent report by Sun et al. (2019) speculated that the LJ trait could also result partly from a combination of various dysfunctional alleles at GmFT2a loci. FLOWERING LOCUS T (FT) is an important flowering integrator in Arabidopsis, with homologs in many other plants. The manipulation of FT homologs in soybean can facilitate the creation of LJ soybean germplasm (Cai et al. 2020; Li et al. 2020a, b).
Numerous studies with the model plant Arabidopsis enhanced the understanding of the molecular mechanisms of flowering in many plants. Many of these studies have identified important genes, such as FT and CONSTANS (CO), and their involvement in flowering (Andres and Coupland 2012; Itoh and Izawa 2013; Pin and Nilsson 2012). Up to date, a lot of genes and loci have been identified to play a vital role in regulating photoperiodic flowering in soybean (Lin et al. 2021). Some of the genes/loci include E1 (Xia et al. 2012), E2 (Watanabe et al. 2011), E3 (Watanabe et al. 2009), E4 (Liu et al. 2008), GmFT2a (Kong et al. 2010; Sun et al. 2011), GmFT4 (Zhai et al. 2014; Samanfar et al. 2017), J/GmELF3a (Lu et al. 2017; Yue et al. 2017), GmFT5a (Kong et al. 2010), GmFT1a/ GmFT1b (Guo et al. 2015; Liu et al. 2018), GmFT2b (Fan et al. 2014; Chen et al. 2020), GmFT2c (Wu et al. 2017), GmFT3a/ GmFT3b (Fan et al. 2014), GmFT5b (Fan et al. 2014), GmFT6 (Wang et al. 2015), GmFT7 (Zhang et al. 2021), PRR3a/ PRR3b (PRR37) (Wang et al. 2020; Li et al. 2020a, b; Lu et al. 2020), GmLux1, and Lux2 (Fang et al. 2021; Bu et al. 2021).
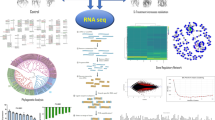
Transcriptome analysis enabled by advances in RNA sequencing (RNA-Seq) and computation biology is a powerful tool to identify gene expression differences and correlations with genetic/developmental cues or environmental conditions (Gillman et al. 2019). RNA-Seq uses deep-sequencing technology that provides a far more precise measurement of the levels of transcripts than other methods (Wang et al. 2009). With the above merits and the low cost nowadays, RNA-Seq has been applied in a lot of studies on flowering in Arabidopsis thaliana and soybeans. In this study, we used RNA-Seq to identify transcriptional variations between HX3, a mutated soybean variety, and the transgenic soybean line overexpressing J gene under SD and long day (LD) condition. This is to help identify the expression of flowering-related genes and the possible pathways which resulted in the delayed flowering in the LJ soybean under SD condition.
Materials and Methods
Soybean Genetic Materials
A natural LJ soybean variety Huaxia-3 and a transgenic line (Joe) overexpressing the J gene were used in the study. The transgenic line was Huaxia-3 transformed with the genome version of the J gene from Zhonghuang-24, a cultivar with conventional juvenility (Yue et al. 2017), and served as the control of the LJ variety. The LJ soybean HX3 was untransformed but has the recessive form of the J gene (j). Each sample for RNA-Seq was collected from the unifoliate leaves (V1 developmental stage) from three randomly selected individual plants, under short- and long-day conditions. All the samples were collected about 9 am and immediately frozen in liquid nitrogen and stored at − 80 °C until RNA extraction. The samples were prepared and coded as HX3L, HX3S, JoeS, and JoeL which refer to Huaxia-3 under long day, Huaxia-3 under short day, transgenic line under short day, and transgenic line under long day, respectively.
Phenotypic Evaluation Under Short- and Long-Day Conditions
The LJ soybean HX3 and the transgenic line were grown under SD (12 h light and darkness) and LD conditions (16 h light and 8 h darkness). The seeds were planted in pots (five plants per pot) on the 16th of May 2019 at the Changping Experimental Station of Institute of Crop Sciences, Chinese Academy of Agricultural Sciences, Beijing, China. Under the SD treatment, the seedlings were placed under natural sunshine for 12 h followed by 12 h in darkness from 7:00 AM to 7:00 PM. For LD treatment, daytime was 5:00 AM–9:00 PM (12 h daylight + 4 h artificial light). The experiments were arranged in a complete randomized design with three replications. Flowering time was recorded at the VE-R1 stage (days from emergence to the first open flower) and maturity was recorded at the VE-R7 stage (days from emergence to physiological maturity) as described by Fehr et al. (1971). For phenotypic evaluation, five individual plants were analyzed per each replicate. SPSS software package was used to analyze the phenotypic data (version 19.0; SPSS Institute Ltd., Armonk, NY, USA).
Total RNA Extraction and cDNA Library Construction
Total RNA was isolated using the TRIzol reagent (Invitrogen) according to the manufacturer’s recommendation. RNA degradation and contamination were verified on 1% agarose gels and the purity checked using the Nano Photometer spectrophotometer. The cDNA library construction and sequencing (on an Illumina Hiseq X ten platform, San Diego, CA, USA) were carried out by Novo gene Co., Ltd. (Beijing, China).
