Abstract
A wide variety of functional regulatory non-coding RNAs (ncRNAs) have been identified as essential regulators of plant growth and development. Depending on their category, ncRNAs are not only involved in modulating target gene expression at the transcriptional and post-transcriptional levels but also are involved in processes like RNA splicing and RNA-directed DNA methylation. To fulfill their molecular roles properly, ncRNAs must be precisely processed by multiprotein complexes. In the case of small RNAs, DICER-LIKE (DCL) proteins play critical roles in the production of mature molecules. Land plant genomes contain at least four distinct classes of DCL family proteins (DCL1–DCL4), of which DCL1, DCL3 and DCL4 are also present in the genomes of bryophytes, indicating the early divergence of these genes. The liverwort Marchantia polymorpha has become an attractive model species for investigating the evolutionary history of regulatory ncRNAs and proteins that are responsible for ncRNA biogenesis. Recent studies on Marchantia have started to uncover the similarities and differences in ncRNA production and function between the basal lineage of bryophytes and other land plants. In this review, we summarize findings on the essential role of regulatory ncRNAs in Marchantia development. We provide a comprehensive overview of conserved ncRNA–target modules among M. polymorpha, the moss Physcomitrium patens and the dicot Arabidopsis thaliana, as well as Marchantia-specific modules. Based on functional studies and data from the literature, we propose new connections between regulatory pathways involved in Marchantia’s vegetative and reproductive development and emphasize the need for further functional studies to understand the molecular mechanisms that control ncRNA-directed developmental processes.
Key message
We provide a comprehensive overview of conserved ncRNA–target modules amongst Marchantia polymorpha, Physcomitrium patens and Arabidopsis thaliana as well as Marchantia-specific modules what contributes to a better understanding of the evolution of ncRNAs in terrestrial plants
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In plants, regulatory non-coding RNAs (ncRNAs) are involved in diverse developmental processes, including shoot apical meristem (SAM) development, leaf morphogenesis, the juvenile-to-adult phase transition, determination of flowering time, fertilization and stress responses (Macintosh et al. 2001; Chen 2005; Mattick et al. 2009; Wang and Chekanova 2017; Hou et al. 2019). Non-coding RNAs are grouped into short (miRNA, siRNA and tasiRNA, among others) and long ncRNAs (Yu et al. 2019; Wierzbicki et al. 2021). MicroRNAs (miRNAs) are a class of short ncRNAs frequently 20–24 nt long that modulate gene expression through a mechanism of post-transcriptional gene silencing: mRNA cleavage or translational inhibition (Bartel 2004; Yu and Wang 2010; Bajczyk et al. 2023). Short interfering RNAs (siRNAs) represent a class of small, double-stranded RNAs (dsRNAs), usually 21–24 nt in length. These are derived from dsRNA precursors that are processed into siRNA duplexes by DICER-LIKE (DCL) activities. They suppress the expression of target mRNAs that correspond to dsRNA or are involved in the silencing and methylation of homologous DNA (Hamilton and Baulcombe 1999; Hamilton et al. 2002; Xie et al. 2004; Axtell 2013). Another class of small ncRNAs, trans-acting small interfering RNAs (tasiRNAs), are secondary siRNAs that silence their target RNAs in a similar manner as miRNAs, via mRNA cleavage or translational inhibition. In contrast, long non-coding RNAs (lncRNAs) are more than 200 nt in length and act in multiple ways to regulate their targets, resulting in chromatin remodeling, transcriptional enhancing or repressing, RNA splicing and target mimicry (F. Liu et al. 2010b; Wang and Chekanova 2017; Palos et al. 2023).
Bryophytes, including liverworts, hornworts and mosses, originated from a group of early diverged land plant lineages (Qiu and Palmer 1999; Shaw et al. 2011; Morris et al. 2018; Donoghue et al. 2021). Marchantia polymorpha (Marchantia) is a complex thalloid liverwort that has gained much attention in recent years as a model system because of its use in advancing genetic and functional research (Ishizaki et al. 2016; Shimamura 2016; Bowman et al. 2017; Bowman 2022). The moss Physcomitrium patens (Physcomitrium) is another model organism used for studies on plant evolution, development, metabolism and genetics (Reski et al. 1994; Cove et al. 2006; Rensing et al. 2008; Prigge and Bezanilla 2010; Falz and Müller-Schüssele 2019). Although most of the knowledge about ncRNAs has arisen from angiosperms, the availability of several genome sequences, along with the use of high-throughput RNA sequencing, has led to initial insights into small RNA and lncRNA machineries in bryophytes (Arif et al. 2012; Alaba et al. 2015; Coruh et al. 2015; Lin et al. 2016; Tsuzuki et al. 2016; Lin and Bowman 2018; Perroud et al. 2018; Meyberg et al. 2020). Research on ncRNAs in Marchantia and Physcomitrium revealed their significant roles in different molecular mechanisms governing the development of these bryophytes and their responses to environmental cues. Functional analysis of miRNAs and their targets in angiosperms and bryophytes indicates that some developmental pathways are regulated by the same conserved modules across all land plants. However, some miRNA–target modules diversified in function during the course of evolution. The presence of various conserved and Marchantia-specific gene regulatory modules in which ncRNAs are involved provides new insights leading to a better understanding of Marchantia development (Fig. 1). In this review, we summarize the essential role of regulatory ncRNAs in Marchantia and propose new putative connections between already known modules that are crucial for life cycle completion in Marchantia.
