Abstract
Purpose
Plant domestication altered leaf litter quality. Since litter traits relate to soil functions and organisms (i.e., litter decomposition and soil decomposer communities), in this study we explore if domestication-induced changes in litter quality have affected their decomposability, and bacterial, fungal, and nematode communities in the soil.
Methods
We collected leaf litter from herbaceous crops and their wild progenitors, and measured litter chemical and physical traits. Then, we performed a litter decomposition assay on a common soil. After three months of litter incubation, we measured mass loss, nematode richness and community composition in ten crops. We also measured soil bacterial and fungal richness and community composition in six crops.
Results
Domesticated litters had less carbon (C) and leaf dry matter content (LDMC), which accelerated decomposition in comparison to wild litters. Fungal richness was higher in microcosms incubated with domesticated litters, while the effects of domestication on bacterial richness differed among crops. Domestication did not affect nematode richness. The effects of domestication on bacterial and fungal community compositions differed among crops. Soils with domesticated litters tended to have nematode communities with a higher abundance of bacterial feeding nematodes, in comparison to soils fed with wild litters.
Conclusion
Domestication altered decomposition at different levels. Leaf litter decomposability increased with domestication, which might alter resource inputs into the soil. Feeding soils with domesticated litters had idiosyncratic effects on soil microbes, but consistent effects on soil nematodes. Overall, domestication altered the linkages between crop residues and soil communities differently for bacteria, fungi, and nematodes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plant domestication is a type of mutualism in which plants produce a service for humans (i.e. food), and humans manage the domesticated plants’ environment and reproduction (Purugganan 2022). Plant evolution in croplands triggered morphological, biochemical and physiological changes from the wild progenitors to the domesticated plants, as the result of two selection forces: natural selection and artificial selection (Doebley et al. 2006; Purugganan and Fuller 2009). Natural selection entails the selection of plant genotypes by conditions that exist in agroecosystems (i.e., climate, nutrient availability, etc.). Artificial selection entails the selection of specific plant traits with desirable characteristics by humans to meet their needs. Plant evolution in agroecosystems has also promoted multiple indirect and unintentional effects on plant traits (Hancock 2012; Milla et al. 2015). For instance, some crops have lost (part of their) chemical defences or have developed softer leaves (Meyer et al. 2012). Domestication probably altered leaf traits, such as carbon (C) and nitrogen (N) contents (Prieto et al. 2017; Robinson et al. 2022; Roucou et al. 2018). Domestication also affected leaf litter traits (García-Palacios et al. 2013). Leaf litter traits influence important soil functions, including litter decomposition and soil communities (Fanin et al. 2014; Freschet et al. 2012). Therefore, domestication might have altered crop litter decomposition and the changes that soil communities undergo during the decomposition process.
Litter decomposition is the process through which soil decomposers and litter-fragmenting soil fauna break down plant residues into smaller pieces and simple molecules (Cotrufo et al. 2010). Litter traits (i.e. chemical composition and physical properties) explain most of the variation in litter decomposition rates (Cornwell et al. 2008), and indicate the quality of litters as a trophic resource for decomposers (Strickland et al. 2009). Leaf litter C, N, and lignin contents usually account for most of the variability in litter decomposability (Cornwell et al. 2008; Freschet et al. 2012). Other nutrients, such as P, Mg or Ca, also explain part of the variance observed during decomposition (García-Palacios et al. 2016a; Pichon et al. 2020). Litter from leaves that are tougher or have low water contents (high leaf-dry matter content; LDMC) tend to decompose more slowly (Kazakou et al. 2009; Pakeman et al. 2011; Pérez Harguindeguy et al. 2015; Rawlik et al. 2022). In general, leaf litter of herbaceous crops have higher quality and thereby decomposes faster than that of their wild progenitors (García-Palacios et al. 2013). This can be explained by shifts in litter chemical properties. Leaf litter of domesticated crops have less C and lignin contents than their wild progenitors, and decomposes faster (García-Palacios et al. 2013; González-Paleo et al. 2022). In a study, cultivated accessions of Silphium integrifolium Michx. had thinner leaves with higher N contents than wild accessions (González-Paleo et al. 2022). Changes in litter quality are linked to higher concentrations of nitrate in soils, which may have implications for the management of agroecosystems (García-Palacios et al. 2013). Despite this, studies that consider domestication-induced changes in physical and chemical leaf litter traits and decomposition rates in several crop species are lacking. Moreover, it is unknown if and how these changes in litter quality and decomposition rates also impact soil microbial decomposers and the microfauna that feed on them.
The litter layer shapes the diversity and composition of soil microbial communities (Fanin et al. 2014). Litter traits are an environmental filter for microbial succession (Kraft et al. 2015). For instance, the microbial community underneath forest litter layers are specific to the plant species contributing litter inputs (Prescott and Grayston 2013). Decomposers can change and adapt to specific litter inputs (Strickland et al. 2009). Such adaptation of decomposers feedbacks positively on the decomposition of subsequent litters of similar identity (Veen et al. 2021). Also, bacterial and fungal communities’ richness and community structure correlate with litter chemical properties and to each other (Purahong et al. 2016), which suggest that the shifts they undergo during the decomposition process are coordinated. The nematode community might also be shaped by litter traits (García-Palacios et al. 2016b), but their relationship with plant litter is mostly indirect because many nematodes feed on bacteria and fungi, but do not act as primary decomposers (Yeates 1999; Yeates et al. 1993). However, even if as an indirect effect, soil nematode communities respond differently to litters of different plant species (Wardle et al. 2006). Also, litters with disparate traits support nematode communities with different abundances of bacterial and fungal feeders (García-Palacios et al. 2016b). Relationships between litter traits and soil microbial and nematode communities have also been found in croplands, where maize and wheat litters determine the establishment of different bacterial, fungal, and nematode communities during decomposition (Banerjee et al. 2016; Sauvadet et al. 2016). However, we ignore if domesticated and wild genotypes of crops influence those processes differently. Understanding the capabilities of crop residues to influence the underneath soil communities through decomposition can help to manage nutrient mineralization and disease suppression, and thereby harness plant growth, pest resistance and the stability of plant production (Angulo et al. 2022; Compant et al. 2019; Liu et al. 2022).
In this study, we did not have a directional hypothesis. We compared the decomposability of litters of ten crops and their wild progenitors, and tested if domestication-induced changes in litter traits shape the soil bacterial, fungal, and nematode communities in the soil beneath the litter layer. We addressed the following questions: (1) Did plant domestication change litter traits and litter mass loss, and if so, are those changes consistent across crop species? (2) Do soils underneath domesticated leaf litters develop different bacterial, fungal, and nematode communities than those incubated with leaf litters from their wild progenitors?
Materials and methods
We carried out a leaf litter decomposition assay in a set of pairs of crops and wild progenitors. After incubating the leaf litters for 88 days we measured litter mass loss, nematode richness and number of individuals per taxa (ten crops), and OTU counts of bacteria and fungi (six crops). We tested the influence of plant domestication and species identities by using fixed-effects models of analysis of variance (ANOVA; for decomposability and richness), permutational multivariate analysis of variance (PERMANOVA; for litter quality and community composition), and analysed which litter traits explained better crop decomposability with a stepwise linear multiple regression. Finally, we visualized the magnitude and direction of domestication effects on soil microbes and nematodes community composition as the distance between the centroids of the samples grouped by species and domestication status in NMDS plots.
