Abstract
Background and Aims
Inoculation of legumes with effective N2-fixing rhizobia is a common practice to improve farming profitability and sustainability. To succeed, inoculant rhizobia must overcome competition for nodulation by resident soil rhizobia that fix N2 ineffectively. In Kenya, where Phaseolus vulgaris (common bean) is inoculated with highly effective Rhizobium tropici CIAT899 from Colombia, response to inoculation is low, possibly due to competition from ineffective resident soil rhizobia. Here, we evaluate the competitiveness of CIAT899 against diverse rhizobia isolated from cultivated Kenyan P. vulgaris.
Methods
The ability of 28 Kenyan P. vulgaris strains to nodulate this host when co-inoculated with CIAT899 was assessed. Rhizosphere competence of a subset of strains and the ability of seed inoculated CIAT899 to nodulate P. vulgaris when sown into soil with pre-existing populations of rhizobia was analyzed.
Results
Competitiveness varied widely, with only 27% of the test strains more competitive than CIAT899 at nodulating P. vulgaris. While competitiveness did not correlate with symbiotic effectiveness, five strains were competitive against CIAT899 and symbiotically effective. In contrast, rhizosphere competence strongly correlated with competitiveness. Soil rhizobia had a position-dependent numerical advantage, outcompeting seed-inoculated CIAT899 for nodulation of P. vulgaris, unless the resident strain was poorly competitive.
Conclusion
Suboptimally effective rhizobia can outcompete CIAT899 for nodulation of P. vulgaris. If these strains are widespread in Kenyan soils, they may largely explain the poor response to inoculation. The five competitive and effective strains characterized here are candidates for inoculant development and may prove better adapted to Kenyan conditions than CIAT899.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Legumes are a versatile group of plants that provide food for human consumption, pastures for grazing, and act as disease break crops in farming systems in many parts of the world (Angus et al. 2015; Broughton et al. 2003; Howieson et al. 2000). Legumes also boost agricultural productivity by supplying nitrogen to soils through a symbiotic relationship formed between the plant and a group of soil bacteria called rhizobia. The symbiosis is established when rhizobia infect legume roots, resulting in the formation of root nodules. Inside legume root nodules, rhizobia fix atmospheric N2 into NH3, which is secreted to the host plant and assimilated. Some of this fixed N2 makes its way into the soil in legume residues and can therefore benefit subsequent crops (Brophy and Heichel 1989; Maluk et al. 2022; Peoples et al. 2009). By harnessing symbiotic N2 fixation, farmers can reduce their reliance on synthetic nitrogen fertilizers, lower production costs, and contribute to global efforts to reduce carbon emissions.
To maximize the benefits of symbiotic N2 fixation in agriculture, legumes are often inoculated with highly effective strains of rhizobia that fix large amounts of N2 (Herridge et al. 2008; Howieson et al. 2000; Vanlauwe et al. 2019). However, rhizobia are common members of soil microbial communities and as such there can be great variation in the numerical abundance and genotypic composition of these soil communities (Brockwell et al. 1995; Kumar et al. 2015). When more than one genotype of rhizobia is capable of nodulating the target legume within the rhizosphere, competition for nodulation of the host plant may arise between the inoculant strain and these resident strains (Yates et al. 2011). When the inoculant strain is displaced in legume nodules by a more competitive resident strain that fixes substantially less N2 than the inoculant, then the full benefit of symbiotic N2 fixation cannot be realized, resulting in a diminished impact on farming systems.
In cases where resident soil rhizobia outcompete the inoculant strain for nodulation of the host, resident soil rhizobia are considered to create a “barrier” to inoculation response. Several studies have investigated overcoming this barrier. Weaver and Frederick (1974) working with soybean reported that > 103 rhizobia were required on the seed relative to the soil population, to achieve a response to inoculation. Other investigators subsequently showed that soil with populations as low as 102 rhizobia.g soil−1 (Singleton and Tavares 1986; Vargas et al. 2000) or even 50 rhizobia.g soil−1 (Thies et al. 1991a) were sufficient to provide a barrier to the inoculation response. In contrast, inoculation responses were achieved for Phaseolus vulgaris when inoculant was delivered into soils with 103 rhizobia.g soil−1 (Hungria et al. 2003; Vlassak et al. 1996) and as many as 104 and 105 rhizobia.g soil−1 (Hungria et al. 2000). Although the reason why an inoculation response can be inhibited by such vastly different numbers of resident soil rhizobia is not clearly understood, competitiveness is a genetic trait (Mendoza-Suarez et al. 2021) which is known to vary widely across individual strains (Aguilar et al. 2022). Thus, while the number of resident rhizobia is a key consideration in nodulation outcomes following inoculation, the competitive ability of resident strains in nodulating the target legume is also an important consideration.
Phaseolus vulgaris (common bean) is a grain legume that is widely grown as a source of dietary protein. Originating from two centers of domestication in Mesoamerica and the Andes (Schmutz et al. 2014), P. vulgaris is now cultivated in many areas of the tropics and sub-tropics. P. vulgaris is considered a promiscuous legume, since it is nodulated by a broad range of rhizobial species and while most of these are in the genus Rhizobium, species of Sinorhizobium and Bradyrhizobium as well as Paraburkholderia and Cupriavidis have also been reported to nodulate this host (da Silva et al. 2012; Dall’Agnol et al. 2016; Martínez-Romero 2003). In Kenya, where P. vulgaris has been cultivated for 400–500 years, grain yields remain low due to a range of issues such as low yielding cultivars, disease pressure, drought and low soil fertility, including low soil nitrogen levels (Katungi et al. 2009; Margaret et al. 2014). Although Kenyan soils contain native rhizobia capable of nodulating P. vulgaris, a substantial proportion of these rhizobia appear to be suboptimally effective at fixing N2 (Anyango et al. 1995; Kawaka et al. 2014). Therefore, P. vulgaris is sometimes inoculated with Rhizobium tropici CIAT899, a highly effective N2-fixing strain isolated from P. vulgaris in Colombia (Martínez-Romero et al. 1991). However, the yield of inoculated plants remains low, possibly due to poor adaptability of the inoculant strain to local environmental conditions and the presence of reasonably high (i.e. 104 cells.g soil−1) populations of resident rhizobia in soils that presumably compete for nodulation of P. vulgaris (Anyango et al. 1998; Musandu and Ogendo 2001). It is therefore possible that the lack of an inoculation response is due to a high population of competitive and ineffective N2 fixing rhizobia in Kenyan soil, creating a barrier to the successful nodulation of the host plant by the commercial inoculant strain.
Previously, we characterized the genotypic diversity of 36 rhizobia strains isolated from cultivated P. vulgaris sampled from 16 sites across five different counties in Kenya (Mwenda et al. 2018). We showed that these strains represented at least five different known Rhizobium spp. and at least one novel Rhizobium spp. Importantly, a relatively high proportion of these strains (22 of 36 or 61%) were effective at fixing N2, with 10 strains fixing N2 at rates equivalent to that of CIAT899. Thus, diverse and effective indigenous rhizobia nodulate cultivated P. vulgaris in Kenya. However, the infrequency with which inoculation with CIAT899 yields a positive response suggests the presence of competitive yet poorly effective strains in the soil. Here, we adapted a celB and gusA dual marker system (Sanchez-Canizares and Palacios 2013) to investigate competitiveness of this diverse range of P. vulgaris nodulating rhizobia from Kenya, in controlled glasshouse co-inoculation experiments with CIAT899.