Schematic illustration of the role of miRNA–target modules that have been studied to date in M. polymorpha development. Modules that are involved in gametophyte development are shown in red, whereas modules involved in the vegetative development program are shown in blue. Since MpRKD has roles in both vegetative and reproductive development, it is shown in a gradient of blue to red. DCLs, master producers of sRNAs, are shown in light green. Arrows and T-bars represent activation and repression, respectively. The question mark on the miR390–TAS3 module indicates that the exact role of this module needs to be determined in future functional studies. Abbreviations: miR, microRNA; ARF, AUXIN RESPONSIVE FACTOR; SPL, SQUAMOSA PROMOTER BINDING PROTEIN-LIKE; KNOX, KNOTTED1-like Homeobox; SUK, SUPPRESSOR of KNOX1; FGMYB, FEMALE GAMETOPHYTE MYB; SUF, SUPPRESSOR OF FEMINIZATION; C3HDZ, C-III HOMEODOMAIN-LEUCINE ZIPPER; DCL, DICER-LIKE; RSL1, ROOT HAIR DEFECTIVE SIX-LIKE 1; FRH1, FEW RHIZOIDS1; RKD, RWP-RK DOMAIN-CONTAINING (Tsuzuki et al. 2016, 2019; Flores-Sandoval et al. 2018; Lin and Bowman 2018; Hisanaga et al. 2019a; Thamm et al. 2020; Dierschke et al. 2021; Csicsely et al. 2023; Futagami et al. 2023; Streubel et al. 2023; Yelina et al. 2023)
DCLs, producers of small RNAs
DCL proteins have the critical function of producing small RNAs from long dsRNA precursors. Arabidopsis thaliana (Arabidopsis) has four DCL proteins (AtDCL1–AtDCL4), and their molecular functions have been characterized by genetic studies (Bologna and Voinnet 2014) as well as biochemical assays (Kurihara and Watanabe 2004; Fukudome et al. 2011; Nagano et al. 2014). Now we know that AtDCL1 is involved in miRNA biogenesis from primary microRNA (pri-miRNA) transcripts (Park et al. 2002; Kurihara and Watanabe 2004), and the other proteins, AtDCL2–AtDCL4, function in the biogenesis of siRNAs with specific sizes of 22, 24 and 21 nt, respectively (Xie et al. 2004; Akbergenov et al. 2006; Henderson et al. 2006). Reflecting the importance of miRNAs for plant development, AtDCL1 knockout mutants are embryonic lethal (Schauer et al. 2002). Notably, 24 nt siRNAs (heterochromatic siRNAs, hc-siRNAs) are produced by AtDCL3 from precursors generated by RNA polymerase IV from heterochromatic genomic regions. They are involved in transcriptional gene silencing (TGS) via RNA-directed DNA methylation of transposable elements. In contrast, 21 nt siRNAs produced by AtDCL4 are important for post-transcriptional gene silencing (PTGS) or RNA interference to regulate endogenous gene expression. Moreover, critical targets of AtDCL4 include phased tasiRNAs derived from the TAS gene loci, including TAS3 (Xie et al. 2005; Yoshikawa et al. 2005). The abundance of 24 nt siRNAs relative to 21 nt siRNAs can be explained by the relative ratio of DCL3 to DCL4 dicing activities in plant tissues (Tabara et al. 2018). Moreover, the activity of DCL4 is post-translationally regulated in response to the cellular redox state (Seta et al. 2017; Tabara et al. 2018). This view is also supported by the dcl mutant analysis of Physcomitrium, as the DCL4 knockout mutant ΔPpdcl4 shows severe developmental abnormalities; however, such a ΔPpdcl4 phenotype can be fully suppressed by the additional introduction of a PpDCL3 knockout mutation (Arif et al. 2012). This indicates that the activity ratio of DCL3 to DCL4, i.e., which type of siRNAs are produced from the precursor RNAs, is also critical to development in mosses. It is important to note that, in Arabidopsis, the occurrence of PTGS or TGS depends on the downstream AGO-guided steps (changes in the ratio AGO1-siRNA target cleavage, an example of PTGS, versus AGO4-siRNA-guided DNA methylation, an example of TGS) (Wierzbicki et al. 2008, 2009). It is highly probable that a similar dependence might be present in bryophytes. In other words, during land plant evolution, plants would have acquired a fine-tuning system for growth and development by balancing PTGS and TGS according to environmental situations.
Interestingly, AtDCL2 and AtDCL4 have redundant functions: to produce viral- and transgene-derived siRNAs (Blevins et al. 2006; Deleris et al. 2006; Fusaro et al. 2006; Henderson et al. 2006). Recently, it has also been suggested that suppression of AtDCL4 activity promotes the biosynthesis of 22 nt siRNAs by AtDCL2, which, in turn, triggers a stress response (Wu et al. 2020). Thus, AtDCL2 and AtDCL4 are key DCLs for the regulation of plant growth and morphogenesis in response to the environment. However, comprehensive genomic analyses revealed that the DCL2 family genes can be found only in seed plants (Axtell et al. 2007; Huang et al. 2015; Bowman et al. 2017; Belanger et al. 2023), suggesting that the DCL2-derived 22 nt siRNAs could have emerged in the seed plant lineage (You et al. 2017; Belanger et al. 2023). Additional groups of DCL proteins, such as monocot-specific DCL5 (formerly DCL3b), have also emerged during seed plant evolution, increasing the complexity of the small RNA system in plants (Patel et al. 2021).
In the Physcomitrium genome, there are four genes encoding canonical DCL proteins: PpDCL1a and PpDCL1b, which are most closely related to AtDCL1, and PpDCL3 and PpDCL4, which are most closely related to AtDCL3 and AtDCL4, respectively (Axtell et al. 2007). In addition, a specific DCL protein MINIMAL DICER-LIKE (mDCL) lacking an N-terminal helicase and a Dicer dimerization domain has been identified in addition to canonical types of PpDCL proteins (Coruh et al. 2015). All Physcomitrium mutants of canonical DCL genes are defective for developmental processes; the juvenile-to-adult transition, in particular, is accelerated in the ΔPpdcl3 mutant (Cho et al. 2008). A similar phenotype was shown in the mdcl mutant, indicating that the mDCL protein, like PpDCL3, also contributes to the hc-siRNA pathway. Interestingly, the mdcl mutation uniquely affected the accumulation of 23 nt and 24 nt siRNAs: the 23 nt and 24 nt siRNAs were increased and decreased, respectively, in mdcl mutants (Coruh et al. 2015), indicating Physcomitrium-specific mechanistic complexity of hc-siRNA biogenesis. In addition to such species-specific aspects of DCL proteins, the Physcomitrium DCL data have also revealed the evolutionarily conserved system of PpDCL1 autoregulation (Arif et al. 2022). The intron region of DCL1 genes in Arabidopsis and Physcomitrium encodes miRNAs, which are distinct in sequence and position in the two species. Thus, the processing of intronic miRNAs and pre-mRNA splicing of host DCL1 would compete mechanistically with each other. Since DCL1 functions in pri-miRNA processing, intronic miRNA biogenesis depends on the level of DCL1, resulting in the autoregulation of DCL1 levels (Rajagopalan et al. 2006; Axtell et al. 2007). Arif et al. (2022) demonstrated increased accumulation of the intronic miRNA miR1047 during abiotic stress and abscisic acid (ABA) treatment in Physcomitrium, but there was no increase in the PpDCL1 transcript itself. The deletion mutant ΔmiR1047 showed hypersensitivity to abiotic stress (Arif et al. 2022). Additionally, Arabidopsis miR838 hosted in an AtDCL1 intron was also upregulated during heat stress (Baev et al. 2014). Therefore, these data suggest the importance of conserved DCL1 autoregulation for stress tolerance in land plants.
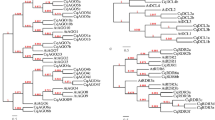
The Marchantia genome encodes four DCL genes: MpDCL1a, MpDCL1b, MpDCL3 and MpDCL4 (Lin et al. 2016; Lin and Bowman 2018; Pietrykowska et al. 2022; Belanger et al. 2023; Figs. 1 and 2). The MpDCL1a, MpDCL3 and MpDCL4 proteins contain basically the same functional domains as Arabidopsis, suggesting the functional conservation of these MpDCL proteins for small RNA biogenesis (Pietrykowska et al. 2022; Csicsely et al. 2023). Recently, Csicsely et al. (2023) reported the phenotype of Mpdclge mutants (genome-edited Mpdcl mutants) generated by the CRISPR/Cas9 system. This study revealed severe developmental defects in the Mpdcl1age mutant, whereas no detectable abnormalities were found in Mpdcl1bge or Mpdcl4ge mutants, apart from the less frequently produced and smaller antheridiophores (Fig. 1). Importantly, the obtained Mpdcl1age mutant plant had two amino acid substitutions and two amino acid insertions within the genomic region encoding the RES III domain, which probably affects the secondary protein structure, impairing the endonucleolytic activity of the MpDCL1a protein. The authors could not find large genomic rearrangements or early stop codons within the MpDCL1a locus. All these data indicate that MpDCL1a has a dominant function in small RNA production in Marchantia (Fig. 1) and that its complete knockout is lethal, as in other plant species. Interestingly, the Mpdcl1bge mutant showed salt tolerance, while the three other kinds of Mpdclge mutants displayed ABA hypersensitivity. Moreover, Mpdcl3ge showed 1-naphthaleneacetic acid insensitivity for thallus growth inhibition and rooting enhancement (Csicsely et al. 2023). These phenotypes of Mpdclge mutants strongly suggest the functional diversity of MpDCL proteins during Marchantia development, such as the involvement of MpDCL1b and MpDCL3 in small RNA biogenesis for salt tolerance and auxin signaling, respectively.