Study system and gathering of leaf litters
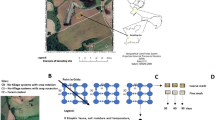
We selected a set of ten herbaceous crops of six plant families including grasses and forbs (red amaranth, borage, cabbage, millet, artichoke, sunflower, lettuce, tobacco, sorghum, and corn), and gathered seeds of a domesticated and a wild progenitor accession for each crop (Table 1, see also Supporting Information, Table S1, for wild progenitor assignment and seed donors). To obtain leaf litter from each pair of domesticated and wild progenitor, we grew 10–20 individuals of each accession in 2010 at the plant growth facilities of the Universidad Rey Juan Carlos, located in Móstoles, central Spain (40º18′48′′ N, 38º52′57′′ W, 632 m.a.s.l.; MAT: 15ºC; MAP: 450 mm). After seedlings emerged in the greenhouse, we transplanted plants outdoors into planting beds with a 30 centimetres depth layer of topsoil (soil pH = 7.6, measured in water; total N = 0.37%; organic C = 5.12%). We grew the plants until senescence, which took a slightly different time for each accession. We collected three samples of naturally senesced fresh leaf litter from three different plant individuals per accession. We discarded leaf litter with signs of herbivory or disease; we air-dried the remaining material for one month, and stored it at room temperature until we performed the decomposition assay.
Measurement of leaf litter traits
Prior to plant litter chemical analyses, we grounded the air-dried litter samples in a mill (IKA MF10; IKA-Werke, Staufen, Denmark) to pass a 1-mm screen. We measured C and N in an elemental analyser (varioMAx N/CN; Elementar, Hanau, Germany). We measured leaf litter fibres (cellulose, hemicellulose, and lignin) by the Van Soest method (Van Soest et al. 1991). We measured ash content through pyrolysis at 550ºC. To bring the ash into solution we dissolved it in aqua regia. Then, we evaluated P using vanadomolybdic colourimetry (Fiske and Subbarow 1925). We measured K and Ca using complexometric titration. We corrected for moisture, and expressed all litter chemistry variables as % of dry weight. We measured leaf litter dry matter content by dividing the mass of oven dried leaf litter at 60ºC by its water-saturated fresh mass (LDMC, g dry mass * g− 1 mass at full hydration; Pérez-Harguindeguy et al. 2016). We measured leaf toughness on air dried litter using a purpose-built penetrometer by breaking the leaf litter lamina (N * mm− 1; Pérez-Harguindeguy et al. 2016) .
Leaf litter decomposition assay and soil respiration
In September 2015, we bulked together the three litter samples of each domesticated and wild progenitor accession to set up microcosms for each of the ten crops. We built five microcosms for each accession and five control microcosms with no litter, totalling 105 microcosms. To focus on how different litters decompose and influence soil microbial and nematode communities, we incubated all litters on the same soil which we collected at 0–10 cm depth in a nearby permanent Mediterranean grassland (soil pH 7.5, measured in water; organic C 2.04%; total N 0.07%; coordinates 30T 0424133 / 4,469,923 N). We selected this soil to avoid the existence of home-field advantages in the decomposition of our litters. We sieved the soil at 2 mm, homogenised it, and stored it at 4 ºC for two weeks while setting up the microcosms. Although soil sieving might damage some soil nematodes, they are much smaller than 2 mm in diameter and sieving enabled appropriate homogenisation of the soil.
To set-up the microcosms we weighted 60 g of sieved fresh soil and introduced it into 250 mL plastic Mason jars (9 cm high, 6 cm diameter), with moisture adjusted to 60% water-holding capacity, which is favourable to microbial activity. We cut the litter into 2–3 cm long fragments. Before placing the litter in the microcosms, we placed 0.75 g of litter into Petri dishes and covered these with a soil inoculum for 24 h to promote the colonization of litter by soil microorganisms. This inoculum consisted of sieved fresh soil from the grassland mixed with distilled water (10 kg of soil to 75 L of water proportion; García-Palacios et al. 2013). To simulate a natural soil layer, we placed the soil inoculum-drenched leaf litter from the Petri dishes (0.75 g per microcosm) on top of the soil surface. We closed the microcosms with parafilm, then placed them in five trays randomly (comprising one experimental block); we included one “no-litter” control microcosm per tray to track changes in soil microbial and nematode communities during soil incubation independently of litter. We set the trays in a growth chamber (J.P. Selecta 4,000,699) over 88 days under optimal conditions for the decomposition process (darkness, 20ºC and 95% air humidity). During the incubation period, we randomised the location of trays every two weeks to minimize the effects of potential temperature and moisture gradients within the chamber. We corrected soil moisture every two weeks. We could collect all litter material remaining after the incubation period in each microcosm, since our microcosms lacked litter-fragmenting fauna. We dried it at 60 °C for 48 h, and then we weighed it to calculate litter mass loss (%). To analyse the soil microbial biomass, we followed a substrate-induced respiration (SIR) method using D-glucose and a MicroResp™ device (Campbell et al. 2003).
Soil microbial and nematode communities
After litter harvest, we collected soil samples to investigate the soil microbial and nematode community responses to the different litters. We analysed soil bacterial and fungal communities in six out of the ten crop-wild progenitor pairs (red amaranth, artichoke, lettuce, sunflower, sorghum and corn). For these analyses, we randomly chose three of the five replicate microcosms per accession of each of the six domesticated crop-wild progenitor pairs, and extracted soil DNA using the DNeasy PowerSoil Kit (QIAGEN GmbH). After extraction, we stored the DNA samples at − 20 ºC and sequenced them at the Illumina MiSeq platform of the Next Generation Genome Sequencing Facility of Wageningen University using the 341 F/805R (bacterial 16 S-rDNA; Herlemann et al. 2011) and FITS7/ITS4 (fungal Internal Transcribed Spacer, ITS, Ihrmark et al. 2012) primer sets.
To obtain an annotated OTU table from the raw MiSeq paired-end reads we performed the following steps. First, we merged the raw reads with a minimum overlap of 25 bp and at least a PHRED score of 25 using the RDP extension to PANDASeq (Masella et al. 2012) named Assembler (Cole et al. 2014). This ensures a base call accuracy of 99.5%. We used Flexbar version 2.5 (Dodt et al. 2012) to remove the primer sequences from the FASTQ files, then we converted the sequences to FASTA format and concatenated them into a single file. We used VSEARCH version 1.0.10 (Rognes et al. 2016) to cluster the sequences into OTUs, using the UPARSE strategy of de-replication, sorting by abundance (with at least two sequences) and clustering using the UCLUST smallmem algorithm (Edgar 2010). Hereafter, we detected chimeric sequences using the UCHIME algorithm (Edgar et al. 2011) implemented in VSEARCH, and we removed them. Finally, we obtained the taxonomic classification for each OTU by using the RDP Classifier version 2.10 (Cole et al. 2014). We implemented all steps in a workflow made with Snakemake (Köster and Rahmann 2012).
We used the rest of the fresh soil (50 g) to extract nematodes in Baermann funnels for 72 h (Baermann 1917). We counted all nematodes under a dissecting microscope and we identified at least 100 individuals to the genus or family level. We measured the water gravimetric soil content by drying soil subsamples at 105ºC for 24 h, and we expressed nematode abundances as number of individuals per 100 g of dry soil. We assigned nematode taxa to trophic groups: bacterivores, fungivores, herbivores, omnivores, and predators (Yeates et al. 1993).
Data analyses
To test for the effects of crop species identity, domestication status, and their interaction on litter mass loss (%) we used a two-way ANOVA model with fixed-effects (lm and anova of package Base; R Development Core team 2021). To test which crop species were different from each other, we ran post hoc analyses by using the model’s adjusted marginal means (emmeans and contrast of package emmeans; Lenth et al. 2022). We corrected P-values for multiple testing using the false discovery rate method (FDR; Benjamini and Hochberg 1995).
