Abstract
Background and aim
Rice and duckweed are two monocotyledonous plants that naturally coexist in paddy fields. While the presence of duckweed in paddy fields significantly improves rice productivity, the interplay between soil microbes and the two plant hosts in this agroecosystem remains unexplored.
Methods
We compared the bacterial community structure between duckweed, rice and soil from multiple rice paddies. We also isolated bacteria from these communities and characterized their modes of bacterial colonization and plant growth-promotion using model plants.
Results
Our data indicate that host-specific and tissue-dependent factors reproducibly orchestrate the bacterial community structure associated with their plant hosts. This is corroborated by results from culture-dependent approaches in which the dominant genus Pantoea isolated from rice aerial tissues can strongly attach and colonize gnotobiotic duckweed in spite of the low representation of this bacterial genus in the natural duckweed microbiome. Our study identified a core of 254 bacterial taxa that are consistently found in all rice and duckweed tissue samples from rice paddy fields. Furthermore, characterizing auxin-producing bacteria isolates from both plant species identified potential plant growth-promoting bacteria that may improve growth for both duckweed and rice in paddy fields.
Conclusions
Results from this work provide evidence for the importance of the host tissue and species context in determining plant colonization by microbes in the paddy field system. The resources generated in this study could facilitate the agronomic deployment of microbes for more sustainable rice production.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Paddy field farming became the dominant form of rice cultivation across the globe from Asia to South America during the 20th Century. However, the large amount of pesticides and fertilizers typically used in modern intensive rice production not only increases the cost of agriculture, but also leads to soil compaction, pesticide residues, environmental pollution and a decline in farmland biodiversity (Benton et al. 2003; Gu et al. 2015). Recent studies demonstrate that integration of multiple species with complementary roles into traditional agricultural systems could minimize the environmental problems associated with intensive rice production (Feng et al. 2016; Zhang et al. 2016; Sha et al. 2017; Yang et al. 2018). The presence of duckweed in flooded rice fields is a common phenomenon (Kumura 2005) and inclusion of duckweed (Lemna minor) in rice paddy agroecosystems has been reported to reduce nitrogen loss from 20–54% (Li et al. 2009). Furthermore, urea combined with the duckweed Spirodela polyrhiza increased rice yields by 9–10%, indicating that duckweed can serve as a green fertilizer to increase yield while simultaneously reducing nitrogen loss in rice production (Yao et al. 2017). In addition to enhancing rice production, duckweed may also suppress rice diseases, reduce the greenhouse gas footprint associated with rice cultivation and remediate heavy metal contaminants in paddy fields (Wang et al. 2015; Ng et al. 2017).
Rice is surrounded by complex microbiota that can influence its growth, nutrition, and health. As ecological environments that combine aquatic and terrestrial components, rice paddy fields contain multiple microbial niches that coexist in one ecologically linked system comprising soil, water, and plant tissues. In addition, each of these niches may also display dynamic behaviors by changing in response to the environment or other biotic factors or both. The composition and structure of the rice microbiome can be affected by factors including drought stress, cultivation patterns and fertilizer applications (Zhu et al. 2016; Santos-Medellín et al. 2017). Microbiomes also vary between different rice tissues and their developmental stages (Edwards et al. 2015). Only limited information is available on how the presence of duckweed can affect community assembly of the rice microbiome. The potential beneficial interplay between multiple plant species, together with the associated microbiomes, provides a basis for the integrated management of traditional agricultural systems such as rice cultivation and raises the possibility that duckweed, together with its associated microbiota, could minimize the environmental problems associated with intensive rice production (Lansing and Kremer 2011; Finkel et al. 2017; Pascale et al. 2020). Bacterial communities of duckweed (Lemna and Spirodela) have recently been characterized (Acosta et al. 2020), and 37 of 47 culturable bacterial strains tested by the Salkowski method were revealed to be potential IAA (indole-3-acetic acid) producing bacteria (Gilbert et al. 2018). These studies indicate that duckweed and its associated bacterial microbiome could be important components of the rice paddy field ecosystem and may impact rice health and productivity.
In this work, we set out to systematically characterize and compare the bacterial microbiota in both duckweed and rice plants from multiple paddy fields in order to gain a better understanding of how host tissues and the environment impact their respective microbial communities. The questions that we wish to address are: 1) How do rice and duckweed bacterial communities compare with their shared environment in paddy field soils? 2) Which bacterial taxa are common between the two plant hosts within and between different paddy environments? 3) Does host selection operate at the stage of bacterial attachment to the plant or does it occur at the level of more complex community dynamics within a particular host? 4) Can microbes isolated from a paddy field environment modulate physiological traits of the plant hosts? Answers to these questions provide a foundation for understanding microbiota dynamics in paddy fields while the characterized resources from this work could pave the way for designing new plant co-cultivation strategies to manipulate and manage the microbial component of paddy field systems for improving rice production.
Materials and methods
Sample collection, processing and duckweed identification
Samples of soil, rice plants (landraces of rice, Oryza sativa) and duckweed were collected randomly from traditional rice paddy fields located in each of three counties in the province of Guizhou, China (Fig. 1). Rice pants were sampled for aerial tissues consisting of leaf blade and stalk pieces far away from the water surface (RA) and roots (RR); duckweed (DW) samples used combined whole fronds with roots due to their small size. Soil samples (SL) from flooded fields around rice roots were also collected from each of the sites to survey the microbial diversity of the local environment. To maintain all samples fresh while transporting to the lab for further process, duckweed was stored in 50 mL sterile Falcon tubes along with paddy water, and rice was stored in a sterile pot containing paddy soil. Nine samples were collected for each type of material (three replicates at each of three locations), with some of the plant tissue samples rinsed with either water alone or with a salt and detergent solution as well, thus resulting in 63 total samples (Table S1). Plant samples treated with two different washes separately included one treatment that was washed with sterile water while the other was washed with salt/detergent solution (137 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, 1.8 mM KH2PO4, 0.5 mM MgSO4, 1 mM CaCl2, 0.1% Triton-X100, pH 7.4), followed by a second wash with sterile water. Species identification of the duckweed samples was carried out by the two-barcode (atpF-atpH and psbK-psbI) approach described in Borisjuk et al. (2014).
Map of field sites. a Guizhou Province (red), China. b Qiandongnan field site locations in Guizhou Province: Leishan (Site 1, square), Congjiang (Site 2, triangle), and Liping county (Site 3, circle). c Typical environment appearance. Rice aerial parts, rice roots, duckweed and soil were collected from each site (Table S1).
