Abstract
Aims
Numerous organisms show range expansions in response to current climate change. Differences in expansion rates, such as between plants and soil biota, may lead to altered interactions in the new compared to the original range. While plant-soil interactions influence plant performance and stress tolerance, the roles of specific soil organisms driving these responses remain unknown.
Methods
We manipulated the abundances of nematodes and arbuscular mycorrhizal fungi (AMF), collected from original and new range soils, and examined their effects on the biomass of range-expanding Centaurea stoebe and native Centaurea jacea. In the first approach, nematode and AMF communities were extracted from field soils, and inoculated to sterilized soil. In the second approach, the abundance of soil organisms in soil inocula was reduced by wet sieving; at first, plants were grown to condition the soil, and then plant-soil feedback was determined under ambient and drought conditions.
Results
The origin of soil communities did not influence the biomass production of range-expanding or native plant species, neither by addition nor by (partial) removal. However, after conditioning and under drought, range expanding C. stoebe produced more biomass with soil communities from the original range while C. jacea, native to both ranges, produced more biomass with new range soil communities.
Conclusions
We show that nematode and AMF communities from original and new range have similar effect on the growth of range expanding C. stoebe. Our results highlight that the effect of soil communities on plant growth increases after soil conditioning and under drought stress.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Current human-induced climate change has enabled many species to expand their range to higher altitudes and latitudes (Parmesan 2006). Species differ in their expansion rates (Berg et al. 2010), so that naturally co-evolved biotic interactions may be replaced by new interactions in the new range. Plants are expected to show higher expansion rates than that exhibited by soil organisms (Berg et al. 2010; Álvarez-Garrido et al. 2019). As the diversity and composition of soil organisms, such as soil-borne fungi and bacteria, changes with latitude (Bahram et al. 2018; Ramirez et al. 2019; Benning and Moeller 2020), range-expanding plant species are expected to establish interactions with different soil communities in their new range than in their original range.
Range-expanding plant species (or neonatives, sensu Essl et al. 2019), which spread within a continent without human-assistance, have shown to experience different interactions with soil communities in their original and new range. More specifically, range-expanding plant species have shown to be less negatively influenced by soil biota from their new, northern range than by soil biota from their original, southern range (van Grunsven et al. 2010; De Frenne et al. 2014). Previous studies have estimated how soil communities that are modified by a plant species subsequently change plant growth. This approach where soils first are conditioned by certain plants, after which the feedback effects to plants of the same or other species are tested is known as plant-soil feedback (Bever et al. 1997; Kulmatiski et al. 2008; van der Putten et al. 2013). In such plant-soil feedback experiments, range-expanding plant species have found to be less negatively affected by soil biota than congeneric plant species that are native in the expansion range (van Grunsven et al. 2007; Engelkes et al. 2008; Dostálek et al. 2016). These findings suggest that release from pathogenic components in soil communities may benefit the performance of range-expanding plant species in the new range, which is also known for introduced exotic plant species (Keane and Crawley 2002; Bardgett and van der Putten 2014). Alternatively, improved growth in the new range might be explained by a more successful association with beneficial mutualists (Callaway et al. 2011). These enemy and mutualist-based mechanisms are not mutually exclusive and can occur simultaneously. At the same time, it is expected that the interactions between plants and mutualistic organisms have weaker effects than those of pathogens due to the relatively low host-specificity of mutualistic symbionts (Pugnaire et al. 2019), but empirical evidence for this is sparse.
A recent study has demonstrated that the composition of nematode communities differs between the original and new range of range-expanding plant species (Wilschut et al. 2019a). Although nematode communities are commonly dominated by taxa that feeds on bacteria, root-feeding nematodes can be equally abundant (van den Hoogen et al. 2019). Hence, nematode communities can directly and indirectly, by modifying plant species interactions with other soil organisms, influence plant growth (Wilschut and Geisen 2020). In their new range, range-expanding plant species have shown to accumulate fewer root-feeding nematodes than their closely related natives, although this pattern can vary between plant species (Morriën et al. 2012; Wilschut et al. 2017). As root-feeding nematodes may play a crucial role in negative plant-soil feedback effects (Wilschut et al. 2019b), they can represent one group of soil-borne enemies from which range-expanding plant species may become released in their new range (Keane and Crawley 2002).
At the same time, range-expanding plant species are effectively associating with arbuscular mycorrhizal fungi (AMF) in their new range (van Grunsven et al. 2014). These fungi from subphylum Glomeromycotina (Spatafora et al. 2016) are obligate root symbionts that receive carbon compounds from the plant and in return improve plant nutrient acquisition but also increase host plant resistance to biotic and abiotic stress (Smith and Read 2008). Consequently, they can influence plant performance directly, but also indirectly, via altered interactions between the host plant and other soil organisms. For example, AMF can increase plant resistance to nematodes and thereby reduce the abundance of these antagonists (Veresoglou and Rillig 2012; Vos et al. 2013). Although a recent study has suggested that there is no significant change in the abundance of AMF associated to range-expanding plant species between the original and new range (Ramirez et al. 2019), the functioning of these fungi associated to range-expanding plant species has not yet been evaluated.
Plant-soil interactions are also influenced by changes in the abiotic environment, and may therefore be sensitive to climate change. Drought events, which are expected to become longer and more frequent (Knapp et al. 2008; Jimenez et al. 2011), can directly trigger changes in soil communities and also indirectly modify soil communities through changes in plant physiology (Meisner et al. 2013; Kaisermann et al. 2017; de Vries et al. 2020). As such, droughts are proposed to amplify the importance of AMF and decrease the effect of root herbivores in determining plant-soil feedback (Pugnaire et al. 2019). However, these effects can be ecosystem and organism group specific: for example, the populations of root-feeding nematodes in mesic grasslands are expected to increase when extremely dry years become more common (Franco et al. 2019).