Transcriptome Sequencing
The processing of raw reads was accomplished through in-house PERL scripts. This involves pruning raw data by removing some reads containing adaptor sequences, ploy-N-containing reads, and low-quality sequences in order to have clean reads. The quality scores for assessing sequencing accuracy were computed for the clean reads, Q20, Q30, and the GC content. The high-quality clean data was used in all the downstream analyses. The clean reads were mapped to the soybean reference genome sequence using a spliced aligner Tophat 2.0.12 software (Kim et al. 2013). Only reads with a perfect match or one mismatch were further analyzed and annotated based on the reference genome. Add to that, HTSeq v 0.6.1 software (Anders et al. 2015) was used to count the read numbers that were mapped to each gene. The quantification of gene expression levels was estimated by reads per kilobase of transcript per million mapped reads for each gene and log2 transformed (Trapnell et al. 2010). After the alignment of the reads using Cufflinks v2.2.1 (Trapnell et al. 2012), the assembled transcripts were further cleaned using the fragments per kilobase per million mapped reads (FPKM) value > 1. The sequence data sets were deposited at the National Centre for Biotechnology Information (NCBI) Sequence Read Archive (SRA) repository, [http://www.ncbi.nlm.nih.gov/sra with the accession PRJNA663421].
Analysis of the RNA-Seq Data
Differential expression analyses of the LJ soybean HX3 and transgenic line (Joe) (three biological replications per treatments) were performed using the DESeq R package (1.18.0) (Anders et al. 2015). DESeq software enables statistical procedures for determining differential expression in digital gene expression data based on the negative binomial distribution (Wang et al. 2010). The resulting p-values were adjusted using Benjamini and Hochberg’s approach to control the false discovery rate (Benjamini and Hochberg 1995). Genes with an adjusted p-value < 0.05 found by DESeq were assigned as significantly differentially expressed only when the fold change was at least one (> 1 or < − 1 in log2 fold change value). We used the Kyoto Encyclopedia of Genes and Genomes (KEGG) database (Kanehisa et al. 2008) to analyze the biological pathways of the identified differentially expressed genes (DEGs). Also, the KOBAS 2.0 web server (http://kobas.cbi.pku.edu.cn/) (Xie et al. 2011) was used to test the statistical enrichment of DEGs in the KEGG pathways. Using the hypergeometric test, based on the Student’s t-test, all KEGG pathways that had a p-value < 0.05 were considered to be significantly enriched. Gene Ontology (GO) enrichment analysis of differentially expressed genes was done using the GOseq R package, in which gene length bias was corrected. GO terms with corrected p value less than 0.05 were considered significantly enriched by differential expressed genes.
Real-Time (RT)–Quantitative PCR
Twelve RNA samples for the RNA seq experiment were used in the RT-qPCR experiment. First-strand cDNA was synthesized from 1 µg of the total RNA using a Fast Quant RT Kit (Tiangen Biotech, Beijing, China). RT-qPCR was performed on Quant Studio 7 Flex (Applied Biosystems, Waltham, MA, USA) using KAPA SYBR® FAST qPCR Kits (KAPA Biosystems, Wilmington, MA, USA) Master Mix. Three biological replications and three technical replications were analyzed for each sample. Data were analyzed using the 2 − ∆∆Ct method (Schmittgen and Livak 2008) with the GmActin (Glyma.18g290800) gene used as the reference gene. The primers used for RT-qPCR in this study are listed in Table S1.
Results
Phenotypic Analysis of the Long Juvenile (LJ) Soybean and Complementary Transgenic Line Under Different Photoperiods
There were significant differences (p ≤ 0.05) between the LJ soybean variety Huaxia-3 (HX3) and Joe (J overexpression), a complementary transgenic line in days to flowering and maturity (Table 1). The LJ soybean had delayed flowering and maturity for about 10 and 6 days respectively later than the transgenic line under SD condition. Furthermore, under LD condition, both the LJ and the transgenic line did not flower up to 90 days after emergence. Thus, the LD experiment was terminated because of the onset of frost in autumn.
RNA-Seq Analysis
Twelve samples comprising of LJ and transgenic soybeans under SD and LD conditions with 3 biological replications totaling 36 samples were profiled at the V1 (unifoliate) growth stage. The total number of clean reads generated for the 12 samples in the RNA-Seq experiment was 674,800,494 of which 626,517,161 (96.2%) was uniquely mapped to the soybean reference genome (i.e., Gmax_275) (Table 2). Average GC content was 45.13% and an average of 97.74% of clean reads had quality scores at Q20 and 93.38% at Q30 (Table 2). To be sure of the similarity of the replications, we conducted a Pearson correlation test. The correlation coefficients among the samples were very high, r2 ≥ 0.90 (Fig. 1) indicative of the similarity of the replications.
