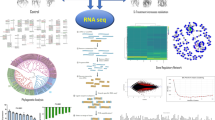
Abstract
Lead (Pb) is a heavy metal with high toxicity to plants. Root is the major organ to respond to Pb stress. However, little is known about how plant roots perceive Pb stress signaling. Here, we describe the transcriptome of Arabidopsis root tips under Pb stress using the RNA-seq assay. A total of 703 differentially expressed genes (DEGs) were identified and expressed at every time points. Some early-responsive DEGs (1 h) were predicted to be negatively involved in cell elongation and cell expansion, while some late-responsive DEGs (24 h) positively participated in defense of oxidative stress. Hydrogen peroxide (H2O2) and superoxide (O2−) were increased significantly in root tips under Pb stress. Cell wall extension related gene XYLOGLUCAN ENDOTRANSGLUCOSYLASE/HYDROLASE 18 (XTH18) was induced in root tips, and xth18 showed reduced root growth inhibition by Pb stress. Our results revealed the potential mechanism of root growth inhibition by Pb stress and shed light for the further study.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lead (Pb), a heavy metal with high toxicity to plants, is regarded as the first one smelted by humans (Kalisińska 2019). In daily life, Pb is everywhere because of its corrosion resistance and plasticity, such as fertilizers, pesticides, and pigments (Hadi and Aziz 2015). The contamination of soil by Pb is mainly caused by mining, chimneys of factories using Pb, the storage battery, and smelting. In accordance with ATSDR (Agency for Toxic Substances and Disease Registry) Priority Substance List published in 2019, Pb was labeled as the second most dangerous environment poison due to its toxicity (https://www.atsdr.cdc.gov/SPL/index.html#2019spl). In terms of plants, Pb is not essential for plants’ growth and development, but Pb toxicity sharply occurs even in very low concentration. Pb toxicity brings morpho-physiological and biochemical changes and disturbs normal functioning of the plants from cell to organ level (Ashraf and Tang 2017).
Roots, the first line of defense against Pb stress, secrete exudates into the rhizosphere to chelate Pb and prevent their uptake into root cells. In addition, roots also uptake large amounts of Pb through the NRAMP (natural resistance-associated macrophage protein), ZIP (Zrt/IRT-like protein), and cation transporters, and immensely restrict its translocation to aerial parts. Simultaneously, Pb particles can be fixed on cell wall by extracellular carbohydrates such as callose and mucilage. It is found that callose synthesis could be induced by Pb stress, which formed an efficient barrier for Pb penetration (Samardakiewicz et al. 2012). In cytoplasm, the compartmentalization and chelation of Pb proceeds with a faster kinetics to preclude their participation in toxic reactions. Nevertheless, the burst of reactive oxygen species (ROS) attributed to Pb toxicity results in cell damage and reduced root length. To confront oxidative stress, antioxidant system was activated, including non-protein thiols (cysteine and glutathione), ascorbic acid, proline, as well as antioxidant enzymes, such as superoxide dismutase (SOD), guaiacol peroxidase (GPX), ascorbate peroxidase (APX), glutathione reductase (GR), and catalase (CAT).
In this study, root tips of Arabidopsis treated without and with Pb stress for different time points were used for RNA-seq assays. Root growth under Pb stress was apparently reduced, and reactive oxygen species (ROS) was significantly increased by Pb stress in root tips. RNA-seq data showed that a total of 703 differentially expressed genes (DEGs) were expressed at all the time points and mainly implicated in transporter activity, ion binding, catalytic activity, plant growth and development, cell wall modification, and abiotic stimulus. Further results demonstrated that the cell wall extension related gene XTH18 was up-regulated by Pb and xth18 showed reduced sensitivity to Pb stress. These results will enrich our understanding of how plant roots perceive Pb stress signaling and provide a solid foundation for breeding of Pb-tolerant crop varieties.
Materials and Methods
Plant Material and Root Imaging
The Arabidopsis wild genotype (Columbia ecotype, Col-0) was served as experimental materials in the current study. Seeds were surface-sterilized in 10% bleach for 10 min and washed 6 times with double-distilled water to remove residual bleach. The pre-treated seeds were plated on half-strength Murashige and Skoog (MS) medium. Plants were stratified at 4 °C for 2 days in darkness and then transferred to a phytotrone set at 22 °C with a 16-h-light/8-h-dark photoperiod (light intensity 100 μmol m−2 s−1), which were vertically placed in a plant incubator (22 °C, day/night). The ImageJ was applied to measure root length of 6-day-old seedlings from images captured with a Canon EOS 760D camera. The assay had three biological replicates with ten seedlings per replicate.
