Abstract
Background and aims
Crop yield and quality are generally limited by poor soils, which is a key limiting factor for sustainable development in modern agriculture. Wild soybean (Glycine soja) is an excellent wild resource, with tolerance to adverse environments, especially poor soil. This study aimed to reveal the key molecular mechanism of wild soybean to resist phosphorus deficiency in soil.
Methods
Differences in the types, amounts and metabolic pathways of small molecule metabolites and gene expression were compared and multi-omics integration analysis was performed between wild and cultivated soybean (Glycine max) seedling roots under sufficient and artificially simulated low-phosphorus in this study.
Results
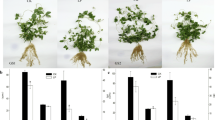
Under low-phosphorus stress, wild soybean seedlings experienced less growth inhibition and root-specific growth compared with cultivated soybean. Genes encoding sulfoquinovosyl transferase (SQD2), catechol O-methyltransferase (COMT), glutathione S-transferase (GST) and peroxidase (POD) were up-regulated; levels of glutamic acid, glycine, putrescine, phenylalanine, tyrosine, catechol and neohesperidin were increased; and levels of glycerol-3-phosphate decreased. Integrated analysis showed that the above genes and metabolites were involved in glutathione metabolism, glycerolipid metabolism and phenylpropane biosynthesis.
Conclusions
These metabolic pathways are involved in phosphorus reuse, while membrane lipid remodelling and reactive oxygen species scavenging are carried out to maintain membrane stability and ensure plant survival under phosphorus deficiency. This study provides new ideas for the study of mechanism of tolerance to phosphorus deficiency in wild soybean and lays the theoretical foundation for developing varieties of cultivated soybean that tolerate poor soils.






Similar content being viewed by others
References
Abelson PH (1999) A Potential Phosphate Crisis. Science, 283(5410):2015–2015. https://doi.org/10.1126/science.283.5410.201
Byrne SL, Foito A, Hedley PE, Morris J, Stewart D, Barth S (2011) Early response mechanisms of perennial ryegrass (Lolium perenne) to phosphorus deficiency. Ann Bot-London 107(2):243–254. https://doi.org/10.1093/aob/mcq234
Correll DL (1998) The role of phosphorus in the eutrophication of receiving waters: a review. J Environ Qual 27:261–266. https://doi.org/10.2134/jeq1998.00472425002700020004x
Chen S, Zhao H, Ding G et al (2015) Genotypic differences in antioxidant response to phosphorus deficiency in Brassica napus. Plant Soil 391:19–32. https://doi.org/10.1007/s11104-015-2395-7
Drevon JJ, Hartwig U (1997) Phosphorus deficiency increases the argon-induced decline of nodule nitrogenase activity in soybean and alfalfa. Planta 201:463–469. https://doi.org/10.1007/s004250050090
Ding N, Huertas R, Torres-Jerez I, Liu W, Watson B, Scheible WR, Udvardi M (2021) Transcriptional, metabolic, physiological and developmental responses of switchgrass to phosphorus limitation. Plant Cell Environ 44(1):186–202. https://doi.org/10.1111/pce.13872
Dong NQ, Lin HX (2021) Contribution of phenylpropanoid metabolism to plant development and plant-environment interactions. J Integr Plant Biol 63(1):180–209. https://doi.org/10.1111/jipb.13054
Deepika D, Singh A (2021) Plant phospholipase D: novel structure, regulatory mechanism, and multifaceted functions with biotechnological application. Crit rev Biotechnol, 1–19. Advance online publication. https://doi.org/10.1080/07388551.2021.1924113
Fu YQ, Yang XJ, Shen H (2014) The physiological mechanism of enhanced oxidizing capacity of rice (Oryza sativa L.) roots induced by phosphorus deficiency. Acta Physiol Plant 36:179–190. https://doi.org/10.1007/s11738-013-1398-3