Materials and methods
Bacterial strains, plasmids and media
Strains and plasmids used in the study are listed in Table 1. Escherichia coli strains were routinely cultured in Lysogeny Broth (LB) media at 37˚C (Bertani 1951) and rhizobia in Tryptone Yeast (TY) media at 28˚C (Beringer 1974). Where appropriate, media was supplemented with antibiotics at the following concentrations (μg ml−1): Spectinomycin (200), streptomycin (200), chloramphenicol (20), tetracycline (20) and nalidixic acid (75).
Marking of CIAT899 with gusA reporter gene
R. tropici CIAT899 was marked with mTn5SSgusA31, a mini-transposon carrying gusA expressed by the symbiotically active nifH promoter (Wilson et al. 1995). The mini-transposon was transferred from E. coli S17.1 strain (MUE254) by bi-parental mating into CIAT899 (Reeve et al. 2016) and transconjugants selected on TY media supplemented with spectinomycin, streptomycin, chloramphenicol and nalidixic acid. Selected transconjugants were assessed for growth rate, N2 fixation efficiency and competitiveness for nodulation on P. vulgaris.
The integration site of mTn5SSgusA31 in CIAT899 transconjugants was identified by a three-step nested PCR and subsequent sequencing approach. In the first step, amplification of single stranded DNA from gusA was achieved with primer GUS134 (Table 1) and 5 ng μl−1 genomic DNA, using cycling conditions of 94 °C for 2 min, followed by 10 cycles of 94 °C for 30 s, 60 °C for 30 s and 70 °C for 90 s. The reaction products were diluted fivefold with water and a 1 μl aliquot used as template for the second PCR, primed with GUS134 paired separately with CEKGRNB1, CEKGRNB2, CEKGRNB3 or CEKGRNB4 (Table 1). The products of this second round of PCR were again diluted fivefold, and 1 μl aliquots used as template for the third PCR, primed with WIL3 and CEKG4. Cycling conditions for the second and third round of PCR were as per Chun et al. (1997). The products of the third round of PCR were separated in 1% (w/v) agarose, excised, purified and Sanger sequenced using WIL3 and CEKG4 primers. Homology searches and sequence comparisons were carried out using the NCBI BLASTN search algorithm and transposon sequences distinguished from chromosomal sequences. The identified insertion site was further confirmed by PCR and sequencing of the DNA at both ends of the inserted mini-transposon using primer pairs IE-F with IE-R and OE-F with OE-R. The N2 fixation ability of CIAT899gusA was compared to that of wild-type CIAT899 on P. vulgaris cv. Tamu 42 days post-inoculation, as previously described (Mwenda et al. 2018).
Marking strains with the celB reporter gene
All 30 rhizobial strains used in this study (Table 1) were marked with a celB-expressing plasmid. The celB-expressing plasmid consisted of a celB and mNeonGreen cassette cloned into the stable broad-host range plasmid pJP2 (Prell et al. 2002). The insert was designed in Geneious (Biomatters Ltd, NZ), using celB (Sessitsch et al. 1996) and mNeonGreen (Shaner et al. 2013) sequences obtained from GenBank (GenBank Accession 11847805 and AGG56535.1). The sequences were fused, silent mutations introduced to remove XbaI and PstI restriction sites, and the constitutive tac promoter and terminal XbaI and PstI restriction sites added. After in silico assembly, the construct was chemically synthesised (GeneArt, Thermo Fisher Scientific) and cloned into the PstI-XbaI site of pJP2, after excision of gusA, resulting in the constitutive expression vector pGM01. This vector was introduced into E. coli ST18 (Thoma and Schobert 2009) by electroporation and into rhizobia by bi-parental conjugation (Reeve et al. 2016). Transconjugants were selected on TY media supplemented with tetracycline, verified first by PCR with primers PGM949F (binding to the oriV origin of replication in pGM01) and PGM1537R (binding to celB) and expression was subsequently confirmed by a β-galactosidase assay (Sessitsch et al. 1996), modified to include a heat-inactivation step (70 °C for 45 min) to inactivate endogenous β-galactosidase activity. The celB-marked strains were subsequently assessed for similarity to wildtype colony morphology and, for a subset, growth rate and competitiveness for nodulation of P. vulgaris was also assessed.
Root staining and scoring of nodule occupants
GUS staining and clearing of roots was performed as described previously (Wilson et al. 1995) with the exception that roots were first vacuum-infiltrated for 30 min in the GUS staining buffer containing 200 µg mL−1 of 5-bromo-4-chloro-3-indolyl-β-D-glucuronic acid (X-Glc) before incubation on a shaker (200 rpm) at 37 °C for 24 h. The CelB staining process was the same as that for GUS, but with 200 µg mL−1 5-Bromo-6-chloro-3-indolyl β-D-galactopyranoside (Magenta-Gal) substituted for X-Glc. To double stain, roots were first stained with X-Glc, incubated at 70 °C for two hours to destroy endogenous β-galactosidases, then stained with Magenta-Gal. Finally, roots were de-stained in 4% (w/v) sodium hypochlorite and rinsed thoroughly. Nodules formed by CIAT899gusA stained blue while those formed by celB-marked strains were magenta. Mixed nodules had various patterns of both colours.
Competition assays
To measure the competitiveness of strains, P. vulgaris cv. Kenya Tamu was co-inoculated with suspensions of liquid cultures of celB-marked strains and the CIAT899gusA reference strain. First, cells in logarithmic phase were harvested by centrifugation and washed free of antibiotics with sterile deionised water and resuspended to 1 × 104 cells ml−1. Each of the celB-marked strains was pre-mixed with CIAT899gusA (at 1:1 volume ratio) and one ml of this suspension inoculated onto surface-sterilised P. vulgaris seeds and sown into pots containing steam-sterilised sand. At the time of seed inoculation, viable cell counts and the proportion of each strain in the bacterial suspensions were confirmed by the Miles and Misra technique (O’Hara et al. 2016). Treatments had three pot replicates, each containing two seeds. Plants were maintained in a naturally-lit glasshouse as described previously (Mwenda et al. 2018), harvested after 21 days following inoculation and roots double-stained with X-Glc and Magenta-Gal. Competitiveness index (CI) was then calculated as follows: CI of Y = \((nodulesY/nodulesX)/(cfuY/cfuX)\), where cfuY and cfuX are the number of colony forming units of co-inoculated strains Y and X per ml of inoculant suspension applied to plants, and nodulesY and nodulesX denotes the number of nodules occupied by the respective strains.
Rhizosphere colonization assays
Four celB-marked strains (NAK120celB, NAK210celB, NAK287celB and CIAT899celB) plus CIAT899gusA were cultured in TY broth until mid-logarithimic phase (OD600 = 0.5) and a 1 ml aliquot of each was centrifuged (10,000 g, 1 min), washed and resuspended in sterile water. Each suspension was serially diluted to give a final viable cell count of 2 × 103 cells ml−1, and then each celB-marked strain was individually mixed in a 1:1 ratio with the CIAT899gusA suspension. For each combination, a 1 ml aliquot was removed for viable cell counts on TY agar with tetracycline (for enumeration of celB strains) and TY agar with spectinomycin (for enumeration of CIAT899gusA) and a further 1 ml aliquot was inoculated onto surface-sterilised pre-germinated seeds of P. vulgaris as described above. Treatments were replicated in eight pots, with two plants per pot which were thinned to one plant per pot at emergence. Seven days after inoculation, four plants from each treatment were carefully removed from their pots and roots gently shaken to remove excess soil. The roots, together with lightly adhering soil, were transferred to 50 ml tubes and resuspended in sterile water before shaking for 30 min at medium speed on an Analite wrist shaker (Ananlite Pty, Australia), after which viable cell counts were determined (O’Hara et al. 2016). Plants in the remaining pots were harvested 17 days after inoculation and roots double stained with X-Glc and Magenta-Gal and counted as described above.