Recent comprehensive genomic analysis of plant DCL proteins demonstrated that the origin of DCL1 family proteins can be traced back to green algae, whereas the DCL3 and DCL4 families would have been acquired after the terrestrialization of plants (Belanger et al. 2023). Thus, the plant small RNA system is hypothesized to have evolved in the following order: the miRNA pathway (by DCL1) then the hc-siRNA pathway (by DCL3) and 21 nt tasiRNA/phasiRNA (phased secondary small interfering RNAs) pathways. Consistent with this suggestion, Tsuzuki et al. (2016) showed that the miR390–TAS3 regulatory module is conserved in Marchantia (Fig. 2). Additionally, the proportion of siRNAs within total small RNAs is lower in Marchantia vegetative tissues than in Arabidopsis, possibly reflecting the relatively simple DCL system (i.e., lacking DCL2) in this liverwort. Further profiling of small RNAs in Mpdclge mutants can provide additional clues of the mechanistic aspects of DCL proteins in Marchantia, leading to further insights into the evolution of the small RNA regulatory system for plant development.
Role of selected small RNA and long non-coding RNA modules in plant development
As mentioned in the introduction, regulatory RNA species participate in many processes related to plant growth and development. For example, in Marchantia, miRNA–mRNA target modules, as well as long ncRNAs, influence thallus development, phase transitions and sexual/asexual reproduction. In this chapter, we review and compare reports on ncRNA regulatory modules in Marchantia, Physcomitrium and Arabidopsis. Figure 2 summarizes sRNA and lncRNA modules described to date that are unique to Marchantia and those that are conserved among these three plant species.
Graphical representation of miRNA–target modules conserved across three plant species, M. polymorpha, P. patens and A. thaliana. The modules conserved among all land plants are shown in red, and modules unique to Marchantia are shown in green. One of the unique modules, miR319–MpRKD, consists of conserved miRNA, but the target is Marchantia specific; therefore, it is shown distinctly in yellow (Tsuzuki et al. 2016, 2019; Flores-Sandoval et al. 2018; Lin and Bowman 2018; Hisanaga et al. 2019a; Thamm et al. 2020; Dierschke et al. 2021; Csicsely et al. 2023; Futagami et al. 2023; Streubel et al. 2023; Yelina et al. 2023)
Conserved miRNA–target modules across land plants
The miR390–TAS3 module is involved in the regulation of ARF clade B transcription factor functions
Auxins are involved in many aspects of plant growth and development and in plant defensive responses to various stresses (Ludwig-Muller 2011; Balzan et al. 2014; Gomes and Scortecci 2021). AUXIN RESPONSIVE FACTORS (ARFs) represent a family of transcription factors (TFs) that regulate the expression of downstream auxin-regulated genes (Liu et al. 1993; Liscum and Reed 2002; Chapman and Estelle 2009). There are three evolutionary lineages of ARF genes in land plants: clades A, B and C. ARF transcription activators (ARF5, ARF6, ARF7, ARF8 and ARF19 in Arabidopsis) are all clustered in the A clade, whereas ARF transcription repressors are split into the B and C clades (Finet and Jaillais 2012). In Arabidopsis, ARF genes of clade B (ARF2, ARF3 and ARF4) are preferentially targeted by trans-acting ARF-targeting small interfering RNAs (TAS3-derived tasiARFs) (Finet and Jaillais 2012; Xia et al. 2017). Arabidopsis miR390 triggers the production of tasiRNAs from the TAS3 locus, which, in turn, target and downregulate mRNA levels of ARF3 and ARF4 (Montgomery et al. 2008; Marin et al. 2010; Rubio-Somoza and Weigel 2011; Endo et al. 2013). Furthermore, ARF3 and ARF4 activity promotes the juvenile-to-adult vegetative phase transition. ARF3 is required for floral meristem determinacy, and its gene is a direct target of APETALA 2 (AP2), which represses ARF3 expression (Liu et al. 2014). Thus, AP2 and miR390 prolong the juvenile phase by repressing ARF3 and/or ARF4 activity (Fahlgren et al. 2006; Garcia et al. 2006; Rubio-Somoza and Weigel 2011). In Arabidopsis, miR390 is mis-paired exactly at the cleavage site of the TAS3 RNA 5′ target site, which results in a non-cleavable target that is critical for TAS3-derived tasiRNA biogenesis.
In contrast, miR390 at the TAS3 RNA 5′ target site is perfectly matched in bryophytes (Axtell et al. 2006; Xia et al. 2017; Lin and Bowman 2018). As in Arabidopsis, TAS3 genes in bryophytes generate tasiRNAs that target ARF class B mRNA; however, they also generate tasiRNAs that target mRNA of AP2 orthologs. In Physcomitrium, the miR390–TAS3 module acts in the gametophytic stages of development to modulate gametophore initiation, protonemal branch determinacy and caulonemal differentiation (Plavskin et al. 2016). TAS3 genes in Physcomitrium include two miR390 target sites and generate two tasiRNAs: tasiAP2 and tasiARF (Axtell et al. 2007; Krasnikova et al. 2013). It is known that, in Physcomitrium, the overexpression of miR390 results in slower gametophore formation, increased miR390-triggered tasiRNA accumulation and a decrease in the number of tasiRNA targets (Cho et al. 2012).
In Marchantia, there is one TAS3 gene that contains two miR390 target sites (Krasnikova et al. 2013). MpTAS3 RNA generates tasiAP2, which regulates MpAP2 expression. However, whether a functional TAS3–ARF module is present in Marchantia is not clear. Generally, in plants, tasiARFs target ARF mRNA regions that encode a highly conserved amino acid sequence: K/RVLQGQE (Xia et al. 2017). Interestingly, a similar sequence is present in the MpARF2 protein but not in MpARF3. Arabidopsis ARF3 and Marchantia ARF2 (but not MpARF3) belong to the same clade, clade B (Finet and Jaillais 2012; Xia et al. 2017). Consequently, there is no evidence that tasiARF targets MpARF3 mRNA, but an expected cleavage site has been found for MpARF2 mRNA (Lin and Bowman 2018). Despite the presence of TAS3, ARF clade B and AP2 genes and miR390 in Marchantia, there still is no report indicating the involvement of these elements in the juvenile-to-adult phase transition in this liverwort.
In summary, the miR390–TAS3 module regulates the juvenile-to-adult phase transition in Arabidopsis and directs proper developmental timing in Physcomitrium. Meanwhile, the exact functions of the miR390–TAS3 module in Marchantia still need to be determined (Fig. 1, left lower side).