Gupta K, Dey A, Gupta B (2013) Plant polyamines in abiotic stress responses. Acta Physiol Plant 35:2015–2036. https://doi.org/10.1007/s11738-013-1239-4
George TS, Hinsinger P, Turner BL (2016) Phosphorus in soils and plants – facing phosphorus scarcity. Plant Soil 401:1–6. https://doi.org/10.1007/s11104-016-2846-9
Gao Y, Jiang Z, Du M et al (2019) Photosynthetic and metabolic responses of eelgrass Zostera marina L. to short-term high-temperature exposure. J Ocean Limnol 37:199–209. https://doi.org/10.1007/s00343-019-7319-6
Gupta, S., Yadav, B. S., Raj, U., Freilich, S., & Varadwaj, P. K (2017) Transcriptomic Analysis of Soil Grown T aestivum cv Root to Reveal the Changes in Expression of Genes in Response to Multiple Nutrients Deficiency Front Plant Sci, 8, 1025. https://doi.org/10.3389/fpls.2017.01025
Hinsinger P (2001) Bioavailability of soil inorganic P in the rhizosphere as affected by root-induced chemical changes: a review. Plant Soil 237:173–195. https://doi.org/10.1023/A:1013351617532
Hosseini P, Matthews BF (2014) Regulatory interplay between soybean root and soybean cyst nematode during a resistant and susceptible reaction. BMC Plant Biol 14:300. https://doi.org/10.1186/s12870-014-0300-9
Hasanuzzaman M, Nahar K, Anee TI, Fujita M (2017) Glutathione in plants: biosynthesis and physiological role in environmental stress tolerance. Physiol Mol Biol Pla 23(2):249–268. https://doi.org/10.1007/s12298-017-0422-2
Idborg H, Edlund PO, Jacobsson SP (2004) Multivariate approaches for efficient detection of potential metabolites from liquid chromatography/mass spectrometry data. Rapid Commun Mass Sp, 18(9):944–954. https://doi.org/10.1002/rcm.1432
Juszczuk IM, Malusa M, Rychter AM (2001) Phosphate deficiency induced oxidative stress in bean (Phaseolus vulgaris L.). in: Horst W.J. et al. (eds) plant nutrition. Developments in plant and soil sciences, vol 92. Springer, Dordrecht. https://doi.org/10.1007/0-306-47624-X_71
Jinglong Z et al (2019) Transcriptome analysis reveals candidate genes related to phosphorus starvation tolerance in sorghum. BMC Plant Biol 19:1. https://doi.org/10.1186/s12870-019-1914-8
Jiao B, Zhao X, Lu W, Guo L, Luo K (2019) The R2R3 MYB transcription factor MYB189 negatively regulates secondary cell wall biosynthesis in Populus. Tree Physiol 39(7):1187–1200. https://doi.org/10.1093/treephys/tpz040
Kirkby EA, Johnston AE (2008) Soil and fertilizer phosphorus in relation to crop nutrition. In: white PJ, Hammond JP (eds) the ecophysiology of plant-phosphorus interactions. Plant ecophysiology, vol 7. Springer, Dordrecht. https://doi.org/10.1007/978-1-4020-8435-5_9
Kofsky J, Zhang H, Song BH (2018) The untapped genetic reservoir: the past, current, and future applications of the wild soybean (Glycine soja). Front Plant Sci 9:949. https://doi.org/10.3389/fpls.2018.00949
Lynch JP (2011) Root phenes for enhanced soil exploration and phosphorus acquisition: tools for future crops. Plant Physiol 156(3):1041–1049. https://doi.org/10.1104/pp.111.175414
Li MX, Guo R,Jiao Y, Jin XF, Zhang HY, Shi LX (2017) Comparison of salt tolerance in Soja based on metabolomics of seedling roots. Front plant Sci, 2017,8: https://doi.org/10.3389/fpls.2017.01101
Liu DP, Li MX, Liu Y, Shi LX (2020) Integration of the metabolome and transcriptome reveals the resistance mechanism to low nitrogen in wild soybean seedling roots. Environ Exp bot, 175. 104043.https://doi.org/10.1016/j.envexpbot.2020.104043
Luo B, Ma P, Nie Z, Zhang X, He X, Ding X, Feng X, Lu Q, Ren Z, Lin H, Wu Y, Shen Y, Zhang S, Wu L, Liu D, Pan G, Rong T, Gao S (2019) Metabolite profiling and genome-wide association studies reveal response mechanisms of phosphorus deficiency in maize seedling. Plant J 97(5):947–969. https://doi.org/10.1111/tpj.14160
