Abstract
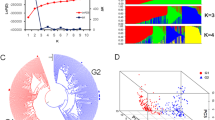
Jatropha (Jatropha curcas) and castor bean (Ricinus communis) possess several taxonomic similarities, and their seeds contain a high proportion of oil (up to 40%) which has been used in various industrial products, including diesel oil. Thirty-two candidate genes responsible for fatty acid biosynthesis were identified in the castor bean genome sequence. Testing of 48 primer pairs from candidate gene regions, including 12 SSRs from castor bean on 54 genotypes of J. curcas, 65% amplified successfully on Jatropha out of which 20% showed polymorphisms. Jatropha genotypes, categorized for oil content, were used in association analysis of candidate gene regions with high oil content. One marker–trait association for the oil trait was identified. Stearoyl desaturase amplicon (700 bp) consisting of intron and exon (P = 0.00013) showed association with high oil content in Jatropha genotypes. Sequencing of the 1.3-kb amplicon, including the 700-bp fragment of stearoyl desaturase, which had shown association with the high oil content, revealed SNPs in the exonic region. The SNPs resulted in substitution of leucine with glutamine in the open reading frame of stearoyl desaturase of low oil content genotypes. The molecular marker is expected to be useful in marker-assisted breeding of high oil content genotypes in Jatropha.



Similar content being viewed by others
References
Akbar E, Yaakob J, Kamarudin SK, Ismail M, Salimon J (2009) Characteristic and composition of Jatropha Curcas oil seed from Malaysia and its potential as biodiesel feedstock. Eur J Sci Res 29:396–403
Aranzana MJ, Kim S, Zhao K, Bakker E, Horton M et al (2005) Genome-wide association mapping in Arabidopsis identifies previously known flowering time and pathogen resistance genes. PLoS Genet 1:e60
Basha SD, Sujatha M (2007) Inter and intra-population variability of Jatropha curcas (L.) characterized by RAPD and ISSR markers and development of population-specific SCAR markers. Euphytica 156:375–386
Bonaventure G, Salas JJ, Pollard MR, Ohlrogge JB (2003) Disruption of the FATB Gene in Arabidopsis Demonstrates an Essential Role of Saturated Fatty Acids in Plant Growth. The Plant Cell 15:1020–1033
Breseghello F, Sorrells ME (2006) Association mapping of kernel size and milling quality in wheat (Triticum aestivum L.) cultivars. Genetics 172:1165–1177
Casa AM, Pressoir G, Brown PJ, Mitchell SE, Rooney WL, Tuinstra MR, Franks CD, Kresovich S (2008) Community resources and strategies for association mapping in sorghum. Crop Sci 48:30–40
Chen Z, Wang ML, Barkley NA, Pittman RN (2010) A simple allele-specific PCR assay for detecting FAD2 alleles in both A and B genomes of the cultivated peanut for high-oleate trait selection. Plant Mol Biol Rep 28(3):542–548
Chi X, Yang Q, Lu Y, Wang J, Zhang Q, Pan L, Chen M, He Y, Yu S (2011) Genome-wide analysis of fatty acid desaturases in soybean (Glycine max). Plant Mol Biol Rep 29(4):769–783
Cuesta-Marcos A, Szőcs P, Close TJ, Filichkin T, Muehlbauer GJ, Smith KP, Hayes PM (2010) Genome-wide SNPs and re-sequencing of growth habit and inflorescence genes in barley: implications for association mapping in germplasm arrays varying in size and structure. BMC Genomics 11:707
Divakara BN, Upadhyaya HD, Wani SP, Laxmipathi Gowda CL (2010) Biology and genetic improvement of Jatropha curcas L.: a review. Appl Energy 87:732–742
Ellwood SR, Phan HTT, Jordan M, Hane J, Torres AM, Avila CM, Serafín Cruz-Izquierdo S, Oliver RP (2008) Construction of a comparative genetic map in faba bean (Vicia faba L.); conservation of genome structure with Lens culinaris. BMC Genomics 9:380–391
Goudet J (1995) FSTAT (Version 1.2): a computer program to calculate F-statistics. J Hered 86:485–489
Guan LL, Xu YW, Wang YB, Chen L, Shao JF, Wu W (2011) Isolation and characterization of a temperature-regulated microsomal oleate desaturase gene (CtFAD2-1) from safflower (Carthamus tinctorius L.). Plant Mol Biol Rep. doi:10.1007/s11105-011-0349-7
Heesacker A, Venkata KK, Gao W, Tang S, Kolkman JM, Gingle A, Matvienko M, Kozik A, Michelmore RM, Lai Z, Rieseberg LH, Knapp SJ (2009) SSRs and INDELs mined from the sunflower EST database: abundance, polymorphisms, and cross-taxa utility. Theor Appl Genet 117:1021–1029