Meristem Size Measurements and GUS Staining
The length of root meristem was measured on a ZEISS Axio Imager Z2 microscope using six-day-old seedlings treated with or without 1 mM Pb. Histochemical GUS staining was performed as described previously (Zhang et al. 2018). These experiments were replicated three times for each of ten seedlings.
RNA-Seq Assay
Five-day-old wild-type Seedlings cultured on 1/2 MS medium were transferred to medium supplemented with l mM Pb(NO3)2 for 1 h, 6 h, 12 h, and 24 h. Root tips (0.5 cm) were collected and quickly frozen with liquid nitrogen, and seedlings on 1/2 MS medium were used as control. Total RNA was isolated using TRIzol reagent (Invitrogen, 15596026). Sequencing libraries were constructed with a total of 3 μg RNA and implemented on Illumina HiSeq 2500 platform (Berry Genomics). All samples were run in three biological replicates. The Kallisto (v 0.44.0) was applied to quantify the transcripts expression levels for each read. The HISAT2 (v 2.0.5) was used to map reads to the Arabidopsis genome. DEGs were analyzed with the DESeq2 package, and genes (Padj < 0.01 and |Log2FoldChange| > 0.1) of which were regarded as the significant DEGs and picked for gene ontology (GO) enrichment analysis.
Reverse Transcription Quantitative PCR
Reverse transcription quantitative PCR (RT-qPCR) assays were performed to validate the dependability of RNA-seq. All RT-qPCR analyses were conducted with three biological replicates and 3 technical replicates within each biological replicate on a Bio-RAD CFX96™ optics module. The relative expression level was computed by the method of comparative △Ct, and Actin7 (At5g09810) was picked to normalize all results. Specific primers are listed in Supplementary Table 1.
Histochemical Detection of O2 − and H2O2 in Root
The 4-day-old seedlings were transferred to the medium with or without 1 mM Pb(NO3)2 for 24 h. The seedlings were immersed in 3,3′-diaminobenzidine (DAB) or nitroblue tetraxolium (NBT) staining solution for detection of H2O2 or O2−in light of the methods described in the former study, respectively (Dunand et al. 2007). Three biological replicates with ten seedlings per replicate.
Results
Root Growth Inhibition by Pb Stress
In this study, seedlings of Col-0 were constantly fostered with or without 1 mM Pb(NO3)2 for 6 days. The results showed that root elongation was apparently inhibited, and root length was decreased by 64.37% under Pb stress compared with that of mock treatment (1/2 MS), while shoot growth was relatively less affected (Fig. 1 a and b). Postembryonic root growth of higher plants is maintained by the root meristem. The reduction of root meristem size by Pb could result from its negative effect on cell division (Fig. 1c). The CYCB1;1 encodes a cyclin-dependent protein kinase and plays an important role in the G2/M phase of the cell cycle. Therefore, the expression of CYCB1;1pro: GUS in roots can be used to mark mitotic activity in meristems (Colon-Carmona et al. 1999). The CYCB1;1pro:GUS was expressed in the actively dividing cells of the root meristem, and the activity was markedly reduced by Pb stress (Fig. 1d), suggesting that Pb stress represses the cell division activity of the transit amplifying cells in the root meristem.
Lead (Pb) inhibits root growth. a Root length was greatly reduced by Pb stress. Seedlings were grown with or without 1 mM Pb for 6 days. Bar = 0.25 cm. b Comparison of root length under the Mock (left) and Pb stress (right) condition. Data are shown as the average (± SD) (n = 10). c The root meristem size decreased sharply under Pb stress. Bar = 50 μm. d Six-day-old transgenic Arabidopsis seedlings expressing proCYCB1;1:GUS were germinated on Murashige and Skoog (MS) medium and transferred to medium without Pb (MS, Mock) or medium containing 1 mM Pb for 24 h, and proCYCB1;1:GUS expression was monitored by histochemical GUS staining. Bar = 50 μm
RNA Sequencing and Identification of DEGs Under Pb Stress
To further explore the underlying mechanism of root growth inhibition in response to Pb stress, RNA-seq assays were carried out with Pb treatments for different time points (0 h/1 h/6 h/12 h/24 h). To test the repeatability and reliability of the sequencing data, the principal component analysis (PCA) was implemented and the results demonstrated that all samples could be divided into five groups (Supplementary Fig. 1), representing that the datasets were available for further analysis. Statistically, a total of 443,708,633 raw reads were obtained from RNA sequencing. The amount of clean reads ranged from 22,626,671 to 33,541,052 among 15 root samples, of which between 78.93% and 94.569% mapped to the Arabidopsis genome (Supplemental Table 2). The reads uniquely mapped to the genome varies between 61.37 and 85.09%. In addition, the minimum Q20 and Q30 of those clean reads were 99.86% and 82.77%, respectively.