Culture-independent bacterial 16S rRNA gene profiling of duckweed, rice and corresponding soil samples
DNA from soil samples and processed plant tissues (duckweed, rice root & rice aerial tissues) were isolated using the Soil Microbe DNA Kit (TIANGEN, China) and the 2x CTAB method, respectively. DNA concentration and purity were quantified using Nanodrop 2000 and were also checked for quality by electrophoresis on 0.5% agarose gels. The V4 region of the bacterial 16S rRNA gene was amplified using primers 515F (5ˊ-GTGCCAGCMGCCGCGGTAA-3ˊ) and 806R (5ˊ-GGACTACHVGGGTWTCTAAT-3ˊ). All PCR reactions were carried out in 30 µL reaction mixture with 15 µL of Phusion® High-Fidelity PCR Master Mix (New England Biolabs, USA), 0.2 µM of forward and reverse primers, and approximately 10 ng template DNA. Thermal cycling consisted of initial denaturation at 98℃ for 1 min, followed by 30 cycles of denaturation at 98℃ for 10 s, annealing at 50℃ for 30 s, elongation at 72℃ for 30 s and final heating at 72℃ for 5 min. An equal volume of 1 × loading buffer (containing SYB green) was mixed with the PCR products and analyzed by electrophoresis on a 2% agarose gel for detection. PCR products were purified using the GeneJETTM Gel Extraction Kit (Thermo Fisher Scientific, USA). Illumina MiSeq DNA sequencing of 16S rRNA gene amplicons was carried out using by the EMSL-JGI (DOE Joint Genome Institute, USA) program and by Novogene Corp. Inc. (CA, USA). Raw reads from this work have been deposited in the Short Read Archive of NCBI under project no. PRJNA545325.
Culturable bacteria isolated from rice and duckweed
Plant-associated bacteria were isolated from surface-sterilized tissues or from plant tissues washed with the salt/detergent solution in an effort to remove loosely bound microbes from the surface. Culturable bacteria associated with rice (labeled Aw and Rt, representing rice aerial parts and rice root tissues, respectively) and duckweed (labeled as Lm) were collected from one location of each site and classified (Table S2). Plant tissues were washed with a salt and detergent containing solution for 20 min and/or a subsequent wash in 10% bleach for 1 min (targeting putative endophytes) and transferred to TSA plates for culture overnight or for up to 2 days at 28 °C to allow bacteria within tissues to grow. The 39 bacterial colonies were then subjected to single colony isolation on subsequent TSA plates. Colonies with different morphological characteristics were purified by repeated streaking and were cryopreserved at -80 °C in 500 µL of LB broth supplemented with 500 µL of 80% glycerol.
Bacterial genotyping and fingerprinting
Cultured bacterial strains were genotyped by standard PCR-based methods using primers targeting the V1-V5 region of 16S rRNA gene (forward primer e9f: 5’-GAGTTTGATCCTGGCTCAG-3’; reverse primer e926r: 5’-CCGTCAATTCCTTTGAGTTT-3’). The PCR reaction mixture contained 2.5 µL of 10 × buffer, 0.2 mM dNTPs, 0.8 µM of each primer, 0.5 µL of 5 U/µL Taq polymerase (Denville Scientific, NJ, USA), ~ 100 ng of template DNA, and filter sterilized water to make a total of 25 µL. The amplification program consisted of 5 min at 95 °C, 25 cycles of 1 min at 95 °C, 30 s at 50 °C, and 1 min at 72 °C, followed by 5 min at 72 °C. PCR products were subjected to electrophoresis in 1% agarose gels for 20 to 30 min and stained with ethidium bromide for visualization. PCR products were purified using ExoSAP-IT™ (Thermo Fisher Scientific, USA) and then subjected to electrophoresis in 1% agarose gels to check for the presence of a single band, before sending to GENEWIZ (NJ, USA) for sequencing. The raw 16S rRNA gene sequences obtained were initially processed by Geneious (https://www.geneious.com/) and Serial Cloner (http://serialbasics.free.fr /Serial_Cloner.html) then compared to sequences available in the Genbank database using BLAST for identification.
The 39 strains of cultured bacteria from this work were further grouped by fingerprinting using PCR amplification of the intergenic spacer (IGS) region in the conserved 16S-23S rRNA operon. Amplification of the 16S-23S rRNA IGS regions was performed using the forward primer 16S-e1390f (5’-TGYACACACCGCCCGTCA-3’) and the reverse primer 23S-e130r (5’-GGGTTBCCCCATTCRG-3’) (Gürtler and Stanisich 1996). The same PCR reaction mixture was used as for the 16S rRNA gene above. The amplification program consisted of 5 min at 95 °C, 30 cycles of 15 s at 95 °C, 30 s at 60 °C, and 90 s at 72 °C, followed by 5 min at 72 °C. PCR products were checked following the methods above.
Screening for potential IAA-producing strains in bacteria collection and validation by LC/MS analysis
Detection of indole-related compounds, such as IAA, was performed using the Salkowski method adopted from Gordon and Weber (1951). In this assay, the free, purified IAA standard results in a compound that has an absorption peak at 530 nm. Bacteria that are found to produce products with similar absorption characteristics are considered to be candidate strains for free IAA production. From a glycerol stock, bacterial cultures were grown in 5 mL of LB supplemented with 5 mM L-tryptophan. They were cultured at 28ºC, shaken at 250 rpm for 2 days or until the bacterial culture became turbid. 1 ml of bacterial culture was centrifuged at 10,000 rpm for 5 min. In duplicate, 100 µL of bacterial supernatant, and 100 µL of each free IAA standard were placed into a 96-well microtiter plate. 200 µL of Salkowski reagent was added to each sample. Samples were incubated for 30 minutes at room temperature. Samples were then read at OD530 nm (optical density) value using a microplate reader (Synergy HT, BioTek, USA), and the absorption peak between 450 nm and 550 nm was recorded. LC/MS was then carried out as previously described (Gilbert et al. 2018) to validate the production of IAA in the bacterial culture medium.
Assay for Arabidopsis root development and for microbial impact
About 60 Arabidopsis seeds (Arabidopsis thaliana, Col-0) in a microcentrifuge tube were sterilized by adding 1 ml of 50% bleach (3% NaClO) and shaken for 1 min, then washed four times with sterile water. Sterilized seeds were spread onto germination plates [1/2 MS (0.5x Murashige & Skoog Modified Basal Medium with Gamborg Vitamins), 1% Sucrose, pH 5.7, 0.25% Phytagel (Sigma, USA)], and stored at 4 °C in the dark for two days, then moved to a growth chamber and grown vertically at 22 °C under 100 µmol m− 2 s− 1 12 h light until use. Candidate strains of IAA-producing bacteria were cultured on liquid LB medium. Controls included culturing without added bacteria, or with the addition of E. coli (Escherichia coli DH5alpha), and strain RU1A. The RU1A bacterial strain was previously isolated from L. minor (strain DWC112, Landolt ID 5576) and showed a strong inhibition of Arabidopsis main root elongation along with a high level of free IAA production in vitro (described in Gilbert et al. (2018)). For the Arabidopsis root development assay, 5 days after plant growth, the bacterial suspension OD600 was adjusted to 0.7 and 100 µL (~ 3.85 × 107 cells) of the culture spread onto a square experimental plate (100 mm x 15 mm) with growth media [1/2 MS, pH 5.7, 0.5% Gellan Gum Powder (PhytoTech Lab, USA)] and dried in a laminar flow hood. For each treatment condition, 6 seedlings were transferred onto a single plate and sealed with micropore tape. After transferring Arabidopsis seedlings onto plates with or without bacteria, the plates were photographed and then placed vertically in the growth chamber under 16 hr light/8 h dark, at 25oC. One week later, the plates were photographed again to compare Arabidopsis root development with and without bacteria inoculation on plates. In this assay, sterile water and E. coli were used as negative controls. Experiments were repeated 3 times with similar results.
