Abstract
Purpose
Soil bacteria comprise the largest number of soil microorganisms and play an important role in moso bamboo (Phyllostachys edulis) stumps decay; however, the characteristics of soil bacterial communities inside and outside these stumps remain unclear.
Methods
The characteristics of soil bacterial communities inside and outside Phyllostachys edulis bamboo stumps were analyzed under three different levels of decay using high-throughput sequencing technology.
Results
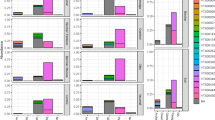
The abundance of operational taxonomic units, diversity index, and richness index inside and outside the bamboo stumps increased as the decay progressed; Proteobacteria, Acidobacteria, Actinobacteria, and Planctomycetes were the most abundant phyla in the soil inside and outside the bamboo stumps showing different levels of decay. Outside the bamboo stumps, there was a very significant positive correlation of Acidobacteria and Planctomycetes with the decaying degree. At the genus level, Rhodanobacter and Singulisphaera played an important role in the process of decay inside and outside the bamboo stumps, respectively. The Tax4Fun function prediction results showed that the relative abundance of the carbohydrate metabolism function was the highest inside and outside the bamboo stumps with different levels of decay. Proteobacteria had a very significant positive correlation with the membrane transport function, while Acidobacteria had a very significant positive correlation with the signal transduction function. Actinobacteria had a very significant positive correlation with the nucleotide metabolism and translation functions.
Conclusion
Our results augment our understanding of the expeditious degradation process of bamboo stumps, and provide a theoretical basis and reference for microbiological research, and sustainable bamboo stump operations.






Similar content being viewed by others
Data availability
Illumina MiSeq amplicon sequences of bacterial 16 S rRNA genes are available in the OMICSMART Sequence Read Archive. (http://www.omicsmart.com).
References
Andrea G, Martina B, Jutta W, Imhoff J (2011) Isolation and characterisation of bacteria from the Eastern Mediterranean deep sea. Antonie van Leeuwenhoek 100:421–435. https://doi.org/10.1007/s10482-011-9599-5
Aßhauer K, Bernd W, Rolf D, Peter M (2015) Tax4Fun: predicting functional profiles from metagenomic 16S rRNA data. Bioinformatics 31:2882–2884. https://doi.org/10.1093/bioinformatics/btv287
Cao Q, Wang S, Chen M, Cao R, Wang P, Zuo Q, Zhao S, Yang B (2021) The structure and diversity of nirS-denitrifying microbial community across three restoration stages of Xishuangbanna tropical forests. Acta Ecol Sin 41:626–636. https://doi.org/10.5846/stxb202004170915 (In Chinese)
Costa O, Hollander M, Pijl A, Liu B, Kuramae E (2020) Cultivation-independent and cultivation-dependent metagenomes reveal genetic and enzymatic potential of microbial community involved in the degradation of a complex microbial polymer. Microbiome 8:76. https://doi.org/10.1186/s40168-020-00836-7
Crawford D (1978) Lignocellulose decomposition by selected streptomyces strains. Appl Environ Microbiol 35:1041–1045. https://doi.org/10.1128/aem.35.6.1041-1045.197
Deng Y, Jiang Y, Yang Y, He Z, Luo F, Zhou J (2012) Molecular ecological network analyses. BMC Bioinf 13:113. https://doi.org/10.1186/1471-2105-13-113
Deng Q, Cheng X, Hui D, Zhang Q, Li M, Zhang Q (2016) Soil microbial community and its interaction with soil carbon and nitrogen dynamics following afforestation in central China. Sci Total Environ 541:230–237. https://doi.org/10.1016/j.scitotenv.2015.09.080
Ding J, Zhang Y, Deng Y, Cong J, Lu H, Sun X, Yang C, Yuan T, Nostrand J, Li D, Zhou J, Yang Y (2015) Integrated metagenomics and network analysis of soil microbial community of the forest timberline. Sci Rep 5:7994. https://doi.org/10.1038/srep07994