The dataset has been uploaded to NCBI database (https://dataview.ncbi.nlm.nih.gov/object/PRJNA593333?reviewer=422k02ihk6skl0tphuhj2oml2r).
Identification and quantification of genes (especially DEGs) expressed under the specific conditions are two vital characters of transcriptome sequencing analysis. In this study, DEGs in response to Pb stress were screened with the threshold of adjusted P value < 0.01. A total of 4284 DEGs were identified at 1 h after Pb stress. With Pb treatments from 0 to 24 h, the number of DEGs reached a peak at 6 h. The up- and down-regulated DGEs reached 2358 and 2739, respectively. However, the amount of DEGs sharply dropped to the minimum value at 12 h. With Pb treatment for 24 h, the amount of up-regulated DEGs was slightly more than that of down-regulated DEGs, and the total number of DEGs was 3498 (Fig. 2a and Supplementary Table 3). Subsequently, a total of 703 expressed DEGs were identified at all the time intervals (Fig. 2b). Hierarchical clustering analysis of 703 DEGs indicated that they could be divided into five groups, and the expression pattern of which was largely different. The result of cluster analyses among four groups with Pb treatments showed that the expression pattern of DEGs at 12 h was similar with that of 24 h, and the difference of which between Pb-1 h and other groups was quite distinct (Fig. 2c). In addition, individual comparisons with a loop design at temporal scale were performed. A total of 3448, 1127, 996, and 3893 DEGs were identified at 1–6 h, 6–12 h, 12–24 h, and 1–24 h, respectively (Supplementary Table 6). For validation of RNA-seq data, a total of 14 DEGs were picked for RT-qPCR assay, and the results of correlation analysis referred that correlation coefficient between RNA-seq and RT-qPCR is 0.751, indicating that the expression trend is consistent (Supplementary Fig. 2).
GO Enrichment Analysis of Commonly Expressed DEGs
RNA-seq provides a quick and feasible method for gene function study. Furthermore, the functional relevance of candidate genes could be analyzed with GO tools to separate gene products into three categories, including molecular function (MF), biological process (BP), and cellular component (CC). Here, the commonly expressed DEGs were also divided into three groups (Supplementary Table 4). In terms of molecular function, the processes represented by the GO terms “oxidoreductase activity”, “transmembrane transporter activity,” “metal ion binding,” and “lipid binding” were significantly enriched (Supplementary Fig. 3), indicating that a variety of detoxification metabolisms were activated after Pb stress for few hours. In the biological process ontology (Supplementary Fig. 4), the major terms were “developmental growth involved in morphogenesis,” “root hair cell development,” “cell wall organization or biogenesis,” and “response to abiotic stimulus,” suggesting that plant growth (especially root development) was affected by Pb stress and genes related to abiotic stimulus were activated. Regarding cellular component (Supplementary Fig. 5), the major terms were “plasma membrane,” “vacuolar membrane,” “cell wall,” and “endomembrane system,” demonstrating that cell membrane system performed vital roles in Pb transport, chelation, and sequestration. Although many GO terms related to Pb stress were enriched, it still needs more information to clarify their relationship in further study.