Hill WG, Robertson A (1968) Linkage disequilibrium in finite populations. Theor Appl Genet 38:226–231
Keller B, Feuillet C (2000) Collinearity and gene density in grass genomes. Trends Plant Sci 5:246–251
Lander ES, Schork NJ (1994) The genetic dissection of complex traits. Science 265:2037–2048
Lewontin RC (1964) The interaction of selection and linkage. I. General considerations; heterotic models. Genetics 49:49–67
Liu K, Muse S (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129
Makkar HPS, Becker K, Sporen F, Wink M (1997) Studies on nutritive potential and toxic constituents of different provenances of Jatropha curcas L. J Agric Food Chem 45:3152–3157
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4325
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in excel population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Sato S, Hirakawa H, Isobe S, Fukai E, Watanabe A, Kato M, Kawashima K, Minami C, Muraki A, Nakazaki N, Takahashi C, Nakayama S, Kishida Y, Kohara M, Yamada M, Tsuruoka H, Sasamoto S, Tabata S, Aizu T, Toyoda A, Shin-I T, Minakuchi Y, Kohara Y, Fujiyama A, Tsuchimoto S, Kajiyama S, Makigano E, Ohmido N, Shibagaki N, Cartagena JA, Wada N, Kohinata T, Atefeh A, Yuasa S, Matsunaga S, Fukui K (2011) Sequence analysis of the genome of an oil-bearing tree, Jatropha curcas L. DNA Res 18:65–76
Sharma A, Chauhan RS (2011) In-silico identification and comparative genomics of candidate genes involved in biosynthesis and accumulation of seed oil in plants. Comparative and Functional Genomics (in press)
Sorrells ME, Rota ML, Bermudez-Kandianis CE et al (2003) Comparative DNA sequence analysis of wheat and rice genomes. Genome Res 13:1818–1827
Tanya P, Taeprayoon P, Hadkam Y, Srinives P (2011) Genetic diversity among Jatropha and Jatropha-related species based on ISSR markers. Plant Mol Biol Rep 29:252–264
Thornsberry JM, Goodman MM, Doebley J, Kresovich S, Nielsen D, Buckler ES (2001) Dwarf8 polymorphisms associate with variation in flowering time. Nat Genet 28:286–289
Tian Z, Qian Q, Liu Q, Yan M, Liu X, Yan C, Liu G, Gao Z, Tang S, Zeng D, Wang Y, Yu J, Gu M, Li J (2009) Allelic diversities in rice starch biosynthesis lead to a diverse array of rice eating and cooking qualities. Proc Natl Acad Sci USA 106:21760–21765
Wang CM, Liu P, Yi C, Gu K, Sun F et al (2011) A first generation microsatellite- and SNP-based linkage map of Jatropha. PLoS One 6(8):e23632
Wilson LM, Whitt SR, Ibanez AM, Rocheford TR, Goodman MM et al (2004) Dissection of maize kernel composition and starch production by candidate gene association. Plant Cell 16:2719–2733
Yu J, Pressoir G, Briggs W, Vroh BI, Yamasaki M, Doebley JF, McMullen MD, Gaut BS, Nielsen DM, Holland J, Kresovich S, Buckler ES (2006) A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38:203–208
Zhao K, Aranzana MJ, Kim S, Lister C, Shindo C, Tang C, Toomajian C, Zheng H, Dean C, Marjoram P, Nordborg M (2007) An Arabidopsis example of association mapping in structured samples. PLoS Genet 31:e4
Zondervan KT, Cardon LR (2004) The complex interplay among factors that influence allelic association. Nat Rev Genet 5:89–100
Acknowledgment
The authors are thankful to the Department of Biotechnology, Ministry of Science and Technology, Government of India for providing a research grant to RSC.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table 1
Primers used to amplify candidate genes for fatty acid biosynthesis in Jatropha (DOCX 42 kb)
Supplementary Table 2
SSRs identified in the candidate genes for fatty acid biosynthesis in castor bean (DOCX 36 kb)
Supplementary Table 3
List of genotypes used in the present study (DOCX 37 kb)
Supplementary Table 4
Annotation of candidate genes for fatty acid biosynthesis in the castor bean genome (DOCX 49 kb)
Rights and permissions
About this article
Cite this article
Sharma, A., Chauhan, R.S. Identification and Association Analysis of Castor Bean Orthologous Candidate Gene-Based Markers for High Oil Content in Jatropha curcas . Plant Mol Biol Rep 30, 1025–1031 (2012). https://doi.org/10.1007/s11105-011-0408-0
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-011-0408-0