Plant-bacteria attachment assay
For the duckweed attachment assay, a baby jar (CultureJar™ G9, cat.# C1770; PhytoTechnology Labs, KS, USA) containing 50 mL of sterile 0.5x SH (Schenk & Hildebrandt Basal Salt Mixture, PhytoTech Lab, USA) was inoculated with bacterial cultures to an OD600 of 0.2 and then covered with a thin layer of duckweed Lm5576. In addition, we set up 50 mL of sterile 0.5x SH media in a baby jar covered with a thin layer of duckweed as a negative control. For the Arabidopsis attachment assay, similar treatments on plates as described for the root development assay above were used.
Duckweed or Arabidopsis samples were harvested after 2- and 7-days post inoculation onto medium with or without bacteria. Plant tissues were then subjected to three different types of chemical treatment: 1) Two washes with sterile water for 1 min; 2) One wash in sterile salt and detergent solution for 20 min, followed by rinsing with sterile water; 3) One wash in sterile salt and detergent solution for 20 min, followed by washing in 5% bleach for 2 min, then Na2S2O3 solution wash. For nucleic acid isolation, 5 ~ 10 washed duckweed fronds or two washed seedlings of Arabidopsis were transferred to 2 mL pre-filled Bead-Beater (OPS Diagnostics LLC, NJ, USA) DNA extraction tubes (0.5 g 100 µm silica beads, 0.5 g 1.7 mm zirconium beads, a single 14 mm silica bead), then mixed with 2x CTAB buffer. Tissues were homogenized by bead beating for 3 min at 4000 rpm using an HT Mini machine (OPS Diagnostics LLC, Lebanon, NJ, USA). Nucleic acid was then collected by ethanol precipitation.
Bacterial attachment detection by PCR and by fluorescence microscopy
To compare the mode of attachment to plants by the isolated bacteria strains, we applied a PCR-based assay targeting the 16S-23S intergenic spacer region of the conserved rRNA gene operon to monitor the presence of bacteria after rinsing the inoculated plants (gnotobiotically grown Lemna minor 5576 (Lm5576) and Arabidopsis thaliana Col-0) with different wash solutions: water, salt and detergent mixture, and bleach. We first used two known strains of Azospirillum PGPBs originally isolated from wheat, epiphyte Sp7 and endophyte Sp245 (Jain and Patriquin 1984), as controls to establish the validity of this method and supported the PCR-based results by fluorescence microscopy. Bacterial attachment PCR was performed on plant-bacteria nucleic acid samples using the protocol of bacterial IGS fingerprinting except that 200 ng was used as template for each reaction mix. For normalization and as a quality control for DNA preparation and integrity, a single-copy plant gene was used in each case. For duckweed, we use a single-copy gene LEAFY (Weigel et al. 1993) as the target from the L. minor genome, and for Arabidopsis we use a unique region of the Metacaspase 7 gene (Lam and Zhang 2012). LEAFY gene amplification of Lm5576 was performed using the forward primer LmLFY2-F (5’-CACCCGTTCATCGTCACAGA-3’) and the reverse primer LmLFY2-R (5’-AGAAGATGTACGCTACGCCG-3’). Metacaspase 7 amplification for Arabidopsis was performed using the forward primer ValMC7A-F (5’-CCCTAATTTCAGAAACAAACC-3’) and the reverse primer ValMC7B-R (5’-GATCCTATAATGGATGTTTTTC-3’). The PCR reaction mixture included 2.5 µL of 10 × buffer, 0.2 mM dNTPs, 0.4 µM of each primer, 0.4 µL of 5 U/µL Taq polymerase, ~ 200 ng of template DNA, and filter sterilized water to make a total of 25 µL. The amplification program for Lm5576 consisted of 1 min at 95 °C, 28 cycles of 15 s at 95 °C, 15 s at 60 °C, and 45 s at 72 °C, followed by 5 min at 72 °C. The amplification program for Arabidopsis consisted of 3 min at 95 °C, 26 cycles of 30 s at 95 °C, 30 s at 50 °C, and 45 s at 72 °C, followed by 5 min at 72 °C. For fluorescence microscopy, bacteria were stained using Syto9 (Gilbert et al. 2018). After co-cultivation with added bacteria as described above, duckweed tissues were harvested for microscopy studies after day 2 or day 7, depending on the attachment time. The fluorescence microscopy method was adopted from that described in Gilbert et al. (2018).
Statistical analysis
DNA sequence analysis using the Next-Generation Sequencing (NGS) methods was performed with the software QIIME (version 1.9.0) (Caporaso et al. 2010). Adaptors and primers were removed using AdapterRemoval (Lindgreen 2012). Phix contamination was removed using the program DeconSeq (Schmieder and Edwards 2011). Reads were merged and filtered by size (according to primer set) and quality (Phred quality score > 2). The sequences were then clustered into operational taxonomic units (OTUs) using an open reference strategy based on 97% identity with GreenGenes Database (13_5 release) (DeSantis et al. 2006) as reference. Taxonomy was assigned with the RDP classifier (Wang et al. 2007) retrained with GreenGenes 16S rRNA database (13_5 release). OTUs assigned to chloroplast and mitochondria were filtered out. Plots and figures were generated with R (version 3.2.1) using packages vegan (Oksanen et al. 2016), plyr (Wickham 2011), and ggplot (Wickham 2016). For bacterial 16S rRNA gene sequences, phylogenetic analyses were performed using MEGA version 6 after multiple alignments of the data by Clustal W. Clustering was performed using the neighbor-joining method. The statistical confidence of the nodes was estimated using 1000 replications of bootstrap.
Results
Variation in OTU abundance profiles for different plant tissues and soil samples from rice paddy fields
A total of 57 samples (8,788,158 reads) produced sufficiently high-quality reads for further analysis while six samples, including two from soil and four from rice root, failed to produce sufficient quality reads and were not included in downstream analyses. Rarefaction analysis indicated that the sequencing depth for the three plant tissue samples (RA, RR, and DW) was sufficient to capture the bacterial diversity in their respective tissues (Fig. S1c) while the microbial diversity in the SL samples could be significantly higher than our data indicated since microbes present at a low abundance in soil samples might not be detectable at the sequencing depth achieved here. For the four sample types, we note that while RA samples showed the highest number of reads (> 2.9 M) per sample, they had the lowest number of distinct OTUs (Table 1). The opposite is observed for SL, where in spite of the relatively low number of reads (~ 0.69 M) per sample, these samples had the highest number of OTUs. This is extended by a more systematic investigation of within- and between-sample α-diversity for the four sample types from which it was observed that bacterial diversity and relative species abundance decreased from SL, RR, DW, and RA, respectively (Fig. 2, Table 2). Significant differences (p < 0.0001), using one-way ANOVA with Tukey’s HSD post-hoc tests, were found between most samples except between DW and RR samples. Taking geographical effects into consideration, the Shannon diversity of each sample type between the three sites was also compared and revealed significant differences (ANOVA; p < 0.05) between RA, DW, RR, and SL samples of site 3 (Fig. 2c). Significant differences (p < 0.05) were also found between RA, DW or RR, and SL samples within site 1 and site 2, respectively.