The miR160–ARFss module is involved in the regulation of ARF clade C transcription factor functions
In angiosperms, the MIR160 family is involved in the regulation of class C ARF genes, which have been implicated in various biological processes during plant development. In Arabidopsis, miR160 downregulates ARF10, ARF16 and ARF17 gene expression (Rhoades et al. 2002; Mallory et al. 2005; Li et al. 2016). It was discovered that, in Arabidopsis, a 3′ regulatory region located downstream of the MIR160a gene contains auxin-response elements that enhance MIR160 gene expression and upregulate miR160 levels (Liu et al. 2010b). Floral organs in carpels (foc) Arabidopsis mutant plants have a transposon insertion in the 3′ regulatory region of MIR160. This insertion results in reduced expression of MIR160 and upregulated expression of ARF10, ARF16 and ARF17 genes in flowers. The foc mutant displays defects in floral organ formation, such as reduced fertility, the appearance of floral organs inside siliques and irregular flower shape, as well as aberrant seeds and viviparous seedlings (Liu et al. 2010b). Defects observed in the foc mutant are the result of an aberrant response to auxin since ARF10, ARF16 and ARF17 are involved in auxin signaling. Plants expressing miR160-resistant ARF17 (35 S:5mARF17 lines) with increased ARF17 mRNA levels display male sterility (Shi et al. 2015). A proper ratio of the miR160–ARF module components also has a significant impact on vegetative development. Arabidopsis plants expressing miR160-resistant ARF17 displayed reduced plant size, leaf shape defects and reduced lateral root number (Mallory et al. 2005). Therefore, miR160-guided post-transcriptional gene regulation plays a pivotal role in the realization of successful vegetative and generative developmental programs.
In Physcomitrium, mRNAs of ARFs belonging to class C are also regulated by miR160 (Plavskin and Timmermans 2012; Plavskin et al. 2016). However, there is a lack of functional studies on the Physcomitrium miR160–ARF module. Similarly, in Marchantia, miR160 regulates the expression of MpARF3, a class C ARF (Lin et al. 2016; Tsuzuki et al. 2016). MpARF3 is a negative regulator of cell proliferation and differentiation and inhibits the developmental transition from asexual to sexual reproduction (Flores-Sandoval et al. 2018; Tsuzuki et al. 2019). Pleiotropic defects caused by both loss-of-function and gain-of-function mutations in MpARF3 were visible throughout the gametophytic stage of the life cycle. Loss-of-function Mparf3 alleles exhibited several developmental effects, including smaller cells, more distinct air pores, more pegged rhizoids, more gametophores per thallus area, ectopic antheridia and fused archegoniophores. Mutants overexpressing MpARF3 exhibited improper thallus shape, with a lack of air chambers and pores. Likewise, Mpmir160 knockout mutants were characterized by improper differentiation of the thallus, including air pores, and also of the gemma cups and gametophores (Fig. 1, left upper side) (Flores‐Sandoval et al. 2018b). In summary, miR160–ARFs derived from clade C modules are evolutionarily conserved between bryophytes and angiosperms. In both cases, these modules act as crucial players for the coordination of both the vegetative and reproductive phases of development.
The miR166–C3HDZ1 (HD-ZIPIII) module regulates plant architecture
In angiosperms, the MIR165/166 family contains numerous members, which have diverged in function and specificity (Coruh et al. 2015; Merelo et al. 2016; Yan et al. 2016; Y. Li et al. 2022). Arabidopsis miR165/166 regulates the expression of the cognate C3HDZ TF gene family (HD-ZIPIII) members: REVOLUTA (REV), PHABULOSA (PHB), PHAVOLUTA (PHV) and ARABIDOPSIS THALIANA HOMEOBOX 8 and 15 (ATHB8, ATHB15) (McConnell et al. 2001; Otsuga et al. 2001; Tang et al. 2003; Kim et al. 2005). These modules regulate transitions during organ development (Rhoades et al. 2002; Floyd and Bowman 2004; Tsuzuki et al. 2016; Yip et al. 2016; Ma et al. 2020). They are known to regulate adaxial/abaxial leaf polarity, shoot meristem maintenance, root meristem size and growth, vascular patterning and flower development (McConnell et al. 2001; Juarez and Timmermans 2004; Kidner and Martienssen 2004; Leibfried et al. 2005; Williams et al. 2005; Husbands et al. 2009; Sakaguchi and Watanabe 2012; Singh et al. 2014; Tsuzuki et al. 2016). Mutations that disrupt the pairing of Arabidopsis REV, PHB or PHV mRNAs with miR165/166 have similar effects on leaf phenotype, including reduced leaves with adaxial characteristics all around the circumference of the leaves (McConnell and Barton 1998; Mallory et al. 2004). These observations imply that related HD-ZIPIII TFs (REV/PHB/PHV) regulated by miR165/166 play similar roles in leaf morphogenesis.
In Physcomitrium, there are six MIR166 genes (Kozomara et al. 2019). Ectopic expression of miR166, which targets HD-ZIPIII mRNAs, causes shorter stature of gametophores and altered leaflet shape and size (Yip et al. 2016). In Marchantia, the miR165/166 family is represented by two members: miR166a and miR166b (Tsuzuki et al. 2016). miR166a regulates MpC3HDZ1 mRNA levels (Lin et al. 2016; Tsuzuki et al. 2016; Lin and Bowman 2018). MpC3HDZ1 is upregulated in archegoniophores and antheridiophores compared to Marchantia vegetative thalli (Bowman et al. 2017). Furthermore, ectopic overexpression of the MpMIR166a gene resulted in unregulated dorsoventral morphogenesis of the vegetative thallus (ventral curling) and downregulation of MpC3HDZ1 expression. Moreover, gemma cup structure was strongly reduced while gemmae were generated (Fig. 1, middle right side) (Tsuzuki et al. 2016).
Although the Marchantia thallus, Physcomitrium leaflets and Arabidopsis leaves represent different types of organs with unrelated origins (the Marchantia thallus and Physcomitrium leaflets represent gametophytes, while leaves represent a part of the Arabidopsis sporophyte), their proper polarity is optimal to perform the functions adapted to each plant’s requirements and is governed by the evolutionarily conserved miR165/166–HD-ZIPIII module.
The miR171–GRAS module controls plant growth during the vegetative phase of life
The MIR171 family is considered one of the oldest miRNA families and was probably already present in the common ancestor of all embryophytes (Han and Zhou 2022). miR171 regulates the expression of members of the GRAS TF gene superfamily. GRAS is named after the first three GRAS proteins discovered in Arabidopsis: GIBBERELLIC ACID INSENSITIVE (GAI), REPRESSOR OF GA (RGA) and SCARECROW (SCR) (Di Laurenzio et al. 1996; Peng et al. 1997; Silverstone et al. 1998). In Arabidopsis, miR171 specifically targets three HAIRY MERISTEM (HAM1–3) genes belonging to the GRAS superfamily. These genes are also called LOST MERISTEM (LOM) or SCARECROW-LIKE (SCL) genes (Han and Zhou 2022). Specifically, miR171 downregulates the expression of several SCL TFs: SCL6/SCL6-IV, SCL22/SCL6-III and SCL27/SCL6-II (Ma et al. 2014). HAM family genes are required for the maintenance of indeterminate growth in angiosperms. Arabidopsis ham mutants exhibit a remarkably unique phenotype of shoot meristem and differentiation arrest (Stuurman et al. 2002; Engstrom 2012). In the case of SCL genes, it was also shown that they are involved in chlorophyll biosynthesis in Arabidopsis (Bolle 2004; M. H. Lee et al. 2008). Plants overexpressing miR171-resistant SCL27 turn yellow and show a significant decrease in chlorophyll content. In contrast, plants overexpressing MIR171c and scl6 scl22 scl27 triple mutants produce dark green leaves, with 40% more chlorophyll than wild-type plants (Ma et al. 2014). These results indicate that, in Arabidopsis, miR171-targeted SCLs are negative regulators of chlorophyll biosynthesis.