Ding X, Huang Y, Jing R, Ma F, An R, Tian Q, Chen B (2018) Bacterial structure and diversity of four plantations in the Yellow River Delta by high-throughput sequencing. Acta Ecol Sin 38:5857–5864. https://doi.org/10.5846/stxb201703260522 (In Chinese)
Ding Y, Du Y, Gao G, Zhang Y, Gao H, Zhu B, Yang S, Zhang J, Qiu Y, Liu H (2021) Soil bacterial community structure and functional prediction of Pinus sylvestris var. Mongolica plantations in the Hulun Buir Sandy Land. Acta Ecol Sin 41:4131–4139. https://doi.org/10.5846/stxb202005251335 (In Chinese)
Ed-Haun C, Chiou-Pin C, Guanglong T, Chih-Yu C (2018) Replacement of natural hardwood forest with planted bamboo and cedar in a humid subtropical mountain affects soil microbial community. Appl Soil Ecol 124:146–154. https://doi.org/10.1016/j.apsoil.2017.11.006
Fang T, Li Y, Yao Z, Li Y, Wang X, Wang Y, Yu Y (2021) Effects of planting broadleaf trees and moso bamboo on soil carbon mineralization and microbial community structure. Chin J Appl Ecol 32:82–92. https://doi.org/10.13287/J.1001-9332.202101.033
Feng D (2018) Dynamic changes of nutrient elements and their effects on the microbial diversity of sediments during the decaying of Nelumbonucifera detritus. Jiang Su University, Jiangsu (In Chinese)
Fuchs B, Zubkov M, Sahm K, Burkill P, Amann R (2000) Changes in community composition during dilution cultures of marine bacterioplankton as assessed by flow cytometric and molecular biological techniques. Environ Microbiol 2:191–201. https://doi.org/10.1046/j.1462-2920.2000.00092.x
Guo Y, Ren C, Yi J, Doughty R, Zhao F (2020) Contrasting responses of rhizosphere bacteria, fungi and arbuscular mycorrhizal fungi along an elevational gradient in a temperate montane forest of China. Front Microbiol 11:2042. https://doi.org/10.3389/fmicb.2020.02042
Hervé V, Ketter E, Pierrat J, Gelhaye E, Frey-Klett P (2016) Impact of Phanerochaete chrysosporium on the Functional Diversity of Bacterial Communities Associated with Decaying Wood. PLoS ONE 11:e0147100. https://doi.org/10.1371/journal.pone.0147100
Hong X, Hong Y, Chen Z, Zhao C, Yang S (2015) Phylogenetic diversity of nirs-type denitrifying bacteria in jiulong river estuary. Microbiology (Beijing, China) 42:1639–1650. https://doi.org/10.13344/j.microbiol.china.140953 (In Chinese)
Hu W, Xiang J, Xiang Y, Chen Y (2020) Effects of nitrogen-doped carbon nanoparticles on bacterial community in paddy rhizosphere soil. Chin J Appl Ecol 31:3842–3850. https://doi.org/10.13287/j.1001-9332.202011.033 (In Chinese)
Hua W, Wu L (2000) Preliminary report on the research of Phyllostachys edulis decay promotion technology. J For Eng 14:38–39. https://doi.org/10.13360/j.issn.1000-8101.2000.01.015 (In Chinese)
Kielak A, Barreto C, Kowalchuk G, van Veen J, Kuramae E (2016) The ecology of Acidobacteria: Moving beyond genes and genomes. Front Microbiol 7:744. https://doi.org/10.3389/fmicb.2016.00744
Li M (2012) Microstructure and chemical composition changes of Betula luminifera, Moso Bamboo and Phyllostachys praecox in decay. Zhejiang A&F University, Zhengjiang (In Chinese)
Li C, Li L, Yang K, Zhang L, Peng Z (2008) Isolating and screening of microorganisms for decomposing bamboo stump. For Res 21:253–257. https://doi.org/10.13275/j.cnki.Iykxyj.2008.02.023 (In Chinese)
Li M, Ai W, Yang M, Meng Y, Tu J, Pu X, Liu X (2014) The changes of microorganism in the bamboo stump with different decomposition degree. J For Eng 28:64–67. https://doi.org/10.13360/j.issn.1000-8101.2014.05.015 (In Chinese)
Li M, Lu J, Ai W, Yang M, Meng Y, Tu J, Pu X, Liu X (2015) Changes of microorganisms in Mao bamboo stumps at different decomposition degrees. Agr Sci Tech 16:1073–1077. https://doi.org/10.16175/j.cnki.1009-4229.2015.05.049
Li W, Tian X, Sheng H, Ekawati D, Zhou Y, Zhang R (2020) Response of bacterial compositions to soil biochemical properties under mulching-intensive management in a Phyllostachys edulis forest. Appl Soil Ecol 150:103436. https://doi.org/10.1016/j.apsoil.2019.103436
