Abstract
Aims
Typha angustifolia is a heavy metal tolerant plant that grows in a uranium mine tailings highly contaminated with iron. In this study three iron oxidizing microbes (FeOBs) isolated from Typha rhizoplane were investigated for their role in plant growth promotion (PGP). Their effect on iron nutrition in Typha under iron replete and excess condition was also evaluated.
Methods
The PGP activities of the FeOBs were studied by measuring their influence on plant growth. To investigate the mechanism of growth promotion their ability to solubilize phosphate, and to produce Indole acetic acid and siderophores were studied. The influence of the FeOBs on root to shoot partitioning of iron was tested by measuring total iron content in roots and shoots treated with microbes.
Results
The FeOBs were named as Paenibacillus cookii JGR8, (MTCC12002), Pseudomonas jaduguda JGR2 (LMG25820) and Bacillus megaterium JGR9 (MTCC12001). The siderophore producers, influenced iron accumulation in the plant root. Additionally P. pseudoalcaligenes JGR2 increased shoot iron content overcoming the root- shoot barrier that allows Typha to exclude metals from its shoot. Among the PGP mechanisms tested, ability to solubilize phosphate appeared to be most significant for increasing the plant biomass.
Conclusion
FeOBs that produce siderophore increased iron content in plant and therefore can be of immense biotechnological importance. However Biomass increase was directly correlated with increased phosphate acquisition and not with enhanced iron accumulation in Typha.







Similar content being viewed by others
Abbreviations
- CAS:
-
Chrome azurol S
- FeOB:
-
Iron oxidizing bacteria
- IAA:
-
Indole acetic acid
- ICPOES:
-
Inductive coupled plasma optical emission spectroscopy
- NIST SRM:
-
National institute of standards and technology standard reference materials
- PGP:
-
Plant growth promoting
- PGPR:
-
Plant growth promoting rhizobacteria
- ppm:
-
Parts per million (mg.kg−1)
References
Anzai Y, Kim H, Park JY, Wakabayashi H, Oyaizu H (2000) Phylogenetic affiliation of the pseudomonads based on 16S rRNA sequence. Int J Syst Evol Microbiol 50(Pt 4):1563–1589
Bar-Ness E, Hadar Y, Chen Y, Shanzer A, Libman J (1992) Iron uptake by plants from microbial siderophores : a study with 7-nitrobenz-2 oxa-1,3-diazole-desferrioxamine as fluorescent ferrioxamine B analog. Plant Physiol 99:1329–1335
Barriuso J, Ramos Solano B, Lucas JA, Lobo AP, García-Villaraco A, Gutiérrez Mañero FJ (2008) Ecology, genetic diversity and screening strategies of plant growth promoting rhizobacteria (PGPR). In: I Ahmad, J Pichtel, S Hayat (eds) Plant-bacteria interactions: strategies and techniques to promote plant growth. Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, Germany. Chapter 1. pp 1–17
Below JF Jr, Connick RE, Coppel CP (1957) Kinetics of the formation of the ferric thiocyanate complex. J Am Chem Soc 80:2961–2967
Benardini JN, Vaishampayan PA, Schwendner P, Swanner E, Fukui Y, Osman S, Satomi M, Venkateswaran K (2011) Paenibacillus phoenicis sp. nov., isolated from the Phoenix Lander assembly facility and a subsurface molybdenum mine. Int J Syst Evol Microbiol 61:1338–1343. doi:10.1099/ijs.0.021428-0
Berendsen RL, Pieterse CM, Bakker PA (2012) The rhizosphere microbiome and plant health. Trends Plant Sci 17:478–486. doi:10.1016/j.tplants.2012.04.001
Bouizgarne B (2013) Bacteria for plant growth promotion and disease management. In: DK Maheshwari (ed) Bacteria in agrobiology: disease management. Springer, Berlin Heidelberg, pp 15–47