Here we use a combination of experimental approaches to disentangle the effects of specific groups of soil organisms from the original and new ranges on the growth of a range-expanding plant species, Centaurea stoebe; we also test if these interactions change in response to soil conditioning and drought. More specifically, we focus on the interactions between plants, nematodes, AMF and other microbes. In a first experiment, we estimated the influence of nematodes and AMF on plant growth by inoculating sterilized soil with nematode and AMF communities. The nematodes and AMF were obtained from field soil in both the original and new range of the range expander. In a second experiment, we used sieves of different mesh sizes (Wagg et al. 2014) to reduce the abundance of AMF and nematodes in original and new range soil. Both these approaches enable us to evaluate the role of nematode and AMF communities from the original and new ranges on the biomass production of range-expanding plant species and thereby examine how these soil organisms may contribute to the changes in plant-soil interactions that have been observed for range-expanding plant species (van Grunsven et al. 2010). Addressing community-level interactions enables to also involve AMF taxa that cannot be grown in cultures (Ohsowski et al. 2014) but that may influence plant growth in a species-specific way (Klironomos 2003; Kiers et al. 2011). Examining nematode impacts on plant performance using nematode communities, instead of single root-feeding nematode taxa, is important, as the susceptibility of root-feeding nematodes to AMF may depend on their mode of parasitism (Hol and Cook 2005) and root-feeding nematodes can show distinctive responses to host plant identity (Wilschut et al. 2017).
We used Centaurea stoebe to test the effects of nematode and AMF communities from its original and new range as this plant species has expanded its range from Southern-Central Europe to North-Western Europe during the past 50 years (NDFF 2019). Centaurea stoebe has been shown to repel root-feeding nematodes, presumably due to its distinctive profile of root volatiles (Wilschut et al. 2017). Such a distinctive profile of root chemicals may also influence other groups of soil organisms when they lack a co-evolutionary history with the plant species. We tested the hypotheses that: i) nematode communities from the original range reduce the biomass of C. stoebe more than nematode communities from the new range; ii) AMF communities from the new range have a more positive effect on C. stoebe than AMF communities from the original range; iii) AMF reduce the negative effect of nematodes and this effect is stronger in the original range; iv) the positive effect of AMF communities from new range increases after soil conditioning and during drought. We compared the growth of C. stoebe with the growth of Centaurea jacea, which is a congeneric species of the range-expander that is native in both the original and new range of the range-expander. We expected C. jacea to have co-evolutionary history with soil communities from both the original and new range, leading to similar interactions with nematodes and AMF from both ranges, also after conditioning and during drought.
Methods
Origin of seeds and soil
Seeds of both C. stoebe and C. jacea were collected from wild populations in the Netherlands. Seeds were surface-sterilized in a 0.5% hypochlorite solution and germinated in sterilised medium. Emerged seedlings were planted in pots directly after the inoculation of soil organisms and dead seedlings were replaced only during the first two weeks.
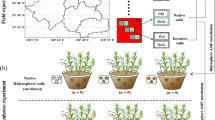
We collected soil from Slovenia, where the range expanding C. stoebe is native (representing the original range), and from the Netherlands where the species recently has established (representing the new range). In both countries, we collected soil from 3–15 cm below the soil surface from a riverine grassland, avoiding the microsites where either of the plant species occurred at that moment. Soil was collected from Slovenia and the Netherlands at the end of May 2014, stored cool and transported to the laboratory of NIOO-KNAW in Wageningen, the Netherlands. Soil collected from each range (either original or new range) was mixed before the experiment to create uniform soil communities per range. Thereafter specific groups of soil organisms were isolated, or the abundances of soil organisms were reduced, using the approaches described below. Additionally, soil from a similar riverine grassland in the Netherlands was collected, sieved using 4 mm mesh, homogenized and gamma-sterilized (> 25 KGray) at Steris AST (Ede, The Netherlands). This sterilized soil was used as a background substrate for adding all specific organism groups or soil fractions as described below.
Overview of experimental approach
We used two approaches to estimate separate and combined effects of nematode and AMF communities from the original and new range on the growth of range-expanding plant species. In Experiment I, we first isolated nematodes and AMF from the field-collected soil and added them, separately or combined, to the sterilized background soil. After 16 weeks, we determined the biomass production of C. stoebe and C. jacea grown in these inoculated soils. In Experiment II, we first reduced the abundance of nematodes and AMF from field-collected soil by a wet-sieving method (Wagg et al. 2014). Then we inoculated the sterilized soil with these wet-sieved soil communities and determined biomass production of the test plants grown in these soils after 13 weeks. Because isolation of nematode and AMF communities and manipulation of soil communities with wet sieving may also influence the communities of small-size non-target microbes, the effect of non-target microbes was also tested in both experiments. During plant growth, soil biota had the opportunity to increase in abundance, similarly to a conditioning phase in a plant-soil feedback experiment (Bever et al. 1997). After harvesting the plants in Experiment II, the soil in the pots was split into two halves: one half was subjected to drought and the other was kept under regular water regime. As in the feedback phase of a plant-soil experiment, a second generation of plants was grown in the conditioned soils and biomass production was determined after 8 weeks.
Experiment I: Addition of soil organisms
The design of the first experiment consisted of a factorial combination of five treatments: plant species (range expanding C. stoebe or native C. jacea), origin of soil organisms (original or new range of the range expander), inoculation of nematodes (yes or no), inoculation of AMF (yes or no) and inoculation of non-target microbes (yes or no). Each treatment combination was replicated 5 times. In addition, there were 5 replicate pots with each plant species grown in sterilized soils, serving as a control. This resulted in 170 one-litre pots in total.