In Marchantia, MpSCARECROW-LIKE (MpGRAS10/MpSCL) mRNA has been shown to be a target of miR171 (Lin et al. 2016; Lin and Bowman 2018; Yelina et al. 2023). However, neither the Mpscl nor the Mpmir171 mutants showed any observable changes in chlorophyll accumulation. Thus, it seems unlikely that the miR171–MpSCL module controls chlorophyll biosynthesis. However, some effects on thallus morphology in Mpscl mutants, such as a stunted thallus with inward-curling edges, were observed (Fig. 1, lower right side) (Yelina et al. 2023). In Physcomitrium, PpGRAS12 is a validated target for miR171 (Axtell et al. 2007; Hiss et al. 2017). Deletion of the PpGRAS12 gene has a negative effect on sporophyte production, as fewer sporophytes were formed in ppgras12 knockout plants compared to wild-type plants. Additionally, at the protonema stage, highly selective and unique growth arrest was seen in plants with inducible PpGRAS12 overexpression (Beheshti et al. 2021). Although GRAS family members regulated by miR171 play important roles in shoot meristem differentiation and protonema proliferation, respectively, in Arabidopsis and Physcomitrium, more functional studies are required to determine the role of the Marchantia miR171–MpSCL module.
The miR156/529–SPL module is involved in the vegetative-to-reproductive phase transition
It has been shown that Arabidopsis miR156(a-h) targets transcripts of several SQUAMOSA PROMOTER BINDING PROTEIN-LIKE TF mRNAs (SPL2/3/4/5/9/10/11/13/15) and regulates the timing of flowering (Wang et al. 2009; Wang 2014; Wang and Wang 2015; Zheng et al. 2019; Jerome Jeyakumar et al. 2020). Marchantia miR529 shares 17 of the 21 nucleotides of Arabidopsis miR156 and could be considered an miRNA of the same family. Additionally, computational analyses found that miR529c has a complementary sequence to MpSPL2 mRNA in a similar context to the Arabidopsis miR156–SPL mRNA module. It was hypothesized that miR529c targets the SPL gene mRNA in Marchantia and might have a conserved/common biological function with Arabidopsis although liverwort will not set flowers. Once Mpmir529c-null mutants (Mpmir529cge mutants) were established with the use of the CRISPR/Cas9 system, surprisingly, the mutant thalli immediately started to transform into sexual-like reproductive organs (gametangia) and produced gametes. This development was observed even in the absence of far-red light and long-day conditions, both of which are applied normally to induce reproductive development on the bench. This result suggested that loss of miR529c function caused gametangium formation and gamete production without the environmental changes normally needed for the reproductive transition (Fig. 1, upper group). Next, an miR529c-resistant MpSPL2 transgene (mMpSPL2 gene) was established and introduced into wild-type plants. The transgene producing miR529c-resistant mRNA induced a reproductive transition in the whole area of thalli without far-red light conditions, as was seen in Mpmir529cge mutants. The MpSPL2 mRNA levels were elevated in both Mpmir529cge and mMpSPL2 mutant plants. It was thus suggested that the transition to reproductive development in liverworts is controlled by the miR156/529–SPL module in a similar manner to angiosperms, even though reproductive processes differ greatly between angiosperms and liverworts (Tsuzuki et al. 2019).
Interestingly, in the moss P. patens, both miR156 and miR529 are present. However, they differ in expression patterns, with miR156 mainly expressed in the protonema and miR529 mainly expressed in gametophores with developed sporophytes (Xie et al. 2021). Of 13 members of the Physcomitrium SPL family, only transcripts of three SPL genes are proven targets of miR156. These three moss miR156-targeted SPLs are in the same phylogenetic group as the miR156-targeted SPL genes in Arabidopsis (Riese et al. 2007, 2008; Streubel et al. 2023). Functional studies indicated that miR156 promotes the vegetative developmental transition from the protonema to the leafy gametophore in P. patens, while PpSPL3 acts in the opposite direction (Cho et al. 2012). The role of miR529 in sporophytes is still unknown.
The miR156/529–SPL module influences developmental transitions in the vegetative phase of the life cycle in both bryophytes and flowering plants. Interestingly, the directionality of miR156/529 action is conserved between liverworts and angiosperms, as the miRNAs repress the vegetative-to-reproductive phase transition. In moss, miR156 promotes the vegetative transition from young filamentous gametophytic tissue to the adult leafy shoot form.
The miR319–TCPs/MYB/MpRKD module governs vegetative and generative developmental pathways
In angiosperms, the conserved miR319/159 family is involved in various processes, such as leaf development, hormone signaling, organ curvature and cell proliferation (Palatnik et al. 2003; Allen et al. 2007; Rubio-Somoza et al. 2014; Schommer et al. 2014). Nearly half of TEOSINTE BRANCHED/CYCLOIDEA/PCF (TCP) mRNAs and R2R3-MYB TF mRNAs are targeted by miR319 in Arabidopsis. It has been accepted that miR159 and miR319 are both essential for the regulation of the developmental process in Arabidopsis. The jaw-D mutation was first identified by genetic screening in Arabidopsis; the mutant plants have serrated rosette leaves, epinastic cotyledons and crinkled leaves (Palatnik et al. 2003). At that time, it was revealed that miR319 is coded in the JAW locus and targets the mRNAs of some TCP TFs. The jaw-D mutant phenotype was caused by the overexpression of miR319a (Palatnik et al. 2003). In addition, mutations in the MIR319a and MIR319b genes caused a moderate reduction in the size of leaf serrations, while the MIR319a/b mutation almost completely suppressed their formation. This suggests that both these genes play a mostly quantitative role in the formation of leaf serrations (Koyama et al. 2017). Moreover, miR319 regulates senescence and jasmonate biosynthesis in Arabidopsis by targeting JAZ genes. It was shown that overexpression of miR319 delayed leaf senescence and reduced jasmonate levels, while inhibition of miR319 accelerated leaf senescence and increased jasmonate levels (Schommer et al. 2008). Based on several studies, it is generally accepted that Arabidopsis MIR319 and MIR159 were derived from a common ancestor miRNA gene, and they are often regarded as constituting one miRNA family due to high sequence similarity and common non-canonical processing steps from the loop-to-base direction before maturation (Palatnik et al. 2007; Bologna et al. 2009; Li et al. 2011). Based on degradome data, loop-to-base processing of pri-miR319 was also reported in the moss Physcomitrium (Addo-Quaye et al. 2009). Moreover, MIR159 is absent in mosses.
