Abstract
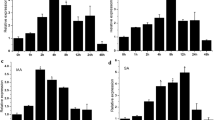
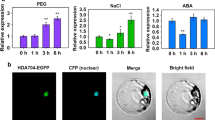
Histone deacetylation catalyzed by histone deacetylases is an important type of histone modification. Histone deacetylases affect various processes of plant development and involve in responding to hormones and biotic and abiotic stresses. Here, we report a tomato PRD3/HDA1 histone deacetylase gene, SlHDA5, which is expressed ubiquitously in different tissues and development stages. Expression profiles in hormone treatments showed that SlHDA5 was induced by abscisic acid (ABA) and methyl jasmonate (MeJA). Seedlings growth of SlHDA5-RNAi lines were more inhibited on the medium containing salt compared with wild type (WT). Under salt stress, chlorophyll in mature leaves degraded earlier in transgenic leaves than that in WT, and transgenic plants displayed wilting earlier and more severe than WT. After drought treatment, transgenic plants wilted and dehydrated earlier than WT, which was confirmed by lower water and chlorophyll content, and higher malondialdehyde (MDA) content in transgenic plants manifesting that the tolerance of transgenic plants to drought receded. Under the treatment of ABA, root length of transgenic seedlings was more strongly repressed by contrast with WT, suggesting repression of SlHDA5 increased seedling sensibility to ABA. Our study indicated that silencing of SlHDA5 resulted in decreasing tolerance to salt, drought, and ABA.





Similar content being viewed by others
References
Alinsug MV, CW Y, Wu K (2009) Phylogenetic analysis, subcellular localization, and expression patterns of RPD3/HDA1 family histone deacetylases in plants. BMC Plant Biol 9(1):37. https://doi.org/10.1186/1471-2229-9-37
Benedetti CE, Costa CL, Turcinelli SR, Arruda P (1998) Differential expression of a novel gene in response to coronatine, methyl jasmonate, and wounding in the Coi1 mutant of Arabidopsis. Plant Physiol 116(3):1037–1042. https://doi.org/10.1104/pp.116.3.1037
Berger SL (2007) The complex language of chromatin regulation during transcription. Nature 447(7143):407–412. https://doi.org/10.1038/nature05915
Bourque S, Dutartre A, Hammoudi V, Blanc S, Dahan J, Jeandroz S, Pichereaux C, Rossignol M, Wendehenne D (2011) Type-2 histone deacetylases as new regulators of elicitor-induced cell death in plants. New Phytol 192(1):127–139. https://doi.org/10.1111/j.1469-8137.2011.03788.x
Chen ZJ, Tian L (2007) Roles of dynamic and reversible histone acetylation in plant development and polyploidy. Biochim. Biophys. Acta Gene Struct. Expr. 1769(5-6):295–307. https://doi.org/10.1016/j.bbaexp.2007.04.007
Chen LT, Wu K (2010) Role of histone deacetylases HDA6 and HDA19 in ABA and abiotic stress response. Plant Signal Behav 5(10):1318–1320. https://doi.org/10.4161/psb.5.10.13168
Chen GP, Hackett R, Walker D, Taylor A, Lin ZF, Grierson D (2004) Identification of a specific isoform of tomato lipoxygenase (TomloxC) involved in the generation of fatty acid-derived flavor compounds. Plant Physiol 136(1):2641–2651. https://doi.org/10.1104/pp.104.041608
Chen LT, Luo M, Wang YY, KQ W (2010) Involvement of Arabidopsis histone deacetylase HDA6 in ABA and salt stress response. J Exp Bot 61(12):3345–3353. https://doi.org/10.1093/jxb/erq154
Cigliano RA, Cremona G, Paparo R, Termolino P, Perrella G, Gutzat R, Consiglio MF, Conicella C (2013) Histone deacetylase AtHDA7 is required for female gametophyte and embryo development in Arabidopsis. Plant Physiol 163(1):431–440. https://doi.org/10.1104/pp.113.221713