Competition for nodulation of P. vulgaris between seed- and soil-borne resident strains
Sterile soil was separately inoculated with NAK120celB, NAK210celB, NAK287celB and CIAT899celB at 102 and 105 cells.g soil−1 to establish these rhizobia as soil-borne resident strains. To achieve this, bacteria were cultured to mid-log phase in TY broth and aliquots containing 2.72 × 105 or 2.72 × 108 cells re-suspended in 80 ml of sterile water. The suspensions were individually mixed into 3.2 kg steam-sterilised soil resulting in 102 and 105 rhizobial cells.g soil−1. Pots were covered with plastic wrap and kept in a shaded glasshouse maintained at 22 °C for four days before sowing. Viable cell counts of rhizobia were performed at sowing, 10 days and 30 days post sowing, from a depth of 5–10 cm by the Miles and Misra technique on TY medium supplemented with tetracycline (20 µg mL−1).
P. vulgaris cv. Kenya Tamu seed was inoculated at two rates with CIAT899gusA and sown into soils containing the pre-established soil-borne resident strains. Firstly, an inoculant carrying 109 cells.g peat−1 of CIAT899gusA was prepared using standard techniques (Yates et al. 2016) and cured for 10 days at 28 °C. This inoculant was diluted 100-fold by mixing with sterile peat and adjusted to 25% moisture with deionised water, resulting in a second inoculant carrying 107 cells.g peat−1. Surface-sterilised seed (Mwenda et al. 2018) was then inoculated with each peat preparation, using established techniques (Yates et al. 2016), with 40% (w/v) gum arabic as the sticker, to obtain a low and high inoculation rate. Rhizobia on seeds were counted by the Miles and Misra method. Two seeds inoculated with CIAT899gusA-containing peat, were sown into each pot containing the pre-established soil-borne resident strains. For the uninoculated treatment, uninoculated surface-sterilised seed was sown into pots with steam sterilised soil. Plants were maintained with N-free nutrient solutions and sterile deionised water as described previously (Mwenda et al. 2018) and harvested after 30 days, roots double-stained with X-Glc and Magenta-Gal, and nodules counted.
Data analysis
Data were analyzed by means with standard errors, Pearson’s correlation and where applicable, an analysis of variance (ANOVA) using SPSS version 22 (IBM Corp, released 2013). ANOVA was preceded by a test for normality and equal variances (Levene’s test). Tukey’s HSD was then used when ANOVA was found to be significant (P ≤ 0.05).
Results
Construction and validation of marked strains
To determine the competitiveness of R. tropici CIAT899, a highly effective N2-fixing symbiont of P. vulgaris and a common inoculant for this legume worldwide (Bala et al. 2011; Hungria et al. 2003), to P. vulgaris-nodulating rhizobia from Kenya, a differential staining approach was used to identify nodule occupants. The reference strain CIAT899 was marked with gusA (β-glucuronidase), while the strains from Kenya were marked with a plasmid-borne celB, encoding a thermostable β-glucosidase. The gusA gene under the control of the nifH promoter, which drives the expression of the genes encoding the nitrogenase complex, was transferred into CIAT899 using a mini-transposon (mTn5SSgusA31), which inserted 6-bp from the 3' end of universal stress protein A gene (uspA, locus tag RTCIAT899_CH07670 in the CIAT899 chromosome). In TY liquid culture, the mean generation time of gusA-marked CIAT899 (CIAT899gusA) did not differ significantly (P > 0.01) to that of wild-type CIAT899 [112 ± 5 min (standard error of the mean) and 111 ± 7 min, respectively]. Similarly, P. vulgaris plants inoculated with CIAT899gusA and cultivated in N-free conditions, yielded mean shoot dry weights that were indistinguishable from wild-type CIAT899 inoculated plants (3.54 ± 0.39 g vs 3.28 ± 0.23 g; P > 0.01). When co-inoculated onto plants in a 1:1 ratio of CIAT899gusA and wild-type CIAT899, 50.3% of the nodules were occupied by CIAT899gusA (stained blue with X-Glc) and 49.7% by the wild-type CIAT899. This indicates that gusA-marked CIAT899 is as competitive at nodulating P. vulgaris as CIAT899, so insertion of gusA into uspA in CIAT899gusA did not compromise the free-living growth, N2 fixation or competitiveness of this strain.
Plasmid pGM01, carrying celB under the control of the constitutive tac promoter, was conjugated into the 28 P. vulgaris-nodulating Kenyan Rhizobium strains, plus the commercial inoculant CIAT899 and R. leguminosarum sv. phaseoli 8002, a well-studied and effective P. vulgaris nodulating strain from the UK (Mwenda et al. 2018; Johnston et al. 1982). The growth rate of five selected strains carrying pGM01 (CIAT899, NAK120, NAK210, NAK239, NAK103) in TY broth (without antibiotic selection) did not differ to the growth rate of the respective unmarked wild-type strains, indicating the presence of the plasmid did not affect strain growth. Furthermore, when a 1:1 ratio of CIAT899gusA and CIAT899celB were co-inoculated onto P. vulgaris and stained with X-Glc and Magenta-Gal, both blue and magenta nodules were observed (Fig. 1A and B) in equal numbers (48 ± 5.6 mean number blue nodules.plant−1 vs 46 ± 6.5 mean number magenta nodules.plant−1), indicating that both marked strains were equally competitive and there was no impact on competition from harboring a chromosomal or plasmid-borne chromogenic marker gene.
P. vulgaris root nodules stained with X-Glc (blue) and Magenta-Gal (magenta) co-inoculated with (A and B) CIAT899gusA (blue nodules) or CIAT899celB (magenta nodules). (C) NAK104 and CIAT899gusA (D) NAK387 and CIAT899gusA. (E) Mixed nodule containing both CIAT899gusA and CIAT899celB. (F) P. vulgaris root nodules formed following inoculation with CIAT899gusA and NAK407celB, showing white nodules formed by the loss of the celB-harboring plasmid pGM01 for NAK407. Scale bar is 1 cm
Nodulation competitiveness
Next, we determined how capable each of the 30 celB-marked strains were at nodulating P. vulgaris when approximately 104 cells of each strain were individually co-inoculated at a 1:1 ratio on plants with CIAT899gusA. On post-harvest staining, 23 strains yielded nodules that were either blue, indicating the presence of CIAT899gusA in the nodule, or magenta, indicating the presence of the celB-marked strain (Fig. 1C and D). In 9% of nodules across treatments, both blue and magenta staining of a single nodule was observed, suggesting mixed infection of the nodule by both gusA and celB marked strains (Fig. 1E). For seven strains (NAK315, NAK334, NAK368, NAK407, NAK287, NAK103 and NAK299) carrying the celB marker, partially unstained or completely unstained nodules were observed following treatment with Magenta-Gal (Fig. 1F). These unstained nodules were observed alongside blue stained nodules, indicative of the presence of CIAT899gusA. The lack of staining, or partial staining, suggested instability of the celB-harboring plasmid (pGM01) in these strains. Re-isolation from stained nodules to confirm loss of plasmid from these strains in planta was not possible, as the CelB staining procedure requires heat-treatment to inactivate background β-galactosidase activity. Therefore, three of the seven strains (NAK287celB, NAK334celB and NAK407celB) and three strains that did not show unstained or partially stained nodules (CIAT899celB, NAK120celB and NAK210celB) were selected and the stability of pGM01 in these strains was assessed in free-living culture in the absence of antibiotic selection. After 20 generations, 100% of the CIAT899, NAK120 and NAK210 cells tested retained pGM01. In contrast, for the three strains that showed evidence of unstained nodules, all showed loss of the plasmid, with only 50% of NAK334, 78% of NAK287 and 72% NAK407 cells retaining the plasmid after 20 generations. This therefore indicates that loss of the plasmid in planta was the likely cause of partially or completely unstained nodules from these seven strains. Consequently, unstained nodules from plants co-inoculated with these seven strains were scored as magenta.