Within- and between-sample microbiome diversity measurements. a Shannon diversity, tissues with same letter are not significantly different (ANOVA; p < 0.0001). b Shannon diversity, tissues and treatments with same letter are not significantly different (ANOVA; p < 0.05). c Shannon diversity, unique tissues at different sites with the same letter are not significantly different (ANOVA; p < 0.05). Abbreviations throughout are as follows: rice aerial tissue (RA), rice root (RR), duckweed (DW), soil (SL), sterile water (W), salt and detergent (SD). Boxes are first and third quartile (box bar is median), whiskers are 1.5x interquartile range, outliers plotted independently (dot)
Comparing the distribution of bacterial phyla between the three sites revealed a highly reproducible pattern that corresponded with sample types rather than their site of origin (Fig. S2). In all three sites, the soil microbiota is the most complex community with the majority of the bacteria represented by Proteobacteria (50–60%) and Acidobacteria (10–15%) (Figs. S2 and S4). Nevertheless, while Proteobacteria become even more dominant in rice roots (up to 80% in site 1), there are fewer OTUs for Acidobacteria, Nitrospirae, Chloroflexi and Verrucomicrobia in rice roots than in the soil, with the last three phyla being almost absent from rice roots collected from two of the three sites. In contrast, the Firmicutes and Actinobacteria are present in both roots and soil samples, indicating that they can associate more effectively with rice roots than the other four phyla of bacteria. While Proteobacteria is still the dominant phylum in the other plant tissue types, fewer distinct OTUs were found in DW and RA tissues when compared to RR. The remaining phyla present in these plant samples are members of Bacteroidetes and Actinobacteria, with a few remaining OTUs classified as members of the phylum Firmicutes (Figs. S2 and S3). Compared to the SL and RR samples, there appears to be a significant enrichment of these four bacterial phyla in RA and DW that is qualitatively similar to the known core microbiota found in model plants (Lundberg et al. 2012; Durán et al. 2018; Walters et al. 2018).
Bacterial community structure in plant hosts appears to be more strongly determined by plant tissue type than the local environment of paddy fields
Hierarchical clustering using the Bray-Curtis distance (Bray and Curtis 1957) revealed that host tissue is the major source of variation in determining microbial community structure (Fig. 3a). Clustering all 57 sets of microbiota data from various samples collected in multiple locations within the three sites revealed the existence of four predominant clusters, defined by their sample type: SL, DW, RR, and RA. While the replicate samples from a single site usually cluster closer together, indicating reproducibility of our experimental system, differences between sites are less significant than sample type. Rinsing the plant tissues with either water or a salt and detergent solution also produced a smaller but noticeable difference in the data obtained (Fig. S3).
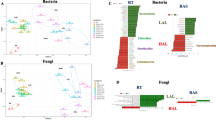
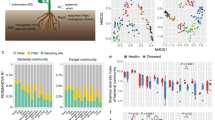
Beta-diversity of rice paddy environmental samples. a Bray-Curtis distance metric phylogeny. Symbol shape indicates site and symbol color indicates sample type (rice aerial tissue, RA; rice root, RR; duckweed, DW; and soil, SL). b Weighted Unifrac distance Principle Component Analysis. Symbol color as in panel a indicates sample type. Diamonds, rice aerial tissues; squares, rice roots; triangles, duckweed; inverted triangles, soil. Each point represents a sample
PCoA (principal coordinate analysis) using the weighted UniFrac distance showed that the microbial communities of SL, RR, RA, and DW were separated from each other with no significant correlation to the particular sampling site (Fig. 3b). The difference in microbial phyla distribution between SL and DW was smaller, compared to that in RR, and the difference was even larger between those of SL and RA. The β-diversity analysis results revealed that the phylogenetic data obtained for the four sample types is robust, since the deduced community structure is highly congruent both between and within sites. We thus grouped all data from similar tissue samples obtained at each of the three sites for subsequent analyses to increase the depth of data for comparative analysis.
Enrichment of common and distinct bacterial taxa between soil, duckweed (Lemna aequinoctialis), rice roots, and rice aerial tissues from paddy fields
As noted above, the most dramatic changes observed between soil and plant tissue types were an increase in the percentage of Proteobacteria along with a large decrease in Acidobacteria, Crenarchaeota, Euryarchaeota, Bacteroidetes, Verrucomicrobia and Chloroflexi, in some cases to almost undetectable levels (Fig. 4a, Fig. S4a). At the family level (Fig. 4b, Fig. S4b), DW had the highest relative abundance in Comamonadaceae (recently merged with the Burkholderiaceae (Parks et al. 2018)) at 32% of the OTUs identified, followed by Sphingomonadaceae (3%) and Rhodospirillaceae (2%). In contrast, Enterobacteriaceae was the dominant family in RA with relative abundance of about 25% (Fig. 4b). In addition, RR samples were found to be rich in Burkholderiaceae (5%), Rhodospirillaceae (5%), and Neisseriaceae (14%), consistent with their known association with root microbiota in plants (Santos-Medellín et al. 2017) and their potential function in nutrient acquisition. Interestingly, the family of Methylobacteriaceae (12%) was enriched in RA compared to either RR or DW from the same paddy fields. The overall distribution of microorganisms at the family level was relatively even in the SL samples, where no dominant family with high relative abundance was evident (Fig. 4b, Fig. S4b). It is worth noting that nine out of the top 10 families are in the Proteobacteria phylum, with Micromonosporaceae representing the only family from the Actinobacteria phylum. Since Proteobacteria comprised more than 80% of the bacterial microbiota in plant tissue samples, we carried out a more detailed comparative analysis between our sample types at the genus level for this phylum (Fig. 4c). Comparing the top three genera among Proteobacteria that appear to dominate each of the three plant sample types, we found a distinct representation in each case: Rubrivivax, Methylibium and Novosphingobium in DW; Pantoea, Sphingomonas and Methylobacterium for RA; and Bradyrhizobium, Azospirillum and Burkholderia in RR (Fig. 4c). From the apparent enrichment of distinct bacterial genera in each of these sample types, there appears to be both plant species-specific and tissue-specific enrichment of these genera of Proteobacteria in the paddy field ecosystem.
Taxonomic differences in microbial community for rice aerial tissue (RA), rice root (RR), duckweed (DW) and soil (SL). a Relative abundance of bacteria OTUs in the top nine ranked phyla from the total OTUs. b Relative abundance of the top 10 bacterial families. All families shown are in the phylum Proteobacteria except for Micromonosporaceae, which is in the phylum Actinobacteria. c Relative abundance of the top 10 ranked OTUs of Proteobacteria at the genus level
As a first step in estimating the possible differences in metabolic potential between the microbiome of duckweed and rice, we used PICRUSt as an algorithm that leveraged the large database of microbial genomes to predict gene functions/pathways that each microbiome likely encodes (Langille et al. 2013). The results indicate that the bacterial microbiota from duckweed tissues may be more enriched in metabolic pathways associated with xenobiotic biodegradation and amino acid metabolism whereas for the bacterial community present in rice aerial tissues, the metabolic pathways for energy as well as vitamins and enzyme cofactors are significantly more abundant (Fig. S5). These results suggest that the distinct microbial communities in these plants may reflect potentially different roles that they play in providing metabolic capabilities for their associated host plants in the rice paddy ecosystem.