Bric JM, Bostock RM, Silverstone SE (1991) Rapid in situ assay for indoleacetic acid production by bacteria immobilized on a nitrocellulose membrane. Appl Environ Microbiol 57:535–538
Brizzi M, Betti L (2010) Statistical tools for alternative research in plant experiments. Metodološki Zvezki 7:59–71
Bulgarelli D, Schlaeppi K, Spaepen S, Loren V, van Themaat E, Schulze-Lefert P (2013) Structure and functions of the bacterial microbiota of plants. Annu Rev Plant Biol 64:807–838. doi:10.1146/annurev-arplant-050312-120106
Cappucino JG, Sherman N (2004) Microcbiology: a laboratory manual. Pearson Education. 6th Edition, pp 319–321
Chakraborty D, Abhay Kumar S, Sen M, Apte SK, Das S, Acharya R, Das T, Reddy AVR, Roychaudhury S, Rajaram H, Seal A (2011) Manganese and iron both influence the shoot transcriptome of Typha angustifolia despite distinct preference towards manganese accumulation. Plant Soil 342:301–317
Charest MH, Beauchamp CJ, Antoun H (2005) Effects of the humic substances of de-inking paper sludge on the antagonism between two compost bacteria and Pythium ultimum. FEMS Microbiol Ecol 52:219–227. doi:10.1016/j.Femsec.2004.11.017
Chen YP, Rekha PD, Arun AB, Shen FT, Lai WA, Young CC (2006) Phosphate solubilizing bacteria from subtropical soil and their tricalcium phosphate solubilizing abilities. Appl Soil Ecol 34:33–41. doi:10.1016/j.apsoil.2005.12.002
Dakora FD, Phillips DA (2002) Root exudates as mediators of mineral acquisition in low nutrient environments. Plant Soil 245:35–47
de Souza MP, Chu D, Zhao M, Zayed AM, Ruzin SE, Schichnes D, Terry N (1999) Rhizosphere bacteria enhance selenium accumulation and volatilization by Indian mustard. Plant Physiol 119:565–574
Dethier Rogers S, Beech J, Sarma KS (1998) Shoot regeneration and plant acclimatization of the wetland monocot Cattail (Typha latifolia). Plant Cell Rep 18:71–75
Devaiah BN, Madhuvanthi R, Karthikeyan AS, Raghothama KG (2009) Phosphate starvation responses and gibberellic acid biosynthesis are regulated by the MYB62 transcription factor in Arabidopsis. Mol Plant 2:43–58. doi:10.1093/mp/ssn081
Dimkpa CO, Svatos A, Dabrowska P, Schmidt A, Boland W, Kothe E (2008) Involvement of siderophores in the reduction of metal-induced inhibition of auxin synthesis in Streptomyces spp. Chemosphere 74:19–25. doi:10.1016/j.chemosphere.2008.09.079
Emerson D, Moyer C (1997) Isolation and characterization of novel iron oxidizing bacteria that grow at circumneutral pH. Appl Environ Microbiol 63:4784–4792
Figueiredo MVB, Seldin L, Araujo F, Mariano RLR (2010) Plant growth promoting rhizobacteria: fundamentals and applications plant growth and health promoting bacteria. In: DK Maheshwari (ed) Plant growth and health promoting bacteria, vol 18. Springer, Berlin Heidelberg, pp 21–43
García JL, Probanza A, Ramos B, Mañero FJG (2001) Ecology, genetic diversity and screening strategies of plant growth promoting rhizobacteria. J Plant Nutr Soil Sci 164:1–7
Gerhardt P, Murray RGE, Wood WA, Krieg NR (1994) Methods for general and molecular bacteriology. American Society for Microbiology, Washington
Glick BR (2003) Phytoremediation: synergistic use of plants and bacteria to clean up the environment. Biotechnol Adv 21:383–393. doi:10.1016/s0734-9750(03)00055-7
Glickmann E, Dessaux Y (1995) A critical examination of the specificity of the salkowski reagent for indolic compounds produced by phytopathogenic bacteria. Appl Environ Microbiol 61:793–796
Grandlic CJ, Mendez MO, Chorover J, Machado B, Maier RM (2008) Plant growth-promoting bacteria for phytostabilization of mine tailings. Environ Sci Technol 42:2079–2084