Correlation coefficients among the read count of the twelve samples comprising a long juvenile soybean (HX3) and the transgenic (Joe) with 3 biological replications under SD and LD conditions. The lower diagonal represents the correlation coefficient; *** represents significance at p < 0.05. The upper diagonal plots show the distribution for each of the 12 replications.1, 2, 3 refers to the 3 replications
Numbers and Expression Patterns of Differentially Expressed Genes (DEGs) and Transcription Factors in the LJ Soybean Under Different Photoperiods
Fragments per Kilobase Million (FPKM) values were used to standardize the expression levels of the genes to enable the comparison of the LJ soybean to the transgenic line. Figure 2 shows the numbers and the expression patterns of all the DEGs under LD and SD conditions. A total of 31 genes were differentially expressed in the LJ soybean HX3 relative to the transgenic line under SD condition. Eighteen of the DEGs were upregulated and 13 were downregulated (Table S2). Under LD condition, however, 2311 DEGs were observed, comprising of 1065 upregulated and 1246 downregulated genes (Table S3). Nine genes were common under both SD and LD conditions with 22 and 2302 being exclusive to SD and LD, respectively (Fig. 2b). A total of 105 transcription factors (TF) were differentially expressed in the LJ soybean HX3 under both photoperiod and classified into 37 families (Table S4). However, only 2 TFs WRKY and SRM were differentially expressed under SD (Table S5). The commonest of the TFs under LD are bHLH, MYB, ERF, and WRKY (Table S5).
Numbers and expression pattern of differentially expressed genes (DEGs). a Volcano plots of DEGs between the long juvenile (LJ) soybean and the transgenic under short day (SD) (HX3S vs. JoeS). b LJ and transgenic under LD condition (HX3L vs. JoeL). c Venn diagrams showing the DEGs shared between LJ under SD (HX3S vs. JoeS) compared to LJ under LD condition (HX3L vs. JoeL)
Annotation of Differentially Expressed Genes (DEGs) in the LJ Soybean
To identify the genes that are differentially expressed in the LJ soybean HX3, a functional categorization was carried out by GO analysis to identify the potential use of these genes. The GO analysis grouped DEGs into three broad categories, biological function, cellular function, and molecular function. Eighteen DEGs under SD were assigned, and 37 GO terms were over represented with none of them significant at a corrected p ≤ 0.05 (Fig. S1a, Table S6). Under LD condition, however, 1767 DEGs were assigned various GO terms of which 55 were over represented at a corrected p ≤ 0.05. (Fig. S1b, Table S7). Most of the DEGs under LD condition were involved in biological processes (Fig. S1bB).
Kyoto Encyclopedia of Genes and Genomes (KEGG) Pathway Analysis of the DEGs in the LJ Soybean
A KEGG database resource for identifying high-level functions of biological systems, especially large-scale molecular datasets generated by genome sequencing was used to test the statistical enrichment of DEGs in the KEGG pathway. According to the KEGG analysis, 21 of the DEGs under SD condition were assigned to various pathways. By comparing the LJ soybean HX3 with the transgenic line under SD condition, the top most enriched pathways are shown in Fig. 3a. The circadian pathway and ether lipid metabolism were the most enriched pathway (p ≤ 0.05) (Table S8) and the circadian pathway included six genes. Under LD condition, 1020 out of the 2311 DEGs were assigned to various pathways. The top most enriched pathways are presented in Fig. 3b. Ribosome, protein processing in endoplasmic reticulum, plant hormone signal transduction, sulfur metabolism, linoleic acid metabolism, sesquiterpenoid, triterpenoid biosynthesis, and circadian rhythm were the most enriched pathway (p ≤ 0.05) (Table S9).
Statistics of pathway enrichment between the long juvenile (LJ) and the transgenic soybean. a LJ soybean under LD condition compared to transgenic soybean under SD (HX3S vs. JoeS). b LJ soybean under LD condition compared to transgenic under LD (HX3L vs. JoeL). The left Y-axis indicates the KEGG pathway. The X-axis indicates the Rich factor. The color gradient denotes the size of the q value; the color is from red to green; the red represents the smaller the q value, and the higher the significant level of enrichment of the matching KEGG pathway. The rich factor shows the ratio of the number of DEGs to the total gene number in certain pathways
Effect of Photoperiod on the Expression of Flowering Related GenesInvolved in the Circadian Clock Pathway in the LJ Soybean
Flowering related genes involved in the circadian clock pathway in the transcriptome showed that transcripts associated with soybean circadian rhythm pathway differed significantly between LJ soybean HX3 relative to the transgenic under SD condition. Table 3 and Fig. 4 shows the list and the expression pattern of circadian rhythms DEGs in the LJ soybean under both SD and LD conditions. The circadian clock pathway had one downregulated genes in the LJ soybean HX3 under SD which was the overexpressed gene Glyma.04G050200 (GmELF3a). Two FT homologs, Glyma.16G044100 (GmFT5a) and Glyma.16G150700 (GmFT2a) and two uncharacterized novel genes, Novel02840 and Novel02485 were also differentially expressed (Table 3).