Strain competitiveness was calculated from the ratio of nodules occupied by the test (i.e. celB-marked strain) and reference (CIAT899gusA) strains, normalized for the proportion of viable cells of each strain inoculated onto plants. Mixed nodules were excluded from this calculation. Analysis of competitiveness for nodulation among the strains showed a wide range of phenotypes compared to CIAT899gusA (Fig. 2). Eight of the strains were more competitive (competitiveness > 1.2) at nodulating P. vulgaris than CIAT899, with the most competitive strain being NAK103 yielding 12-fold more nodules on this host. Five strains, including CIAT899 harboring the celB plasmid, were equally competitive (competitiveness between 0.8 and 1.2) at nodulating P. vulgaris. Four strains (NAK387, NAK254, NAK210, NAK334) were partially competitive compared to CIAT899gusA, yielding ratios of between 0.8 and 0.5. A further ten strains were poorly competitive, yielding fewer than half the number of nodules on P. vulgaris compared to the reference strain (competitiveness < 0.5). Three strains (NAK242, NAK299 and NAK284) did not nodulate P. vulgaris when co-inoculated with CIAT899 and were therefore categorized as uncompetitive.
Competitiveness of 30 P. vulgaris-nodulating strains carrying celB when co-inoculated with CIAT899gusA. Strain competitiveness index was calculated from the ratio of nodules occupied by the test (i.e. celB-marked strain) and reference (CIAT899gusA) strains, normalized for the proportion of viable cells of each strain inoculated onto plants. Strains were classed as more competitive (ratio > 1.2), equally competitive (1.2 to 0.8), partially competitive (0.8 to 0.5) and poorly competitive (< 0.5). NAK242, NAK299 and NAK284 did not nodulate P. vulgaris when co-inoculated with CIAT899 and were therefore classed as uncompetitive. CIAT899celB, which was equally competitive with CIAT899gusA, is denoted by a red column
No correlation between competitiveness and symbiotic effectiveness
In a previous study, the capacity of 28 Kenyan strains plus 8002 to fix N2 with P. vulgaris was compared to CIAT899 (Mwenda et al. 2018). To investigate whether a link between competitiveness and effectiveness of these strains existed, we analyzed the correlation between their observed competitiveness and N2 fixation capacity, expressed as a percentage of biomass achieved by plants inoculated with the test strain compared to CIAT899. We found that competitiveness did not correlate to effectiveness for N2 fixation (r2 = 0.0114) (Fig. 3). For example, of the 17 strains that were effective N2 fixers (> 75% of N2 fixed by CIAT899), only seven were either more competitive or equally competitive with CIAT899, while a further seven strains were poorly competitive, which included the least competitive strains NAK295 and 8002. The two remaining effective strains (NAK299 and NAK284) were uncompetitive (i.e. did not nodulate P. vulgaris). These data indicate that the traits of competitiveness and symbiotic N2 fixing effectiveness are not correlated in the strains analyzed.
Correlation of the competitiveness of 30 P. vulgaris nodulating strains with their symbiotic effectiveness. Data for N2 fixation effectiveness from Mwenda et al. (2018) expressed as a percentage of biomass achieved by plants inoculated with the test strain compared to CIAT899. Strain competitiveness index was calculated as indicated earlier. CIAT899 (defined as 100% effective and equally competitive) denoted by a red diamond. Dotted line denotes the linear correlation between the two traits, with r2 = 0.0114
Rhizosphere competence correlates with nodule occupancy
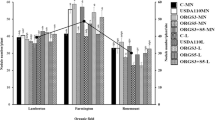
We next investigated the role of rhizosphere competence in nodulation outcomes when strains were co-inoculated with CIAT899gusA onto P. vulgaris. Four celB-harboring strains were chosen, representing strains that were more competitive (NAK287), equally competitive (CIAT899), partially competitive (NAK210) and poorly competitive (NAK120). Strains were co-inoculated with CIAT899gusA at ~ 103 cells seed−1, yielding approximately equal proportions of each strain at time of inoculation (Table 2). For the CIAT899celB and CIAT899gusA treatment, this equal proportion of strains was maintained in host rhizospheres sampled seven days after inoculation and in nodules on P. vulgaris plants harvested at 17 days post inoculation (Table 2), which supports there being no differences in competitiveness in these two strains, as was previously observed (Fig. 2). For NAK210, shown to be partially competitive from the earlier co-inoculation experiment with mature nodules at 21 days post inoculation, the proportion of cells declined from 43% at inoculation to 2% in the rhizosphere, and this low proportion was maintained through to nodulation, with only 5% of nodules occupied by this strain. While a similar, but less severe, decline in rhizosphere cell numbers was observed for NAK120 (41% to 14%), this strain did not occupy any P. vulgaris nodules. Therefore, both partially (NAK210) and poorly (NAK120) competitive strains were outcompeted by CIAT899 in the rhizosphere, and this was mirrored in later nodulation outcomes.
A decline in the proportion of NAK287 from 44% at inoculation to 14% in the rhizosphere was also observed, with this proportion then increasing to 50% occupancy in P. vulgaris root nodules (Table 2). As with the previous co-inoculation experiment, unstained nodules were also observed on plants of this treatment (caused by the instability of pGM01 plasmid in NAK287), so all unstained nodules from this treatment were scored as harboring NAK287. However, strain numbers in the rhizosphere were determined by addition of antibiotics to media to distinguish between CIAT899gusA (streptomycin resistant) and celB-harboring strains (tetracycline resistant) strains. The loss of plasmid pGM01 from some of the NAK287 cells would therefore mean the true density of NAK287 in the rhizosphere would be higher than recorded, as NAK287 cells that had lost pGM01 could not be enumerated on tetracycline plates. These data therefore suggest that NAK287 was likely able to maintain relatively equal numbers in the rhizosphere, leading to competitive nodulation of P. vulgaris compared to CIAT899.
Inoculation success in soils with established resident rhizobia
Having determined the competitiveness phenotypes of P. vulgaris nodulating rhizobia, we next investigated the outcome of strain competition in a system designed to mirror inoculation of P. vulgaris in the field, where a seed-borne inoculant is introduced at the time of sowing, into a soil with a pre-existing rhizobial population. To create the pre-existing populations, we chose the same test strains as the rhizosphere experiment (i.e. NAK287, CIAT899, NAK210 and NAK120) and inoculated them separately into sterile soil at 102 and 105 rhizobia.g soil−1. After a week in soil devoid of organic matter, added nutrients and host plants, these densities increased to (6.2 ± 0.7) × 104 and (1.8 ± 0.4) × 106 cells.g soil−1, respectively, creating two rhizobial densities similar to those often reported in soils where P. vulgaris is cultivated. P. vulgaris seeds inoculated with CIAT899gusA at low [(7.4 ± 0.4) × 104 cells.seed−1] and high [(6.6 ± 0.4) × 106 cells.seed−1] rates, representing rates below and above the recommended 105 cells seed−1 (Bullard et al. 2005; Lupwayi et al. 2000), were sown into pots containing these pre-existing rhizobial populations. The resultant plants were harvested 30 days after sowing and nodule occupants assessed using the dual marker system.