Grebe M (2011) Plant biology: unveiling the Casparian strip. Nature 473:294–295. doi:10.1038/473294a
Guang-Can, Shu-Jun, Miao-Ying, Guang-Hui (2008) Phosphate-solubilizing and -mineralizing abilities of bacteria isolated from soils. Pedosphere 18:515–523
Gupta A, Gopal M, Tilak KV (2000) Mechanism of plant growth promotion by rhizobacteria. Indian J Exp Biol 38:856–862
Gutiérrez Mañero FJ, Probanza A, Ramos B, Colón Flores JJ, Lucas García JA (2003) Ecology, genetic diversity and screening strategies of plant growth promoting rhizobacteria (PGPR). J Plant Nutr 26:1101–1115
Hameeda B, Harini G, Rupela OP, Wani SP, Reddy G (2008) Growth promotion of maize by phosphate-solubilizing bacteria isolated from composts and macrofauna. Microbiol Res 163:234–242. doi:10.1016/j.micres.2006.05.009
Hanert HH (2006) The genus siderocapsa and other iron and manganese oxidizing eubacteria. Prokaryotes 7:1005–1015. doi:10.1007/0-387-30747-8_50
Hiltner L (1904) Uber neuere erfahrungen und probleme auf dem gebiet der boden sbakteriologie und unter besonderer berucksichtigung det grundungung und branche. Arb Deut Landw Ges 98:59–78
Hrynkiewicz K, Baum C (2012) The potential of rhizosphere microorganisms to promote the plant growth in disturbed soils. In: A Malik, E Grohmann (eds) Environmental protection strategies for sustainable development. Springer, Netherlands, pp 35–64
Illmer P, Barbato A, Schinner F (1995) Solubilization of hardly-soluble AlPO4 with P-solubilizing microorganisms. Soil Biol Biochem 27:265–270
Joseph B, Ranjan Patra R, Lawrence R (2007) Characterization of plant growth promoting rhizobacteria associated with chickpea (Cicer arietinum L.). Int J Plant Prod 1:141–152
Jukes TH, Cantor CR (1969) Evolution of protein molecules. In: HN Munro (ed) Mammalian protein metabolism, vol 3. Academic Press, Inc., New York, pp 21–132
Kappler A, Straub KL (2005) Geomicrobiological cycling of iron. Rev Mineral Geochem 59:85–108
Kaymak HC, Yarali F, Guvenc I, Donmez MF (2008) The effect of inoculation with plant growth Rhizobacteria (PGPR) on root formation of mint (Mentha piperita L) cuttings. Afr J Biotechnol 7:4479–4483
Kovacs G, Burghardt J, Pradella S, Schumann P, Stackebrandt E, Marialigeti K (1999) Kocuria palustris sp. nov. and Kocuria rhizophila sp. nov., isolated from the rhizoplane of the narrow-leaved cattail (Typha angustifolia). Int J Syst Bacteriol 49(Pt 1):167–173
McLaughlin BE, Van Loon GW, Crowder AA (1985) Comparison of selected washing treatments on Agrostis gigantea samples from mine tailings near Copper Cliff, Ontario, before analysis for Cu, Ni, Fe and K content. Plant Soil 85:433–436
Naseer S, Lee Y, Lapierre C, Franke R, Nawrath C, Geldner N (2012) Casparian strip diffusion barrier in Arabidopsis is made of a lignin polymer without suberin. Proc Natl Acad Sci U S A 109:10101–10106. doi:10.1073/pnas.1205726109
Navarro L, Dunoyer P, Jay F, Arnold B, Dharmasiri N, Estelle M, Voinnet O, Jones JD (2006) A plant miRNA contributes to antibacterial resistance by repressing auxin signaling. Science 312:436–439. doi:10.1126/science.1126088
Perez-Miranda S, Cabirol N, George-Tellez R, Zamudio-Rivera LS, Fernandez FJ (2007) O-CAS, a fast and universal method for siderophore detection. J Microbiol Meth 70:127–131. doi:10.1016/j.mimet.2007.03.023
Perez-Torres CA, Lopez-Bucio J, Cruz-Ramirez A, Ibarra-Laclette E, Dharmasiri S, Estelle M, Herrera-Estrella L (2008) Phosphate availability alters lateral root development in Arabidopsis by modulating auxin sensitivity via a mechanism involving the TIR1 auxin receptor. Plant Cell 20:3258–3272. doi:10.1105/tpc.108.058719