In Physcomitrium, miR319 is known to control the levels of MYB gene expression. Although there is a low degree of complementarity between miR319 and the MYB gene, experiments have shown that miR319 can guide the cleavage of the MYB mRNA (Xia et al. 2016). The intricate and multifaceted biological function of miR319, as well as relatively lesser-known functions of this miRNA in liverworts, encouraged research on Marchantia. Marchantia plants overexpressing MIR319b showed abnormal and obvious morphological phenotypes in thallus and gemma cup structure. It was demonstrated that thalli in 35 S:MpMIR319b plants were curled to the dorsal side. Additionally, this ectopic expression triggered the absence of visible gemma cups while gemmae were generated (Tsuzuki et al. 2016). Moreover, Mpmir319age and Mpmir319bge null mutants were established using the CRISPR/Cas9 system. They showed normal thallus development, which was intriguing, but they could produce a smaller number of gemmae and gemma cups on thalli with sizes similar to those of the wild type. Interestingly, Mpmir319a/bge double knockout plants showed even fewer gemma cups and gemmae than single mutants. Importantly, double mutant plants produced sexual reproductive organs (gametangia), but at a much-reduced frequency compared to wild-type plants. Additionally, these male and female gametangia showed irregular shapes, possibly reflecting asymmetric organ formation during their development (Futagami et al. 2023). RNA degradome analysis found a complementary sequence to miR319 in MpRKD (RWP-RK DOMAIN-CONTAINING) mRNA and a cleavage site (Lin et al. 2016). The complementary sequences are localized just downstream of the termination codon of the MpRKD mRNA. It was previously reported that MpRKD is preferentially expressed in the egg and sperm cells and that it is involved in germ cell formation (Koi et al. 2016). In addition, Mprkd mutants were described to show slightly asynchronous thallus growth and lacked outgrowths at the edges of gemma cups, although gemmae were produced. To determine whether the observed defects in mir319a and b mutants were caused by unregulated expression of the RKD gene, an miR319-resistant RKD gene was constructed under the direction of the RKD gene promoter and introduced into a wild-type liverwort. Plants expressing the resistant RKD gene showed no gemma cups or gemmae and phenocopied mir319a, mir319b and mir319a/b mutants quite well (Futagami et al. 2023). Marchantia has two TCP and 22 R2R3-MYB genes. Of these, a complementary sequence to miR319 could be found only in the MpMYB21 gene. RNA degradome and 5′ RACE analysis detected R2R3-MYB21 mRNA cleavage (Lin et al. 2016; Tsuzuki et al. 2016). As in the RKD gene, the mir319-resistant form of the MYB21 gene was established and placed under the direction of the MYB21 promoter then introduced into liverwort to obtain transgenic lines. The transgenic plants grew thalli of smaller size with gemma cups/gemmae at a similar density. The phenotype was almost identical to that of transgenic liverwort plants to which mir319-sensitive MYB21 was similarly introduced. The available data suggested that miR319a or miR319b downregulates MpRKD function, possibly in a spatial manner, and induces the pathway for gemma cup/gemma formation (Futagami et al. 2023). The implications of the presented findings are vast, as miR319 has been shown to regulate complex and diverse developmental pathways in plants, including Arabidopsis, Physcomitrium and Marchantia. As such, further investigation of the biological function of miR319 is crucial for a better general understanding of plant growth and development.
Marchantia-specific non-coding RNA–target modules
The MpFRH1–MpRSL1 module is involved in the development of various thallus structures
MpFRH1–MpRSL1 is an example of a unique module identified only in Marchantia so far. RSL (ROOT HAIR DEFECTIVE SIX-LIKE 1) class 1 genes encoding basic helix–loop–helix TFs are known to positively regulate the development of root hairs in vascular plants and rhizoids in liverwort and moss, respectively (Proust et al. 2016; Honkanen et al. 2018; Thamm et al. 2020). In Marchantia, a liverwort-specific miRNA called FEW RHIZOIDS1 (MpFRH1; also called Mp-miR11861; Tsuzuki et al. 2016; Thamm et al. 2020) was found to negatively regulate a single RSL class 1 gene, MpRSL1. In contrast, no FRH1 recognition sequence was detected in the RSL class 1 transcripts in the moss P. patens, the lycophyte Selaginella kraussiana or the angiosperm representative A. thaliana (Honkanen et al. 2018). Moreover, FRH1 miRNA was not found in the moss, lycophyte or angiosperm representatives. Hence, the FRH1 miRNA is thought to have originated only in the liverwort lineage. It is possible that this miRNA is also present in other liverwort species since a conserved target site has been identified in orthologs of the MpRSL1 gene in representatives from the classes Marchantiopsida and Jungermanniopsida (Honkanen et al. 2018). Mpfrh1 loss-of-function plants showed more rhizoid cell groups than wild-type plants. A similar phenotype of more groups of rhizoid cells was observed in Mprsl1 gain-of-function transgenic plants. Moreover, higher levels of MpRSL1 mRNA were present in Mpfrh1 loss-of-function mutants (Thamm et al. 2020). Hence, MpFRH1 miRNA was shown to act as a repressor of rhizoid cell development in Marchantia. Additionally, Mpfrh1 gain-of-function and Mprsl1 loss-of-function transgenic plants had no rhizoids or rarely developed rhizoids, produced empty gemma cups or were characterized by lower production of gemmae and lacked multicellular papillae. Therefore, it seems that MpRSL1 positively regulates the development of Marchantia thallus structures derived from a single epidermal cell: gemmae, rhizoids and papillae (Fig. 1, bottom left first group) (Honkanen et al. 2018; Thamm et al. 2020).
Although the function of RSL class 1 genes has been shown to be conserved, the mechanisms that regulate their expression differ between liverworts and angiosperms. In Arabidopsis, the GLABRA2 (GL2) TF negatively regulates RSL class 1 genes. The AtGL2 protein directly interacts with L1 box motifs present in the promoters of two RSL class 1 genes, AtRHD6 and AtRSL1 (Lin et al. 2015). Through this interaction, the AtGL2 protein inhibits the initiation of root hair development by suppressing the transcription of RSL class 1 genes. However, no miRNA–RSL1 modules have been identified thus far in angiosperms or mosses. Therefore, distinct land plant lineages have developed diverse and independent negative regulators to govern the expression of conserved genes responsible for the root or rhizoid development essential for terrestrial life.
The Mpo-MR-13–MpSPL1 module is involved in meristem dormancy control
SPL genes are found in all green plants, ranging from unicellular green algae to angiosperms, where they act as important regulators of diverse developmental processes (Chen et al. 2010; Preston and Hileman 2013; Xu et al. 2016). In angiosperms, many members of the SPL gene family are post-transcriptionally regulated by miR156/157, miR529 and miR535 (Xu et al. 2016; Li et al. 2022a; Tregear et al. 2022; Zhang et al. 2022). miR156/157–SPL regulatory pathways have been found to be conserved in most land plants. In Arabidopsis, 10 of 16 SPL genes have been found to be targets of miR156/157 (Wang et al. 2009; Preston and Hileman 2013; Xu et al. 2016). In M. polymorpha, four SPL genes have been identified: MpSPL1–MpSPL4 (Tsuzuki et al. 2016, 2019; Bowman et al. 2017; Streubel et al. 2023). Mpo-MR-13 (also called Mp-miR11671) was identified as a unique miRNA-targeting MpSPL1 gene in Marchantia (Lin et al. 2016; Tsuzuki et al. 2016; Streubel et al. 2023). Mpo-MR-13 might be a liverwort-specific miRNA, as its target sites were found to be conserved in SPL sequences of Jungermanniopsida and Marchantiopsida representatives, respectively (Streubel et al. 2023).