To identify common bacterial OTUs among samples from the 3 tissue types, we carried out a Venn analysis of the 397 OTUs that resided within phylum Proteobacteria and were found among the microbial samples derived from the various plant samples. Of these, 254 OTUs were present in all plant sample types, and the core OTU collection we found in all three tissue types comprised mainly members of the Burkholderiaceae (> 20%), which were also highly enriched in RA, DW, and RR samples (Fig. 5). We found about 10% of the Sphingomonadaceae and Rhodocyclaceae were significantly underrepresented in RA compared with those in RR and DW (p < 0.05) (Fig. 5b). In contrast, the genera that are preferentially enriched in each plant sample types that we noted above may reflect their differential importance in each particular plant species or tissue type.
Identification of common Proteobacteria OTUs from rice aerial tissue (RA), rice root (RR) and duckweed (DW). a Unique Proteobacteria OTU Venn diagram. b Depleted or enriched OTUs (%) at the family level for all samples. Only taxa with more than 10 OTUs are shown. c Core OTUs (family level) distributed within the phylum Proteobacteria (total = 254)
Characterization of bacterial isolates from rice and duckweed tissues
To characterize the mode of growth-promoting action for plant-associated bacteria in the paddy field ecosystem, we isolated 39 bacteria strains from the three plant tissue types used in this study. Using 16S rRNA gene sequencing, we first determined the most likely genus for each of the isolated bacteria (Fig. 6d, Table S2). Consistent with our culture-independent analysis of the microbiome composition in these samples, most of the isolated bacteria from plants in the paddy fields belong to the Proteobacteria phylum (Pantoea, Enterobacter, Aeromonas, Acinetobacter and Pseudomonas) and some from the Firmicutes (Bacillus, Lysinibacillus and Staphylococcus). Perhaps the most striking observation is that all the isolated Pantoea strains were from RA tissues, where it was also most abundant, representing about 20% of all the Proteobacteria OTUs (Fig. 4c). This observation indicates that Pantoea species may be present at a higher percentage in terms of total bacterial counts in RA. In contrast, under our culture conditions RR appears to have a higher abundance of Bacillus and Enterobacter species, while there is a relatively even distribution of three different genera (Enterobacter, Aeromonas, Pseudomonas) among strains isolated from DW (Fig. 6).
Structural and functional characteristics of culturable bacteria isolated from duckweed and rice. a Neighbor Joining tree constructed based on bacterial 16S rRNA gene sequence. Symbol shape indicates site and symbol color indicates sample type (rice aerial tissue, RA; rice root, RR; duckweed, DW; and soil, SL). b Bacterial producers of indole related compounds where tick (√) indicates high and cross (×) indicates low or undetectable levels (Table S3). LC-MS mass spectrum validation is shown in Fig. S6 and Arabidopsis root growth assay is shown in Fig. S12. c Duckweed attachment assay attachment strength: strong (+++), moderate (++), weak (+) and no significant attachment (-); absence of symbol indicates strain was not tested. Attachment results for the Pantoea genus are shown in Fig. 7; all others are shown in Fig. S10. d Bacteria classification at genus level. The scale bar indicates the average number of substitutions per site
To gain more information about the potential modes of interaction between the isolated bacteria and plant hosts, we screened for bacteria strains that can produce the phytohormone IAA and also compared their relative attachment strength to duckweed and the model plant Arabidopsis thaliana. Out of the 39 strains that we isolated in this study, five resulted in a color change to pink, indicating production of indole-related compounds at a sufficiently high level to produce an absorption peak at 530 nm (Fig. 6b, Table S3). We corroborated this initial result with more careful chemical analysis using liquid chromatography-mass spectrometry (LC/MS) and validated this conclusion by positively identifying free IAA in the culture supernatant from these five bacterial strains (Fig. S6). Interestingly, these five IAA-producing strains are distributed among four different genera. Our result highlights the importance of strain-specific characteristics between different members of the same genus. In addition, fourteen (36%) of the isolated bacterial strains from this study are brown type, as observed by a color change from yellow to brown with a maximum absorbance increase at around 480 nm (Gilbert et al. 2018). The major molecule in these "brown-type" bacteria was identified as indole in our previous study (Gilbert et al. 2018), although this observation needs further validation by mass spectrometry using the strains isolated in the present study.
Using two well-characterized strains of Azospirillum PGPBs, we found that the epiphyte Sp7 could be washed off from the duckweed Lm5576 by a salt and detergent mixture. In contrast, the endophyte Sp245 binds much more tightly to duckweed and remains detectable in a PCR-based assay, even after washing the tissue with a 5% bleach solution (Figs. S8 & S9). We then applied this assay to a selected set of 19 bacterial strains in our collection (Fig. 7 and S10), covering all five of the IAA-producing strains. The results indicate that nine of the tested strains may be characterized as possible endophytes (i.e. strong attachment (+++) even after bleach treatment), while the other ten bacterial strains are potentially epiphytes that bind onto the surface of duckweed with varying degrees of tightness. Our data revealed that most of our isolated Pantoea strains display relatively strong binding to the duckweed strain Lm5576 while varying levels of interaction were observed for the other bacterial genera tested. Comparing strains with a moderate (strain Aw4a) and strong (Aw4d) interaction with duckweed, we observed that the former can be removed from duckweed by bleach rinsing while the latter cannot (Fig. 7). Interestingly, when these two bacterial strains were inoculated onto gnotobiotic Arabidopsis seedlings, we found that both strains colonized this model land plant very strongly and resisted bleach treatment (Fig. S11). Thus, there may be some quantitative difference in how the various Pantoea strains interact with different plant hosts. However, our finding that multiple Pantoea strains isolated from RA can interact strongly with gnotobiotic duckweed (Fig. 6c) indicates that the low representation of this bacterial genus in the ambient duckweed microbiota collected from paddy fields is not due to its inherently low attachment to this host.
Attachment of two Pantoea strains isolated from rice on duckweed. aPantoea strain Aw4a attachment profile harvested on day 2 and 7. bPantoea strain Aw4d attachment profile harvested on day 2 and 7. Abbreviations are as follows: Lemna minor 370-DWC112 (Lm5576), bacterial 16S-23S rRNA gene intergenic spacer region (IGS, ~1.5 kb), Lm5576 LEAFY gene (LFY, ~500 bp), sterile water (W), salt and detergent (SD), SD with 5% bleach (BL), gnotobiotic Lm5576 (-). Controls used for PCR were No TC: no template control, (+ strain ID): bacteria DNA, and Lm5576: gnotobiotic Lm5576
Effects of potential PGPBs from duckweed and rice on root development of Arabidopsis
Focusing on the five IAA-producing strains in the bacterial collection described in this study (Fig. 6b), we determined if they could alter plant development in a model plant system. Using the dicot A. thaliana as a host, we found that addition of IAA-producing bacteria such as the Microbacterium RU1A (Gilbert et al. 2018), but not E. coli, can increase lateral root proliferation while inhibiting extension of the main root of the seedlings. Similar phenotypes, which mimic those conferred by exogenous IAA addition (Fukaki et al. 2005), were also observed with the five IAA-producing bacteria described in this study (Fig. 8 & S12, Table S4). While the root phenotypes conferred by the five bacterial strains isolated from the paddy field were quantitatively similar in terms of root shortening (Fig. 8), there may be qualitative differences such as the degree of root branching between the different strains (Fig. S12) that could correlate with varying levels of IAA sensed by the plant. These five IAA-producing bacteria and two characterized endophytes were also tested for their effects on chlorophyll content of duckweed. Strains Aw3a and Rt8a significantly increased the amount of chlorophyll in duckweed, which is commonly associated with improved plant health under ambient conditions (Fig. S13).