Laure G, Reisdorf-Cren M, Rothstein SJ, Chardon F, Suzuki A (2010) Biological functions of asparagine synthetase in plants. Plant Sci 179(3):141–153, ISSN 0168-9452. https://doi.org/10.1016/j.plantsci.2010.04.010
Laure G, Rothstein SJ, Suzuki A (2016) Asparagine metabolic pathways in arabidopsis. Plant Cell Physiol 57(4):675–689. https://doi.org/10.1093/pcp/pcv184
Liu JH, Wang W, Wu H, Gong X, Moriguchi T (2015) Polyamines function in stress tolerance: from synthesis to regulation. Front Plant Sci 6:827. https://doi.org/10.3389/fpls.2015.00827
Lam HM, Xu X, Liu X, Chen W, Yang G, Wong FL, Li MW, He W, Qin N, Wang B, Li J, Jian M, Wang J, Shao G, Wang J, Sun SS, Zhang G (2010) Resequencing of 31 wild and cultivated soybean genomes identifies patterns of genetic diversity and selection. Nat Genet 42(12):1053–1059. https://doi.org/10.1038/ng.715
Li M, Xu J, Wang X, Fu H, Zhao M, Wang H, Shi L (2018) Photosynthetic characteristics and metabolic analyses of two soybean genotypes revealed adaptive strategies to low-nitrogen stress. J Plant Physiol 229:132–141. https://doi.org/10.1016/j.jplph.2018.07.009
Ma QP, Hao S, Chen X et al (2016) Validation of reliability for reference genes under various abiotic stresses in tea plant. Russ J Plant Physiol 63:423–432. https://doi.org/10.1134/S1021443716030080
Meena SK, Pandey R, Sharma S et al (2021) Cross tolerance to phosphorus deficiency and drought stress in mungbean is regulated by improved antioxidant capacity, biological N2-fixation, and differential transcript accumulation. Plant Soil 466:337–356. https://doi.org/10.1007/s11104-021-05062-0
Mo X, Zhang M, Liang C, Cai L, Tian J (2019) Integration of metabolome and transcriptome analyses highlights soybean roots responding to phosphorus deficiency by modulating phosphorylated metabolite processes. Plant Physiol Bioch 139:697–706. https://doi.org/10.1016/j.plaphy.2019.04.033
Müller J, Gödde V, Niehaus K, Zörb C (2015) Metabolic adaptations of white Lupin roots and shoots under phosphorus deficiency. Front Plant Sci 6:1014. https://doi.org/10.3389/fpls.2015.01014
Nianiou-Obeidat I, Madesis P, Kissoudis C, Voulgari G, Chronopoulou E, Tsaftaris A, Labrou NE (2017) Plant glutathione transferase-mediated stress tolerance: functions and biotechnological applications. Plant Cell Rep 36(6):791–805. https://doi.org/10.1007/s00299-017-2139-7
Omer S, Kumar S, Khan BM (2013) Over-expression of a subgroup 4 R2R3 type MYB transcription factor gene from Leucaena leucocephala reduces lignin content in transgenic tobacco. Plant Cell Rep 32(1):161–171. https://doi.org/10.1007/s00299-012-1350-9
Okazaki Y, Otsuki H, Narisawa T, Kobayashi M, Sawai S, Kamide Y, Kusano M, Aoki T, Hirai MY, Saito K (2013) A new class of plant lipid is essential for protection against phosphorus depletion. Nat Commun 4:1510. https://doi.org/10.1038/ncomms2512
Pontigo S, Ulloa M, Godoy K, Nikolic N, Nikolic M, Mora ML, Cartes P (2018) Phosphorus efficiency modulates phenol metabolism in wheat genotypes. J Plant Nutr Soil Sc 18:904–920. https://doi.org/10.4067/S0718-95162018005002603
Qu B, He X, Wang J, Zhao Y, Teng W, Shao A, Zhao X, Ma W, Wang J, Li B, Li Z, Tong Y (2015) A wheat CCAAT box-binding transcription factor increases the grain yield of wheat with less fertilizer input. Plant Physiol 167(2):411–423. https://doi.org/10.1104/pp.114.246959
Rahantaniaina MS, Tuzet A, Mhamdi A, Noctor G (2013) Missing links in understanding redox signaling via thiol/disulfide modulation: how is glutathione oxidized in plants? Front Plant Sci 4:477. https://doi.org/10.3389/fpls.2013.00477