To explore how crop species identities and status of domestication influenced litter traits, we used principal components analysis (PCA). Some crops produced little leaf litter, so some trait values were missing. We applied a multiple imputation approach to gain statistical power (Nakagawa and Freckleton 2008). First, we removed two out of 60 replicates because all trait values were missing. We imputed missing values for individual traits (0.79%) by using predictive mean matching (mice of package mice, with parameters: m = 100, method = “pmm”, include = c(“sp”, “dom_status”), exclude = “id”; Buuren and Groothuis-Oudshoorn 2011), because this method preserves the structure of the data and has high accuracy when there is a low percentage of missing values (Goretzko et al. 2020). Then, we carried out a PCA to visualize how domesticated and wild progenitor accessions cluster according to litter quality variables (PCA of package FactoMineR; Lê et al. 2008). To test if crop species identities, domestication status, and their interaction affeted the litter traits that we used in the PCA, we performed a two-way PERMANOVA analysis (adonis function in package vegan, setting method = “euclidean”; Oksanen et al. 2020).
To explore whether and which litter quality parameters explained litter mass loss, we ran a multiple linear regression in which litter N, C, lignin, hemicellulose, cellulose, ash content, P, Ca, K, toughness, and LDMC were explanatory variables of litter mass loss rates (%) (lm of package Base; R Development Core team 2021). We tested the significance of explanatory variables (i.e., litter traits) using ANOVA (anova of Package base; R Development Core team 2021). Moreover, we calculated Pearson correlations of all traits with litter mass loss individually, and tested their significance by performing Pearson correlation tests (cor.test. of package Base;R Development Core team 2021). As we did not measure litter traits and mass loss data at the same level of observation (three replicates for trait analyses in the plant growing stage, and five replicates per bulked litter mass loss in the microcosm incubation stage), we calculated for each plant accession trait averages among the replicates. Then, we checked for collinearity between explanatory variables using the variance inflation factor (VIF). We used a linear model in which litter mass loss was the dependent variant, and all litter traits were independent variables. We removed litter ash content because it had a high VIF score (≥ 10) whenever we included litter C in the model (Vittinghoff et al. 2012). To identify the litter traits that accounted for most of the variation in litter mass loss across plant accessions, we selected the best-fitting linear models using Akaike’s information criterion (AIC) by running a model selection algorithm (dredge of package MuMIn; Bartoń 2022). We selected the model with the lowest AIC as the best model. Then, we used models within a 2 ΔAIC for model averaging and estimated the weighted coefficients, confidence intervals, and relative importance of each litter trait (model.avg and importance of package MuMIn; Bartoń 2022).
We measured alpha diversity of bacterial and fungal communities as rarefied OTU richness. We calculated rarefied diversity (rarefy of package vegan; Oksanen et al. 2020) after rarefaction to the median read count across all samples (Aguirre de Cárcer et al. 2011). The median of read counts was 7539 sequences for bacteria, and 31,610 sequences for fungi. We tested the effects of crop species identity, domestication status, and their interaction on the alpha diversity of soil bacterial and fungal communities (OTU richness; measured in six of the ten crops of the study system), and nematode community (taxa richness; measured in all ten crops of the study system) using two-way ANOVA models (lm and anova of package Base, (R Development Core team 2021)). We performed post hoc analyses based on the model’s adjusted marginal means (emmeans and contrast of package emmeans; Lenth et al. 2022), and we corrected P-values using FDR (Benjamini and Hochberg 1995). We also investigated the effects of crop species identity, domestication status, and their interaction on the beta diversity (i.e., community composition) of soil bacterial, fungal and nematode communities. We calculated beta diversity as the Bray-Curtis dissimilarity among samples in the matrices of square root weighted OTU (bacteria and fungi) or individuals per 100 g of dry soil (nematodes) data. To visualize how crop species identity and status of domestication of litters influenced beta diversity, we performed a non-metric multidimensional scaling (NMDS) by using the Bray-Curtis matrices (metaMDS function in package vegan; Oksanen et al. 2020). We tested the effect of crop species identity, domestication status, and their interaction on beta diversity (Bray-Curtis dissimilarity) by performing two-way PERMANOVAs with 10,000 permutations (adonis function of package vegan; Oksanen et al. 2020). PERMANOVAs can yield significant differences among groups through differences among centroids, variances or both (Anderson 2001). Thus, when necessary, we ran PERMDISP tests to test if centroids or variance were causing differences among groups in PERMANOVAs (betadisper function of package vegan; Oksanen et al. 2020).
To investigate how microbial and nematode richness correlated to community composition, and how the nematode community correlated to soil basal respiration, microbial biomass (SIR), and mass loss, we performed pairwise Mantel tests with 10,000 permutations using Bray-Curtis matrices for community composition data and Euclidean matrices for quantitative data (mantel.rtest function of package ade4; Dray and Dufour 2007). We carried out all statistical analyses and visualization in R 4.1.1 (R Development Core team 2021). Significance level used for all analyses was 0.05. We made all the figures using the ggplot2 package (Wickham 2016) and Paint.NET 4.3.11.
Results
Plant domestication effects on litter mass loss
Domesticated litters decomposed faster than their wild counterparts (Table 2, Domestication status, F1, 95 = 15.69, P = 0.002). Mass loss rates were 10% higher on average in domesticated litters (ranging from 0.4 to 30%) (Fig. 1). This was consistent across crops, except for Sorghum and Cenchrus, which decomposed at similar rates regardless of their status of domestication (Table S2, Fig. 1). Moreover, litters from different crop species decomposed at different rates, independently of their domestication status (Table 2, Crop species, F9, 95 = 50.74, P < 0.001). Brassica, Lactuca, and Nicotiana litters decomposed the fastest, and those of Sorghum and Cenchrus the slowest (Fig. 1).
Plant domestication effects on litter quality traits
Two axes of our ordination analysis explained 61% of the variance we observed in leaf litter traits. The analysis separated the crop species in two groups, grasses and non-grasses (Fig. 2). Grasses had higher leaf toughness, litter cellulose and hemicellulose, and lower litter N, Ca and K contents than non-grasses. Domestication had a consistent effect on most leaf litter traits (Table S3, Domestication status, F9, 95 = 12.90, P < 0.001), resulting in more decomposable leaf litter. In general, litter of domesticated crops had higher hemicellulose, ash content, P, and K, but had lower C, lignin, toughness, and LDMC (Table S4, Fig. S1). However, the magnitude of the effects of domestication on litter traits differed among crop species (Table S3, Crop species * Domestication status interaction, F9, 95 = 5.92, P < 0.001). The effect of domestication on N, Ca, and cellulose was idiosyncratic among crop species (Table S4, Fig. S1).
Principal component analysis (PCA) of leaf litter traits differences between crop species identities and their domestication status. Leaf litter samples are plotted over the two PCA axes with the highest variance explained (a). Loads of the leaf litter traits inputted for the PCA ordination plot are plotted (b). Domestication-induced changes in all leaf litter traits appear in Table S4 and Fig. S1
Litter quality as a driver of litter mass loss
Litter traits explained up to 84% of the variance observed in litter mass loss. Higher litter C, N, lignin, cellulose, Ca, K, and toughness associated with increased decomposition rate, while higher levels of hemicellulose, P, and LDMC related negatively to litter decomposition rate (Table S5). LDMC alone explained up to 70% of the variance in litter mass loss (Table S6, β = -140.84). The best-fitting model included LDMC and litter C as predictors, and explained 74% of the variance in litter mass loss (Table S6, β = -143.78, β = 1.10). LDMC was the most important predictor for litter decomposability with slower decomposition rate the higher the LDMC (Table S7, P < 0.001).