Demetriou K, Kapazoglou A, Tondelli A, Francia E, Stanca MA, Bladenopoulos K, Tsaftaris AS (2009) Epigenetic chromatin modifiers in barley: I. Cloning, mapping and expression analysis of the plant specific HD2 family of histone deacetylases from barley, during seed development and after hormonal treatment. Physiol. Plant. 136(3):358–368. https://doi.org/10.1111/j.1399-3054.2009.01236.x
Earley K, Lawrence RJ, Pontes O, Reuther R, Enciso AJ, Silva M, Neves N, Gross M, Viegas W, Pikaard CS (2006) Erasure of histone acetylation by Arabidopsis HDA6 mediates large-scale gene silencing in nucleolar dominance. Genes Dev 20(10):1283–1293. https://doi.org/10.1101/gad.1417706
Exposito-Rodriguez M, Borges AA, Borges-Perez A, Perez JA (2008) Selection of internal control genes for quantitative real-time RT-PCR studies during tomato development process. BMC Plant Biol 8(1):131. https://doi.org/10.1186/1471-2229-8-131
Fuchs J, Demidov D, Houben A, Schubert I (2006) Chromosomal histone modification patterns—from conservation to diversity. Trends Plant Sci 11(4):199–208. https://doi.org/10.1016/j.tplants.2006.02.008
Fujita M, Fujita Y, Maruyama K, Seki M, Hiratsu K, Ohme-Takagi M, Tran LSP, Yamaguchi-Shinozaki K, Shinozaki K (2004) A dehydration-induced NAC protein, RD26, is involved in a novel ABA-dependent stress-signaling pathway. Plant J 39(6):863–876. https://doi.org/10.1111/j.1365-313X.2004.02171.x
Guo J-E, Hu Z, Guo X, Zhang L, Yu X, Zhou S, Chen G (2017) Molecular characterization of nine tissue-specific or stress-responsive genes of histone deacetylase in tomato (Solanum lycopersicum). J Plant Growth Regul 36(3):566–577. https://doi.org/10.1007/s00344-016-9660-8
Heath RL, Packer L (1968) Photoperoxidation in isolated chloroplasts .I. Kinetics and stoichiometry of fatty acid peroxidation. Arch Biochem Biophys 125(1):189–198. https://doi.org/10.1016/0003-9861(68)90654-1
Hollender C, Liu Z (2008) Histone deacetylase genes in Arabidopsis development. J Integr Plant Biol 50(7):875–885. https://doi.org/10.1111/j.1744-7909.2008.00704.x
Jang IC, Pahk YM, Song SI, Kwon HJ, Nahm BH, Kim JK (2003) Structure and expression of the rice class-I type histone deacetylase genes OsHDAC1-3: OsHDAC1 overexpression in transgenic plants leads to increased growth rate and altered architecture. Plant J 33(3):531–541. https://doi.org/10.1046/j.1365-313X.2003.01650.x
Kim KC, Lai ZB, Fan BF, Chen ZX (2008a) Arabidopsis WRKY38 and WRKY62 transcription factors interact with histone deacetylase 19 in basal defense. Plant Cell 20(9):2357–2371. https://doi.org/10.1105/tpc.107.055566
Kim JM, To TK, Ishida J, Morosawa T, Kawashima M, Matsui A, Toyoda T, Kimura H, Shinozaki K, Seki M (2008b) Alterations of lysine modifications on the histone H3 N-tail under drought stress conditions in Arabidopsis thaliana. Plant & Cell Physiology 49(10):1580–1588. https://doi.org/10.1093/pcp/pcn133
Kim JM, To TK, Ishida J, Matsui A, Kimura H, Seki M (2012) Transition of chromatin status during the process of recovery from drought stress in Arabidopsis thaliana. Plant & Cell Physiology 53(5):847–856. https://doi.org/10.1093/pcp/pcs053
Kouzarides T (2007a) Chromatin modifications and their function. Cell 128(4):693–705. https://doi.org/10.1016/j.cell.2007.02.005
Kouzarides T (2007b) SnapShot: histone-modifying enzymes. Cell 131(4):822–822.e1. https://doi.org/10.1016/j.cell.2007.11.005