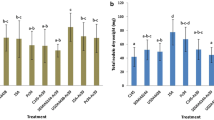
At both low and high seed and soil inoculation rates, seed-borne CIAT899gusA was outcompeted in soils with resident populations of celB-marked CIAT899 (equally competitive to CIAT899gusA), with 91% or more of nodules occupied by the strain resident in the soil (Fig. 4). A similar outcome was observed when NAK287 (more competitive) and NAK210 (partially competitive) were inoculated into soil, with 98% or more (for NAK287) and 85% or more (for NAK210) of nodules occupied by these resident strains. In contrast, when the poorly competitive strain NAK120 was resident in the soil at high densities (i.e. 106 cells.g soil), only 35% and 21% of nodules on P. vulgaris were formed by this strain at both low and high CIAT899gusA seed inoculation rates (Fig. 4b and d). This proportion of nodules occupied by NAK120 dropped to 28% and 12% when the same rates of seed-borne CIAT899gusA were inoculated at low NAK120 soil densities (i.e. 104 cells.g soil−1). Therefore, these results indicate that in soils with rhizobial densities of 104–106 cells.g soil−1, superior inoculation rates are inadequate to overcome nodulation of the legume by strains resident in the soil, unless the resident strains compete poorly with the inoculant strain.
Percentage of P. vulgaris nodules occupied by seed-borne CIAT899gusA versus percentage occupied by resident soil-borne CIAT899 (equally competitive), NAK 287 (more competitive), NAK210 (partially competitive) or NAK120 (poorly competitive), each individually harboring celB on pGM01. The seed-borne strain was applied at 7.4 ± 0.4 × 104 cells seed−1 and 6.6 ± 0.4 × 106 cells seed−1, while the soil-borne strain densities were 6.2 ± 0.7 × 104 and 1.8 ± 0.4 × 106 cells g−1 of soil at planting. Data are means of eight plant replicates per treatment
Discussion
A wide range of competitiveness phenotypes were observed in the 28 P. vulgaris-nodulating rhizobia isolated from Kenya, with only 27% of strains more competitive at nodulating P. vulgaris than CIAT899. This result is consistent with the long-standing use of CIAT899 in Africa and South America as a competitive and highly effective inoculant strain for P. vulgaris, with the strain also tolerant to high temperature and soil acidity (Bala et al. 2011; Hungria et al. 2000, 2003). Crucially, the competitiveness of the strains examined in this work did not correlate with strain effectiveness, with many instances of highly effective strains being poorly competitive, including the well-studied and highly effective P. vulgaris reference strain R. leguminosarum sv. phaseoli 8002, while some strains were even unable to nodulate P. vulgaris when co-inoculated with CIAT899. Only five strains (NAK103, NAK287, NAK227, NAK288 and NAK378) were more competitive than CIAT899 and capable of fixing N2 effectively with P. vulgaris. This emphasizes that competitiveness and effectiveness are separate genetic traits (Mendoza-Suarez et al. 2020) and reinforces the importance of measuring both characteristics when evaluating strains for suitability as inoculants.
Plasmid pGM01 proved an effective vector to express the celB reporter gene, and the plasmid was easily conjugated into all target strains. (Sanchez-Canizares and Palacios 2013; Sessitsch et al. 1996). While the plasmid was stable in free-living conditions under tetracycline selection, it was unstable in a small number of strains in planta (7 strains out of 30) and in a subset of these strains tested in the absence of antibiotic selection. pGM01 is derived from the expression vector pJP2, which itself was constructed from the stable broad-host range parent plasmid RK2 (Prell et al. 2002). Plasmids pJP2 and pGM01 harbor the parCBA plasmid partitioning system and the parDE toxin-antitoxin plasmid stability system, and pJP2 has been widely used as a stable expression system across a range of Alphaproteobacteria (Haskett et al. 2018; Mendoza-Suarez et al. 2020; Mulley et al. 2011; Prell et al. 2009; Tett et al. 2012). Why pGM01 was unstable in some strains assessed in this study is unclear, although it has previously been noted that the level of stability provided by parCBA can be strain dependent (Easter et al. 1998). Given that the 28 Kenyan strains investigated here represent at least six species of Rhizobium (Mwenda et al. 2018), it is possible that different expression levels of plasmid stability determinants in some of these diverse backgrounds could have resulted in plasmid loss.
Previous studies using gusA and celB dual markers have typically delivered these reporter genes via mini-transposons (Sessitsch et al. 1996) or via insertion into the bacterial genome by homologous recombination (Sanchez-Canizares and Palacios 2013). While these approaches generate stable marked strains, they can also generate variants with altered symbiotic phenotypes or require detailed genome sequence analysis to perform, which tends to limit their applicability to smaller number of strains. In contrast, although pGM01 appears unstable in some strain backgrounds, a majority of test strains in this study were able to stably maintain pGM01, making the plasmid a useful tool for dual strain competition experiments where screening of a relatively large numbers of strains is required.
Strains CIAT899, NAK287, NAK210 and NAK120 differed in their ability to colonize the rhizosphere of P. vulgaris when co-inoculated with CIAT899, with the proportion of nodules occupied by the strains strongly mirrored by the proportion of cells in the rhizosphere. Only NAK287 did not show this trend, however it is highly likely this was due to the instability of pGM01 in this strain, rather than an actual decrease in numbers of NAK287 in the soil. The ability of strains to colonize and thrive in the rhizosphere is related to a range of factors, including the survival of inoculant strains on seed (Deaker et al. 2004), survival and growth of strains under prevalent environmental conditions such as pH (Anyango et al. 1998), strain growth rates (Li and Alexander 1986), tolerance to microbial antagonism (Mrabet et al. 2006), chemotaxis and motility (Cooper 2007), root attachment (Janczarek et al. 2015), and ability to metabolize plant root exudates (Streit et al. 1992). In fact, a recent INSeq analysis of Rhizobium leguminosarum bv. viceae 3841 in the Pisum sativum rhizosphere showed 170 genes were required for rhizosphere growth (Wheatley et al. 2020). It is therefore likely that one or more genetic differences in rhizosphere competence between strains tested in this study may have caused a relatively enhanced ability of one strain to nodulate P. vulgaris ahead of another. Rhizosphere competence appears to be a significant factor in nodulation outcomes and therefore critical when considering new inoculant strains.
When seed inoculated CIAT899 was sown into soils with pre-existing rhizobial populations of 104 or 106 cells.g soil−1, nodulation of P. vulgaris was dominated in most cases by rhizobia resident in the soil at both low and high inoculation densities. Only when the resident strain was poorly competitive (NAK120), was seed-borne CIAT899 able to occupy a majority of root nodules. Most strikingly, when CIAT899 was both the seed- and soil-borne resident strain, more than 95% of nodules were formed by CIAT899 pre-existing in the soil. This indicates that the resident strain has a strong advantage in nodulating the target legume and only when the soil harbors a very poorly competitive strain, can the seed-borne inoculant dominate in the nodule. A likely key difference between seed and resident soil rhizobia was their physiological state, in that the inoculant strain was shifted from a relatively nutrient-rich and high moisture environment of the peat to the nutrient-poor and drier environment of the soil. In contrast, the resident strain had increased in number over a seven-day period in the soil prior to sowing with the inoculant-carrying seed and was presumably habituated to this environment. This physiological difference between inoculant and resident strains would likely lead to large differences in the ratio of the two strains during seedling germination. In fact, in a system which compared nodulation of Glycine max by two isogenic and equally competitive antibiotic resistant derivatives of Bradyrhizobium japonicum E11, the resident strain outnumbered the seed inoculated strain in the root infection zone four- to seven-fold (López-García et al. 2002). This type of position-dependent numerical advantage, which sees the resident strain dominate the infection zone of the host legume, would represent a significant hurdle for an inoculant strain to overcome, unless resident rhizobia are poorly competitive, such as was shown with NAK120 in this work.