Domestication and species identity effects on the richness of soil microbial and nematode communities
The domestication status of the litters left to decompose on the soil surface of the microcosms affected the soil bacterial OTU richness, but the magnitude and direction of this effect differed among the crop species identities of the litter (Table 3, Crop species * Domestication status interaction, F5, 95 = 7.05, P < 0.001, Fig. 3). Soils incubated with litter of domesticated Sorghum had higher OTU richness than those receiving litter of its wild progenitor (Table S8, P < 0.001, Fig. 3). In contrast, soils incubated with litter of domesticated Helianthus had lower OTU richness than those with litter of its wild progenitor (Table S8, P < 0.008, Fig. 3). Soils incubated with litters of Amaranthus, Cynara, Lactuca, and Zea exhibited a similar OTU richness regardless of their domestication status (Table S8, Fig. 3). Fungal community OTU richness increased in soils incubated with litters of domesticated accessions (Table 3, Domestication status, F1, 95 = 4.44, P = 0.048, Fig. 3). This effect was consistent across crop species (Table 3, Crop species * Domestication status interaction F5, 95 = 0.59, P = 0.707). In fact, crop species identity of the litters did not change fungi OTU richness (Table 3, Crop species, F5, 95 = 2.42, P = 0.072). Nematode genera richness varied between microcosms incubated with litters of different crops (Table 3, Crop species identity, F9, 95 = 10.30, P < 0.001): soils incubated with litters of Lactuca and Nicotiana had the lowest, while soils incubated with Sorghum litter had the highest nematode richness (Fig. 3). Nematode richness did not differ between microcosm with domesticated or wild litters (Table 3, Domestication status, F1, 95 = 0.71, P = 0.402; Crop species * Domestication status interaction F9, 95 = 0.85, P = 0.570). In general, mean abundances of each nematode feeding group were similar between microcosms with litters of domesticated and wild accessions (Fig. S2). Bacterivores were the most abundant feeding type of nematodes in all soils.
Plant domestication and species identity effects on the soil microbial and nematode community composition
Soils incubated with litters of the same crop species had similar bacterial and fungal communities (Table 4, Crop species, F5, 95 = 1.76, P = 0.003; F5, 95 = 1.53, P = 0.001, respectively, Fig. 4). Within each crop, the status of domestication of the litters shaped the composition of bacterial and fungal communities in different directions (Table 4, Crop species * Domestication status interaction, F5, 95 = 1.67, P = 0.008; F5, 95 = 1.25, P = 0.030, respectively, Fig. 4). These changes may arise from differences in the abundances of the most abundant groups. For bacteria, these groups were Acidobacteria, Actinobacteria, Bacteroidetes, Proteobacteria, Gemmatimonadetes and Verrucomicrobia (Fig. S3). Regarding fungal communities, the most abundant groups were Ascomycota, Basidiomycota and Zygomycota (Fig. S4). The strength of domestication-induced shifts in soil bacterial and fungal community compositions was different between crops (Table S10). Soils fed with litters of the same crop species developed similar nematode communities (Table 4, Crop species, F9, 95 = 2.46, P < 0.001, Fig. 4). Soils with litters of Nicotiana, Lactuca, Brassica, Cenchrus, Zea and Cynara associated to higher abundances of generalist bacterivores (Prismatolaimus and Rhabditis); soils incubated with litters of Amaranthus, Borago, Sorghum and Helianthus associated to increased abundances of generalist bacterivores (Eumonhystera, Teratocephalus and Mesorhabditis), and omnivores/predators (Clarkus and Thornia); microcosms with Cynara, Sorghum and Healianthus litters also presented higher abundances of plant parasitic nematodes (Tylenchorhynchus and Helicotylenchus) (Figs. 4 and 5). Domestication had a consistent effect across crop species (Table 4, Domestication status, F1, 95 = 2.05, P = 0.047), with a similar effect size among crops (Table S10). In general, domestication increased the abundance of bacterial feeders in the community (Figs. 4 and 5).
Non-metric multidimensional scaling (NMDS) analyses of community composition based on the Bray-Curtis dissimilarity matrices of the weighted relative abundances of soil bacterial (a = zoom-in version; all points are displayed in Fig. S5), fungal (b = zoom-in version; all points are displayed in Fig. S6), and nematodes communities (c). The direction and magnitude of domestication-induced changes in community composition of bacteria (d), fungi (e), and nematodes (f) are represented by arrows that cover the distance between centroids of the points belonging to the wild progenitors and the domesticated crops for each of the plant species
Loadings of the most influential nematode taxa in the NMDS analysis displayed in Fig. 4
Bacterial richness correlated positively with bacterial community composition (r = 0.971, P < 0.001), but fungal richness did not correlate with fungal community composition (r = 0.203, P = 0.140). Nematode richness correlated positively with nematode community composition (r = 0.253, P < 0.001). Nematode richness correlated positively to litter mass loss (r = 0.253, P < 0.001), but did not correlate to any soil respiration measure or microbial biomass (Table S11). Nematode community composition correlated to litter mass loss (r = 0.240, P < 0.001) and soil basal respiration (Table S11, r = 0.067, P = 0.042), but did not correlate to microbial biomass (Table S11, r = 0.011, P = 0.400).
Discussion
Our results showed that leaf litter of domesticated crops decomposes faster than litter of their wild progenitors. This effect was mediated by domestication-induced changes in leaf litters, which were softer, and had less C concentrations and LDMC than their wild counterparts. The faster litter decomposition in the domesticated microcosms impacted the soil bacterial, fungal and nematode communities in different ways. Our results show that domestication-induced changes in crop traits altered the linkages between plant residues and soil organisms in different ways for bacteria, fungi and nematodes.
Domestication impacts on leaf litter decomposability
The litter of domesticated plants decomposed faster than those of their wild progenitors among all crop species. Our results suggest that this is related to an increment in leaf litter quality after domestication, such as higher P, and lower C, lignin, and LDMC. García-Palacios et al. (2013) and Delgado-Baquerizo et al., (2016) also found that domesticated accessions of herbaceous crops decomposed faster than their wild counterparts, and that these increments correlated with changes in C, N, P, and lignin contents. In another study, leaf litter of domesticated accessions of the perennial herb Silphium integrifolium Michx. (Asteraceae) also exhibited higher decomposability when compared to wild accessions, which correlated with lower LDMC in domesticated varieties (González-Paleo et al. 2022). This is similar to previous findings from natural ecosystems, where chemical (i.e., C, N, and lignin contents) and physical (i.e., LDMC) traits explain most of the variation in litter mass loss (Cornwell et al. 2008; Dias et al. 2017; Freschet et al. 2012; García-Palacios et al. 2016a). Therefore, our findings align well with established hypothesis that an increase in litter quality, induced in our study system by plant domestication, speeds up decomposition.
Effects of domestication on soil microbes
Soil fungal community richness increased consistently in soils incubated with domesticated litters, as compared to soils incubated with litters of wild progenitors. This effect was inconsistent (i.e., increasing for some crops and decreasing for others) in bacterial richness. In contrast, previous research reported that bacterial and fungal richness increase or decrease similarly during decomposition (Purahong et al. 2016). A possible explanation for this mismatch would be that our litters lean towards the highly decomposable end of the global spectrum (Zhang et al. 2008), promoting the proliferation of bacteria over fungi. An increment in resource accessibility could downregulate common species of fungi that dominate in the early stages of decomposition, that usually digest not so accessible carbon sources such as cellulose or hemicellulose. This would make room for a wider variety of fungi species, since the resources in our labile litters might have been easily accessible by most fungal species through the use of generalist enzymes like laccase or β-glucosidase (Boer et al. 2005; Eichlerová et al. 2015).