Lagace M, Chantha SC, Major G, Matton DP (2003) Fertilization induces strong accumulation of a histone deacetylase (HD2) and of other chromatin-remodeling proteins in restricted areas of the ovules. Plant Mol Biol 53(6):759–769. https://doi.org/10.1023/B:PLAN.0000023665.36676.89
Lim MY, Pulla RK, Park JM, Harn CH, Jeong BR (2012) Over-expression of l-gulono-gamma-lactone oxidase (GLOase) gene leads to ascorbate accumulation with enhanced abiotic stress tolerance in tomato. In Vitro Cell. Dev. Biol. Plant 48:453–461
Liu XC, CW Y, Duan J, Luo M, Wang KC, Tian G, Cui YH, KQ W (2012a) HDA6 directly interacts with DNA methyltransferase MET1 and maintains transposable element silencing in Arabidopsis. Plant Physiol 158(1):119–129. https://doi.org/10.1104/pp.111.184275
Liu X, Luo M, Wu K (2012b) Epigenetic interplay of histone modifications and DNA methylation mediated by HDA6. Plant Signal Behav 7(6):633–635. https://doi.org/10.4161/psb.19994
Liu C, Li LC, Chen WQ, Chen X, ZH X, Bai SN (2013) HDA18 affects cell fate in Arabidopsis root epidermis via histone acetylation at four kinase genes. Plant Cell 25(1):257–269. https://doi.org/10.1105/tpc.112.107045
Liu XC, Yang SG, Zhao ML, Luo M, CW Y, Chen CY, Tai R, KQ W (2014) Transcriptional repression by histone deacetylases in plants. Mol Plant 7(5):764–772. https://doi.org/10.1093/mp/ssu033
Loidl P (2004) A plant dialect of the histone language. Trends Plant Sci 9(2):84–90. https://doi.org/10.1016/j.tplants.2003.12.007
Luger K, Mader AW, Richmond RK, Sargent DF, Richmond TJ (1997) Crystal structure of the nucleosome core particle at 2.8 angstrom resolution. Nature 389(6648):251–260. https://doi.org/10.1038/38444
Luo M, Wang YY, Liu XC, Yang SG, Lu Q, Cui YH, KQ W (2012) HD2C interacts with HDA6 and is involved in ABA and salt stress response in Arabidopsis. J Exp Bot 63(8):3297–3306. https://doi.org/10.1093/jxb/ers059
Luo M, Tai R, CW Y, Yang SG, Chen CY, Lin WD, Schmidt W, KQ W (2015) Regulation of flowering time by the histone deacetylase HDA5 in Arabidopsis. Plant J 82(6):925–936. https://doi.org/10.1111/tpj.12868
Lusser A, Kolle D, Loidl P (2001) Histone acetylation: lessons from the plant kingdom. Trends Plant Sci 6(2):59–65. https://doi.org/10.1016/S1360-1385(00)01839-2
Pandey R, Muller A, Napoli CA, Selinger DA, Pikaard CS, Richards EJ, Bender J, Mount DW, Jorgensen RA (2002) Analysis of histone acetyltransferase and histone deacetylase families of Arabidopsis thaliana suggests functional diversification of chromatin modification among multicellular eukaryotes. Nucleic Acids Res 30(23):5036–5055. https://doi.org/10.1093/nar/gkf660
Pei ZM, Kuchitsu K, Ward JM, Schwarz M, Schroeder JI (1997) Differential abscisic acid regulation of guard cell slow anion channels in Arabidopsis wild-type and abi1 and abi2 mutants. Plant Cell 9(3):409–423. https://doi.org/10.1105/tpc.9.3.409
Pipal A, Goralik-Schramel M, Lusser A, Lanzanova C, Sarg B, Loidl A, Lindner H, Rossi V, Loidl P (2003) Regulation and processing of maize histone deacetylase Hda1 by limited proteolysis. Plant Cell 15(8):1904–1917. https://doi.org/10.1105/tpc.013995
Schmid M, Davison TS, Henz SR, Pape UJ, Demar M, Vingron M, Scholkopf B, Weigel D, Lohmann JU (2005) A gene expression map of Arabidopsis thaliana development. Nat Genet 37(5):501–506. https://doi.org/10.1038/ng1543
Sendra R, Rodrigo I, Salvador ML, Franco L (1988) Characterization of pea histone deacetylases. Plant Mol Biol 11(6):857–866. https://doi.org/10.1007/BF00019525