Lian C, Liu R, Zhang S, Yuan J, Luo J, Yang F, Fei B (2020) Ultrastructure of parenchyma cell wall in bamboo (Phyllostachys edulis) culms. Cellulose 27:7321–7329. https://doi.org/10.1007/s10570-020-03265-9
Lin L (2011) A preliminary study of distribution and rotting speed of bamboo stump in phyllostachys heterocycla var. pubescens forest. World Bamboo Ratt 9:57–59. https://doi.org/10.13640/j.cnki.wbr.2011.03.020 (In Chinese)
Liu Q (2016) Absorption transformation and utilization in Phyllostachys edulis forests. Chinese Academy of Forest, Beijing (In Chinese)
Liu B, Zhang S, Zhou Y, Lin J, Sun F (2015) Mildew in bamboo flour treated with different solvents. J Zhejiang A F Univ 32:11–17. https://doi.org/10.11833/j.issn.2095-0756.2015.01.002 (In Chinese)
Liu X, Siemann E, Cui C, Liu Y, Guo X, Zhang L (2019a) Moso bamboo (Phyllostachys edulis) invasion effects on litter, soil and microbial PLFA characteristics depend on sites and invaded forests. Plant Soil 438:85–99. https://doi.org/10.1007/s11104-019-04010-3
Liu G, Chen L, Shi X, Yuan Z, Yuan L, Lock T, Kallenbach R (2019b) Changes in rhizosphere bacterial and fungal community composition with vegetation restoration in planted forests. Land Degrad Dev 30:1147–1157. https://doi.org/10.1002/ldr.3275
Lu Y, Li K, Liang Q, Li C, Zhang C (2019) Effects of leaf litter decomposition on bacterial community structure in the leaf litter of four dominant tree species in Mount Tai. Acta Ecol Sin 39:3175–3186. https://doi.org/10.5846/stxb201806211365 (In Chinese)
Lucas-Borja M, Candel D, Jindo K, Moreno J, Andrés M, Bastida F (2012) Soil microbial community structure and activity in monospecific and mixed forest stands, under Mediterranean humid conditions. Plant Soil 354:359–370. https://doi.org/10.1007/s11104-011-1072-8
Megumi Y, Satoshi I, Shigeto O, Keishi S (2009) Temporal shifts in diversity and quantity of nirS and nirK in a rice paddy field soil. Soil Biol Biochem 41:2044–2051. https://doi.org/10.1016/j.soilbio.2009.07.012
Rahman M, Quadir Q, Rahman A, Asha M, Chowdhury M (2015) Screening and characterization of phosphorus solubilizing bacteria and their effect on rice seedlings. Res Agric Livest Fish 1:27–35. https://doi.org/10.3329/ralf.v1i1.22353
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett WS, Huttenhower C (2011) Metagenomic biomarker discovery and explanation. Genome Biol 12:R60. https://doi.org/10.1186/gb-2011-12-6-r60
Shen C, Ni Y, Liang W, Wang J, Chu H (2015) Distinct soil bacterial communities along a small-scale elevational gradient in alpine tundra. Front Microbiol 6:582. https://doi.org/10.3389/fmicb.2015.00582
Song J (2018) Study on the change of components of bamboo stumps in bamboo forest of Phyllostachys edulis and artificially promoting decomposition technology. Central South University of Forestry and Technology, Hunan (In Chinese)
Tang W (2013) Effect of bamboo stump rotting on soil physicochemical properties. World Bamboo Ratt 11:31–33. https://doi.org/10.13640/j.cnki.wbr.2013.02.008 (In Chinese)
Wang S, Wang B, Fan S, Xiao X, Xia W, Guan F (2021a) Influence of strip cutting management on soil bacterial community structure and diversity in Phyllostachys edulis stands. J Nanjing For Univ 45:60–68. https://doi.org/10.12302/j.issn.1000-2006.202004038 (In Chinese)
Wang X, Luo Z, Zhang R, Niu Y, Li L, Tian J, Sun P, Liu J (2021b) Soil bacterial community characteristics and ecological function prediction of alfalfa and crop rotation systems in the Loess Plateau, Northwest China. Chin J Appl Ecol. https://doi.org/10.13287/j.1001-9332.202204.028 (In Chinese)
Wei P (2016) Study on the main chemical constitutes distribution of Phyllostachys edulis (Carr.) J. Houz cell wall. Chinese Academy of Forestry, Beijing (In Chinese)