Phylogenetic studies have shown that, of the Arabidopsis SPL family, AtSPL8 is the closest relative of MpSPL1 (Flores-Sandoval et al. 2018; Streubel et al. 2023). However, AtSPL8 mRNA is not regulated by miR156. In Arabidopsis, AtSPL8, along with miR156-targeted SPLs, regulates anther development (Unte et al. 2003). However, there are no functional studies showing the involvement of MpSPL1 in reproductive organ development in Marchantia. Meristem dormancy is a feature of the shade-avoidance response observed in angiosperms, which results in repressing the outgrowth of lateral meristems. To investigate the possible roles of the Mpo-MR-13–MpSPL1 module in the shade-avoidance response in Marchantia, several mutant plants were generated. Mpspl1 loss-of-function plants showed a more branched phenotype than wild-type plants and no dormant meristems. On the contrary, Mpspl gain-of-function plants developed fewer active meristems and, consequently, had increased dormant meristems compared to wild-type plants. As expected, Mpo-mr-13 loss-of-function plants showed a similar phenotype to Mpspl1 gain-of-function plants, i.e., more dormant meristems (Streubel et al. 2023). Hence, the Mpo-MR-13–MpSPL1 module was shown to be important for meristem dormancy and branching during the shade-avoidance response in the thallus of Marchantia (Fig. 1, upper group). Additionally, this study demonstrated that the Mpo-MR-13–MpSPL1 module is controlled by PIF-mediated phytochrome signaling (Streubel et al. 2023). A similar dependence was proposed in Arabidopsis, in which PIF-mediated repression of the expression of several MIR156 genes releases the miR156-targeted SPLs from post-transcriptional control, enabling them to pursue the shade-avoidance mechanism (Xie et al. 2017). However, which AtSPL gene is involved in shade avoidance is still unknown. Therefore, distinct miRNAs but similar mechanisms of SPL-regulated meristem dormancy important for the shade-avoidance mechanism evolved independently in the liverwort and angiosperm lineages.
The MpFGMYB–MpSUF lncRNA module governs sex determination in Marchantia
Like 70% of other liverwort species, sex in Marchantia is determined by the sex chromosomes U (female) and V (male) (Renner et al. 2017; Hisanaga et al. 2019b). The U-chromosome has a “Feminizer” locus, which has been shown to be dominant, since plants possessing both U and V chromosomes ultimately develop female sexual organs. This U-linked gene from the Feminizer locus, named MpBPCU, is a member of the plant-specific Basic Pentacysteine (BPC)/Barley B Recombinant (BBR) TF family. MpBPCU was shown to be necessary for archegoniophore development, as Mpbpcu mutants do not produce sex organs. A genomic fragment containing the MpBPCU gene was sufficient to convert from a male to a female phenotype (archegoniophore development) when introduced into the background of male individuals; however, egg cells could not develop because of the lack of other U-linked genes essential for the differentiation of egg cells. An MpBPCU gametolog, MpBPCV, was found on the male chromosome. BPCU and BPCV genes function in reproductive phase induction in females and males, respectively, but only BPCU was reported to be the master sex determinant for female sex determination in Marchantia (Iwasaki et al. 2021). The Feminizer, an MpBPCU TF, was shown to regulate the autosomal sex-determining locus, MpFGMYB–SUF. The MpFGMYB–SUF module consists of the FEMALE GAMETOPHYTE MYB (MpFGMYB) TF gene and SUF (SUPPRESSOR OF FEMINIZATION), which encodes an lncRNA. MpFGMYB encodes an R2R3-MYB–type TF specifically expressed in archegoniophores to promote female sexual program progression. SUF is transcribed in an antisense direction from the promoter located at the 3′ end of the MpFGMYB locus. SUF is expressed mainly in antheridiophores, but it is also present in the male vegetative thalli. Expression of the SUF gene inhibits transcription of the MpFGMYB gene, thereby suppressing the female gametogenesis program in males. In females, the MpBPCU TF negatively regulates SUF expression by binding to GAGA motifs enriched in the SUF promoter region. Consequently, SUF is not transcribed, resulting in the successful expression of MpFGMYB, which further promotes feminization in plants (Hisanaga et al. 2019a) (Fig. 1, upper right side).
Comparing the genetic elements involved in sexual organ development in Physcomitrium and Marchantia, it is important to note that sex chromosomes are not present in P. patens. However, two homologs of MpFGMYB (Pp3c6_23360 and Pp3c5_7650) were found to be highly expressed in moss archegonia (Pu et al. 2020). Whether these two homologs are essential for female sexual organ development in mosses is not clear. On the other hand, in Arabidopsis, there are seven orthologs of MpFGMYB, of which three proteins, namely, MYB64, MYB98 and MYB119, have been reported to function in female gametogenesis (Hisanaga et al. 2019a, 2019b). AtMYB98 has been shown to be expressed specifically in synergid cells and is essential for their differentiation (Dubos et al. 2010). AtMYB64 and AtMYB119 are expressed transiently during embryo sac development and are especially crucial for embryo sac cellularization. AtMYB119 expression was shown to be regulated by histidine kinase CYTOKININ-INDEPENDENT 1 (CKI1). CKI1 expression was found mainly in the central cell nucleus and, to a lesser extent, in the egg cell nucleus (Pischke et al. 2002; Hejatko et al. 2003). CKI1 activity is necessary for the specification of the developmental program of the central cell and antipodal cells, with concomitant suppression of egg cell and synergid cell program development in central and antipodal cells (Yuan et al. 2016). Therefore, the role of FGMYBs in female gametophyte differentiation in Marchantia and Arabidopsis is retained, but their regulatory pathways are different.
The MpKNOX1–MpSUK1 lncRNA module is critical for sporophyte development
The eukaryotic life cycle alternates between the haploid and diploid phases. In phylogenetically diverse eukaryotes, paralogous homeodomain (HD) protein families are implicated in the haploid-to-diploid reprogramming of gene expression upon fertilization (Herskowitz 1989; Zhao et al. 2001; J. H. Lee et al. 2008; Hedgethorne et al. 2017). In the green plant lineage, they are represented by the KNOTTED1-like homeobox (KNOX) and BEL1-like homeodomain (BELL) protein families. Importantly, to fulfill their functions, KNOX and BELL genes need to be expressed in gametes to enable KNOX/BELL protein heterodimerization just after gamete fusion (Bowman et al. 2016). This HD protein system is also functional in Marchantia. Notably, gametic expression of class I MpKNOX1 and MpBELL2/3/4 genes is necessary for diploid sporophyte development, as revealed by the analysis of loss-of-function mutant plants. In the case of the MpKNOX1 gene, a similar mode of expression regulation by lncRNA was identified, as in the case of the MpFGMYB–MpSUF module. The MpKNOX1 gene is expressed specifically in mature egg cells, and, following fertilization, its expression is observed during sporophyte development until the future sporogenous cells become distinct. Interestingly, an antisense lncRNA SUPPRESSOR OF KNOX1 (MpSUK1) is transcribed from an intergenic promoter and overlaps with the last two exons of the MpKNOX1 locus. Unlike the expression profile observed for MpKNOX1, MpSUK1 expression is predominantly observed in antheridia, suggesting that this lncRNA may act in cis to repress MpKNOX1 expression during antheridium development (Fig. 1, upper right side) (Dierschke et al. 2021). Although similar expression activation of the three Physcomitrium class I KNOX genes (MKN2, MKN4 and MKN5) was observed, an lncRNA regulatory module was not found in any of these genes (Singer and Ashton 2007; Sakakibara et al. 2008). Instead, polycomb-mediated repressive chromatin modification is required to maintain gametophyte identity through repression of the sporophyte genetic program (Mosquna et al. 2009; Okano et al. 2009). In the case of triple loss-of-function of class I KNOX moss mutant plants (MKN2-4-5dis-1), sporophytes exhibited defects in cell proliferation during development (Singer and Ashton 2007; Sakakibara et al. 2008). Therefore, in both bryophytes, the class I KNOX genes retained the ancestral function of controlling the haploid-to-diploid transition, but the sporophytic expression pattern suggests their neofunctionalization as stimulators of cell proliferation in the diploid generation.