As we were interested in biotic effects, all soil communities were inoculated into 630 g of background soil, which had been sterilized as described above. To test the effect of AMF, we extracted AMF spores from soils of both origins following the protocol of International Culture Collection of (Vesicular) Arbuscular Mycorrhizal Fungi (INVAM). We aimed to apply spores from 70 g of soil per pot, but as the numbers of spores in inocula were variable, we applied a fixed amount (100 spores) to each pot. Collected spores were stored at 4 ˚C in 1 ml distilled water until they were pipetted next to seedling roots during planting. To test the effect of the non-target microbial community, we suspended 70 g of soil in 200 ml of demineralized water per pot, and left it settle for half an hour so that large particles sank. Then, the supernatant was sieved using a 20 μm mesh. The treatment combination with the addition of only non-target microbes was prepared as the small fraction treatment in Experiment II (below). All pots that did not receive this non-target microbial community received 200 ml of distilled water when seedlings were planted in order to ensure that all pots had received the same amount of water. Plants, non-target microbial communities and AMF communities were allowed to establish for two weeks before nematodes were added to avoid AMF spores being damaged by nematodes before germination. Nematode communities used in the experiment were collected from soil samples (using 70 g of soil per pot) using an Oostenbrink elutriator (Oostenbrink 1960). Nematodes were added to the pots assigned to nematode treatment using a volume of 200 ml of tap water while other pots received the same amount of tap water.
Experiment II: Reduction of the abundance of soil organisms
Conditioning phase
The design of the second experiment consisted of factorial combination of three treatments: plant species identity (range expanding C. stoebe or native C. jacea), origin of soil organisms (original or new range of range expander) and fraction size of soil organisms (full, medium and small). Each treatment combination was replicated five times. In addition, 5 replicates of each species were grown in sterilized soil, resulting into 70 pots in total. As in the first experiment, we were interested only in biotic effects, and we established pots with 560 g (dry weight) of sterilized background soil and 70 g of alive soil inocula from either original or new range. Additionally, 70 g of sterilized soil inocula from the opposite range of alive soil inocula was added to test the effect of soil communities in uniform abiotic conditions. To obtain sterilized soil inocula, range-specific soil inocula were gamma-sterilized as indicated above. Plants in control treatments were grown in 100% gamma-sterilized soil. We decreased the abundance of soil organisms in an experimental unit by sieving 70 g of live inoculum soil (mixed in 200 ml water) with either 1000 µm, 50 µm or 20 µm size mesh. The soil that did not pass selected mesh size was sterilized by two autoclaving cycles (30 min at 121ºC with 24 h between the cycles) and mixed with the sterilized soil in pots before seedlings were planted. Although such sieving method influences soil communities in general, selected mesh sizes have shown to have greatest effect on specific groups of soil organisms (Wagg et al. 2014). Specifically, we expect 1000 µm (full) fraction to consist of the total soil community, 50 µm (medium) fraction to contain decreased abundance of AMF, and 20 µm (small) fraction to have non-target microbes and decreased abundance of AMF and nematodes (Wagg et al. 2014). Seedlings were planted on 26th of May 2014. After above- and belowground plant biomass was harvested, soil from each pot was kept separately in a plastic bag at 4 ºC and used in the feedback phase described below.
Feedback phase
The experimental design of the feedback phase consisted of factorial combinations of four treatments: drought (yes or no), plant species identity that was constant in the conditioning and feedback phase (range expanding C. stoebe or native C. jacea), origin of soil organisms (original or native range of the range expander) and fraction size (full, medium, small and sterilized). In addition, 20 pots (5 replicates for each plant species for original and new range) were included as a control. To maintain similar abiotic conditions for control plants, these plants were grown in soil collected from pots where plants were growing with non-target microbes in the Experiment I and this was sterilised by two cycles of autoclaving as indicated above before the start of the feedback phase. Each treatment combination had five replicates, resulting into 160 pots in total.
Soil from each pot from the conditioning phase of the Experiment II was divided into two halves of 350 g each. One-litre pots were thereafter filled with the equal parts (350 g) of inocula from the first phase and sterilized soil (collected and sterilized with gamma irradiation as indicated above). To evaluate whether the interactions between plant species and soil organisms change in extreme weather conditions, we subjected half of the pots to drought. By weighing the pots and providing water up to the pre-specified weight, the water level in drought treatments was kept at 30% of dry weight while water level at regular conditions was kept at 60%.
AMF colonisation
The root colonisation by AMF was estimated after harvesting Experiment I and after harvesting the conditioning phase of the Experiment II using magnified grid-line intersection method (McGonigle et al. 1990). Stained roots were mounted on microscope slides and the percentage of root length colonised by AMF was estimated by scoring the presence or absence of AMF structures at each intersection of root and the vertical crosshair for 120 intersections per plant individual. An intersection was considered mycorrhizal if the vertical crosshair intersected a hypha, an intercellular coil, an arbuscule or a vesicle. Counting was performed using an Olympus CH20 microscope at 400 × magnification.
Statistical analyses
All plant biomass was dried at 70 ˚C to constant weight and weighed. To analyse the effect of soil organisms on plant growth, we compared the growth response of each plant in a treatment with growth in sterilized conditions, using random pairing of treatment and control plants; more specifically calculating ln(treatment/control) following Brinkman et al. (2010). This paired approach was used in all experiments. Using this paired approach, negative values indicate growth reduction by soil organisms compared to sterilized conditions while positive values indicate growth promotion. Aboveground and belowground biomass response to treatments was analysed using ANOVA with all experimental treatments as fixed factors. More specifically, fixed factors in Experiment I were plant species identity (C. jacea, C. stoebe), the origin of soil organisms (original and new range), inoculation with nematodes (yes or no), AMF (yes or no), non-target microbes (yes or no). In the conditioning phase of the Experiment II the fixed factors were plant species identity, origin of soil organisms and fraction size (full, medium, small). In the feedback phase of the Experiment II, linear mixed-effects models, using package “lmerTest” (Kuznetsova et al. 2017), were used to analyse plant biomass response. In these models, plant species identity, origin of soil organisms, the fraction size of soil communities and drought (yes or no) served as fixed factors and pot number from the conditioning phase as random factor (Brinkmann et al. 2010).