Identification of Early- and Late-Responsive Genes Under Pb Stress
To investigate the early- and late-responsive genes related to Pb defense, GO enrichment analysis of DEGs at 1 h and 24 h was performed, and the top 20 GO items (biological process) are shown in Supplementary Figs. 6 and 7, respectively. Interestingly, the terms named “regulation of cell size” and “cell growth” were specifically enriched at 1 h. This result suggests that the DEGs at the early stage (1 h) of Pb stress were negatively involved in cell elongation/expansion and root growth, which demonstrated that cell elongation inhibition in root is a fast response to Pb stress. Furthermore, “root development,” “ion transport,” and “response to abiotic stress” related genes were significantly enriched at 1–6 h (Supplemental Fig. 8). Genes (Expansin, etc.) related to cell elongation were also isolated as DEGs and involved in root development, indicating that cell elongation consistently participated in defense responses of Pb stress at the early stage. At 6–12 h, “cell cycle,” “cell division,” and “response to metal ion” related genes were enriched, suggesting that Pb stress still affected cell division (Supplemental Fig. 9). Moreover, “oxidation-reduction process,” “response to oxidative stress,” and “response to metal ion” related genes were enriched at 12–24 h (Supplemental Fig. 10), demonstrating that oxidation-reduction process was progressively activated by Pb stress. “Response to oxidative stress” and “plant-type cell wall organization” related genes were uniquely enriched at 24 h (Fig. 3a). Similarly, “oxidation-reduction process” and “response to oxidative stress” related genes were also enriched at 1–24 h (Supplemental Fig. 11), implying that late-responsive DEGs were positively participated in defense against oxidative stress. Furthermore, heavy metal stress could result in the overproduction of ROS inside plant cells (Fahr et al. 2013). Accordingly, 4-day-old seedlings were used to estimate the content of hydrogen peroxide (H2O2) and superoxide (O2−). As shown in Fig. 3 b and c, the staining was significantly strengthened after Pb treatment for 24 h, indicating that Pb indeed greatly promotes ROS production in root tips.
Expression patterns of early- and late-responsive DEGs and physiological changes of Arabidopsis roots in response to lead (Pb) stress. a Hierarchical clustering heatmap of partial DEGs at 1 h and 24 h. b Detection of hydrogen peroxide in roots stained with DAB. Bar = 50 μm. c Detection of superoxide in roots stained with NBT. Bar = 50 μm
XTH18 Plays an Important Role in Pb-Induced Root Growth Inhibition
Xyloglucan endotransglucosylase/hydrolases (XTHs), one kind of cell wall-modifying enzymes, are correlates with hemicellulose modification and influences cellular expansion and cell wall loosening. In general, there are 33 members of the XTH gene family in Arabidopsis, and 10 of them exhibited root-specific expression (XTH-5, XTH-12, XTH-13, XTH-14, XTH-17, XTH-18, XTH-19, XTH-20, XTH-26, and XTH-31). Our RNA-seq data showed that the expression of XTH5, XTH18, XTH20, and XTH31 was quickly and consistently induced by Pb stress (Fig. 4a). According to previous study, XTH18 is also considered as one of major contributors to root growth and for normal root development (Vissenberg et al. 2005). Subsequently, we examined the effect of Pb on XTH18 expression by expressing the GUS reporter gene under the control of the XTH18 promoter (proXTH18::GUS) in 6-day-old Col-0 seedlings. The proXTH18::GUS/GFP was mainly expressed in root meristem and differentiation zone, and its expression was significantly enhanced by Pb treatments for 6 h, 12 h, and 24 h, compared with the mock (0 h) treatment (Fig. 4 b and c). We also compared Pb-induced root growth inhibition between Col-0 (wild-type) and xth18 mutant. Root growth of xth18 showed reduced sensitivity to Pb stress compared with that of Col-0 (Fig. 4 d and e). These results indicated XTH18-mediated cell wall modification was involved in inhibition of root growth under Pb stress.
Phenotype of the xth18 mutant and the expression pattern of XTH18 under mock and Pb stress. a Heatmap of ten root-specific expressed XTHs at four time points. b, c Six-day-old transgenic proXTH18::GUS/GFP seedlings were exposed to 0 (Mock) and 1 mM Pb for 6 h, 12 h, and 24 h. Bar = 100 μm. d, e Root growth of wild-type and xth18 plants after 6-day exposure to 0 (Mock) and 1 mM Pb. Data are shown as the average (± SD) (n = 10). Bar = 0.5 cm
A Putative Working Model of Pb Stress
To better understand the molecular mechanism of plant defense against Pb stress, a supposed model was constructed based on our RNA-seq data (Fig. 5). Perceiving Pb stress signal was the first principle step for plants subjected to Pb stress. In this study, many signaling molecules were identified (Supplementary Table 5). Two calmodulin genes (At2g41110 and At3g56800) were identified and up-regulated after Pb stress for 1 h, while they were constantly down-regulated at 6, 12, and 24 h, indicating that they could be rapidly induced for responding to Pb stress and probably function as the sensor of Pb stress. In addition, three mitogen-activated protein kinase kinase kinase (MPKKK, At2g30040, At2g32510, and At4g36950) were commonly up-regulated by Pb stress at all time intervals and proved to be involve in stress-activated protein kinase signaling cascade in Arabidopsis. Thereafter, Pb was transported from rhizosphere to root and shoot through transporters. Nramp3 (At2g23150), one of the