Sun T, Busta L, Zhang Q, Ding P, Jetter R, Zhang Y (2018) TGACG-BINDING FACTOR 1 (TGA1) and TGA4 regulate salicylic acid and pipecolic acid biosynthesis by modulating the expression of SYSTEMIC ACQUIRED RESISTANCE DEFICIENT 1 (SARD1) and CALMODULIN-BINDING PROTEIN 60g (CBP60g). New Phytol 217(1):344–354. https://doi.org/10.1111/nph.14780
Srivalli S, Khanna-Chopra R (2008) Role of glutathione in abiotic stress tolerance. In: Khan NA, Singh S, Umar S (eds) Sulfur assimilation and abiotic stress in plants. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-540-76326-0_10
Schmittgen T, Livak K (2008) Analyzing real-time PCR data by the comparative CT method. Nat Protoc 3:1101–1108. https://doi.org/10.1038/nprot.2008.73
Schachtman DP, Reid RJ, Ayling SM (1998) Phosphorus uptake by plants: from soil to cell. Plant Physiol 116(2):447–453. https://doi.org/10.1104/pp.116.2.447
Tamagnone L, Merida A, Parr A, Mackay S, Culianez-Macia FA, Roberts K, Martin C (1998) The AmMYB308 and AmMYB330 transcription factors from antirrhinum regulate phenylpropanoid and lignin biosynthesis in transgenic tobacco. Plant Cell 10(2):135–154. https://doi.org/10.1105/tpc.10.2.135
Vanholme R, Morreel K, Ralph J, Boerjan W (2008) Lignin engineering. Curr Opin Plant Biol 11(3):278–285. https://doi.org/10.1016/j.pbi.2008.03.005
Vance CP, Uhde-Stone C, Allan DL (2003) Phosphorus acquisition and use: critical adaptations by plants for securing a nonrenewable resource. New Phytol 157(3):423–447. https://doi.org/10.1046/j.1469-8137.2003.00695.x
Wang Y, Lysøe E, Armarego-Marriott T, Erban A, Paruch L, van Eerde A, Bock R, Liu-Clarke J (2018) Transcriptome and metabolome analyses provide insights into root and root-released organic anion responses to phosphorus deficiency in oat. J Exp Bot 69(15):3759–3771. https://doi.org/10.1093/jxb/ery176
Wu P, Shou H, Xu G, Lian X (2013) Improvement of phosphorus efficiency in rice on the basis of understanding phosphate signaling and homeostasis. Curr Opin Plant Biol 16(2):205–212. https://doi.org/10.1016/j.pbi.2013.03.002
Xiao Z, Sun X, Liu X, Li C, He L, Chen S, Su J (2016) Selection of reliable reference genes for gene expression studies on Rhododendron molle G. Don Front Plant Sci 7:1547. https://doi.org/10.3389/fpls.2016.01547
Yang Xu, Shi Q, Gong B (2020) Characterization of COMT1-mediated low phosphorus resistance mechanism by metabolomics in tomato plants. Environ Exp Bot 179:104187. https://doi.org/10.1016/j.envexpbot.2020.104187
Yu B, Xu C, Benning C (2002) Arabidopsis disrupted in SQD2 encoding sulfolipid synthase is impaired in phosphate-limited growth. P Natl Acad Sci USA 99(8):5732–5737. https://doi.org/10.1073/pnas.082696499
Zhu SN, Chen ZJ, Xie BX, Guo Q, Chen MH, Liang CY, Bai ZL, Wang XR, Wang HC, Liao H, Tian J (2021) A phosphate starvation responsive malate dehydrogenase, GmMDH12 mediates malate synthesis and nodule size in soybean (Glycine max). Environ Exp Bot, Volume 189:104560, ISSN0098-8472. https://doi.org/10.1016/j.envexpbot.2021.104560
Zhao M, Guo R, Li M, Liu Y, Wang X, Fu H, Wang S, Liu X, Shi L (2020) Physiological characteristics and metabolomics reveal the tolerance mechanism to low nitrogen in Glycine soja leaves. Physiol Plantarum 168:819–834. https://doi.org/10.1111/ppl.13022
Zhang S, Tang D, Korpelainen H, Li C (2019) Metabolic and physiological analyses reveal that Populus cathayana males adopt an energy-saving strategy to cope with phosphorus deficiency. Tree Physiol 39(9):1630–1645. https://doi.org/10.1093/treephys/tpz074
Zhang J, Yang D, Li M, Shi L (2016) Metabolic profiles reveal changes in wild and cultivated soybean seedling leaves under salt stress. PLoS One 11(7):e0159622. https://doi.org/10.1371/journal.pone.0159622
Acknowledgments
We thank International Science Editing (http://www.internationalscienceediting.com) for editing this manuscript.