AMF colonisation in control pots was always 0 (data not shown), therefore we analysed the change of AMF colonization to the addition of soil organisms (Experiment I) and to the wet-sieving treatments (Experiment II). As the abundances of different AMF structures were highly correlated, the analyses of total AMF colonisation (presence of hyphae, arbuscules or vesicules) is presented. To meet the assumptions of general linear models, AMF colonisation was arcsine-transformed prior to the analyses, while measured colonisation % is presented as mean values. The colonisation of AMF in plant roots was analysed using ANOVA with all experimental factors as fixed factors as for the analyses of plant biomass response. In case of a significant interaction between the fixed factors, the difference between all treatment combinations was estimated using Tukey’s HSD test. The composition of AMF and general fungal communities were also screened with molecular methods (see Supplementary information). All statistical analyses were performed in R (version 3.6.3, R Core Development Team 2020).
Results
Experiment I: Addition of soil organisms
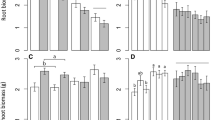
The addition of soil organisms reduced the aboveground biomass of range expanding Centaurea stoebe more than that of the native Centaurea jacea (Table 1, mean ± SE: -0.29 ± 0.04 and -0.23 ± 0.03 respectively). Plant above- and belowground biomass were not influenced by the origin of soil organisms (Table 1). The addition of non-target microbes reduced above-and belowground biomass of both plant species (Table 1, Fig. 1). Plant belowground biomass was also influenced by the interaction between nematodes and AMF (Table 1) but Tukey HSD test did not detect significant differences between treatment combinations (Fig. 1S-a).
Above- and belowground biomass response (left and right panel, respectively) of native plant species Centaurea jacea and range-expanding plant species Centaurea stoebe to the inoculation of nematode (a, b), arbuscular mycorrhizal fungal (c, d) and non-target microbial (e, f) communities. Biomass response to the presence of soil organisms was calculated as ln(treatment/control) following Brinkman et al. (2010), so that values above zero indicate positive and below zero negative response. Boxplot solid line represents the mean, box represent the values between 25 and 75th percentile. Letters indicate significant difference between treatments according to ANOVA analyses (Table 1)
AMF colonisation differed significantly between plant species (Table S1), being higher in range expanding C. stoebe than in native C. jacea (mean % of colonized roots ± SE being 27.45 ± 11.82 and 22.93 ± 11.17, respectively). AMF colonisation in plant roots was influenced by the interaction between the addition of AMF spores and non-target microbes (Table S1). The addition of AMF spores resulted in increased AMF colonisation in the absence but not in the presence of non-target microbes (Fig. 2S-a). The origin of non-target microbes also influenced AMF colonisation (Table S1). In the absence of non-target microbes, there was more AMF colonisation in soil with inocula from the new range than from the original range, but this difference disappeared when non-target microbes were added (Fig. 2S-b). There was also significant interaction between the addition of AMF spores and nematodes (Table S1). The addition of AMF spores resulted in similar AMF colonisation in the presence and absence of nematodes but when AMF spores were not added, AMF colonisation in plant roots was significantly lower in the presence than in the absence of nematodes (Fig. 1S-b).
Above- and belowground biomass response (left and right panel, respectively) to the presence of soil communities from different size fractions, displayed separately for native Centaurea jacea and range expanding Centaurea stoebe. The size of the fraction in wet sieving treatments is < 1000 µm (full fraction), < 50 µm (medium fraction) and < 20 µm (small fraction). Biomass response was calculated as ln(treatment/control) following Brinkman et al. (2010), so that values above zero indicate positive and below zero negative response. Boxplot solid line represents the mean, box represent the values between 25 and 75th percentile. Letters indicate significant difference between treatments in Table 2 according to Tukey HSD test
Experiment 2: Reducing the abundance of soil organisms
Conditioning phase
Aboveground biomass of plants did not differ significantly by species, soil origin or the size of the soil community fraction (Table 2, Fig. 2). Plant belowground biomass was significantly influenced by the size of the soil community fraction (Table 2). Belowground biomass of plants was significantly lower in the presence of soil communities from full (< 1000 µm) compared to medium (< 50 µm) or small-sized (< 20 µm) fraction (Fig. 2).
AMF colonisation of plant roots was influenced by the interaction between plant species and the size of the soil community fraction (Table S2). Soil communities from the small fraction reduced AMF colonisation more in the roots of C. jacea than in the roots of C. stoebe (Fig. 3S-a). AMF colonisation was also influenced by the interaction between range and the size of the soil community fraction (Table S2), as in the original range, plants growing with medium size soil communities had significantly lower AMF colonisation than plants growing with full soil communities and there was no significant difference between medium and small-size soil community fraction. In the new range soil, plants growing with full and medium size soil communities had similar AMF colonization but this was significantly lower in small-size soil community fraction (Figure 3S-b).
Feedback phase
In the feedback phase, aboveground biomass was influenced by the three-way interaction between drought, plant species identity and soil origin (Table 3). Tukey post-hoc comparison revealed that range expanding C. stoebe produced more biomass under drought when growing with original range soil communities (Fig. 3). On the other hand, C. jacea, that is native in both ranges, produced more biomass under drought with soil communities from the new range of C. stoebe (Fig. 3).
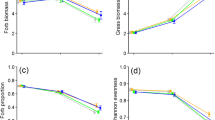
Above-and belowground biomass response of native Centaurea jacea (left panels) and range expanding Centaurea stoebe (right panels) to the presence of conditioned soil organisms from different ranges in ambient conditions and in drought. Biomass response to the presence of soil organisms was calculated as ln(treatment/control) following Brinkman et al. (2010), so that values above zero indicate positive and below zero negative response. Boxplot solid line represents the mean, box represent the values between 25 and 75th percentile. Letters indicate significant difference between treatments according to Tukey HSD test (p < 0.05)
There was a significant effect of drought on plant belowground biomass (Table 3). This indicated that soil communities promoted belowground biomass more under drought than under the regular water regime (mean ± SE 0.58 ± 0.07 and 0.37 ± 0.07 for drought and ambient conditions respectively). There was a significant interaction between plant species and soil origin (Table 3). This was because C. stoebe produced more belowground biomass when growing with soil communities from the original range than with soil communities from the new range, whereas there was no significant difference in the effect of soil community origin on the belowground biomass of native C. jacea (Fig. 3).