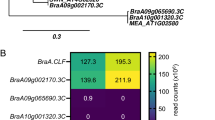
Heat map analysis of flowering related genes involved in the circadian clock pathway in the long juvenile (LJ) soybean. The values used to draw heat map are Log2 (fold change) of expression levels of the DEGs. HX3L vs. JoeL refers to LJ soybean under LD compared to the transgenic under LD, HX3S vs. JoeS refers to LJ soybean compared to transgenic soybean under SD
Under LD condition, two circadian rhythm pathway flowering-related genes were downregulated in the LJ soybean HX3. This included Glyma.16G027200 (SPA, SUPRESSOR OF PHYA), and a TF Glyma.19G224700 (PIF3, phytochrome-interacting factor 3) (Table 3). However, seven other circadian flowering related clock genes were upregulated (Table 3). Analysis of flowering related genes involved in the circadian clock pathway in the LJ soybean HX3 under both photoperiods shows that the overexpressed gene in the transgenic (Joe) Glyma.04G050200 (GmELF3a) was downregulated under both photoperiods (Fig. 4) but was not differentially expressed under LD. Figure S2 shows the soybean circadian clock genes in a biological network highlighting the DEGs between the LJ soybean HX3 and the transgenic under SD condition.
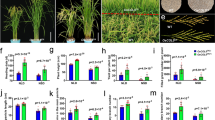
Confirmation of Gene Expression Profiles Using RT-qPCR
Eleven flowering related DEGs were selected to confirm the RNA-Seq results. Four out of the five genes under SD condition showed the same trends with the RNA-Seq data. Under LD condition, five out of the six genes showed a similar trend with the RNA-Seq data. Glyma.08G363100 (GmFT4) was downregulated under SD and upregulated under LD condition showing the same trend as the RNA-Seq result. These lend credence to the validity of the RNA-Seq experiment. Nine genes having similar trends as RNA-Seq are presented in Fig. 5.
Discussion
Our study focused on the profile of the transcriptome of a natural LJ soybean with the recessive j gene relative to the transgenic soybean overexpressing the J/GmELF3a gene. This gave us an insight into the clock related flowering genes differentially expressed in the LJ soybean HX3 compared to the transgenic soybean under different photoperiods. RNA-Seq is a high-throughput sequencing technology that enables the study of comprehensive transcriptional profile and has been widely used to study molecular responses to abiotic and biotic stresses and for comparing transcriptomes under different treatments (Zeng et al. 2019; Luo et al. 2019; Gillman et al. 2019; Wang et al. 2009).
The LJ soybean had delayed flowering for about 10 days and matured later than the transgenic under SD (Table 1). This is attributed to the recessive allele (j) in the LJ soybean since the functional J gene results in early flowering under SD condition and is in consonance with a previous study by Yue et al. (2017) where transgenic complementation of the J gene in the LJ soybean HX3 resulted in early flowering. This was also confirmed by the down-regulation of the overexpress gene GmELF3a in the HX3 (Table 3). Under LD condition, interestingly, both the transgenic and the LJ soybeans did not flower more than 90 days after emergence, even though the transgenic line had the functional J gene. The ortholog of J gene in Arabidopsis ELF3 has been reported to trigger early flowering under LD condition (Zagotta et al. 1996). The non-flowering of both the transgenic line and the LJ soybean HX3 could be as a result of their genetic background since LJ varieties with either functional or the recessive j gene hardly flower under LD conditions.
We observed more DEGs under LD than SD. The higher number of DEGs under LD compared to SD may be as a result of the differences between the growth and development under the different photoperiods. Soybean is a SD crop; therefore, the LD may have imposed stress on it. It should be noticed that many phenotypes such as flowering day exhibit a small variance under SD than under LD. A study by Jiang et al. (2017) involving wheat also reported few numbers of DEGs under its normal growth conditions compared to salt stress condition.
The KEGG analysis of the transcriptome data revealed that the circadian clock pathway was the most significantly enriched pathway in the LJ soybean under SD condition (Fig. 3a). This is indicative that the circadian pathway played a role in the delayed flowering of the LJ soybean under SD. The circadian clock in plants regulates seasonal changes in day length and temperature to control flowering time. Circadian clock genes have been reported to be involved in regulating the photoperiod of flowering in plants (Imaizumi and Kay 2006; Hudson 2010). This ensures that the transition from vegetative to reproductive stages happens under optimum physiological and environmental conditions, thereby enhancing reproductive success (Piggins et al. 2011).
In Arabidopsis, FT is an important flowering promoter, and also serves as integrators of internal and external flowering indicators and is widely conserved in many plant species (Pin and Nilsson 2012). Two of the downregulated FT homologs in the LJ soybean under SD condition GmFT2a and GmFT5a play significant roles in the induction of flowering under SD. GmFT2a and GmFT5a were reported to cause delayed flowering when downregulated by RNA interference (RNAi) (Guo et al. 2015). Also, recent studies of CRISPR/Cas9-generated loss-of-function mutants of these genes revealed these genes were involved in flowering induction, with a delayed flowering observed under SD for GmFT2a mutants (Cai et al. 2018, 2020). These are in consonance with our study where the down regulation of these two genes seems to trigger delayed flowering in the LJ soybean.