Effects on Arabidopsis root development by treatment with auxin-producing bacteria from rice and duckweed on paddy fields. Increase in primary root length in millimeters (mm) after 7 days. Bacteria include E. coli as a negative control and Microbacterium sp. RU1A (1A) as a positive control. Arabidopsis inoculated with different bacteria were compared to sterile group using Wilcoxon test with results indicated on the top (p-value ≤ 0.05 = *, p-value ≤ 0.0001 = ****)
Discussion
Focusing on microbes tightly associated with two plant species collected from the same paddy fields (rice and duckweed), we show here the presence of reproducible bacterial community structures under field conditions and between sites located more than 200 km apart. While there is a gross similarity at the phylum level to the core microbiota described for other plants, in which Proteobacteria dominates as the major constituent (Lundberg et al. 2012; Edwards et al. 2015), there are clearly distinct preferences for bacteria classes at the family and genus levels in the three types of plant tissues examined here. The differential enrichment of particular genera of bacteria in duckweed, as well as between aerial and root tissues of rice, likely reflects the varied roles that the associated microbiota plays in the particular plant species or rice tissues. While distinct plant microbiota community structures have been reported before (Larousse et al. 2017; Walters et al. 2018), the mechanism for the differential enrichment of particular genera in various species or tissue context remains unclear. There are at least two levels of selective enrichment that can operate to generate the apparent diversity from a common microbial reservoir: 1) differences in microbe-plant tissue interaction may affect the initial steps of attachment and subsequent colonization by the microbe, and 2) differences in host cell physiology and response to microbial colonization can exert constraints on the types of bacteria that can thrive in a particular niche.
We examined the first possibility by testing Pantoea strains cultured from the aerial tissues of rice on duckweed for their ability to attach to plant tissues under laboratory conditions. Using a PCR-based attachment assay, we showed that the majority of the Pantoea strains isolated can attach strongly to duckweed. Consequently, the near absence of this genus of bacteria in the ambient duckweed microbiota from paddy fields is unlikely to be a result of their inability to attach to this plant host. Similarly, the very low levels of Pantoea species in the rice root microbiota is also unlikely to be due to their inability to interact with root tissues, since two IAA-producing Pantoea strains from rice are able to bind strongly to Arabidopsis and alter root development under laboratory conditions. Thus, at least for the Pantoea genus, our data would be more consistent with a post-attachment selection model in which the local tissue-specific context of the host would determine its steady state abundance relative to other microbes present in the same niche. This could involve both plant-microbe as well as microbe-microbe interactions within each niche that will ultimately determine the success of colonization by a particular microbe.
One possible way that plant-associated bacteria may promote improved plant functions is through the production of phytohormones, such as the auxin IAA, that can help reprogram host-encoded pathways for abiotic stress tolerance, nutrient acquisition and disease resistance (Berg et al. 2015). In addition to phytohormones that could be produced by plant-associated bacteria, there are likely additional signaling molecules that mediate communication between the host plant and different microbes that can further modify the phenotypic outcome of the interaction (Hassani et al. 2018; Stringlis et al. 2018). Nevertheless, the results from our functional characterization of five IAA-producing bacteria strains in this study are consistent with their potential role as PGPBs that could enhance lateral root development in land plants, as well as forming stable associations either as epiphytes or endophytes on duckweed.
In addition to the distinctive bacterial community structures described here for duckweed and the two rice tissues from multiple paddy fields, we also identified 254 shared OTUs that were found in all tissue samples examined. Interestingly, the large majority of these highly conserved OTUs maps to the Comamonadaceae family, which has recently been merged with the Burkholderiaceae family (Parks et al. 2018). While this group of microbes is dominant in duckweed, they are less prominent in rice tissues (Fig. 4b, Fig. S4b). Nevertheless, a recent functional study with synthetic communities in Arabidopsis has identified this family as one of the key players in orchestrating anti-fungal activities in association with more robust plant health (Durán et al. 2018). As we have found in this study, the conserved association and prominent representation of Comamonadaceae genera within plant tissues of both rice and duckweed from paddy fields suggest the possible importance of this bacteria family for plant health in the field as well. For duckweed, the high representation of this family of bacteria may indicate its increased importance for the lifestyle of this aquatic macrophyte and may correlate with the predicted increase of metabolic potential in the bacteria microbiota of duckweed for xenobiotic degradation and amino acid metabolism (Fig. S5).
One potential confounding issue that may influence the interpretation of our results from soil is the necessity of using a specialized DNA extraction kit for nucleic acid isolation from these environmental samples, which may potentially result in differential extraction of DNA from the different bacterial phyla present. The difficulty in performing DNA isolation from soil, relative to plant tissues, is exemplified by the low yields in sequence reads from these samples. Consequently, the derived ratios between the respective bacterial phyla within the soil are likely to be estimates at best. Nevertheless, the similar patterns observed from soil samples of the three sites would indicate that the relative levels for the major phyla revealed from our data analysis are likely to be reasonable approximations.
Microbial attachment to plants was investigated in this study using a generic bacterial primer set for 16S-23S intergenic spacer region to detect the presence of bacteria on the plant tissue while a single-copied LEAFY gene of the plant was used as a reference gene to normalize for the quantity of DNA used. This provides a relatively scalable assay that can be systematically deployed to examine plant-microbe interactions under a variety of conditions. In contrast, the most commonly used approaches for measuring microbial attachment to plant tissues involve microscopy and/or colony counting, both of which are labor intensive and time consuming. Previous studies have shown that certain concentrations of ethanol, NaClO (i.e. bleach) or Tween-20 can sterilize the surface of roots and seeds (Koeppel and Wu 2013). Our PCR-based method uses gnotobiotic duckweed and Arabidopsis as model plants and uses chemical treatments to define surface accessibility and differentiate between plant-microbe interactions of different strengths. With this approach, our present work has provided evidence that the selection for stable colonization of Pantoea species in rice aerial tissues and not in duckweed most likely occurs after the attachment of members of this bacterial genus onto the plant and may be the result of the host tissue context as well as competition with other microbes in the same niche.
In this work, we have begun to establish rice root-and leaf-derived microbiota culture collections with strains representing the majority of bacterial species that are reproducibly detected by culture-independent community sequencing. This will augment the collection of duckweed-associated bacteria described previously (Gilbert et al. 2018). In parallel, we are systematically cataloguing their potential for IAA production and ability to attach to plants, using gnotobiotic Arabidopsis and duckweed as our model systems. While it would have been more ideal to use rice plants as a test system for the microbes isolated from rice tissues, the small size and rapid growth of the model plant systems such as duckweed and Arabidopsis make them more amenable for large-scale studies. Nevertheless, it is important to further test our model in the future by using rice plants grown under laboratory conditions to directly examine the tissue-dependent attachment and colonization efficiency of these strains. As the recent studies with synthetic communities in Arabidopsis have illustrated (Bai et al. 2015), a robust bacterial community that can be stably established in the target plant would set the stage for testing whether their presumed beneficial roles can be reproduced under laboratory conditions and eventually transferred to their hosts under field conditions. The resources and data produced by our work here laid the foundation for this approach to improve rice production via management of the microbiota in paddy fields.