Discussion
The possible role of soil organisms in the success of range-expanding plant species is well acknowledged (Engelkes et al. 2008; van der Putten et al. 2013; Dostálek et al. 2016; van Nuland et al. 2017). However, most of the available evidence suggesting the important role of soil organisms is obtained by observational studies comparing the composition of soil organisms between ranges (Ramirez et al. 2019; Wilschut et al. 2019a), or by experimental studies that examine plant performance in response to complete soil communities (van Grunsven et al. 2010; Manrubia et al. 2019). Here we have taken a step further by studying the impacts of natural nematode and AMF communities, important groups of plant-associated organisms, from the original and new range on the growth of range-expanding C. stoebe in an experimental setting. In the present study, we found that biomass production of the range-expanding plant species did not differ, irrespective of the addition or reduced abundance of nematodes or AMF.
The results of the present study did not provide evidence for our first hypothesis that nematodes from the original range will suppress the growth of range expanding C. stoebe more than nematodes from the new range. In its original range, C. stoebe has been shown to associate with higher numbers of root-feeding nematodes compared to its new range (Wilschut et al. 2019a, 2020). Perennial plants, such as C. stoebe, occupy the same location for years and therefore may have well-developed interactions with soil organisms in natural habitats. The use of field soil where none of the study species was present during soil collection can be one of the reasons explaining the lack of different effects of nematode communities from the original and new range in this study. Alternatively, the lack of the effect of nematodes on plant growth might be driven by low abundances of (root-feeding) nematodes in our treatments as our selected methods enabled to test the interactions between plants and nematode communities, but did not enable to control for the abundance of nematodes in the soil. Low nematode pressure on plant growth is additionally supported by the results of our second experiment that demonstrated similar plant growth with soil communities from medium (< 50 µm) and small (< 20 µm) size fraction, although the latter treatment has been shown to reduce nematode abundance (Wagg et al. 2014). At the same time, our findings are in line with previous studies that have reported no change in the biomass of common plant species in response to the addition of nematode communities (Wurst et al. 2009, 2010). To enhance our understanding of the role of nematodes in the success of range-expanding plant species, we need more studies recording plant performance (growth, seed production) and nematode community composition simultaneously, for example by using high-throughput amplicon sequencing (Wilschut and Geisen 2020).
The absence of a positive effect of AMF spore addition on the range expander C. stoebe does not provide support for our second hypothesis that AMF from the new range enhance plant growth more than AMF from the original range. Also, the simultaneous addition of AMF and nematodes did not affect plant biomass production, providing no support for our third hypothesis that AMF reduce a negative effect of nematodes on plant growth. However, such an effect would only be possible to detect if nematode addition would have led to a decrease in plant performance. The addition of AMF spores enabled 16% of the roots of C. stoebe to become colonized by these fungi, which is comparable to AMF colonization recorded in the roots of the range-expanding plant species Tragopogon dubius in the greenhouse experiment with whole-soil inocula (van Grunsven et al. 2014). Higher AMF colonisation in the roots of C. stoebe than in the roots of C. jacea, confirms that range-expanding plant species are as effective as native plant species in associating with AMF (van Grunsven et al. 2014). At the same time, in the first experiment there was lower AMF colonization in the roots of C. stoebe than in the second experiment (where it was between 49–69%). Therefore, we cannot exclude that the lack of the effect of AMF on plant growth in our first experiment is at least partly caused by the low abundance of available propagules or by the absence of specific AMF that do not produce spores (Ohsowski et al. 2014). However, in our second experiment plants showed most biomass reduction when growing with soil communities from full fraction where there was highest AMF colonisation. This pattern suggests that under our experimental conditions, the costs of delivering carbon compounds to AMF are greater for the plant than the benefits they obtain in return (Johnson et al. 1997). Future studies should manipulate not only the presence but also the abundance of AMF and nematodes in soil to examine if the effects of AMF, as well as nematodes and other soil organisms during range expansion depends on their quantity.
There was no difference in the effect of soil communities from the original and new ranges in the first experiment and in the conditioning phase of the second experiment. However, the effect of the origin of soil communities became evident in the feedback phase of Experiment II, in line with a recent experiment that demonstrated that it can take between four and six months to reveal the effect of soil microbes on plant biomass (Wang et al. 2019). Moreover, the origin of soil communities determined plant biomass production under drought, being in line with a recent meta-analyses, that suggests intensified plant-soil interactions under drought (Beals et al. 2020). At the same time, a recent study has reported no differences in community-level biomass of range expanders between drought and ambient conditions (Manrubia et al. 2019), which suggests that in communities, neighbouring plant species might reduce the intensity of plant-soil interactions during drought.