Contrarily to most florigen proteins, GmFT4 and GmFT1a inhibit flowering (Liu et al. 2018). GmFT4 was slightly upregulated in the LJ soybean under SD condition. GmFT4 was reported as a flowering repressor by previous studies and a candidate for the E10 gene in soybean (Samanfar et al. 2017). A study by Zhai et al. (2014) reported that the LJ soybean HX3 showed a relatively high GmFT4 expression under SD condition. This aligns with our study in which the GmFT4 was upregulated under SD in the LJ soybean. Under LD, only two FT genes were differentially expressed in the LJ soybean HX3, and this includes GmFT4 which was slightly downregulated and GmFT1b which was slightly upregulated (Table 3). GmFT1a and GmFT4 were reported to be strongly induced under LD conditions (Liu et al. 2018; Zhang et al. 2016).
The other circadian clock genes such as PRR7, PRR5, CRY1, CRY1a, and Novel 03,244 (Gigantea like protein) were slightly upregulated in the LJ soybean HX3 under LD (Table 3). PRRs are important parts of transcriptional circadian networks in plants (Farre and Liu 2013). Li et al. (2020a, b) reported that the PRR3 and its homologue PRR7 may be involved in flowering regulation through the circadian pathway. CRY is the photolyase-related blue-light receptors that regulate plant’s responses to light and the circadian clock major evolutionary lineages (Schmitz et al. 2005). During the shift from the vegetative to reproductive phases, the biological circadian clock genes PRR5, PRR7, and SPA1, act on the downstream gene CRY2 to affect the flowering process in Arabidopsis (Nakamichi et al. 2010; Weidler et al. 2012). CRY2 regulates photoperiodic flowering in Arabidopsis; however, in soybean, CRY1a is reported to be one of the major regulators of photoperiodic flowering in soybean (Jung et al. 2012). GIGANTEA protein in Arabidopsis was reported to play multiple roles in the circadian clock and flowering (Watanabe et al. 2011) but its role in soybean has not been fully understood.
Conclusion
We compared changes in the expression of circadian clock flowering related genes in a natural LJ soybean variety with a loss of function allele for the J gene and a transgenic line with the overexpressed gene under different photoperiods. Analysis of the RNA-Seq data shows that the delayed flowering under SD may be as a result of the down-regulation of GmELF3a together with some key FT genes in the LJ soybean under SD condition. Our study adds to the evidence emerging that FT genes may also be implicated in the LJ trait in soybean under SD. Future studies should explore the non-flowering of both the LJ soybean HX3 and transgenic line under LD condition to identify the genes involved. This will contribute to the knowledge gap in flowering time regulation in soybean under tropical conditions.
Abbreviations
- SD:
-
Short day
- LD:
-
Long day
- LJ:
-
Long juvenile
- HX3:
-
Huaxia-3
- FT:
-
FLOWERING LOCUS T
- PRR:
-
Pseudo-response regulator
- TF:
-
Transcription factor
References
Andres F, Coupland G (2012) The genetic basis of flowering responses to seasonal cues. Nat Rev Genet 13(9):627–639. https://doi.org/10.1038/nrg3291
Anders S, Pyl PT, Huber W (2015) HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31(2):166–169
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate : a practical and powerful approach to multiple testing. J R Stat Soc B 57(1):289–300. https://doi.org/10.1111/j.2517-6161.1995.tb02031.x
Bu T, Lu S, Wang K, Dong L, Li S, Xie Q, Xu X, Cheng Q, Chen L, Fang C, Li H, Liu B, Weller JL, Kong F (2021) A critical role of the soybean evening complex in the control of photoperiod sensitivity and adaptation. Proc Natl Acad Sci U S A 118(8):e2010241118
Cai Y, Chen L, Liu X, Guo C, Sun S, Wu C, Jiang B, Han T, Hou W (2018) CRISPR/Cas9-mediated targeted mutagenesis of GmFT2a delays flowering time in soya bean. Plant Biotechnol J 16(1):176–185. https://doi.org/10.1111/pbi.12758
Cai Y, Wang L, Chen L, Wu T, Liu L, Sun S, Wu C, Yao W, Jiang B, Yuan SJPbj (2020) Mutagenesis of GmFT2a and GmFT5a mediated by CRISPR/Cas9 contributes for expanding the regional adaptability of soybean. Plant Biotechnol J 18(1):298–309. https://doi.org/10.1111/pbi.13199
Carter TE, Nelson RL, Sneller CH, Cui Z (2004) Genetic diversity in soybean. Soybeans: Improvement, Production, and Uses. 2004 (soybeansimprove):303–416
Chen L, Cai Y, Qu M, Wang L, Sun H, Jiang B, Wu T, Liu L, Sun S, Wu C, Yao W, Yuan S, Han T, Hou W (2020) Soybean adaption to high-latitude regions is associated with natural variations of GmFT2b, an ortholog of FLOWERING LOCUS T. Plant Cell Environ 43:934–944