Data Accessibility
Raw reads have been deposited in the Short Read Archive of NCBI under project no. PRJNA545325.
References
Acosta K, Xu J, Gilbert S, Denison E, Brinkman T, Lebeis S, Lam E (2020) Duckweed hosts a taxonomically similar bacterial assemblage as the terrestrial leaf microbiome. PLoS One 15(2):e0228560. https://doi.org/10.1371/journal.pone.0228560
Bai Y, Müller DB, Srinivas G, Garrido-Oter R, Potthoff E, Rott M, Dombrowski N, Münch PC, Spaepen S, Remus-Emsermann M, Hüttel B, McHardy AC, Vorholt JA, Schulze-Lefert P (2015) Functional overlap of the Arabidopsis leaf and root microbiota. Nature 528:364–369. https://doi.org/10.1038/nature16192
Berg G, Rybakova D, Grube M, Köberl M (2015) The plant microbiome explored: implications for experimental botany. J Exp Bot 67:995–1002. https://doi.org/10.1093/jxb/erv466
Benton TG, Vickery JA, Wilson JD (2003) Farmland biodiversity: is habitat heterogeneity the key? Trends Ecol Evol 18:182–188. https://doi.org/10.1016/s0169-5347(03)00011-9
Borisjuk N, Chu P, Gutierrez R, Zhang H, Acosta K, Friesen N, Sree KS, Garcia C, Appenroth KJ, Lam E (2014) Assessment, validation and deployment strategy of a two-barcode protocol for facile genotyping of duckweed species. Plant Biol 17:42–29. https://doi.org/10.1111/plb.12229
Bray JR, Curtis JT (1957) An ordination of the upland forest communities of southern Wisconsin. Ecol Monogr 27:326–349. https://doi.org/10.2307/1942268
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336. https://doi.org/10.1038/nmeth.f.303
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72:5069–5072. https://doi.org/10.1128/AEM.03006-05
Durán P, Thiergart T, Garrido-Oter R, Agler M, Kemen E, Schulze-Lefert P, Hacquard S (2018) Microbial interkingdom interactions in roots promote Arabidopsis survival. Cell 175:973–983. https://doi.org/10.1101/354167
Edwards J, Johnson C, Santos-Medellín C, Lurie E, Podishetty NK, Bhatnagar S, Eisen JA, Sundaresan V (2015) Structure, variation, and assembly of the root-associated microbiomes of rice. Proc Natl Acad Sci USA 112:E911–E920. https://doi.org/10.1073/pnas.1414592112
Feng J, Li F, Zhou X, Xu C, Fang F (2016) Nutrient removal ability and economical benefit of a rice-fish co-culture system in aquaculture pond. Ecol Eng 94:315–319. https://doi.org/10.1016/j.ecoleng.2016.06.002
Finkel OM, Castrillo G, Herrera Paredes S, Salas González I, Dangl JL (2017) Understanding and exploiting plant beneficial microbes. Curr Opin Plant Biol 38:155–163. https://doi.org/10.1016/j.pbi.2017.04.018
Fukaki H, Nakao Y, Okushima Y, Theologis A, Tasaka M (2005) Tissue-specific expression of stabilized SOLITARY-ROOT/IAA14 alters lateral root development in Arabidopsis. Plant J 44:382–395. https://doi.org/10.1111/j.1365-313x.2005.02537.x
Gilbert S, Xu J, Acosta K, Poulev A, Lebeis S, Lam E (2018) Bacterial production of indole related compounds reveals their role in association between duckweeds and endophytes. Front Chem 6:265. https://doi.org/10.3389/fchem.2018.00265
Gordon SA, Weber RP (1951) Colorimetric Estimation of Indoleacetic Acid. Plant Physiol 26:192–195. https://doi.org/10.1104/pp.26.1.192
Gu BJ, Ju XT, Chang J, Ge Y, Vitousek PM (2015) Integrated reactive nitrogen budgets and future trends in China. Proc Natl Acad Sci USA 112:8792–8797. https://doi.org/10.1073/pnas.1510211112
Gürtler V, Stanisich VA (1996) New approaches to typing and identification of bacteria using the 16S-23S rDNA spacer region. Microbiology 142:3–16. https://doi.org/10.1099/13500872-142-1-3
Hassani MA, Durán P, Hacquard S (2018) Microbial interactions within the plant holobiont. Microbiome 6(1):58. https://doi.org/10.1186/s40168-018-0445-0
Jain DK, Patriquin DG (1984) Root hair deformation, bacterial attachment, and plant growth in wheat-azospirillum associations. Appl Environ Microbiol 48:1208–1213. https://doi.org/10.1016/0141-4607(84)90089-1
Koeppel AF, Wu M (2013) Surprisingly extensive mixed phylogenetic and ecological signals among bacterial operational taxonomic units. Nucleic Acids Res 41:5175–5188. https://doi.org/10.1093/nar/gkt241
Kumura M (2005) Populations, community composition and biomass of aquatic organisms in the flood water of rice fields and effects of field management. Soil Sci Plant Nutr 51:159–181. https://doi.org/10.1111/j.1747-0765.2005.tb00021.x
Lam E, Zhang Y (2012) Regulating the reapers: activating metacaspases for programmed cell death. Trends Plant Sci 17:487–494. https://doi.org/10.1016/j.tplants.2012.05.003
Langille MGI, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes JA, Clemente JC, Burkepile DE, Vega Thurber RL, Knight R, Beiko RG, Huttenhower C (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31:814–821. https://doi.org/10.1038/nbt.2676
Lansing JS, Kremer JN (2011) Rice, fish, and the planet. Proc Natl Acad Sci U S A 108(50):19841–19842. https://doi.org/10.1073/pnas.1117707109
Larousse M, Rancurel C, Syska C, Palero F, Etienne C, Industri B, Nesme X, Bardin M, Galiana E (2017) Tomato root microbiota and Phytophthora parasitica-associated disease. Microbiome 5:56. https://doi.org/10.1186/s40168-017-0273-7
Li H, Liang XQ, Lian YF, Xu L, Chen YX (2009) Reduction of ammonia volatilization from urea by a floating duckweed in flooded rice fields. Soil Sci Soc Am J 73:1890. https://doi.org/10.2136/sssaj2008.0230
Lindgreen S (2012) AdapterRemoval: easy cleaning of next-generation sequencing reads. BMC Res Notes 5:337. https://doi.org/10.1186/1756-0500-5-337
Lundberg D, Lebeis SL, Paredes SH, Yourstone S, Gehring J, Malfatti S, Tremblay J, Engelbrektson A, Kunin V, Del Rio TG, Edgar RC, Eickhorst T, Ley RE, Hugenholtz P, Tringe SG, Dangl JL (2012) Defining the core Arabidopsis thaliana root microbiome. Nature 488:86–90. https://doi.org/10.1038/nature11237
Ng CA, Wong LY, Lo PK, Bashir MJK, Chin SJ, Tan SP, Chong CY, Yong LK (2017) Performance of duckweed and effective microbes in reducing arsenic in paddy and paddy soil. AIP Conference Proceedings 1828:020031. https://doi.org/10.1063/1.4979402
Oksanen JF, Blanchet G, Friendly M, Kindt R, Legendre P, McGlinn PR, O’Hara RB, Simpson GL, Solymos P, Stevens M, Wagner H (2016) Vegan: Community Ecology Package, Version 2.5–3