Differences in bacterial and fungal community compositions with domestication were crop species-specific. This finding agrees with previous work, which shows that plant identity (here crop identity) has a strong influence on the composition of leaf litter and soil bacterial and fungal communities (Fanin et al. 2014; Veen et al. 2021). Previous studies have attributed these changes in bacterial and fungal community composition to differences in litter quality (Alfaro et al. 2017; García-Palacios et al. 2016b; Purahong et al. 2016; Rummel et al. 2020). Specifically, bacterial community composition would depend of N, Ca, and cellulose content (Purahong et al. 2016; Rummel et al. 2020), and fungal community composition would depend of N, Ca, and lignin (Alfaro et al. 2017; Purahong et al. 2016). Our results partly support these claims, since we found that domestication had a crop species-specific effect on N, Ca, and cellulose contents (increased for some crops and decreased for others). Therefore, N, Ca, and cellulose might be the drivers of the variations in bacterial and fungal community compositions in our soils because all of them are affected by domestication in a crop species-specific manner. Bacterial richness and community composition were strongly correlated and reacted similarly to domesticated crop litter inputs. However, fungal richness and community composition reacted differently to litter addition of domesticated crops. This is reinforced by the lack of correlation between fungal richness and community composition. Our results contrast with previous studies that suggested that both richness and community composition of bacteria and fungi react similarly to plant litter inputs (Sauvadet et al. 2016). Our findings suggest that domestication caused idiosyncratic shifts in litter quality among crop species, which might have altered the existing dynamics within the microbial communities. This would explain that domestication-induced changes in soil communities are not coordinated between bacteria and fungi in our study system, suggesting that these groups responded differently to changes in litter quality.
Impacts of domestication on soil nematodes
Bacterial feeding nematodes remained the most abundant trophic group among domestication statuses and crop species identities. Nematode community composition mainly changed with the identities of the litters that we added to the soils. Our findings agree with previous studies which reported that bacterivore nematodes are the most abundant trophic group in soils, especially around decomposing plant litter (van den Hoogen et al. 2019; Wasilewska 1991). Even though the nematode community shows specificity to litter types, nematodes do not interact directly with plant residues (Yeates 1999). Rather, nematodes interact with microbes through predation (Jiang et al. 2017). Hence, nematode richness and community composition would be primarily determined by plant litter effects on bacterial and fungal communities (Yeates 1999). Although organic matter decomposition might liberate organic substances with nematicidal activities (i.e., glucosinolate-derived compounds in Brassicas, short-chain fatty acids under anaerobic conditions, or N compounds generated from low C:N organic materials (Oka 2010)), we did not detect clear detrimental effects of decomposition on nematode abundances. For instance, the abundance of nematodes correlates with microbial biomass, whilst both correlate to litter quality and decomposability (Sauvadet et al. 2016). Our data hints that nematode community composition might be related to microbial biomass because we found that plant domestication accelerated decomposition by increasing litter quality. Indeed, nematode community composition correlated positively with litter mass loss. An accelerated resource input into soils would augment microbial biomass. We expect bacteria to be very abundant in this stage of decomposition. Moreover, nematode community composition correlated weakly with soil basal respiration, but did not correlate with microbial biomass. Bacterivore nematodes feed on bacterial cell aggregates (Rønn et al. 2012), so they probably ingest bacteria independently if they are dead or alive. In accordance to our claim, litter addition of domesticated crops impacted soil nematode community composition by increasing the abundance of bacterial feeders across all soils. Adding litter of domesticated accessions of labile species as Brassica, Lactuca, and Nicotiana increased the abundance of opportunistic bacterivore nematodes (Rhabditis, Panagrolaimus), which indicate that soil bacterial biomass increases and decreases quickly over time (Ferris and Bongers 2006). However, domesticated accessions of more recalcitrant litters as Borago, Helianthus, and Shorghum promoted generalist bacterivores (Prismatolaimus, Eumonhystera, Teratocephalus), which are commonly associated with more stable bacterial populations (Ferris and Bongers 2006), and omnivore-predators, which might increase their populations due to the abundance of nematodes of lower trophic levels (Ferris et al. 2012). Litter from domesticated accessions of Cynara and Sorghum seemed to be associated with herbivore nematodes. Although no plants grew in the microcosms during our experiment, it is known that herbivore nematodes might survive in the soil for long periods without plant roots (Ribeiro et al. 2020); the reason behind such association to herbivore nematodes remains unclear and deserves further attention. Our findings suggest that nematode community composition in soils fed with the litters of domesticated plants are different than that of soils fed with litters of their wild progenitors. This disruption might mirror the shifts in the bacterial community observed, which are related to domestication-induced changes in litter quality, which would therefore explain why higher abundance of bacterivore nematodes was found across all crop species identities, while the bacterivore taxa groups with higher abundances were crop species-specific.
Remarks on our experimental design and future studies
The novelty of our experiment lies in analysing the effects of domestication on bacterial, fungal and nematode communities in the same soil samples at the same time. Additionally, we investigated this alongside domestication-induced changes in litter quality and decomposability. Since we pooled leaf litter, we produced pseudo-replicates. Therefore, we needed fewer replicates in the experiment but could not analyse trait variability within accessions. As we used leaf litter only, our results might not extrapolate to other types of plant residues (roots and stems). In particular, roots contribute a high share of cropland residues, and dead roots behave differently from leaf litter during decomposition (Hobbie et al. 2010; Sauvadet et al. 2016). A previous study hinted that traits that control leaf litter decomposability in crops might differ from those that control root litter decomposition (Barel et al. 2019). Also, we surveyed soil communities once for the full duration of our experiment. But soil biotas undergo considerable temporal changes throughout the decomposition process (García-Palacios et al. 2016b; Purahong et al. 2016; Veen et al. 2021; Wang et al. 2004; Wardle et al. 2006). For example, Ascomycota fungi dominate at the beginning stages of litter decay, while Basidiomycota become more abundant later on (Purahong et al. 2016). Future studies should address if and how the effects of domestication on soil biotas, mediated through litter decomposition, vary in the different stages of the process. Temporal variations should also be addressed for other types of plant residues (i.e., stems and roots) because their relationship to microbial and nematode communities remains unknown. We did not find works about the effect of litter leachates on soil nematodes during decomposition. This needs further investigation, since nematodes are important indicators of soil health (Gao et al. 2020). Also, if and how domestication altered litter traits in stem and root residues deserve further exploration since they contribute a high share in litter inputs to agricultural soils.
Conclusion
In this study, we report the impacts of domestication-induced changes in litter traits on soil microbial and nematode communities during the decomposition of plant residues. Domesticated litters generally increased fungal richness while their effects were crop species-specific for bacterial richness (i.e., increasing for some crops and decreasing for others). Bacterial and fungal community composition chiefly depended on the taxonomic identities of the litters added to the soils, and the influence of the status of domestication of the added litters was crop species-specific. This means that the effect of incubating soils with litters of domesticated accessions was quite variable and idiosyncratic among crop species. Incubating soils with litters of domesticated crops increased the abundances of bacterivore nematodes when compared to litters of wild progenitors, but the bacterivore nematode taxa which thrived depended of the plant litter identity. Overall, our results indicate that plant domestication altered the linkages between crop residues and soil organisms in different ways for bacteria, fungi and nematodes.