Destro D, Carpentieri-Pípolo V, Kiihl RdS, de ALMEIDA LA (2001) Photoperiodism and genetic control of the long juvenile period in soybean: a review. Crop Breed Appl Biotechnol 1:72–92
Fan C, Hu R, Zhang X, Wang X, Zhang W, Zhang Q, Ma J, Fu YF (2014) Conserved CO-FT regulons contribute to the photoperiod flowering control in soybean. BMC Plant Biol 14:9. https://doi.org/10.1186/1471-2229-14-9
Fang X, Han Y, Liu M, Jiang J, Li X, Lian Q, Xie X, Huang Y, Ma Q, Nian H, Qi J (2021) Modulation of evening complex activity enables north-to-south adaptation of soybean. Sci China Life Sci 64(2):179–195. https://doi.org/10.1007/s11427-020-1832-2
Farre EM, Liu T (2013) The PRR family of transcriptional regulators reflects the complexity and evolution of plant circadian clocks. Curr Opin Plant Biol 16(5):621–629. https://doi.org/10.1016/j.pbi.2013.06.015
Fehr W, Caviness C, Burmood D, Pennington J (1971) Stage of development descriptions for soybeans, Glycine max (L.) Merrill 1. Crop sci 11(6):929–931
Gillman JD, Biever JJ, Ye S, Spollen WG, Givan SA, Lyu Z, Joshi T, Smith JR, Fritschi FB (2019) A seed germination transcriptomic study contrasting two soybean genotypes that differ in terms of their tolerance to the deleterious impacts of elevated temperatures during seed fill. BMC Res Notes 12(1):522
Guo G, Xu K, Zhang X, Zhu J, Lu M, Chen F, Liu L, Xi ZY, Bachmair A, Chen Q, Fu YF (2015) Extensive analysis of GmFTL and GmCOL expression in northern soybean cultivars in field conditions. PLoS ONE 10(9):e0136601. https://doi.org/10.1371/journal.pone.0136601
Hartwig EE, Kiihl RA (1979) Identification and utilization of a delayed flowering character in soybeans for short-day conditions. Field Crops Res 2:145–151
Hudson KA (2010) The circadian clock-controlled transcriptome of developing soybean seeds. The Plant Genome 3(1):3–13
Hymowitz T (1970) On the domestication of the soybean. Econ Bot 24(4):408–421
Imaizumi T, Kay SA (2006) Photoperiodic control of flowering: not only by coincidence. Trends Plant Sci 11(11):550–558
Itoh H, Izawa T (2013) The coincidence of critical day length recognition for florigen gene expression and floral transition under long-day conditions in rice. Mol Plant 6(3):635–649. https://doi.org/10.1093/mp/sst022
Jiang Q, Niu F, Sun X, Hu Z, Li X, Ma Y, Zhang H (2017) RNA-Seq analysis of unintended effects in transgenic wheat overexpressing the transcription factor GmDREB1. Crop J 5(3):207–218. https://doi.org/10.1016/j.cj.2016.12.001
Jung CH, Wong CE, Singh MB, Bhalla PL (2012) Comparative genomic analysis of soybean flowering genes. PLoS ONE 7(6):e38250. https://doi.org/10.1371/journal.pone.0038250
Kanehisa M, Araki M, Goto S, Hattori M, Hirakawa M, Itoh M, Katayama T, Kawashima S, Okuda S, Tokimatsu T, Yamanishi Y (2008) KEGG for linking genomes to life and the environment. Nucleic Acids Res 36(Database issue):D480–484. https://doi.org/10.1093/nar/gkm882
Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL (2013) TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 14(4):R36. https://doi.org/10.1186/gb-2013-14-4-r36
Kong F, Liu B, Xia Z, Sato S, Kim BM, Watanabe S, Yamada T, Tabata S, Kanazawa A, Harada K, Abe J (2010) Two coordinately regulated homologs of FLOWERING LOCUS T are involved in the control of photoperiodic flowering in soybean. Plant Physiol 154(3):1220–1231
Li C, Li Y, Li Y, Lu H, Hong H, Tian Y, Li H, Zhao T, Zhou X, Liu J, Zhou X, Jackson SA, Liu B, Qiu L (2020a) A domestication-associated gene GmPRR3b regulates the circadian clock and flowering time in soybean. Mol Plant 13:745–759
Li MW, Wang Z, Jiang B, Kaga A, Wong FL, Zhang G, Han T, Chung G, Nguyen H, Lam HM (2020b) Impacts of genomic research on soybean improvement in East Asia. Theor Appl Genet 133(5):1655–1678. https://doi.org/10.1007/s00122-019-03462-6
Lin X, Liu B, Weller JL, Abe J, Kong F (2021) Molecular mechanisms for the photoperiodic regulation of flowering in soybean. J Integr Plant Biol 63:981–994. https://doi.org/10.1111/jipb.13021344
Liu B, Kanazawa A, Matsumura H, Takahashi R, Harada K, Abe J (2008) Genetic redundancy in soybean photoresponses associated with duplication of the phytochrome A gene. Genet 180(2):995–1007
Liu W, Jiang B, Ma L, Zhang S, Zhai H, Xu X, Hou W, Xia Z, Wu C, Sun S, Wu T, Chen L, Han T (2018) Functional diversification of Flowering Locus T homologs in soybean: GmFT1a and GmFT2a/5a have opposite roles in controlling flowering and maturation. New Phytol 217(3):1335–1345. https://doi.org/10.1111/nph.14884