P. vulgaris is often cultivated in soils with between 104 to 106 rhizobia.g soil−1 (Alberton et al. 2006; Anyango et al. 1998; Hungria et al. 2000; Thies et al. 1991b), and the numbers of cells in the soil environment tested in this study reflect these population sizes. Numerous studies investigating the effect of soil rhizobia population size on inoculation success in the field have shown that nodulation can be increased following inoculation in soils with 103 rhizobia.g soil−1 (Bergersen 1970; Hungria et al. 2000; Kyei-Boahen et al. 2017), 104 and 105 rhizobia.g soil−1 (Hungria et al. 2003; Vlassak et al. 1996) but also that nodulation by the inoculant can be inhibited in soils with as few as 102 rhizobia.g soil−1 (Singleton and Tavares 1986) or even less (Thies et al. 1991a). Genotypic variation of rhizobia in these different soil environments is the likely cause of these different responses to inoculation. Thus, if soils are composed mainly of strains that are highly competitive for nodulation of the target legume, it can be expected that even at relatively low numbers, they would dominate the inoculant strain. Conversely, soils composed predominantly by poorly competitive strains would present little barrier to inoculation response.
Five of the 28 Kenyan rhizobia analyzed in this study (NAK103, NAK287, NAK227, NAK288 and NAK378) were more competitive at nodulating P. vulgaris than CIAT899 and these strains are also effective at fixing N2 with P. vulgaris (Mwenda et al. 2018). Thus, Kenyan soils do harbor competitive and effective rhizobia for this important grain legume. These five strains are therefore candidates for further inoculant development evaluations and may prove better adapted to the local environment of east Africa than CIAT899, which originates from Colombia (Martínez-Romero et al. 1991). Indeed, several studies have used a similar approach of identifying locally adapted inoculant strains for cultivation of P. vulgaris in Spain (Mulas et al. 2015; Pastor-Bueis et al. 2019). However, this study also found that two strains that were as competitive (NAK266 and NAK312), and three strains that were more competitive (NAK315, NAK332 and NAK368) than CIAT899, were only partially (NAK315, NAK266 and NAK312) or poorly (NAK332 and NAK368) effective at fixing N2. The presence of these suboptimal yet competitive strains in soils where P. vulgaris is cultivated could inhibit inoculation responses and result in diminished benefits from symbiotic N2 fixation. Future studies should therefore investigate the extent to which suboptimal and competitive rhizobia are present across areas of P. vulgaris cultivation in Kenya. If they prove widespread, the recent development of a reporter plasmid for high-throughput simultaneous measurement of rhizobial competitiveness and N2 fixation (Mendoza-Suarez et al. 2020), may well provide a valuable tool to identify elite inoculant strains for P. vulgaris in Kenya.
Data Availability
All data generated or analysed during this study are included in this published article.
Change history
23 February 2023
Missing Open Access funding information has been added in the Funding Note.
References
Aguilar OM, Collavino MM, Mancini U (2022) Nodulation competitiveness and diversification of symbiosis genes in common beans from the American centers of domestication. Sci Rep 12:4591. https://doi.org/10.1038/s41598-022-08720-0
Alberton O, Kaschuk G, Hungria M (2006) Sampling effects on the assessment of genetic diversity of rhizobia associated with soybean and common bean. Soil Biol Biochem 39:1298–1307
Angus JF, Kirkegaard JA, Hunt JR, Ryan MH, Ohlander L, Peoples MB (2015) Break crops and rotations for wheat. Crop Pasture Sci 66:523–552. https://doi.org/10.1071/CP14252
Anyango B, Wilson KJ, Beynon JL, Giller KE (1995) Diversity of rhizobia nodulating Phaseolus vulgaris L. in two Kenyan soils with contrasting pHs. Appl Environ Microbiol 61:4016–4021. https://doi.org/10.1128/aem.61.11.4016-4021.1995
Anyango B, Wilson K, Giller KE (1998) Competition in Kenyan soils between Rhizobium leguminosarum biovar phaseoli strain Kim5 and R. tropici strain CIAT899 using the gusA marker gene. Plant Soil 204:69–78
Bala A, Karanja N, Mazvita M, Lwimbi L, Abaidoo R, Giller KE (2011) Production and the use of rhizobial inoculants in Africa: N2Africa project report
Bergersen FJ (1970) Some Australian studies relating to the long-term effects of the inoculation of legume seeds. Plant Soil 32:727–736
Beringer JE (1974) R factor transfer in Rhizobium leguminosarum. J Gen Microbiol 84:188–198
Bertani G (1951) Studies on lysogenesis. I. The mode of phage liberation by lysogenic Escherichia coli. J Bacteriol 62:293–300
Brockwell J, Bottomley PJ, Thies JE (1995) Manipulation of rhizobia microflora for improving legume productivity and soil fertility: A critical assessment. Plant Soil 174:143–180
Brophy LS, Heichel GH (1989) Nitrogen release from roots of alfalfa and soybean grown in sand culture. Plant Soil 116:77–84
Broughton WJ, Hernández G, Blair M, Beebe S, Gepts P, Vanderleyden J (2003) Beans (Phaseolus spp.) – model food legumes. Plant Soil 252:55–128. https://doi.org/10.1023/A:1024146710611
Bullard GK, Roughley RJ, Pulsford DJ (2005) The legume inoculant industry and inoculant quality control in Australia: 1953–2003. Aust J Exp Agric 45:127–140
Chun KT, Edenberg HJ, Kelley MR, Goebl MG (1997) Rapid amplification of uncharacterized transposon-tagged DNA sequences from genomic DNA. Yeast 13:233–240. https://doi.org/10.1002/(sici)1097-0061(19970315)13:3%3c233::Aid-yea88%3e3.0.Co;2-e
Cooper JE (2007) Early interactions between legumes and rhizobia: disclosing complexity in a molecular dialogue. J Appl Microbiol 103:1355–1365
da Silva K, Florentino LA, da Silva KB, de Brandt E, Vandamme P, de Souza Moreira FM (2012) Cupriavidus necator isolates are able to fix nitrogen in symbiosis with different legume species. Syst Appl Microbiol 35:175–182. https://doi.org/10.1016/j.syapm.2011.10.005
Dall'Agnol RF, Plotegher F, Souza RC, Mendes IC, Dos Reis Junior FB, Béna G, Moulin L, Hungria M (2016) Paraburkholderia nodosa is the main N2-fixing species trapped by promiscuous common bean (Phaseolus vulgaris L.) in the Brazilian 'Cerradão'. FEMS Microbiol Ecol 92. https://doi.org/10.1093/femsec/fiw108
Deaker R, Roughley RJ, Kennedy IR (2004) Legume seed inoculation technology—a review. Soil Biol Biochem 36:1275–1288. https://doi.org/10.1016/j.soilbio.2004.04.009