Probanza A, Lucas Garca JA, Ruiz Palomino M, Ramos B, Gutierrez Manero FJ (2002) Pinus pinea L. seedling growth and bacterial rhizosphere structure after inoculation with PGPR Bacillus (B. licheniformis CECT 5106 and B. pumilus CECT 5105). Appl Soil Ecol 20(2):75–84, (10) 20: 75–84
Rajkumar M, Ae N, Prasad MN, Freitas H (2010) Potential of siderophore-producing bacteria for improving heavy metal phytoextraction. Trends Biotechnol 28:142–149. doi:10.1016/j.tibtech.2009.12.002
Richardson AE, Simpson RJ (2011) Soil microorganisms mediating phosphorus availability update on microbial phosphorus. Plant Physiol 156:989–996. doi:10.1104/pp.111.175448
Rooney AP, Price NP, Ehrhardt C, Swezey JL, Bannan JD (2009) Phylogeny and molecular taxonomy of the Bacillus subtilis species complex and description of Bacillus subtilis subsp. inaquosorum subsp. nov. Int J Syst Evol Microbiol 59:2429–2436. doi:10.1099/ijs.0.009126-0
Saharan BS, Nehra V (2011) Plant growth promoting rhizobacteria: a critical review. Life Sci Med Res 2011:1–30
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sato A, Miura K (2011) Root architecture remodeling induced by phosphate starvation. Plant Signal Behav 6:1122–1126. doi:10.4161/psb.6.8.15752
Schachtman DP, Reid RJ, Ayling SM (1998) Phosphorus uptake by plants: from soil to cell. Plant Physiol 116:447–453
Scheldeman P, Goossens K, Rodriguez-Diaz M, Pil A, Goris J, Herman L, De Vos P, Logan NA, Heyndrickx M (2004) Paenibacillus lactis sp. nov., isolated from raw and heat-treated milk. Int J Syst Evol Microbiol 54:885–891
Sepe A, Barbieri P, Peduzzi R, Demarta A (2008) Evaluation of recA sequencing for the classification of Aeromonas strains at the genotype level. Lett Appl Microbiol 46:439–444. doi:10.1111/j.1472-765X.2008.02339.x
Sinha S, Mukherjee SK (2008) Cadmium-induced siderophore production by a high Cd-resistant bacterial strain relieved Cd toxicity in plants through root colonization. Curr Microbiol 56:55–60. doi:10.1007/s00284-007-9038-z
Taylor GJ, Crowder AA (1983) Use of the DCB technique for extraction of hydrous iron oxides from roots of wetland plants. Am J Bot 70:1254–1257
Timmusk S, van West P, Gow NA, Huffstutler RP (2009) Paenibacillus polymyxa antagonizes oomycete plant pathogens Phytophthora palmivora and Pythium aphanidermatum. J Appl Microbiol 106:1473–1481. doi:10.1111/j.1365-2672.2009.04123.x
Varvel GE, Peterson GA, Anderson FN (1991) A revised method for determining phosphate-phosphorus levels in sugar beet leaf petioles. J ASS BT 19:138–142
Vyas P, Gulati A (2009) Organic acid production in vitro and plant growth promotion in maize under controlled environment by phosphate-solubilizing fluorescent Pseudomonas. BMC Microbiol 9:174. doi:10.1186/1471-2180-9-174
Xin YH, Zhang DC, Liu HC, Zhou HL, Zhou YG (2009) Pseudomonas tuomuerensis sp. nov., isolated from a bird’s nest. Int J Syst Evol Microbiol 59:139–143. doi:10.1099/ijs.0.000547-0
Acknowledgments
We sincerely acknowledge Council of Scientific and Industrial Research (CSIR) India and University Grants Commission, India for funding. Upal Das Ghosh acknowledges CSIR EMR- I for his fellowship. We thank Dr. SoumaleeBasu, Department of Microbiology C. U and Dr. GeetanjaliSundaram, Department of Biochemistry C.U. for critically reading the manuscript. The ICPOES measurement was performed under DAE V2 5 year plan project TULID.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Responsible Editor: Henk Schat.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1aFig. S1bFig. S1c