metal transporters, was induced by Pb stress at all time points, and expression of which reached a peak at 6 h. We also identified three ABC transporters (At2g39350, At3g47780, and At4g01830), and two genes of which were continuously induced by Pb. Simultaneously, Pb particles could be absorbed by cell wall. In the cytoplasm, the majority of Pb ion, with or without chelation by phytochelatins, were finally compartmented in vacuole. One cation/H+ antiporter (At5g41610) was isolated and upregulated at all time points except stress for 12 h. Meanwhile, the increased Pb in the cell was responsible for generation of ROS (Ashraf et al. 2017). Under Pb stress condition, a large deal of ROS was generated in chloroplast by photosynthetic system and mitochondrion by respiratory electron transport chain (RETC). Three DEGs encoding NADPH related reductase (At1g10310, At2g17420, and At4g30210) were isolated, while expressed pattern of which was different under Pb stress. Two nitrate reductase (NADH, At1g37130 and At1g77760) were consistently induced by Pb stress, and their expression level reached peaks at 1 h. Furthermore, ROS were also generated through respiratory burst oxidase homolog (RBOH) located on cell membrane. To balance the cell metabolism, the activity of oxidoreductases was enhanced to scavenge ROS. Two genes encoding SOD (At2g28190 and At5g23310) were rapidly upregulated at 1 h and their expression level reach peak after Pb stress for 12 h. A protective enzyme POD (At4g08770) was also induced by Pb stress at all time points. Genes encoding CAT (At4g35090) and peroxidase (At2g48150) were upregulated under Pb stress to transform hydrogen peroxide to water. As shown in Supplemental Figs. 12 and 13, genes involved in Pb stress were differentially expressed and their correlation coefficient between RNA-seq and RT-qPCR is 0.805. Meanwhile, the ROS signaling pathway was activated to stimulate the defense responses.
Models of molecular mechanism against Pb stress in Arabidopsis root. Under Pb stress, the root growth was drastically inhibited. Subsequently, Pb particles were transported from rhizosphere into root, and lots of which were absorbed on cell wall or transferred into cell through membrane transporters (such as ABC transporters, ZIP transporters, and Ca2+ channels). While some of them were still stuck in the intercellular space. In the cytoplasm, Pb ion can be chelated by phytochelatins to synthesis Pb-chelates, which reacted with ATP to release the energy for normal life activities under the certain conditions and finally stored in the vacuole. Simultaneously, the majority of unchelated Pb ion was pumped inside the vacuole though the membrane proton pump. In addition, Pb ion also could be transported from root to shoot. With the accumulation of Pb in plant, a large deal of reactive oxygen species (ROS) was produced and regarded as the messenger against Pb stress. In terms of the chloroplast in the shoot, oxygen could be transformed to superoxide ion based on the photosynthetic system. Meanwhile, sugar-P was synthesized with the reaction of PGA and NADPH and transported to root. Under the catalytic action of sugar-P and NADPH, the majority of ROS was generated in intercellular space through RBOH and mitochondrion by respiratory electron transport chain (RECT), respectively. The ROS was transferred as the messenger to stimulate the defense responses against Pb stress. Subsequently, superoxide ion was transformed to hydrogen peroxide with the reaction of SOD, which was resolved into water and oxygen by antioxidant enzymes CAT and GPX. The black spots represent Pb ion. The bold curved lines with marine blue represent cell wall. The thin curved lines with marine blue represent cell membrane. The pansy oval represents ABC transporters. The pansy oval represents ABC transporters. The oval with dark khaki represents ZIP transporters. The oval with cardinal red represents Ca2+ channels. The tangerine oval represents RBOH. The bigger oval with cornflower blue represents vacuole. The circle with crimson represents antiporter. The pink rectangle represents mitochondrion. The oval with deep sky blue represents RECT. The dotted square with green represents chloroplast in shoot. The blue rectangles represent photosynthetic system. The irregular shape with yellow and red words represents ROS signaling. The red words represent their coding genes were u-pregulated under Pb stress. The green words represent the coding genes were down-regulated under Pb stress. Pb lead, ZIP Zrt/IRT-like protein, RBOH respiratory burst oxidase homolog, ATP adenosine triphosphate, ADP adenosine diphosphate, NADPH nicotinamide adenine dinucleotide phosphate, RECT respiratory electron transport chain, Sugar-P sugar phosphate, PSI/II photosystem I/II, PGA 3-phosphoglycerate, RUBP ribulose 1,5-bisphosphate, SOD superoxide dismutase, CAT catalase, GPX guaiacol peroxidase, GSH glutathione, reduced, GSSG glutathione, oxidized, ROS reactive oxygen species
Discussion
Over the years, heavy metal contaminants attributed to anthropogenic activities are becoming more and more severe. As a nonessential element for plant growth and development, the release of Pb into environment results in declined crop yield and threatens human health. Accordingly, it is urgent to understand the underlying molecular mechanism of Pb tolerance for breeding new varieties. In this study, genes and metabolism pathways related to Pb tolerance were identified via RNA-seq, which lay a soil foundation for further study on Pb stress.