Easter CL, Schwab H, Helinski DR (1998) Role of the parCBA operon of the broad-host-range plasmid RK2 in stable plasmid maintenance. J Bacteriol 180:6023–6030. https://doi.org/10.1128/jb.180.22.6023-6030.1998
Haskett TL, Terpolilli JJ, Ramachandran VK, Verdonk CJ, Poole PS, O’Hara GW, Ramsay JP (2018) Sequential induction of three recombination directionality factors directs assembly of tripartite integrative and conjugative elements. PLoS Genet 14:e1007292. https://doi.org/10.1371/journal.pgen.1007292
Herridge DF, Peoples MB, Boddey RM (2008) Global inputs of biological nitrogen fixation in agricultural systems. Plant Soil 311:1–18
Howieson JG, O’Hara GW, Carr SJ (2000) Changing roles fo legumes in mediterranean agriculture; developments from an Australian perspective. Field Crop Res 65:107–122
Hungria M, Campo RJ, Mendes IC (2003) Benefits of inoculation of the common bean (Phaseolus vulgaris) crop with efficient and competitive Rhizobium tropici strains. Biol Fertil Soils 39:88–93
Hungria M, Andrade DS, Chueire LMO, Probanza AG-M, F J, Megias M (2000) Isolation and characterization of new efficient and competitive bean (Phaseolus vulgaris L.) rhizobia from Brazil. Soil Biol Biochem 32
Janczarek M, Rachwał K, Cieśla J, Ginalska G, Bieganowski A (2015) Production of exopolysaccharide by Rhizobium leguminosarum bv. trifolii and its role in bacterial attachment and surface properties. Plant Soil 388:211–227
Johnston AWB, Hombrecher G, Brewin NJ, Cooper CM (1982) Two transmissible plasmids in Rhizobium leguminosarum strain 300. J Gen Microbiol 128:85–93
Katungi E, Farrow A, Chianu J, Sperling L, Beebe S (2009) Common bean in Eastern and Southern Africa: a situation and outlook analysis. International Centre for Tropical Agriculture 61
Kawaka F, Dida MM, Opala PA, Ombori O, Maingi J, Osoro N, Muthini M, Amoding A, Mukaminega D, Muoma J (2014) Symbiotic Efficiency of native rhizobia nodulating common bean (Phaseolus vulgaris L.) in soils of Western Kenya. Int Sch Res Notices 2014:258497. https://doi.org/10.1155/2014/258497
Kumar N, Lad G, Giuntini E, Kaye ME, Udomwong P, Shamsani NJ, Young JP, Bailly X (2015) Bacterial genospecies that are not ecologically coherent: population genomics of Rhizobium leguminosarum. Open Biol 5:140133. https://doi.org/10.1098/rsob.140133
Kyei-Boahen S, Savala CEN, Chikoye D, Abaidoo R (2017) Growth and yield responses of cowpea to inoculation and phosphorus fertilization in different environments. Front Plant Sci 8:646. https://doi.org/10.3389/fpls.2017.00646
Li D-M, Alexander M (1986) Bacterial growth rates and competition affect nodulation and root colonization by Rhizobium meliloti. Appl Environ Microbiol 52:807–811
López-García SL, Vázquez TE, Favelukes G, Lodeiro AR (2002) Rhizobial position as a main determinant in the problem of competition for nodulation in soybean. Environ Microbiol 4:216–224. https://doi.org/10.1046/j.1462-2920.2002.00287.x
Lupwayi NZ, Olsen PE, Sande ES, Keyser HH, Collins MM, Singleton PW, Rice WA (2000) Inoculant quality and its evaluation. Field Crop Res 65:259–270. https://doi.org/10.1016/S0378-4290(99)00091-X
Maluk M, Ferrando-Molina F, Lopez del Egido L, Langarica-Fuentes A, Yohannes GG, Young MW, Martin P, Gantlett R, Kenicer G, Hawes C, Begg GS, Quilliam RS, Squire GR, Young JPW, Iannetta PPM, James EK (2022) Fields with no recent legume cultivation have sufficient nitrogen-fixing rhizobia for crops of faba bean (Vicia faba L.). Plant Soil 472:345–368. https://doi.org/10.1007/s11104-021-05246-8
Margaret N, Tenywa JS, Otabbong E, Mubiru DN, Basamba TA (2014) Development of common bean (Phaseolus vulgaris L.) production under low soil phosphorus and drought in sub-Saharan Africa: A review. J Sustain Dev 7:128–139
Martínez-Romero E (2003) Diversity of Rhizobium-Phaseolus vulgaris symbiosis: overview and perspectives. Plant Soil 252:11–23
Martínez-Romero E, Segovia L, Mercante FM, Franco AA, Graham P, Pardo MA (1991) Rhizobium tropici, a novel species nodulating Phaseolus vulgaris L. beans and Leucaena sp. trees. Int J Syst Bacteriol 41:417–426. https://doi.org/10.1099/00207713-41-3-417
Mendoza-Suarez MA, Geddes BA, Sanchez-Canizares C, Ramirez-Gonzalez RH, Kirchhelle C, Jorrin B, Poole PS (2020) Optimizing Rhizobium-legume symbioses by simultaneous measurement of rhizobial competitiveness and N2 fixation in nodules. Proc Natl Acad Sci U S A 117:9822–9831. https://doi.org/10.1073/pnas.1921225117
Mendoza-Suarez M, Andersen SU, Poole PS, Sanchez-Canizares C (2021) Competition, nodule occupancy, and persistence of inoculant strains: key factors in the rhizobium-legume symbioses. Front Plant Sci 12:690567. https://doi.org/10.3389/fpls.2021.690567
Mrabet M, Mnasri B, Romdhane SB, Laguerre G, Aouani ME, Mhamdi R (2006) Agrobacterium strains isolated from root nodules of common bean specifically reduce nodulation by Rhizobium gallicum. FEMS Microbiol Ecol 56:304–309. https://doi.org/10.1111/j.1574-6941.2006.00069.x
Mulas D, Seco V, Casquero PA, Velázquez E, González-Andrés F (2015) Inoculation with indigenous rhizobium strains increases yields of common bean (Phaseolus vulgaris L.) in northern Spain, although its efficiency is affected by the tillage system. Symbiosis 67:113–124. https://doi.org/10.1007/s13199-015-0359-6
Mulley G, White JP, Karunakaran R, Prell J, Bourdes A, Bunnewell S, Hill L, Poole PS (2011) Mutation of GOGAT prevents pea bacteroid formation and N2 fixation by globally downregulating transport of organic nitrogen sources. Mol Microbiol 80:149–167. https://doi.org/10.1111/j.1365-2958.2011.07565.x
Musandu AAO, Ogendo OJ (2001) Response of common bean to Rhizobium inoculation and fertilizers. J Food Technol Africa 6:121–125
Mwenda GM, O’Hara GW, De Meyer SE, Howieson JG, Terpolilli JJ (2018) Genetic diversity and symbiotic effectiveness of Phaseolus vulgaris-nodulating rhizobia in Kenya. Syst Appl Microbiol 41:291–299. https://doi.org/10.1016/j.syapm.2018.02.001
O’Hara GW, Hungria M, Woomer P, Howieson JG (2016) Counting rhizobia. In: Howieson JG, Dilworth MJ (eds) Working with rhizobia. Australian Centre for International Agricultural Research, Canberra
Pastor-Bueis R, Sánchez-Cañizares C, James EK, González-Andrés F (2019) Formulation of a highly effective inoculant for common bean based on an autochthonous elite strain of rhizobium leguminosarum bv. phaseoli, and genomic-based insights into its agronomic performance. Front Microbiol 10. https://doi.org/10.3389/fmicb.2019.02724