Dendogram of three bacteria generated using PHYLIP software version 3.695. Phylogenetic tree or dendogram for the individual bacteria were generated by aligning the sequence in ClustalX program followed by tree generation using Phylip 3.695 software. Bar represents 10 changes in bases. Numbers at the node indicates the bootstrap values (%). Bootstrap (n = 1000 replicates) values ≤50 % are not shown. a] P. cookie JGR8; Closely related strains of P. cookii JGR8 in BLAST analysis have been used to construct the dendogram. Paenibacillus azoreducens strain D-TSB3 is used as outgroup. (JPEG 5 kb)
b] B. megaterium JGR9; Closely related strains of B. megaterium that shows identity with JGR9 are used for constructing the dendogram. Bacillus aryyabhattai strain S4-248 is used as outgroup. (JPEG 8 kb)
c] P. jadugudaJGR2: The tree has been constructed taking the whole Pseudomonas aeruginosa group in concern (Anzai et al. 2000) Pseudomonas stutzeri was used as outgroup. (JPEG 6 kb)
Table S1
recA gene analysis of these three bacteria. The recA gene was sequenced and analyzed through tBlastx analysis. (DOCX 14 kb)
Table S2
Comparative FAME analysis of three bacteria. FAME analysis of the three bacteria was done from MTCC, IMTECH, India. The result obtained for each bacterium was tabulated and compared with other related bacteria. a] P. cookii JGR8 was compared with P. phoenicis 3PO2SA, P. barengoltzii ATCC BAA 1209, P. timonensis CIP 108005, P. macerans ATCC8244, P. lactis MB1871, P. lautus NRRL NRS666, P. glucanolyticus DSM 5162, P. polymyxa JCM2507, P. cookii LMG18419; (Benardini et al. 2011; Scheldeman et al. 2004). b] B. megateriumJGR9 was compared with B. cereus FO 011, B. pumilus FO 033, B. subtilis sub sp. Subtilis NRS744, B. tequilensis B-41771, B. vallismortis B 14890, B. megaterium DSM32; (Rooney et al. 2009; Venkateswaran et al. 2000) c] P. jaduguda JGR2 was compared with P. tuomuerensis sp. nov. 78-123T, P. mendocina CGMCC 1.1804T, P. pseudoalcaligenes CGMCC 1.1806T, P. alcaliphila JCM 10630T (Xin et al. 2009). (DOCX 24 kb)
Table S3
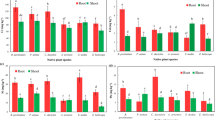
Calculation for ANOVA to determine statistical significance of PGP activity over plant growth. Plants were grown in presence and absence of iron (1,000 ppm) with or without bacteria. Plant lengths were measured after 2 months. Nine plants for each treatment were taken and the experiments were repeated three times. Results obtained for 27 plants per treatment were subjected to statistical analysis by ANOVA. 1. Iron replete condition a] P. cookie JGR8; b] B. megaterium JGR9; c] P. jaduguda JGR2 d] Mixed bacterial population. 2. Iron excess condition a] P. cookie JGR8; b] B. megaterium JGR9; c] P. jaduguda JGR2 d] Mixed bacterial population. (DOCX 28 kb)
Table S4
Calculation of statistical significance of iron accumulation in plant tissues by means of confidence interval. Iron accumulation in plant roots and shoots were determined by digesting the plant tissues in nitric acid and hydrogen peroxide and analyzing the digested samples by ICPOES against standard orchard leaf (NIST SRM 1571). a] Iron replete root; b] Iron replete shoot; c] Iron excess root; d] Iron excess shoot. (DOCX 15 kb)
Rights and permissions
About this article
Cite this article
Ghosh, U.D., Saha, C., Maiti, M. et al. Root associated iron oxidizing bacteria increase phosphate nutrition and influence root to shoot partitioning of iron in tolerant plant Typha angustifolia . Plant Soil 381, 279–295 (2014). https://doi.org/10.1007/s11104-014-2085-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-014-2085-x