Funding
This work was supported by the National Natural Science Foundation of China (No. 32072012) and Natural Science Foundation of Jilin Province, China (No. 20200201134JC).
Author information
Authors and Affiliations
Contributions
Software: Jing Chen and Ji Zhou; Project administration, Methodology, Writing-review & editing, Supervision: Lianxuan Shi and Tao Zhang; Data curation: Jing Chen, Ji Zhou and Mu Li; Formal analysis: Jing Chen, Yunan Hu, and Mingxia Li.
Corresponding authors
Ethics declarations
Competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Responsible Editor: Jiayin Pang.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information

Figure S3
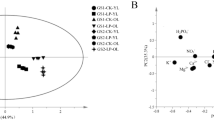
Changes in Additional Outcomes. Line diagrams showing changes in scores on a) the Word List Total Recall, b) the NIH Toolbox Fluid Cognition Composite, c) PROMIS anxiety, d) PROMIS depression, and e) PROMIS Satisfaction with Social Roles in the active and sham tDCS groups across baseline and follow-up (x-axis). Bars represent 95% confidence intervals (PNG 27 009 kbIntegration network of metabolites and genes in cultivated soybean seedling roots under low-phosphorus stress. Genes and metabolites are represented by numbers and letters, respectively. 1, Glyma.17G070500.Wm82.a2.v1; 2, Glyma.11G171400.Wm82.a2.v1; 3, Glyma.06G158700.Wm82.a2.v1; 4, Glyma.10G194800.Wm82.a2.v1; A, glutamic acid; B, proline; C, putrescine; D, pyruvic acid; E, 4-aminobutyric acid; F, alpha-ketoglutaric acid; G, asparagine; H, succinic acid; I, fumaric acid; J, aspartic acid; K, citric acid; L, salicylic acid. The thicker the edge is, the stronger the correlation is. The size of a node is proportional to the correlation between nodes (PNG 359 kb)
Table S1
Primers for qRT-PCR of genes in wild and cultivated soybean seedling roots (DOCX 14 kb)
Table S2
KEGG pathway of DEGs in wild and cultivated soybean seedling roots (DOCX 21 kb)
Table S3
Changes of transcription factors in wild and cultivated soybean seedling roots under low-phosphorus stress (DOCX 18 kb)
Table S4
Contribution rate of wild and cultivated soybean seedling root metabolites to the first and second principal components (PC1 and PC2, respectively) (DOCX 25 kb)
Table S5
DEGs co-expressed in wild and cultivated soybean seedling roots under low-phosphorus stress (DOCX 15 kb)
Table S6
KEGG annotation, GO annotation and log2(LP/CK) for some DEGs in cultivated soybean seedling roots under low-phosphorus stress (DOCX 14 kb)

Supplementary Figure 1.
qRT-PCR of genes in the roots of wild and cultivated soybean seedlings (PNG 57.1 KB)

Supplementary Figure 2.
DEGs of wild and cultivated soybean seedling roots under low-phosphorus stress and control: (a)Venn and (b) volcano diagrams (PNG 132 KB)
Rights and permissions
About this article
Cite this article
Chen, J., Zhou, J., Li, M. et al. Membrane lipid phosphorus reusing and antioxidant protecting played key roles in wild soybean resistance to phosphorus deficiency compared with cultivated soybean. Plant Soil 474, 99–113 (2022). https://doi.org/10.1007/s11104-022-05316-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-022-05316-5