In terms of plant morphology under Pb stress, previous studies found that an inhibitory effect of root growth and shoot growth was pronounced, and the extent of which was enhanced with the increasing concentration of Pb (Seneviratne et al. 2016). In the present study, root length significantly declined under Pb treatment compared with the control, which is consistent with the former studies. However, the shoot growth of Col-0 was not much influenced at the same level of Pb stress, which might be due to the duration and concentration of Pb treatment. Another possible reason is that root is more sensitive than other organs in response to abiotic stresses. Cell division activity and root meristem size were greatly reduced by Pb stress, which implies inhibition of root growth by Pb stress is a complex process.
RNA-seq has been widely applied to identify the DEGs among different conditions in Arabidopsis (Loraine et al. 2013). Nevertheless, there is little study on investigation of Pb stress by transcriptome profiling. In this study, a total of 15 samples were collected for RNA-seq analysis. According to Supplementary Table 2, the raw reads and total mapped rates of Col-Pb-0 h-3 were 22,880,621 and 78.93%, respectively, which were also lower than that of two others in the same group, speculating that these different results could be attributed to substrate depletion during sequencing processes. Regarding DEGs number, the number of down-regulated genes were much more than that of up-regulated genes at 1 h, 6 h, and 12 h, while the contrary situation occurred at 24 h (Fig. 2a). The results of previous study found that the number of up-regulated DEGs at 12 h was less than that at 24 h under Pb stress in maize root, and the number of down-regulated DEGs at 12 h was more than that at 24 h, consistent with the results in our study. These results indicated that the onset of response to Pb stress is early and the metabolism gradually reached a steady state in vivo. Additionally, a total of 703 commonly expressed DEGs were identified at all the time intervals (Fig. 2c). GO annotation analysis is regarded as a valid way for comprehensive investigation of gene function. In the previous study, genes involved in metal ion binding (GO: 0046872) were regarded to bind the harmful metal ions for chelate synthesis (Ortiz et al. 2019). In terms of GO analysis of DEGs under Pb stress, a lot of GO terms (transporter activity, catalytic activity, developmental growth, response to abiotic stimulus, cell wall, etc.) were enriched, and most of which were also identified in the current study, demonstrating that the strategies for confronting Pb stress were similar among different species (Tian et al. 2015). As shown in Fig. 3a, the expression level of partial DEGs at 1 h and 24 h was displayed and the changes were significant, which could be regarded as the marker genes at early and late stages of Pb stress. The accumulation of ROS resulted from heavy metal stress has been reported in previous study (Ashraf et al. 2017). As shown in Fig. 1 c and d, the color of root was markedly deepened after Pb stress, suggesting that large amounts of ROS were produced, which could be an important indicator of Pb stress.
When plants are exposed to environmental stress conditions, cell wall functions as a barrier against stress factors including metals. Under Pb stress, Pb particles can be fixed on cell wall, which affects the cell wall loosening. XTHs correlate with hemicellulose modification and influence cellular expansion and cell wall loosening. In addition, XTH18 is considered as one of major contributors to root growth and for normal root development (Vissenberg et al. 2005). In this study, the expression of XTH18 was induced and the xth18 mutant shows reduced sensitivity under Pb stress; these results indicated that cell wall modification is involved in lead stress response.