Water availability has shown to influence the functioning of soil communities (Hawkes et al. 2017) and the composition of fungal and bacterial communities in soil with soil communities adapted to contemporary water availability being more beneficial for plant growth (Lau and Lennon 2012; Remke et al. 2020). Thus enhanced growth of range expanding C. stoebe with soil communities from the original range during drought suggests that these soil communities are better adapted to drought conditions than soil communities in the new range. This is further supported by climatic data, as the number of days per year without precipitation is higher in Slovenia, where the original range soil communities were collected, than in the Netherlands, where the new range soil communities originated (climatedata.eu). At the same time, the native C. jacea produced more biomass with soil communities from the new range during drought. As seeds of both plant species used in this experiment were collected from the new range, this pattern can suggest that local adaptation, which is absent in range-expanding plant species, is shaping the plant-soil interactions in native plant species. Recent study has shown that local adaptation in annual range-expanding plant species can occur relatively rapidly (Lustenhouwer et al. 2018). Characterization of leaf traits, biomass production, life cycle and reproductive capacity have indicated post-introduction evolution of C. stoebe in its invasive range compared to its original range (Henery et al. 2010; Hahn et al. 2012). Similar process might occur during range expansion as well, but this remains to be studied. The positive effect of soil communities from the original range on the growth of range-expanding plant species during drought as recorded in the present study suggests that the local adaptation can also have a positive effect on plant growth. Thus, although largely overlooked so far, local adaptation in range-expanding plant species may be an additional factor increasing the abundance of range-expanding plant species in their new range.
To manage the high number of treatments in this study, variation in soil communities was compromised by creating one composite soil sample from the original range and one from the new range of the range expander. Future studies should test the effect of soil communities from different locations within both ranges to validate the general nature of the patterns reported here. However, our results are in line with previous studies showing that soil communities from multiple locations in the original and new range have similar effect on the biomass production of range-expanding plant communities during soil conditioning (Koorem et al. 2018) and feedback phase (Koorem et al. 2020). The effect of soil communities reported in experimental studies may at least partly depend on the methodological approach (Wang et al. 2018), in this study demonstrated by the negative effect of non-target soil microbes found in Experiment I that was not confirmed in Experiment II. We therefore encourage the use of combinations of approaches in future studies in order to gain further insights into the interactions between soil communities and range-expanding plant species. For example, a dilution-to extinction approach using sterilized soil in combination with molecular methods for characterizing soil community composition as used in Yang et al. (2020) could help to get further insight into the role of different soil organisms in determining the performance of range-expanding plant species.
We conclude that the interactions between soil communities and range-expanding plant species are dynamic and might be revealed more clearly under stressful conditions such as drought. Therefore, combining the results of studies using different interacting species, growing conditions and approaches will be of key importance to enhance understanding of the effects of specific groups of soil organisms on the growth of range-expanding plant species.
Availability of data and material
Data will be stored at in-house database of NIOO-KNAW.
Code availability
Code will be available upon request.
References
Álvarez-Garrido L, Viñegla B, Hortal S et al (2019) Distributional shifts in ectomycorrhizal fungal communities lag behind climate-driven tree upward migration in a conifer forest-high elevation shrubland ecotone. Soil Biol Biochem 137:107545. https://doi.org/10.1016/j.soilbio.2019.107545
Bahram M, Hildebrand F, Forslund SK et al (2018) Structure and function of the global topsoil microbiome. Nature 560:233–237. https://doi.org/10.1038/s41586-018-0386-6
Bardgett RD, van der Putten WH (2014) Belowground biodiversity and ecosystem functioning. Nature 515:505–511. https://doi.org/10.1038/nature13855
Beals KK, Moore JAM, Kivlin SN et al (2020) Predicting plant-soil feedback in the field: meta-analysis reveals that competition and environmental stress differentially snfluence PSF. Front Ecol Evol 8:191. https://doi.org/10.3389/fevo.2020.00191
Benning JW, Moeller DA (2020) Microbes, mutualism, and range margins: testing the fitness consequences of soil microbial communities across and beyond a native plant’s range. New Phytol 229(5):2886–2900. https://doi.org/10.1111/nph.17102
Berg MP, Toby Kiers E, Driessen G et al (2010) Adapt or disperse: Understanding species persistence in a changing world. Glob Chang Biol 16:587–598. https://doi.org/10.1111/j.1365-2486.2009.02014.x
Bever JD, Westover KM, Antonovics J (1997) Incorporating the soil community into plant population dynamics: The utility of the feedback approach. J Ecol 85:561–573. https://doi.org/10.2307/2960528
Callaway RM, Bedmar EJ, Reinhart KO et al (2011) Effects of soil biota from different ranges on Robinia invasion: Acquiring mutualists and escaping pathogens. Ecology 92:1027–1035. https://doi.org/10.1890/i0012-9658-92-5-1027