Data availability
All data generated or analysed during this study are included in this published article as a supplementary information file.
References
Alfaro FD, Manzano M, Marquet PA, Gaxiola A (2017) Microbial communities in soil chronosequences with distinct parent material: the effect of soil pH and litter quality. J Ecol 105:1709–1722. https://doi.org/10.1111/1365-2745.12766
Anderson MJ (2001) A new method for non-parametric multivariate analysis of variance. Aust Ecol 26:32–46. https://doi.org/10.1111/j.1442-9993.2001.01070.pp.x
Angulo V, Beriot N, Garcia-Hernandez E, Li E, Masteling R, Lau JA (2022) Plant–microbe eco-evolutionary dynamics in a changing world. New Phytol. https://doi.org/10.1111/nph.18015
Baermann G (1917) A simple method for the detection of Ankylostomum (nematode) larvae in soil tests. Simple Method Detect. Ankylostomum Nematode Larvae Soil Tests 41–47
Banerjee S, Kirkby CA, Schmutter D, Bissett A, Kirkegaard JA, Richardson AE (2016) Network analysis reveals functional redundancy and keystone taxa amongst bacterial and fungal communities during organic matter decomposition in an arable soil. Soil Biol Biochem 97:188–198. https://doi.org/10.1016/j.soilbio.2016.03.017
Barel JM, Kuyper TW, de Boer W, De Deyn GB (2019) Plant presence reduces root and shoot litter decomposition rates of crops and wild relatives. Plant Soil 438:313–327. https://doi.org/10.1007/s11104-019-03981-7
Bartoń K (2022) MuMIn: Multi-Model Inference. https://CRAN.R-project.org/package=MuMIn
Benjamini Y, Hochberg Y (1995) Controlling the false Discovery rate: a practical and powerful Approach to multiple testing. J R Stat Soc Ser B Methodol 57:289–300. https://doi.org/10.1111/j.2517-6161.1995.tb02031.x
Campbell CD, Chapman SJ, Cameron CM, Davidson MS, Potts JM (2003) A rapid microtiter plate method to measure carbon dioxide evolved from carbon substrate amendments so as to determine the physiological profiles of soil microbial communities by using whole soil. Appl Environ Microbiol 69:3593–3599. https://doi.org/10.1128/AEM.69.6.3593-3599.2003
Cole JR, Wang Q, Fish JA, Chai B, McGarrell DM, Sun Y, Brown CT, Porras-Alfaro A, Kuske CR, Tiedje JM (2014) Ribosomal database project: data and tools for high throughput rRNA analysis. Nucleic Acids Res 42:D633–D642. https://doi.org/10.1093/nar/gkt1244
Compant S, Samad A, Faist H, Sessitsch A (2019) A review on the plant microbiome: Ecology, functions, and emerging trends in microbial application. J Adv Res 19:29–37. Special Issue on Plant Microbiome. https://doi.org/10.1016/j.jare.2019.03.004
Cornwell WK, Cornelissen JHC, Amatangelo K, Dorrepaal E, Eviner VT, Godoy O, Hobbie SE, Hoorens B, Kurokawa H, Pérez-Harguindeguy N, Quested HM, Santiago LS, Wardle DA, Wright IJ, Aerts R, Allison SD, Van Bodegom P, Brovkin V, Chatain A, Callaghan TV, Díaz S, Garnier E, Gurvich DE, Kazakou E, Klein JA, Read J, Reich PB, Soudzilovskaia NA, Vaieretti MV, Westoby M (2008) Plant species traits are the predominant control on litter decomposition rates within biomes worldwide. Ecol Lett 11:1065–1071. https://doi.org/10.1111/j.1461-0248.2008.01219.x
Cotrufo MF, Galdo ID, Piermatteo D (2010) Litter decomposition: concepts, methods and future perspectives. In: Heinemeyer A, Bahn M, Kutsch WL (eds) Soil carbon dynamics: an integrated methodology. Cambridge University Press, Cambridge, pp 76–90
de Aguirre D, Denman SE, McSweeney C, Morrison M (2011) Evaluation of subsampling-based normalization strategies for tagged high-throughput sequencing data sets from gut microbiomes. Appl Environ Microbiol 77:8795–8798. https://doi.org/10.1128/AEM.05491-11
de Boer W, Folman LB, Summerbell RC, Boddy L (2005) Living in a fungal world: impact of fungi on soil bacterial niche development⋆. FEMS Microbiol Rev 29:795–811. https://doi.org/10.1016/j.femsre.2004.11.005
Delgado-Baquerizo M, Reich PB, García-Palacios P, Milla R (2016) Biogeographic bases for a shift in crop C: N: P stoichiometries during domestication. Ecol Lett 19:564–575. https://doi.org/10.1111/ele.12593
Dias ATC, Cornelissen JHC, Berg MP (2017) Litter for life: assessing the multifunctional legacy of plant traits. J Ecol 105:1163–1168. https://doi.org/10.1111/1365-2745.12763
Dodt M, Roehr JT, Ahmed R, Dieterich C (2012) FLEXBAR—Flexible Barcode and Adapter Processing for Next-Generation sequencing platforms. Biology 1:895–905. https://doi.org/10.3390/biology1030895
Doebley JF, Gaut BS, Smith BD (2006) The molecular genetics of crop domestication. Cell 127:1309–1321. https://doi.org/10.1016/j.cell.2006.12.006
Dray S, Dufour A-B (2007) The ade4 package: implementing the duality diagram for ecologists. J Stat Softw 22:1–20. https://doi.org/10.18637/jss.v022.i04
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. https://doi.org/10.1093/bioinformatics/btr381
Eichlerová I, Homolka L, Žifčáková L, Lisá L, Dobiášová P, Baldrian P (2015) Enzymatic systems involved in decomposition reflects the ecology and taxonomy of saprotrophic fungi. Fungal Ecol 13:10–22. https://doi.org/10.1016/j.funeco.2014.08.002
Fanin N, Hättenschwiler S, Fromin N (2014) Litter fingerprint on microbial biomass, activity, and community structure in the underlying soil. Plant Soil 379:79–91. https://doi.org/10.1007/s11104-014-2051-7
Ferris H, Bongers T (2006) Nematode indicators of organic enrichment. J Nematol 38:3–12
Ferris H, Sánchez-Moreno S, Brennan EB (2012) Structure, functions and interguild relationships of the soil nematode assemblage in organic vegetable production. Appl Soil Ecol 61:16–25. Microorganisms and the Sustainable Management of Soil. https://doi.org/10.1016/j.apsoil.2012.04.006
Fiske CH, Subbarow Y (1925) The colorimetric determination of phosphorus. J Biol Chem 66:375–400. https://doi.org/10.1016/S0021-9258(18)84756-1
Freschet GT, Aerts R, Cornelissen JHC (2012) A plant economics spectrum of litter decomposability. Funct Ecol 26:56–65. https://doi.org/10.1111/j.1365-2435.2011.01913.x
Gao D, Wang F, Li J, Yu S, Li Z, Zhao J (2020) Soil nematode communities as indicators of soil health in different land use types in tropical area. Nematology 22:595–610. https://doi.org/10.1163/15685411-00003325
García-Palacios P, McKie BG, Handa IT, Frainer A, Hättenschwiler S (2016a) The importance of litter traits and decomposers for litter decomposition: a comparison of aquatic and terrestrial ecosystems within and across biomes. Funct Ecol 30:819–829. https://doi.org/10.1111/1365-2435.12589