Signal transduction is a key step for plant defense system. During this process, a great quantity of signaling molecules and stress-related proteins were synthesized to activate specific genes for dealing with the stimulus. Calmodulin and mitogen-activated protein kinase kinase kinase (MAPKKK) have been recognized as essential signal molecules in response to stresses (Loraine et al. 2013). In our study, three MAPKKK genes were always up-regulated after Pb treatment at four time points, while two calmodulin genes were only up-regulated at 1 h and down-regulated at 6 h, 12 h, and 24 h, indicating that signal transduction of stress was coordinated. According to previous study, Pb is likely transported by non-specific transporters (Loraine et al. 2013). Therefore, Pb was absorbed into root and finally transported into vacuoles via membrane transporters, such as genes of ABC, ZIP, NRAMP, zinc transporter families, and antiporters. As shown in Supplementary Table 5, a total of 20 significant DEGs related to Pb transport, chelation, and sequestration were isolated for Pb metabolism. Pectin is considered as one of the Pb-chelating agents, and the expression level of its lyase decreased sharply at four time points, suggesting that synthesis of pectin was activated under Pb stress. Krämer et al. (2007) reported that the NRAMP transporters existed in root and shoot, and played an essential role in transport of metal ions through the plasma membrane and the tonoplast (Krämer et al. 2007), which was up-regulated at four time points after Pb treatment. In addition, the content of ROS increased markedly when exposed to abiotic stress (Huang et al. 2019). In this study, two photosystem elements were up-regulated to form much ROS with the extension of the stress time. Two genes encoding respiratory burst oxidase homolog (RBOH) protein were isolated and also up-regulated at 6 h, 12 h, and 24 h. To improve the adverse circumstance attributed to ROS, a large amount of antioxidant enzymes was compounded to release of ROS, such as SOD, CAT, and APX. Furthermore, the ROS signaling was simultaneously motivated for further response to Pb stress.
Conclusion
In summary, our study is mainly focused on the root growth inhibition of model plant Arabidopsis in response to Pb stress at morphological, physiological, and molecular levels and found that the early-responsive genes and the late-responsive genes, and the content of hydrogen peroxide (H2O2) and superoxide (O2−) increased significantly under Pb stress. In addition, XTH18, which encodes a xyloglucan endotransglucosylase, was induced by Pb and xth18 mutant seedling showed increased resistance to Pb stress, which indicated that cell wall modification was involved in of root growth inhibition under Pb stress. Additionally, a putative model was constructed for better understanding the defense mechanism in response to Pb toxicity.
References
Ashraf U, Tang X (2017) Yield and quality responses, plant metabolism and metal distribution pattern in aromatic rice under lead (Pb) toxicity. Chemosphere 176:141–155
Ashraf U, Kanu AS, Deng Q, Mo Z, Pan S, Tian H, Tang X (2017) Lead (Pb) toxicity; physio-biochemical mechanisms, grain yield, quality, and Pb distribution proportions in scented rice. Front Plant Sci 8:259
Colon-Carmona A, You R, Haimovitch-Gal T, Doerner P (1999) Technical advance: spatio-temporal analysis of mitotic activity with a labile cyclin-GUS fusion protein. Plant J 20:503–508
Dunand C, Crèvecoeur M, Penel C (2007) Distribution of superoxide and hydrogen peroxide in Arabidopsis root and their influence on root development: possible interaction with peroxidases. New Phytol 174:332–341
Fahr M, Laplaze L, Bendaou N, Hocher V, El Mzibri M, Bogusz D, Smouni A (2013) Effect of lead on root growth. Front Plant Sci 4:175
Hadi F, Aziz T (2015) A mini review on lead (Pb) toxicity in plants. J Biol Life Sci 6:91–101
Huang H, Ullah F, Zhou DX, Yi M, Zhao Y (2019) Mechanisms of ROS regulation of plant development and stress responses. Front Plant Sci 10:800
Kalisińska E (2019) Mammals and birds as bioindicators of trace element contaminations in terrestrial environments: an ecotoxicological assessment of the Northern Hemisphere. Springer, Berlin
Krämer U, Talke IN, Hanikenne M (2007) Transition metal transport. FEBS Lett 581:2263–2272
Loraine AE, McCormick S, Estrada A, Patel K, Qin P (2013) RNA-seq of Arabidopsis pollen uncovers novel transcription and alternative splicing. Plant Physiol 162:1092–1109