Brinkman PE, van der Putten WH, Bakker EJ, Verhoeven KJF (2010) Plant-soil feedback: Experimental approaches, statistical analyses and ecological interpretations. J Ecol 98:1063–1073. https://doi.org/10.1111/j.1365-2745.2010.01695.x
De Frenne P, Coomes DA, De Schrijver A et al (2014) Plant movements and climate warming: Intraspecific variation in growth responses to nonlocal soils. New Phytol 202:431–441. https://doi.org/10.1111/nph.12672
de Vries FT, Griffiths RI, Knight CG et al (2020) Harnessing rhizosphere microbiomes for drought-resilient crop production. Science 368:1095–9203. https://doi.org/10.1126/science.aaz5192
Dostálek T, Münzbergová Z, Kladivová A, Macel M (2016) Plant–soil feedback in native vs. invasive populations of a range expanding plant. Plant Soil 399:209–220. https://doi.org/10.1007/s11104-015-2688-x
Engelkes T, Morriën E, Verhoeven KJF et al (2008) Successful range-expanding plants experience less above-ground and below-ground enemy impact. Nature 456:946–948. https://doi.org/10.1038/nature07474
Essl F, Dullinger S, Genovesi P et al (2019) A conceptual framework for range-expanding species that track human-induced environmental change. 69:908–919. https://doi.org/10.1093/biosci/biz101
Franco ALC, Gherardi LA, de Tomasel CM et al (2019) Drought suppresses soil predators and promotes root herbivores in mesic, but not in xeric grasslands. Proc Natl Acad Sci U S A 116:12883–12888. https://doi.org/10.1073/pnas.1900572116
Hahn MA, Buckley YM, Müller-Schärer H (2012) Increased population growth rate in invasive polyploid Centaurea stoebe in a common garden. Ecol Lett 15:947–954. https://doi.org/10.1111/j.1461-0248.2012.01813.x
Hawkes CV, Waring BG, Rocca JD, Kivlin SN (2017) Historical climate controls soil respiration responses to current soil moisture. PNAS 114:6322–6327. https://doi.org/10.1073/pnas.1620811114
Henery ML, Bowman G, Mráz P et al (2010) Evidence for a combination of pre-adapted traits and rapid adaptive change in the invasive plant Centaurea stoebe. J Ecol 98:800–813. https://doi.org/10.1111/j.1365-2745.2010.01672.x
Hol WHG, Cook R (2005) An overview of arbuscular mycorrhizal fungi-nematode interactions. Basic Appl Ecol 6:489–503. https://doi.org/10.1016/j.baae.2005.04.001
Jimenez M, Jaksic F, Armesto J et al (2011) Extreme climatic events change the dynamics and invasibility of semi-arid annual plant communities. Ecol Lett 14(12):1227–1235. https://doi.org/10.1111/j.1461-0248.2011.01693.x
Johnson NC, Graham JH, Smith FA (1997) Functioning of mycorrhizal associations along the mutualism-parasitism continuum. New Phytol 135:575–585. https://doi.org/10.1046/j.1469-8137.1997.00729.x
Kaisermann A, de Vries FT, Griffiths RI, Bardgett RD (2017) Legacy effects of drought on plant – soil feedbacks and plant – plant interactions. New Phyt 215(4):1413–1424. https://doi.org/10.1111/nph.14661
Keane RM, Crawley MJ (2002) Exotic plant invasions and the enemy release hypothesis. Trends Ecol Evol 17:164–170. https://doi.org/10.1016/S0169-5347(02)02499-0
Kiers ET, Duhamel M, Beesetty Y et al (2011) Reciprocal rewards stabilize cooperation in the mycorrhizal symbiosis. Science 333(6044):880–883. https://doi.org/10.1126/science.1208473
Klironomos JN (2003) Variation in plant response to native and exotic arbuscular mycorrhizal fungi. Ecology 84:2292–2301. https://doi.org/10.1890/02-0413
Knapp AK, Beier C, Briske DD et al (2008) Consequences of more extreme precipitation regimes for terrestrial ecosystems. BioSci 58:811–821. https://doi.org/10.1641/B580908
Koorem K, Kostenko O, Snoek LB et al (2018) Relatedness with plant species in native community influences ecological consequences of range expansions. Oikos 1–10. https://doi.org/10.1111/oik.04817
Koorem K, Snoek BL, Bloem J et al (2020) Community-level interactions between plants and soil biota during range expansion. J Ecol 108(5):1860–1873. https://doi.org/10.1111/1365-2745.13409
Kulmatiski A, Beard KH, Stevens JR, Cobbold SM (2008) Plant-soil feedbacks: A meta-analytical review. Ecol Lett 11:980–992. https://doi.org/10.1111/j.1461-0248.2008.01209.x
Kuznetsova A, Brockhoff PB, Christensen RHB (2017) lmerTest package: tests in linear mixed effects models. J Stat Soft 82(13):1–26. https://doi.org/10.18637/jss.v082.i13
Lau JA, Lennon JT (2012) Rapid responses of soil microorganisms improve plant fitness in novel environments. PNAS 109:14058–14062. https://doi.org/10.1073/pnas.1202319109
Lustenhouwer N, Wilschut RA, Williams JL et al (2018) Rapid evolution of phenology during range expansion with recent climate change. Glob Chang Biol 24:e534–e544. https://doi.org/10.1111/gcb.13947
Manrubia M, Putten van der W, Weser C et al (2019) Soil functional responses to drought under range - expanding and native plant communities. Funct Ecol 1–15. https://doi.org/10.1111/1365-2435.13453
McGonigle TP, Miller MH, Evans DG et al (1990) A new method which gives an objective measure of colonization of roots by vesicular- arbuscular mycorrhizal fungi. New Phytol 115:495–501. https://doi.org/10.1111/j.1469-8137.1990.tb00476.x
Meisner A, De Deyn GB, de Boer W, van der Putten WH (2013) Soil biotic legacy effects of extreme weather events influence plant invasiveness. Proc Natl Acad Sci 110:9835–9838. https://doi.org/10.1073/pnas.1300922110
Morriën E, Duyts H, van der Putten WH (2012) Effects of native and exotic range-expanding plant species on taxonomic and functional composition of nematodes in the soil food web. Oikos 121:181–190. https://doi.org/10.1111/j.1600-0706.2011.19773.x
NDFF (2019) Verspreidingsatlas. http://verspreidingsatlas.nl (Accessed 25 Oct 2019)
Ohsowski BM, Zaitsoff PD, Öpik M, Hart MM (2014) Where the wild things are: Looking for uncultured Glomeromycota. New Phytol 204:171–179. https://doi.org/10.1111/nph.12894
Oostenbrink M (1960) Estimating nematode populations by some elected methods. In: Sasser, J.N., Jenkins, W.R. (Eds.), Nematology. Univ. of North Carolina Press 85e102
Parmesan C (2006) Ecological and evolutionary responses to recent climate change. AREES 637–671. https://doi.org/10.1146/annurev.ecolsys.37.091305.110100