Sridha S, Wu K (2006) Identification of AtHD2C as a novel regulator of abscisic acid responses in Arabidopsis. Plant J. 46(1):124–133. https://doi.org/10.1111/j.1365-313X.2006.02678.x
Tanaka M, Kikuchi A, Kamada H (2008) The Arabidopsis histone deacetylases HDA6 and HDA19 contribute to the repression of embryonic properties after germination. Plant Physiol 146(1):149–161. https://doi.org/10.1104/pp.107.111674
Taunton J, Hassig CA, Schreiber SL (1996) A mammalian histone deacetylase related to the yeast transcriptional regulator Rpd3p. Science 272(5260):408–411. https://doi.org/10.1126/science.272.5260.408
Tian L, Chen ZJ (2001) Blocking histone deacetylation in Arabidopsis induces pleiotropic effects on plant gene regulation and development (vol 98, pg 200, 2001). Proc Natl Acad Sci U S A 98:7647–7647
To TK, Kim JM (2014) Epigenetic regulation of gene responsiveness in Arabidopsis. Front Plant Sci 4. https://doi.org/10.3389/fpls.2013.00548
To TK, Kim JM, Matsui A, Kurihara Y, Morosawa T, Ishida J, Tanaka M, Endo T, Kakutani T, Toyoda T, Kimura H, Yokoyama S, Shinozaki K, Seki M (2011) Arabidopsis HDA6 regulates locus-directed heterochromatin silencing in cooperation with MET1. PLoS Genet 7(4):e1002055. https://doi.org/10.1371/journal.pgen.1002055
Wang Z, Cao H, Chen FY, Liu YX (2014) The roles of histone acetylation in seed performance and plant development. Plant Physiol. Biochem. 84:125–133. https://doi.org/10.1016/j.plaphy.2014.09.010
Wu K, Zhang L, Zhou C, CW Y, Chaikam V (2008) HDA6 is required for jasmonate response, senescence and flowering in Arabidopsis. J Exp Bot 59(2):225–234. https://doi.org/10.1093/jxb/erm300
Yang XJ, Seto E (2007) HATs and HDACs: from structure, function and regulation to novel strategies for therapy and prevention. Oncogene 26(37):5310–5318. https://doi.org/10.1038/sj.onc.1210599
Yu CW, Liu XC, Luo M, Chen CY, Lin XD, Tian G, Lu Q, Cui YH, KQ W (2011) HISTONE DEACETYLASE6 interacts with FLOWERING LOCUS D and regulates flowering in Arabidopsis. Plant Physiol 156:173–184
Zhao LM, JX L, Zhang JX, PY W, Yang SG, KQ W (2015a) Identification and characterization of histone deacetylases in tomato (Solanum lycopersicum). Front Plant Sci 5. https://doi.org/10.3389/fpls.2014.00760
Zhao JH, Zhang JX, Zhang W, KL W, Zheng F, Tian LN, Liu XC, Duan J (2015b) Expression and functional analysis of the plant-specific histone deacetylase HDT701 in rice. Front Plant Sci 5. https://doi.org/10.3389/fpls.2014.00764
Zhao JH, Li MZ, Gu DC, Liu XC, Zhang JX, Wu KL, Zhang XH, da Silva JAT, Duan J (2016) Involvement of rice histone deacetylase HDA705 in seed germination and in response to ABA and abiotic stresses. Biochem. Biophys. Res. Commun. 470(2):439–444. https://doi.org/10.1016/j.bbrc.2016.01.016
Zhou CH, Zhang L, Duan J, Miki B, KQ W (2005) HISTONE DEACETYLASE19 is involved in jasmonic acid and ethylene signaling of pathogen response in Arabidopsis. Plant Cell 17(4):1196–1204. https://doi.org/10.1105/tpc.104.028514
Funding
This work was supported by National Natural Science Foundation of China (no. 31572129) and the Natural Science Foundation of Chongqing of China (cstc2015jcyjA80026).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Yu, X., Gao, Q., Chen, G. et al. SlHDA5, a Tomato Histone Deacetylase Gene, Is Involved in Responding to Salt, Drought, and ABA. Plant Mol Biol Rep 36, 36–44 (2018). https://doi.org/10.1007/s11105-017-1057-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-017-1057-8