García-Palacios P, Shaw EA, Wall DH, Hättenschwiler S (2016b) Temporal dynamics of biotic and abiotic drivers of litter decomposition. Ecol Lett 19:554–563. https://doi.org/10.1111/ele.12590
García-Palacios P, Milla R, Delgado-Baquerizo M, Martín-Robles N, Álvaro-Sánchez M, Wall DH (2013) Side-effects of plant domestication: ecosystem impacts of changes in litter quality. New Phytol 198:504–513. https://doi.org/10.1111/nph.12127
González-Paleo L, Ravetta D, Van Tassel D (2022) From leaf traits to agroecosystem functioning: effects of changing resource use strategy during silphium domestication on litter quality and decomposition rate. Plant Soil 471:655–667. https://doi.org/10.1007/s11104-021-05224-0
Goretzko D, Heumann C, Bühner M (2020) Investigating parallel analysis in the context of missing data: a simulation study comparing six missing data methods. Educ Psychol Meas 80:756–774. https://doi.org/10.1177/0013164419893413
Hancock JF (ed) (2012) Plant evolution and the origin of crop species, 3rd edn. CABI, Wallingford
Herlemann DP, Labrenz M, Jürgens K, Bertilsson S, Waniek JJ, Andersson AF (2011) Transitions in bacterial communities along the 2000 km salinity gradient of the Baltic Sea. ISME J 5:1571–1579. https://doi.org/10.1038/ismej.2011.41
Hobbie SE, Oleksyn J, Eissenstat DM, Reich PB (2010) Fine root decomposition rates do not mirror those of leaf litter among temperate tree species. Oecologia 162:505–513. https://doi.org/10.1007/s00442-009-1479-6
Ihrmark K, Bödeker ITM, Cruz-Martinez K, Friberg H, Kubartova A, Schenck J, Strid Y, Stenlid J, Brandström-Durling M, Clemmensen KE, Lindahl BD (2012) New primers to amplify the fungal ITS2 region – evaluation by 454-sequencing of artificial and natural communities. FEMS Microbiol Ecol 82:666–677. https://doi.org/10.1111/j.1574-6941.2012.01437.x
Jiang Y, Liu M, Zhang J, Chen Y, Chen X, Chen L, Li H, Zhang X-X, Sun B (2017) Nematode grazing promotes bacterial community dynamics in soil at the aggregate level. ISME J 11:2705–2717. https://doi.org/10.1038/ismej.2017.120
Kazakou E, Violle C, Roumet C, Pintor C, Gimenez O, Garnier E (2009) Litter quality and decomposability of species from a Mediterranean succession depend on leaf traits but not on nitrogen supply. Ann Bot 104:1151–1161. https://doi.org/10.1093/aob/mcp202
Köster J, Rahmann S (2012) Snakemake—a scalable bioinformatics workflow engine. Bioinformatics 28:2520–2522. https://doi.org/10.1093/bioinformatics/bts480
Kraft NJB, Adler PB, Godoy O, James EC, Fuller S, Levine JM (2015) Community assembly, coexistence and the environmental filtering metaphor. Funct Ecol 29:592–599. https://doi.org/10.1111/1365-2435.12345
Lê S, Josse J, Husson F (2008) FactoMineR: an R package for multivariate analysis. J Stat Softw 25:1–18. https://doi.org/10.18637/jss.v025.i01
Lenth RV, Buerkner P, Herve M, Love J, Miguez F, Riebl H, Singmann H (2022) emmeans: stimated Marginal Means, aka Least-Squares Means. https://CRAN.R-project.org/package=emmeans
Liu S, García-Palacios P, Tedersoo L, Guirado E, van der Heijden MGA, Wagg C, Chen D, Wang Q, Wang J, Singh BK, Delgado-Baquerizo M (2022) Phylotype diversity within soil fungal functional groups drives ecosystem stability. Nat Ecol Evol 1–10. https://doi.org/10.1038/s41559-022-01756-5
Masella AP, Bartram AK, Truszkowski JM, Brown DG, Neufeld JD (2012) PANDAseq: paired-end assembler for illumina sequences. BMC Bioinformatics 13:31. https://doi.org/10.1186/1471-2105-13-31
Meyer RS, DuVal AE, Jensen HR (2012) Patterns and processes in crop domestication: an historical review and quantitative analysis of 203 global food crops. New Phytol 196:29–48. https://doi.org/10.1111/j.1469-8137.2012.04253.x
Milla R, Osborne CP, Turcotte MM, Violle C (2015) Plant domestication through an ecological lens. Trends Ecol Evol 30:463–469. https://doi.org/10.1016/j.tree.2015.06.006
Nakagawa S, Freckleton RP (2008) Missing inaction: the dangers of ignoring missing data. Trends Ecol Evol 23:592–596. https://doi.org/10.1016/j.tree.2008.06.014
Oka Y (2010) Mechanisms of nematode suppression by organic soil amendments—A review. Appl Soil Ecol 44:101–115. https://doi.org/10.1016/j.apsoil.2009.11.003
Oksanen J, Blanchet FG, Friendly M, Kindt R, Legendre P, McGlinn D, Minchin PR, O’Hara RB, Simpson GL, Solymos P, Stevens MHH, Szoecs E, Wagner H (2020) vegan: Community Ecology Package. https://CRAN.R-project.org/package=vegan
Pakeman RJ, Eastwood A, Scobie A (2011) Leaf dry matter content as a predictor of grassland litter decomposition: a test of the ‘mass ratio hypothesis’. Plant Soil 342:49–57. https://doi.org/10.1007/s11104-010-0664-z
Pérez Harguindeguy N, Cortez J, Garnier E, Gillon D, Poca M (2015) Predicting leaf litter decomposability: an exploratory assessment of leaf traits, litter traits and spectral properties in six Mediterranean herbaceous species. Ecol Austral 025:054–064
Pérez-Harguindeguy N, Díaz S, Garnier E, Lavorel S, Poorter H, Jaureguiberry P, Bret-Harte MS, Cornwell WK, Craine JM, Gurvich DE, Urcelay C, Veneklaas EJ, Reich PB, Poorter L, Wright IJ, Ray P, Enrico L, Pausas JG, de Vos AC, Buchmann N, Funes G, Quétier F, Hodgson JG, Thompson K, Morgan HD, Steege H, Sack L, Blonder B, Poschlod P, Vaieretti MV, Conti G, Staver AC, Aquino S, Cornelissen JHC (2016) Corrigendum to: New handbook for standardised measurement of plant functional traits worldwide. Aust J Bot 64:715–716. https://doi.org/10.1071/bt12225_co
Pichon NA, Cappelli SL, Soliveres S, Hölzel N, Klaus VH, Kleinebecker T, Allan E (2020) Decomposition disentangled: a test of the multiple mechanisms by which nitrogen enrichment alters litter decomposition. Funct Ecol 34:1485–1496. https://doi.org/10.1111/1365-2435.13560
Prescott CE, Grayston SJ (2013) Tree species influence on microbial communities in litter and soil: Current knowledge and research needs. For Ecol Manag 309:19–27. Influence of tree species on forest soils: New evidence from field studies. https://doi.org/10.1016/j.foreco.2013.02.034
Prieto I, Litrico I, Violle C, Barre P (2017) Five species, many genotypes, broad phenotypic diversity: when agronomy meets functional ecology. Am J Bot 104:62–71. https://doi.org/10.3732/ajb.1600354
Purahong W, Wubet T, Lentendu G, Schloter M, Pecyna MJ, Kapturska D, Hofrichter M, Krüger D, Buscot F (2016) Life in leaf litter: novel insights into community dynamics of bacteria and fungi during litter decomposition. Mol Ecol 25:4059–4074. https://doi.org/10.1111/mec.13739
Purugganan MD (2022) What is domestication? Trends Ecol Evol. https://doi.org/10.1016/j.tree.2022.04.006