Ortiz J, Soto J, Fuentes A, Herrera H, Meneses C, Arriagada C (2019) The endophytic fungus Chaetomium cupreum regulates expression of genes involved in the tolerance to metals and plant growth promotion in Eucalyptus globulus roots. Microorganisms 7:490
Samardakiewicz S, Krzesłowska M, Bilski H, Bartosiewicz R, Woźny A (2012) Is callose a barrier for lead ions entering Lemna minor L. root cells? Protoplasma 249:347–351
Seneviratne M, Gunaratne S, Bandara T, Weerasundara L, Rajakaruna N, Seneviratne G, Vithanage M (2016) Plant growth promotion by Bradyrhizobium japonicum under heavy metal stress. S Afr J Bot 105:19–24
Tian SQ, Gu CS, Liu LQ, Zhu XD, Zhao YH, Huang SZ (2015) Transcriptome profiling of Louisiana iris root and identification of genes involved in lead-stress response. Int J Mol Sci 16:28087–28097
Vissenberg K, Kris V, Mika O, Yasue O, Ryusuke Y, Jean-Pierre V, Kazuhiko N (2005) Differential expression of AtXTH17, AtXTH18, AtXTH19 and AtXTH20 genes in Arabidopsis roots. Physiological roles in specification in cell wall construction. Plant Cell Physiol 46:192–200
Zhang XY, Zhou WK, Chen Q, Fang MM, Zheng SS, Scheres B, Li CY (2018) Mediator subunit MED31 is required for radial patterning of Arabidopsis roots. P Natl Acad Sci USA 115:5624–5633
Acknowledgments
This study was supported by the Basic Research Program of Shandong (Grant ZR2018ZC08N1), the Ministry of Agriculture-Chinese (2016ZX08009003-001-006), the National Key Research and Development Program of China (2019YFD1000300), and the Tai-Shan Scholars from the Shandong Provincial Government (tsqn20161021 and tsxk20150901).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Compliance with Ethical Standards
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of Interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key Message
Lead inhibits root growth. The early- and late-responsive genes involved in lead stress were identified by RNA-seq analysis with Arabidopsis root tips. XTH18 which is a component in cell wall modification was upregulated, and root growth of xth18 showed reduced sensitivity to lead stress, suggesting that cell wall modification is involved in root growth inhibition under lead stress.
Electronic supplementary material
Supplementary Table 1
Primers for reverse transcription quantitative PCR. (DOC 45 kb) (DOCX 17 kb)
Supplementary Table 2
Quality evaluation of data sequenced on the Illumina platform. (DOCX 16 kb)
Supplementary Table 3
List of DEGs (Padj < 0.01) at four time points. (XLSX 1168 kb)
Supplementary Table 4
List of GO terms enriched with 703 commonly expressed DEGs. (XLSX 119 kb)
Supplementary Table 5
List of key DEGs involved in Pb metabolism. (XLSX 142 kb)
Supplementary Table 6
List of DEGs (Padj < 0.01) at 1 h-6 h, 6 h-12 h, 12 h-24 h and 1 h-24 h. (XLSX 1186 kb)
Supplementary Fig. 1
Principal component analysis (PCA) of 15 samples based on the transcripts expression levels estimated by transcripts per million (TPM) value. (JPG 411 kb)
Supplementary Fig. 2
Correlation analysis of 14 DEGs between RNA-seq and RT-qPCR. (JPG 119 kb)
Supplementary Fig. 3
Gene ontology analysis of 703 commonly expressed DEGs (molecular function). (JPG 3430 kb)
Supplementary Fig. 4
Gene ontology analysis of 703 commonly expressed DEGs (biological process). (JPG 3787 kb)
Supplementary Fig. 5
Gene ontology analysis of 703 commonly expressed DEGs (cellular component). (JPG 1644 kb)
Supplementary Fig. 6
The top 20 terms of GO enrichment analysis at 1 h (biological process). (JPG 649 kb)
Supplementary Fig. 7
The top 20 terms of GO enrichment analysis at 24 h (biological process). (JPG 677 kb)
Supplementary Fig. 8
The significant GO terms at 1 h-6 h. (biological process). (PNG 168 kb)
Supplementary Fig. 9
The significant GO terms at 6 h-12 h. (biological process). (PNG 182 kb)
Supplementary Fig. 10
The significant GO terms at 12 h-24 h. (biological process). (PNG 163 kb)
Supplementary Fig. 11
The significant GO terms at 1 h-24 h. (biological process). (PNG 137 kb)
Supplementary Fig. 12
Heatmap of genes involved in Pb stress at four time points. (JPG 230 kb)
Supplementary Fig. 13
Correlation analysis of 19 genes involved in Pb stress between RNA-seq and RT-qPCR. (JPG 290 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zheng, S., Ren, P., Zhai, M. et al. Identification of Genes Involved in Root Growth Inhibition Under Lead Stress by Transcriptome Profiling in Arabidopsis. Plant Mol Biol Rep 39, 50–59 (2021). https://doi.org/10.1007/s11105-020-01233-y
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-020-01233-y