Lu S, Dong L, Fang C, Liu S, Kong L, Cheng Q, Chen L, Su T, Nan H, Zhang D, Zhang L, Wang Z, Yang Y, Yu D, Liu X, Yang Q, Lin X, Tang Y, Zhao X, Yang X, Tian C, Xie Q, Li X, Yuan X, Tian Z, Liu B, Weller JL, Kong F (2020) Stepwise selection on homeologous PRR genes controlling flowering and maturity during soybean domestication. Nat Genet 52(4):428–436
Lu S, Zhao X, Hu Y, Liu S, Nan H, Li X, Fang C, Cao D, Shi X, Kong L, Su T, Zhang F, Li S, Wang Z, Yuan X, Cober ER, Weller JL, Liu B, Hou X, Tian Z, Kong F (2017) Natural variation at the soybean J locus improves adaptation to the tropics and enhances yield. Nat Genet 49(5):773–779. https://doi.org/10.1038/ng.3819
Luo Q, Teng W, Fang S, Li H, Li B, Chu J, Li Z, Zheng Q (2019) Transcriptome analysis of salt-stress response in three seedling tissues of common wheat. Crop J 7(3):378–392. https://doi.org/10.1016/j.cj.2018.11.009
Nakamichi N, Kiba T, Henriques R, Mizuno T, Chua NH, Sakakibara H (2010) Pseudo-response regulators 9, 7, and 5 are transcriptional repressors in the Arabidopsis circadian clock. Plant Cell 22(3):594–605. https://doi.org/10.1105/tpc.109.072892
Neumaier N, James A (1993) Exploiting the long-juvenile trait to improve adaptation of soybeans to the tropics. Food Legume Newslet 8
Piggins HD, Guilding C, Adams S, Carré IA (2011) Downstream of the plant circadian clock: output pathways for the control of physiology and development. J Essays Biochem 49:53–69
Pin PA, Nilsson O (2012) The multifaceted roles of FLOWERING LOCUS T in plant development. Plant Cell Environ 35(10):1742–1755. https://doi.org/10.1111/j.1365-3040.2012.02558.x
Samanfar B, Molnar SJ, Charette M, Schoenrock A, Dehne F, Golshani A, Belzile F, Cober ER (2017) Mapping and identification of a potential candidate gene for a novel maturity locus, E10, in soybean. Theor Appl Genet 130(2):377–390. https://doi.org/10.1007/s00122-016-2819-7
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc 3(6):1101–1108. https://doi.org/10.1038/nprot.2008.73
Schmitz RJ, Hong L, Michaels S, Amasino RM (2005) FRIGIDA-ESSENTIAL 1 interacts genetically with FRIGIDA and FRIGIDA-LIKE 1 to promote the winter-annual habit of Arabidopsis thaliana. Development 132(24):5471–5478. https://doi.org/10.1242/dev.02170
Sinclair TR, Hinson KJCS (1992) Soybean flowering in response to the long-juvenile trait. Crop Sci 32(5):1242–1248
Spehar CR (1995) Impact of strategic genes in soybean on agricultural development in the Brazilian tropical savannahs. Field Crops Res 41(3):141–146
Sun F, Xu M, Park C, Dwiyanti MS, Nagano AJ, Zhu J, Watanabe S, Kong F, Liu BH, Yamada T, Abe J (2019) Characterization and quantitative trait locus mapping of late-flowering from a Thai soybean cultivar introduced into a photoperiod-insensitive genetic background. PLoS ONE 14(12):e0226116. https://doi.org/10.1371/journal.pone.0226116
Sun H, Jia Z, Cao D, Jiang B, Wu C, Hou W, Liu Y, Fei Z, Zhao D, Han T (2011) GmFT2a, a soybean homolog of FLOWERING LOCUS T, is involved in flowering transition and maintenance. PLoS ONE 6(12):e29238. https://doi.org/10.1371/journal.pone.0029238
Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR, Pimentel H, Salzberg SL, Rinn JL, Pachter L (2012) Differential gene and transcript expression analysis of RNA-Seq experiments with TopHat and Cufflinks. Nat Protoc 7(3):562–578. https://doi.org/10.1038/nprot.2012.016
Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ, Salzberg SL, Wold BJ, Pachter L (2010) Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28(5):511–515. https://doi.org/10.1038/nbt.1621
Wang L, Feng Z, Wang X, Wang X, Zhang X (2010) DEGseq: An R package for identifying differentially expressed genes from RNA-Seq data. Bioinformatics 26(1):136–138. https://doi.org/10.1093/bioinformatics/btp612
Wang L, Sun S, Wu T, Liu L, Sun X, Cai Y, Li J, Jia H, Yuan S, Chen L, Jiang B, Wu C (2020) Hou W and Han T (2020) Natural variation and CRISPR/Cas9-mediated mutation in GmPRR37 affect photoperiodic flowering and contribute to regional adaptation of soybean. Plant Biotechnol J 18:1869–1881. https://doi.org/10.1111/pbi.13346
Wang Z, Gerstein M, Snyder M (2009) RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet 10(1):57–63. https://doi.org/10.1038/nrg2484
Wang Z, Zhou Z, Liu Y, Liu T, Li Q, Ji Y, Li C, Fang C, Wang M, Wu M, Shen Y, Tang T, Ma J, Tian Z (2015) Functional evolution of phosphatidylethanolamine binding proteins in soybean and Arabidopsis. Plant Cell 27(2):323–336