Purugganan MD, Fuller DQ (2009) The nature of selection during plant domestication. Nature 457:843–848. https://doi.org/10.1038/nature07895
R Development Core team (2021) R: A language and environment for statistical computing. R Foundation for Statistical Computing. R Core Team, Vienna. https://www.R-project.org/
Rawlik K, Kasprowicz M, Nowiński M, Jagodziński AM (2022) The afterlife of herbaceous plant species: a litter decomposition experiment in a temperate oak-hornbeam forest. For Ecol Manag 507:120008. https://doi.org/10.1016/j.foreco.2022.120008
Ribeiro LM, Campos HD, Neves DL, Dias-Arieira CR (2020) Survival of Pratylenchus brachyurus under dry soil conditions. Heliyon 6:e05075. https://doi.org/10.1016/j.heliyon.2020.e05075
Robinson ML, Schilmiller AL, Wetzel WC (2022) A domestic plant differs from its wild relative along multiple axes of within-plant trait variability and diversity. Ecol Evol 12:e8545. https://doi.org/10.1002/ece3.8545
Rognes T, Flouri T, Nichols B, Quince C, Mahé F (2016) VSEARCH: a versatile open source tool for metagenomics. PeerJ 4:e2584. https://doi.org/10.7717/peerj.2584
Rønn R, Vestergård M, Ekelund F (2012) Interactions between bacteria, protozoa and nematodes in soil. Acta Protozool 2012:223–235. https://doi.org/10.4467/16890027AP.12.018.0764
Roucou A, Violle C, Fort F, Roumet P, Ecarnot M, Vile D (2018) Shifts in plant functional strategies over the course of wheat domestication. J Appl Ecol 55:25–37. https://doi.org/10.1111/1365-2664.13029
Rummel PS, Pfeiffer B, Pausch J, Well R, Schneider D, Dittert K (2020) Maize root and shoot litter quality controls short-term CO2 and N2O emissions and bacterial community structure of arable soil. Biogeosciences 17:1181–1198. https://doi.org/10.5194/bg-17-1181-2020
Sauvadet M, Chauvat M, Cluzeau D, Maron P-A, Villenave C, Bertrand I (2016) The dynamics of soil micro-food web structure and functions vary according to litter quality. Soil Biol Biochem 95:262–274. https://doi.org/10.1016/j.soilbio.2016.01.003
Strickland MS, Osburn E, Lauber C, Fierer N, Bradford MA (2009) Litter quality is in the eye of the beholder: initial decomposition rates as a function of inoculum characteristics. Funct Ecol 23:627–636. https://doi.org/10.1111/j.1365-2435.2008.01515.x
van Buuren S, Groothuis-Oudshoorn K (2011) Mice: multivariate imputation by chained equations in R. J Stat Softw 45:1–67. https://doi.org/10.18637/jss.v045.i03
van den Hoogen J, Geisen S, Routh D, Ferris H, Traunspurger W, Wardle DA, de Goede RGM, Adams BJ, Ahmad W, Andriuzzi WS, Bardgett RD, Bonkowski M, Campos-Herrera R, Cares JE, Caruso T, de Brito Caixeta L, Chen X, Costa SR, Creamer R, Mauro da Cunha Castro J, Dam M, Djigal D, Escuer M, Griffiths BS Gutiérrez C, Hohberg K, Kalinkina D, Kardol P, Kergunteuil A, Korthals G, Krashevska V, Kudrin AA, Li Q, Liang W, Magilton M, Marais M, Martín JAR, Matveeva E, Mayad EH, Mulder C, Mullin P, Neilson R, Nguyen TAD, Nielsen UN, Okada H, Rius JEP, Pan K, Peneva V, Pellissier L, Silva J, Pitteloud C, Powers TO, Powers K, Quist CW, Rasmann S, Moreno SS, Scheu S, Setälä H, Sushchuk A, Tiunov AV, Trap J, van der Putten W, Vestergård M, Villenave C, Waeyenberge L, Wall DH, Wilschut R, Wright DG, Yang J, Crowther TW (2019) Soil nematode abundance and functional group composition at a global scale. Nature 572:194–198. https://doi.org/10.1038/s41586-019-1418-6
Van Soest PJ, Robertson JB, Lewis BA (1991) Methods for dietary fiber, neutral detergent fiber, and nonstarch polysaccharides in relation to animal nutrition. J Dairy Sci 74:3583–3597. https://doi.org/10.3168/jds.S0022-0302(91)78551-2
Veen GF (Ciska), ten, Hooven FC, Weser C, Hannula SE (eds) (2021) Steering the soil microbiome by repeated litter addition. J Ecol 109:2499–2513. https://doi.org/10.1111/1365-2745.13662
Vittinghoff E, Glidden DV, Shiboski SC, McCulloch CE (2012) Regression methods in Biostatistics, Statistics for Biology and Health. Springer US, Boston. https://doi.org/10.1007/978-1-4614-1353-0
Wang K-H, McSorley R, Marshall AJ, Gallaher RN (2004) Nematode community changes associated with decomposition of Crotalaria juncea amendment in litterbags. Appl Soil Ecol 27:31–45. https://doi.org/10.1016/j.apsoil.2004.03.006
Wardle DA, Yeates GW, Barker GM, Bonner KI (2006) The influence of plant litter diversity on decomposer abundance and diversity. Soil Biol Biochem 38:1052–1062. https://doi.org/10.1016/j.soilbio.2005.09.003
Wasilewska L (1991) Long-term changes in communities of soil nematodes on fen peat meadows due to the time since their drainage. Ekol Pol 39
Wickham H (2016) Package `ggplot2`: elegant graphics for data analysis. Springer-Verl, New York, 1–222
Yeates G (1999) Effects of plants on nematode community structure. Annu Rev Phytopathol 37:127–149. https://doi.org/10.1146/annurev.phyto.37.1.127
Yeates GW, Bongers T, De Goede RGM, Freckman DW, Georgieva SS (1993) Feeding habits in soil nematode families and genera—an outline for soil ecologists. J Nematol 25:315–331
Zhang D, Hui D, Luo Y, Zhou G (2008) Rates of litter decomposition in terrestrial ecosystems: global patterns and controlling factors. J Plant Ecol 1:85–93. https://doi.org/10.1093/jpe/rtn002
Acknowledgements
We thank Nieves Martín Robles and Rocío Vigo for assistance with data gathering and lab work. We also thank all seed providers. This work was supported by MINECO (grants CGL2014-56567-R, CGL2017-83855-R, BES-2012-054356, PCIN-2014-053, PID2020-113021RA-I00), the European Union (Eco-serve project, 2013–2014 BiodivERsA/FACCE-JPI, with the national funders ANR, NWO, FCT, MINECO, FORMAS, and SNSF) and MICROAGRO (Fundación BBVA, Spain). JP was supported by Ministerio de Ciencia e Innovación (FPI fellowship PRE2018-085280), PGP was supported by Ministerio de Ciencia e Innovación (grant PID2020-113021RA-I00). We thank two anonymous reviewers for their comments, which made our work clearer and more complete.
Funding
Open Access funding provided thanks to the CRUE-CSIC agreement with Springer Nature.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Responsible Editor: Luz E. Bashan.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Palomino, J., García-Palacios, P., De Deyn, G.B. et al. Impacts of plant domestication on soil microbial and nematode communities during litter decomposition. Plant Soil 487, 419–436 (2023). https://doi.org/10.1007/s11104-023-05937-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-023-05937-4