Wu Z, Hao Z, Sun Y, Guo L, Huang L, Zeng Y, Wang Y, Yang L, Chen B (2016) Comparison on the structure and function of the rhizosphere microbial community between healthy and root-rot Panax notoginseng. Appl Soil Ecol 107:99–107. https://doi.org/10.1016/j.apsoil.2016.05.017
Xie X, Liu N, Yang B, Yu C, Zhang Q, Zheng X, Xu L, Li R, Liu J (2016) Comparison of microbial community in hydrolysis acidification reactor depending on different structure dyes by Illumina MiSeq sequencing. Int Biodeterior Biodegrad 111:14–21. https://doi.org/10.1016/j.ibiod.2016.04.004
Xiong J, Liu Y, Lin X, Zhang H, Zeng J, Hou J, Yang Y, Yao T, Knight R, Chu H (2012) Geographic distance and pH drive bacterial distribution in alkaline lake sediments across Tibetan Plateau. Environ Microbiol 14:2457–2466. https://doi.org/10.1111/j.1462-2920.2012.02799.x
Xu F, Li S, Su J (2021) Changes of soil bacterial community structure at the secondary successional stages in the pinus yunnanensis forest. Chin J Appl Ecol 32:887–894. https://doi.org/10.13287/J.1001-9332.202103.039 (In Chinese)
Xue X, Wang L, Han X, Chen R, Wang J (2021) Effects of different tree disk mulching on soil microbial community structure and diversity in dwarfing rootstock apple orchard. Acta Ecol Sin 41:1528–1536. https://doi.org/10.5846/stxb201909191955 (In Chinese)
Yang J, Zhou G, Tian Y, Liu Q, Liu C, Yang Q, Zhou J (2015) Differential analysis of soil bacteria diversity in different mixed forests of Dalbergia odorifera. Acta Ecol Sin 35:8117–8127. https://doi.org/10.5846/stxb201409191856 (In Chinese)
Yuan Z, Liu F, Liu Z, Huang Q, Zhang G, Pan H (2021) Structural variability and differentiation of niches in the rhizosphere and endosphere bacterial microbiome of moso bamboo (Phyllostachys edulis). Sci Rep 11:1574. https://doi.org/10.1038/S41598-021-80971-9
Zhang T, Xu F, Huai B, Yang X, Sui W (2020) Effects of land use changes on soil bacterial community diversity in the riparian wetland along the downstream of Songhua River. J Environ Sci (Beijing, China) 41:4273–4283. https://doi.org/10.13227/j.hjkx.202003088 (In Chinese)
Zheng R (2011) Effects of phyllostachys pubescens forest bamboo pockets fertilization on bamboo pockets rot and the bamboo shoot emergence. J Fujian For Sci Technol 38:72–74. https://doi.org/10.3969/j.issn.1002-7351.2011.01.18 (In Chinese)
Zhou L, Li J, Zhao Y, Luo Y, Bai Y, Chen J, Wu Z, Lin W (2020) Variation of bacterial communities in the rhizosphere soils of successive rotations Casuarina equisetifolia plantations based on high-throughput sequencing analysis. Acta Ecol Sin 40:2670–2679. https://doi.org/10.5846/stxb201902250348
Zhu C (2005) Ammonium bicarbonate with salt to remove bamboo stump. Spec Econ Anim Plants 3:27. (In Chinese)
Zhu Y, Long H, Lou C, Gu X, Yue J (2013) The technology to promote the decay of bamboo stumps. J Bamboo Res 32:53–57. (In Chinese)
Acknowledgements
We thank the member of Institute of Forestry and Environment, Fujian Agriculture and Forestry University lively discussions that improved the work.
Funding
This work was supported by the Forestry Science and Technology Extension Demonstration Project financed by the Chinese Central Government [grant number Min (2016) TG14 to L.Cai.].
Author information
Authors and Affiliations
Contributions
The manuscript was drafted and researched by Fengna Liang and Liping Cai. Research and sample collection were completed by Xiao Huang, Huixin Zheng, Na Sun, Xuelong Qin, Cheng Jin, Le Yu and Yonglai Huang. The test plan was completed by Liping Cai and Xiangqing Ma. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interest.
Additional information
Responsible Editor: Timothy J. Fahey.
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Liang, F., Huang, X., Zheng, H. et al. Characteristics of the soil bacterial community in the decomposition process inside and outside moso bamboo stumps. Plant Soil 478, 635–650 (2022). https://doi.org/10.1007/s11104-022-05493-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-022-05493-3