Pugnaire FI, Morillo JA, Peñuelas J et al (2019) Climate change effects on plant-soil feedbacks and consequences for biodiversity and functioning of terrestrial ecosystems. Sci Adv 2375–2548 https://doi.org/10.1126/sciadv.aaz1834
R Core Development Team (2020) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/
Ramirez KS, Snoek LB, Koorem K et al (2019) Range-expansion effects on the belowground plant microbiome. Nat Ecol Evol 3:604–611. https://doi.org/10.1038/s41559-019-0828-z
Remke MJ, Johnson NC, Wright J et al (2020) Sympatric pairings of dryland grass populations, mycorrhizal fungi and associated soil biota enhance mutualism and ameliorate drought stress J Ecol 1–14. https://doi.org/10.1111/1365-2745.13546
Smith S, Read D (2008) Mycorrhizal symbioses. Third Edition. Academic Press
Spatafora JW, Chang Y, Benny GL et al (2016) A phylum-level phylogenetic classification of zygomycete fungi based on genome-scale data. Mycologia 108:1028–1046. https://doi.org/10.3852/16-042
van den Hoogen J, Geisen S, Routh D et al (2019) Soil nematode abundance and functional group composition at a global scale. Nature 572:194–198. https://doi.org/10.1038/s41586-019-1418-6
van der Putten WH, Bardgett RD, Bever JD et al (2013) Plant-soil feedbacks: The past, the present and future challenges. J Ecol 101:265–276. https://doi.org/10.1111/1365-2745.12054
van Grunsven RHA, van der Putten WH, Bezemer TM et al (2007) Reduced plant-soil feedback of plant species expanding their range as compared to natives. J Ecol 95:1050–1057. https://doi.org/10.1111/j.1365-2745.2007.01282.x
van Grunsven RHA, van der Putten WH, Bezemer TM et al (2010) Plant-soil interactions in the expansion and native range of a poleward shifting plant species. Glob Chang Biol 16:380–385. https://doi.org/10.1111/j.1365-2486.2009.01996.x
van Grunsven RHA, Yuwati TW, Kowalchuk GA et al (2014) The northward shifting neophyte Tragopogon dubius is just as effective in forming mycorrhizal associations as the native T. pratensis. Plant Ecol Divers 7:533–539. https://doi.org/10.1080/17550874.2013.824517
Van Nuland M, Bailey J, Schweitzer J (2017) Divergent plant–soil feedbacks could alter future elevation ranges and ecosystem dynamics. Nat Ecol Evol 1:0150. https://doi.org/10.1038/s41559-017-0150
Veresoglou SD, Rillig MC (2012) Suppression of fungal and nematode plant pathogens through arbuscular mycorrhizal fungi. Biol Lett 8:214–217. https://doi.org/10.1098/rsbl.2011.0874
Vos C, Schouteden N, van Tuinen D et al (2013) Mycorrhiza-induced resistance against the root-knot nematode Meloidogyne incognita involves priming of defense gene responses in tomato. Soil Biol Biochem 60:45–54. https://doi.org/10.1016/j.soilbio.2013.01.013
Wagg C, Bender SF, Widmer F, van der Heijden MGA (2014) Soil biodiversity and soil community composition determine ecosystem multifunctionality. Proc Natl Acad Sci 111:5266–5270. https://doi.org/10.1073/pnas.1320054111
Wang M, De Deyn GB, Bezemer TM (2019) Separating effects of soil microorganisms and nematodes on plant community dynamics. Plant Soil 441:455–467. https://doi.org/10.1007/s11104-019-04137-3
Wang M, Ruan W, Kostenko O et al (2018) Removal of soil biota alters soil feedback effects on plant growth and defense chemistry. New Phytol 221:1478–1491. https://doi.org/10.1111/nph.15485
Wilschut RA, Geisen S (2020) Nematodes as drivers of plant performance in natural systems. Trends Plant Sci 26:237–247. https://doi.org/10.1016/j.tplants.2020.10.006
Wilschut RA, Geisen S, Martens H et al (2019a) Latitudinal variation in soil nematode communities under climate warming-related range-expanding and native plants. Glob Chang Biol 25(8):2714–2726. https://doi.org/10.1111/gcb.14657
Wilschut RA, Magnée KJH, Geisen S et al (2020) Plant population and soil origin effects on rhizosphere nematode community composition of a range - expanding plant species and a native congener. Oecologia 194:237–250. https://doi.org/10.1007/s00442-020-04749-y
Wilschut RA, Silva JCP, Garbeva P, van der Putten WH (2017) Belowground plant–herbivore interactions vary among climate-driven range-expanding plant species with different degrees of novel chemistry. Front Plant Sci 8:1–10. https://doi.org/10.3389/fpls.2017.01861
Wilschut RA, van der Putten WH, Garbeva P et al (2019b) Root traits and belowground herbivores relate to plant–soil feedback variation among congeners. Nat Commun 10:1564. https://doi.org/10.1038/s41467-019-09615-x
Wurst S, Beersum S Van, Wagenaar R et al (2009) Community Plant defence against nematodes is not mediated by changes in the soil microbial community. Funct Ecol 23:488–495. https://www.jstor.org/stable/40205556
Wurst S, Wagenaar R, Biere A (2010) Microorganisms and nematodes increase levels of secondary metabolites in roots and root exudates of Plantago lanceolata. Plant Soil 329:117–126. https://doi.org/10.1007/s11104-009-0139-2
Yang G, Roy J, Veresoglou SD, Rillig MC (2020) Soil biodiversity enhances the persistence of legumes under climate change. New Phytol 2945–2956. https://doi.org/10.1111/nph.17065
Acknowledgements
We thank L.B. Snoek for the help with bioinformatics, Branko Vreš and Marta Manrubia for the help with soil collection. This is publication 7207 of the Netherlands Institute of Ecology (NIOO-KNAW).
Funding
This study was supported by the European Research Council (ERC advanced grant ERC-ADV 323020 (SPECIALS) to W.H.v.d.P.) and by the Estonian Research Council (grants PUTJD78 and MOBTP105).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare no conflicts of interest.
Additional information
Responsible Editor: François Teste.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Koorem, K., Wilschut, R.A., Weser, C. et al. Disentangling nematode and arbuscular mycorrhizal fungal community effect on the growth of range-expanding Centaurea stoebe in original and new range soil. Plant Soil 466, 207–221 (2021). https://doi.org/10.1007/s11104-021-05020-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-021-05020-w