Watanabe S, Hideshima R, Xia Z, Tsubokura Y, Sato S, Nakamoto Y, Yamanaka N, Takahashi R, Ishimoto M, Anai T, Tabata S (2009) Map-based cloning of the gene associated with the soybean maturity locus E3. Genet 182(4):1251–1262
Watanabe S, Xia ZJ, Hideshima R, Tsubokura Y, Sato S, Yamanaka N, Takahashi R, Anai T, Tabata S, Kitamura K, Harada K (2011) A map-based cloning strategy employing a residual heterozygous line reveals that the GIGANTEA gene is involved in soybean maturity and flowering. Genet 188(2):395-U260. https://doi.org/10.1534/genetics.110.125062
Weidler G, zur Oven-Krockhaus S, Heunemann M, Orth C, Schleifenbaum F, Harter K, Hoecker U, Batschauer A (2012) Degradation of Arabidopsis CRY2 is regulated by SPA proteins and phytochrome A. Plant Cell 24(6):2610–2623. https://doi.org/10.1105/tpc.112.098210
Wu F, Sedivy EJ, Price WB, Haider W, Hanzawa Y (2017) Evolutionary trajectories of duplicated FT homologues and their roles in soybean domestication. Plant J 90(5):941–953
Xia Z, Watanabe S, Yamada T, Tsubokura Y, Nakashima H, Zhai H, Anai T, Sato S, Yamazaki T, Lü S, Wu H (2012) Positional cloning and characterization reveal the molecular basis for soybean maturity locus E1 that regulates photoperiodic flowering. P Natl Acad Sci USA 109(32):E2155–E2164
Xie C, Mao X, Huang J, Ding Y, Wu J, Dong S, Kong L, Gao G, Li C, Wei L (2011) KOBAS 2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res 39 (Web Server issue): W316–322. https://doi.org/10.1093/nar/gkr483
Yue Y, Liu N, Jiang B, Li M, Wang H, Jiang Z, Pan H, Xia Q, Ma Q, Han T, Nian H (2017) A single nucleotide deletion in J encoding GmMELF3 confers long juvenility and is associated with adaption of tropic soybean. Mol Plant 10(4):656–658. https://doi.org/10.1016/j.molp.2016.12.004
Zagotta MT, Hicks KA, Jacobs CI, Young JC, Hangarter RP, Meeks-Wagner DR (1996) The Arabidopsis ELF3 gene regulates vegetative photomorphogenesis and the photoperiodic induction of flowering. Plant J 10(4):691–702. https://doi.org/10.1046/j.1365-313x.1996.10040691.x
Zeng A, Chen P, Korth KL, Ping J, Thomas J, Wu C, Srivastava S, Pereira A, Hancock F, Brye K, Ma J (2019) RNA sequencing analysis of salt tolerance in soybean (Glycine max). Genomics 111(4):629–635. https://doi.org/10.1016/j.ygeno.2018.03.020
Zhai H, Lü S, Liang S, Wu H, Zhang X, Liu B, Kong F, Yuan X, Li J, Xia Z (2014) GmFT4, a homolog of FLOWERING LOCUS T, is positively regulated by E1 and functions as a flowering repressor in soybean. PLoS ONE 9(2):e89030. https://doi.org/10.1371/journal.pone.0089030
Zhang S, Singh MB, Bhalla PL (2021) Molecular characterization of a soybean FT homologue, GmFT7. Sci Rep 11(1):1–11. https://doi.org/10.1038/s41598-021-83305-x
Zhang X, Zhai H, Wang Y, Tian X, Zhang Y, Wu H, Lu S, Yang G, Li Y, Wang L, Hu B, Bu Q, Xia Z (2016) Functional conservation and diversification of the soybean maturity gene E1 and its homologs in legumes. Sci Rep 6:29548. https://doi.org/10.1038/srep29548
Funding
This research was funded by the National Key R&D Program of China (2017YFD0101400), Major Science and Technology Projects of China (2016ZX08004-003), China Agriculture Research System (CARS-04), and the CAAS (Chinese Academy of Agriculture Sciences) Agricultural Science and Technology Innovation Project. The Chinese Scholarship Council provided the financial support to the first author to study for his doctoral degree in CAAS.
Author information
Authors and Affiliations
Contributions
TH, YY, and BJ conceived and designed the project. ES performed the experimental treatment, sampling, and data analysis. ES, BJ, YY, and TH wrote the manuscript. LL assisted with experimental procedures. SEI revised the manuscript. YY, TW, SY, and SS provided materials and resources for the experiment. All the authors agreed on the final manuscript submitted.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key Messages
• Transcriptome analysis of a long juvenile soybean genotype under short day revealed the circadian rhythm pathways was the most significantly enriched.
• FLOWERING LOCUS T homologs may be implicated in delayed flowering of a long juvenile soybean genotype under short day condition.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sapey, E., Jiang, B., Liu, L. et al. Transcriptome Profile of a Long-Juvenile Soybean Genotype Huaxia-3 Under Short and Long Photoperiod. Plant Mol Biol Rep 40, 433–445 (2022). https://doi.org/10.1007/s11105-021-01332-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-021-01332-4