Peoples MB, Brockwell J, Herridge DF, Rochester IJ, Alves BJR, Urquiaga S, Boddey RM, Dakora FD, Bhattarai S, Maskey SL, Sampet C, Rerkasem B, Khans DF, Hauggaard-Nielsen H, Jensen ES (2009) The contributions of nitrogen-fixing crop legumes to the productivity of agricultural systems. Symbiosis 48:1–17
Prell J, Boesten B, Poole P, Priefer UB (2002) The Rhizobium leguminosarum bv. viciae VF39 gamma-aminobutyrate (GABA) aminotransferase gene (gabT) is induced by GABA and highly expressed in bacteroids. Microbiology 148:615–623
Prell J, White JP, Bourdes A, Bunnewell S, Bongaerts RJ, Poole PS (2009) Legumes regulate Rhizobium bacteroid development and persistence by the supply of branched-chain amino acids. Proc Natl Acad Sci U S A 106:12477–12482. https://doi.org/10.1073/pnas.0903653106
Reeve WG, Tiwari RP, De Meyer SE, Poole PS (2016) Specialised genetic techniques for rhizobia. In: Howieson JG, Dilworth MJ (eds) Working with rhizobia. Centre for International Agricultural Research, Canberra
Sanchez-Canizares C, Palacios J (2013) Construction of a marker system for the evaluation of competitiveness for legume nodulation in Rhizobium strains. J Microbiol Methods 92:246–249. https://doi.org/10.1016/j.mimet.2012.12.022
Schmutz J, McClean PE, Mamidi S, Wu GA, Cannon SB, Grimwood J, Jenkins J, Shu S, Song Q, Chavarro C, Torres-Torres M, Geffroy V, Moghaddam SM, Gao D, Abernathy B, Barry K, Blair M, Brick MA, Chovatia M, Gepts P, Goodstein DM, Gonzales M, Hellsten U, Hyten DL, Jia G, Kelly JD, Kudrna D, Lee R, Richard MM, Miklas PN, Osorno JM, Rodrigues J, Thareau V, Urrea CA, Wang M, Yu Y, Zhang M, Wing RA, Cregan PB, Rokhsar DS, Jackson SA (2014) A reference genome for common bean and genome-wide analysis of dual domestications. Nat Genet 46:707–713. https://doi.org/10.1038/ng.3008
Sessitsch A, Wilson KJ, Akkermans AD, de Vos WM (1996) Simultaneous detection of different Rhizobium strains marked with either the Escherichia coli gusA gene or the Pyrococcus furiosus celB gene. Appl Environ Microbiol 62:4191–4194. https://doi.org/10.1128/aem.62.11.4191-4194.1996
Shaner NC, Lambert GG, Chammas A, Ni Y, Cranfill PJ, Baird MA, Sell BR, Allen JR, Day RN, Israelsson M, Davidson MW, Wang J (2013) A bright monomeric green fluorescent protein derived from Branchiostoma lanceolatum. Nat Methods 10:407–409. https://doi.org/10.1038/nmeth.2413
Singleton PW, Tavares JW (1986) Inoculation response of legumes in relation to the number and effectiveness of indigenous Rhizobium populations. Appl Environ Microbiol 51:1013–1018. https://doi.org/10.1128/aem.51.5.1013-1018.1986
Streit WR, Kosch K, Werner D (1992) Nodulation competitiveness of Rhizobium leguminosarum bv. phaseoli and Rhizobium tropici strains measured by glucuronidase (gus) gene fusion. Biol Fertil Soils 14:140–144. https://doi.org/10.1007/BF00336264
Tett AJ, Rudder SJ, Bourdes A, Karunakaran R, Poole PS (2012) Regulatable vectors for environmental gene expression in Alphaproteobacteria. Appl Environ Microbiol 78:7137–7140. https://doi.org/10.1128/AEM.01188-12
Thies JE, Singleton PW, Bohlool BB (1991a) Influence of the size of indigenous rhizobial populations on establishment and symbiotic performance of introduced rhizobia on field-grown legumes. Appl Environ Microbiol 57:19–28
Thies JE, Singleton PW, Bohlool BB (1991b) Modeling symbiotic performance of introduced rhizobia in the field by use of indices of indigenous population size and nitrogen status of the soil. Appl Environ Microbiol 57:29–37
Thoma S, Schobert M (2009) An improved Escherichia coli donor strain for diparental mating. FEMS Microbiol Lett 294:127–132
Vanlauwe B, Hungria M, Kanampiu F, Giller KE (2019) The role of legumes in the sustainable intensification of African smallholder agriculture: Lessons learnt and challenges for the future. Agric Ecosyst Environ 284:106583. https://doi.org/10.1016/j.agee.2019.106583
Vargas MAT, Mendes IC, Hungria M (2000) Response of field-grown bean (Phaseolus vulgaris L.) to Rhizobium inoculation and nitrogen fertilization in two Cerrados soils. Biol Fertil Soils 32:228–233
Vlassak K, Vanderleyden J, Franco A (1996) Competition and persistence of Rhizobium tropici and Rhizobium etli in tropical soil during successive bean (Phaseolus vulgaris L.) cultures. Biol Fertil Soils 21:61–68. https://doi.org/10.1007/BF00335994
Weaver RW, Frederick LR (1974) Effect of inoculum rate on competitive nodulation of Glycine max L. Merrill I. Greenhouse studies. Agron J 66:229–232. https://doi.org/10.2134/agronj1974.00021962006600020014x
Wheatley RM, Ford BL, Li L, Aroney STN, Knights HE, Ledermann R, East AK, Ramachandran VK, Poole PS (2020) Lifestyle adaptations of Rhizobium from rhizosphere to symbiosis. Proc Natl Acad Sci U S A 117:23823–23834. https://doi.org/10.1073/pnas.2009094117
Wilson KJ, Sessitsch A, Corbo JC, Giller KE, Akkermans AD, Jefferson RA (1995) β-Glucuronidase (GUS) transposons for ecological and genetic studies of rhizobia and other gram-negative bacteria. Microbiology 141(Pt 7):1691–1705
Yates RJ, Howieson J, Reeve W, O’Hara GW (2011) A re-appraisal of the biology and terminology describing rhizobial strain succes in nodule occupancy of legumes in agriculture. Plant Soil 348:255–267
Yates RJ, Abaidoo R, Howieson J (2016) Field experiments with rhizobia. In: Howieson J, Dilworth M (eds) Working with rhizobia. Australian Centre for International Agricultural Research, Canberra
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions. This work was supported by a scholarship to George M. Mwenda by Murdoch University and N2Africa project: Putting Nitrogen Fixation to Work for Smallholder Farmers in Africa (www. N2Africa.org), funded by the Bill & Melinda Gates Foundation [OPP1020032] and by the Grains and Research Development Corporation of Australia (grants UMU1810 and UMU1901).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study concept and design. Experimentation was performed by George M. Mwenda and Yvette J. Hill. The first draft was written by George M. Mwenda and Jason J. Terpolilli and all authors commented on previous version of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Responsible Editor: Euan K. James.
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mwenda, G.M., Hill, Y.J., O’Hara, G.W. et al. Competition in the Phaseolus vulgaris-Rhizobium symbiosis and the role of resident soil rhizobia in determining the outcomes of inoculation. Plant Soil 487, 61–77 (2023). https://doi.org/10.1007/s11104-023-05903-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-023-05903-0