Parks DH, Chuvochina M, Waite DW, Rinke C, Skarshewski A, Chaumeil PA, Hugenholtz P (2018) A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life. Nat Biotechnol 36:996–1004. https://doi.org/10.1038/nbt.4229
Pascale A, Proietti S, Pantelides IS, Stringlis IA (2020) Modulation of the root microbiome by plant molecules: the basis for targeted disease suppression and plant growth promotion. Front Plant Sci 10:1741. https://doi.org/10.3389/fpls.2019.01741
Santos-Medellín C, Edwards J, Liechty Z, Nguyen B, Sundaresan V (2017) Drought stress results in a compartment-specific restructuring of the rice root-associated microbiomes. mBio 8:e00764–e00717. https://doi.org/10.1128/mBio.00764-17
Schmieder R, Edwards R (2011) Quality control and preprocessing of metagenomic datasets. Bioinformatics 27:863–864. https://doi.org/10.1093/bioinformatics/btr026
Sha Z, Chu Q, Zhao Z, Yue Y, Lu L, Yuan J, Cao L (2017) Variations in nutrient and trace element composition of rice in an organic rice-frog coculture system. Sci Rep 7(1):15706. https://doi.org/10.1038/s41598-017-15658-1
Stringlis IA, Zhang H, Pieterse CMJ, Bolton MD, de Jonge R (2018) Microbial small molecules - weapons of plant subversion. Nat Prod Rep 35(5):410–433. https://doi.org/10.1039/C7NP00062F
Walters WA, Jin Z, Youngblut N, Wallace JG, Sutter J, Zhang W, González-Peña A, Peiffer J, Koren O, Shi Q, Knight R, Glavina Del Rio T, Tringe SG, Buckler ES, Dangl JL, Ley RE (2018) Large-scale replicated field study of maize rhizosphere identifies heritable microbes. Proc Natl Acad Sci USA 115:7368–7373. https://doi.org/10.1073/pnas.1800918115
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267. https://doi.org/10.1128/aem.00062-07
Wang C, Li S, Lai DYF, Wang W, Ma Y (2015) The effect of floating vegetation on CH4 and N2O emissions from subtropical paddy fields in China. Paddy Water Environ 13:425–431. https://doi.org/10.1007/s10333-014-0459-6
Weigel D, Alvarez J, Smyth DR, Yanofsky MF, Meyerowitz EM (1993) LEAFY controls floral meristem identity in Arabidopsis. Cell 69:843–859. https://doi.org/10.1007/978-1-4757-9607-0_17
Wickham H (2011) The split-apply‐combine strategy for data analysis. J Stat Softw 40:1–29. https://doi.org/10.18637/jss.v040.i01
Wickham H (2016) ggplot2. Elegant graphics for data analysis, 2nd edn. Springer International Publishing, Cham
Yang H, Yu D, Zhou J, Zhai S, Bian X, Weih M (2018) Rice-duck co-culture for reducing negative impacts of biogas slurry application in rice production systems. J Environ Manage 213:142–150. https://doi.org/10.1016/j.jenvman.2018.02.077
Yao Y, Zhang M, Tian Y, Zhao M, Zhang B, Zhao M, Zeng K, Yin B (2017) Duckweed (Spirodela polyrhiza) as green manure for increasing yield and reducing nitrogen loss in rice production. Field Crop Res 214:273–282. https://doi.org/10.1016/j.fcr.2017.09.021
Zhang J, Hu L, Ren W, Guo L, Tang J, Shu M, Chen X (2016) Rice-soft shell turtle coculture effects on yield and its environment. Agr Ecosyst Environ 224:116–122
Zhu YG, Su JQ, Cao ZH, Xue K, Quensen J, Guo GX, Yang YF, Zhou JZ, Chu HY, Tiedje JM (2016) A buried Neolithic paddy soil reveals loss of microbial functional diversity after modern rice cultivation. Sci Bull 61:1052–1060. https://doi.org/10.1007/s11434-016-1112-0
Acknowledgements
Duckweed research at the Lam Lab is supported in part by a grant from the Department of Energy (DE-SC0018244) and a Hatch project (#12116) from the New Jersey Agricultural Experiment Station at Rutgers University. We are grateful to Prof. Qiyi Lei of Kaili University, and the villagers of three counties in Qiandongnan Prefecture for assisting field surveys and sample collection. The work in the Long Lab was supported by the National Natural Science Foundation of China (31761143001 & 31870316), Ministry of Ecology and Environment of China (2019HJ2096001006), Key Laboratory of Ethnomedicine (Minzu University of China) of the Ministry of Education of China (KLEM-ZZ201906 & KLEM-ZZ201806), Minzu University of China (YLDXXK201819), the Ministry of Education of China and State Administration of Foreign Experts Affairs of China (B08044), and the China Scholarship Council (CSC No. 201606390030). This work was also supported in part by the Guangdong Academy of Sciences’ special project to W.H. (2019GDASYL-0103031). We thank Ilya Raskin (Rutgers University, New Brunswick, NJ, USA) for providing access to the mass spectrometry instrument in his laboratory as well as for providing LC-MS related reagents and also thank Michael Lawton (Rutgers University) for editorial assistance to improve the manuscript. K.A. acknowledges the Rutgers University SGS Excellence Fellowship for providing student support for part of this work. The Lebeis Lab thanks Elizabeth Denison for performing the quality control for the DNA samples prior to 16S rDNA amplicon sequencing and acknowledges the start-up funds provided to S.L. from the University of Tennessee, Knoxville. The contribution by the Facilities Integrating Collaborations for User Science (FICUS) initiative under Contract Nos. DE-AC02-05CH11231 (JGI) and DE-AC05-76RL01830 (EMSL) to the sequencing of the duckweed microbiome work is also gratefully acknowledged.
Author information
Authors and Affiliations
Contributions
E.L. and C.L. designed the project approach; W.H. performed most of the experiments in the study; E.L. isolated the bacteria strains; C.L. led the field surveys and sample collections; S.G. and A.P. performed the mass spectrometry studies; K.A. and S.G. assisted in the set up of the bacteria attachment and Salkowski assays; S.L. assisted in data analysis and drafting of the manuscript; W.H. and E.L. drafted the manuscript with help from all other authors.
Corresponding authors
Additional information
Responsible Editor: Birgit Mitter.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Huang, W., Gilbert, S., Poulev, A. et al. Host-specific and tissue-dependent orchestration of microbiome community structure in traditional rice paddy ecosystems. Plant Soil 452, 379–395 (2020). https://doi.org/10.1007/s11104-020-04568-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-020-04568-3