Further, during land plant evolution, class I KNOX genes were confined to functioning in the maintenance of sporophytic meristematic cells. In Arabidopsis, all four members of the class I KNOX family participate in a different developmental process throughout the sporophytic life phase. However, their expression in specific domains of SAM preserves the pool of indeterminate meristematic cells and specifies the boundaries for the propagation of lateral organs and the stem during plant development (Hay and Tsiantis 2010). For example, SHOOT MERISTEMLESS (STM) is the first class I KNOX gene expressed during early embryogenesis, and its constant expression is critical for shoot meristem formation and maintenance (Barton and Poethig 1993; Long et al. 1996; Scofield et al. 2007, 2008). Additionally, STM plays an essential role in the transient activity of the central floral stem cells and subsequently directs the formation of carpels and the placental tissues from which ovules arise (Long et al. 1996; Long and Barton 2000). The boundaries between class I KNOX-expressing and non-expressing cells need to be precisely established, which requires the repression of KNOX gene expression in developing organs. This repression is achieved by a complex regulatory network, in which proteins like ASYMMETRIC LEAVES1 and 2 (AS1 and AS2) bind directly to the KNOX1 gene promoters to repress their transcription. Also, the combined actions of two polycomb repressive complexes (PRCs) suppress class I KNOX transcription by chromatin modifications (Byrne et al. 2000; Ori et al. 2000; Xu et al. 2003; Katz et al. 2004; Guo et al. 2008; Xu and Shen 2008).
The available data on class I KNOX genes demonstrate that, in the representatives of different land plant branches, class I KNOX genes are crucial factors controlling the proliferation status of the diploid generation. Although class I KNOX genes have the same function in the studied bryophytes, the mechanisms that regulate their expression are different. In fact, the Physcomitrium KNOX gene repression mechanism using the PRC complex resembles the one of Arabidopsis, while in Marchantia the lncRNA MpSUK1 is the major regulator of the MpKNOX1 gene expression profile.
Coordination among non-coding RNA regulatory modules drives Marchantia polymorpha development
Despite the substantial heterogeneity in growth conditions and specialized multicellular body forms, it was recently reported that there is significant conservation of transcription programs across most organs in vascular plants. In the case of the Physcomitrium and Marchantia transcriptomes of leaf-like organs (leaflets and thallus, respectively), similarities but also prominent differences were observed when compared with the transcriptomes of vascular plants (Julca et al. 2021). Additionally, the specificity of regulatory programs is also visible in plant microtranscriptome compositions; for example, in Marchantia, of 265 miRNAs only seven are conserved with those of other land plants (Tsuzuki et al. 2016; Bowman et al. 2017; Pietrykowska et al. 2022).
The gathered data indicate that ncRNAs play crucial roles in controlling various developmental processes during the M. polymorpha life cycle. By targeting major TFs, Marchantia small RNAs coordinate the balance between plant growth and development. However, to properly fulfill their molecular functions, small RNAs must be precisely processed by multiprotein complexes. Mutations in machineries responsible for their biogenesis most often have pleiotropic effects on plant phenotypic features, as has been reported in Arabidopsis and many other plants (Jacobsen et al. 1999; Golden et al. 2002; Liu et al. 2005; Xie et al. 2005; Khraiwesh et al. 2010; Parent et al. 2015). A similar effect was also observed in Marchantia, when each of four DCL genes present in the Marchantia genome was mutated (Csicsely et al. 2023). It has been suggested that MpDCL1a is responsible for miRNA biogenesis since mutants of this gene showed the most severe abnormalities during development. Changes in antheridiophore morphology in Mpdcl1bge and Mpdcl4ge mutant plants and the lack of antheridiophore production in Mpdcl3ge suggest that these three proteins may be involved in the processing of a specific group of small RNAs that are important for the regulation of male reproductive branch formation. Therefore, the cooperative action of all four DCL proteins generates an interwoven network of miRNA and siRNA regulatory pathways that are necessary for successful vegetative and reproductive developmental programs in Marchantia (Fig. 3).
Putative new connections between regulatory pathways involved in vegetative and reproductive development in M. polymorpha. Colored boxes indicate experimentally confirmed non-coding RNA–target modules. Modules shown in blue and red boxes are involved in vegetative and reproductive development, respectively, whereas modules shown in a blue to red gradient are involved in both vegetative and reproductive development. Black bars indicate regulatory pathways that have been experimentally validated, gray arrows and bars indicate regulatory pathways suggested in the cited papers and dashed gray arrows and bars indicate regulatory pathways suggested in this review. The lowercase letters on the right sides of boxes indicate whether the gene or module is involved in the vegetative developmental program (a), the vegetative-to-reproductive phase transition (b) or the reproductive developmental program (c)
The most profound and robust impact on the Marchantia life cycle is exerted by the miR160–ARF clade C module, as nearly all aspects of growth and development were affected when MpARF3 or MpMIR160 activity was disturbed (Flores-Sandoval et al. 2018b). This result suggests that the MpARF3 gene is a superior factor in coordinating the cell proliferation and differentiation processes, resulting in maintaining an accurate balance between the meristematic and differentiated states in a cell or tissue. Interestingly, it was also proposed that MpARF3 forms a negative feedback loop with miR160 to regulate its own expression level. In Mparf3 mutant plants, the MpMIR160 transcript was significantly downregulated, while, in plants expressing an additional copy of the MpARF3 gene under its native promoter, the miR160 level was upregulated compared to in wild-type plants. Therefore, MpMIR160 transcription appears to depend on MpARF3 activity, suggesting that MpARF3 activates its own repressor (Fig. 3) (Flores‐Sandoval et al. 2018b). Gene co-expression analysis in different tissues and developmental stages of Marchantia revealed that MpARF3 forms a co-expression group with several genes that are related to the vegetative-to-reproductive phase transition and are highly active in reproductive structures. Among similarly expressed genes, three TFs were recognized: MpC3HDZ, MpSPL1 and MpSPL2. MpC3HDZ and MpSPL2 are post-transcriptionally regulated by conserved miR166 and miR529c, respectively, and MpSPL1 is targeted by Mpo-MR-13, which is unique to Marchantia (Flores-Sandoval et al. 2018a). Although the reported TF expression dynamics were similar to those of MpARF3, analysis of the transcriptomes of gain- and loss-of-function mutants of the miR160–MpARF3 regulatory module, along with mutant plant phenotype analysis, revealed that MpARF3 in fact inhibits the reproductive transition in M. polymorpha. MpARF3 activates the expression of Mp-MR-13 and Mp-MIR529c genes to downregulate mRNA levels of MpSPL1 and MpSPL2, respectively, resulting in prolonged vegetative growth (Flores-Sandoval et al. 2018a; Flores‐Sandoval 2018b). The relationship between MpARF3 and the miR166–MpC3HDZ module is not clear, as no additional analyses are available. From Arabidopsis studies, it is known that the HD-ZIPIII–miR165/166 signaling pathway, among others, is essential for normal SAM functions (Williams et al. 2005). Marchantia plants overexpressing miR166a revealed deregulated dorsoventral morphogenesis of the thallus, which might be the consequence of MpC3HDZ downregulation. Therefore, a properly balanced interplay of the miR160–MpARF3 and miR166–MpC3HDZ modules might be required for the precise coordination of cell proliferation and differentiation processes in the Marchantia thallus (Fig. 3).
As described in the previous section, the miR529c–MpSPL2 module was shown to control the vegetative-to-reproductive phase transition in Marchantia (Tsuzuki et al. 2019). In contrast, the Mpo-MR-13–MpSPL1 module is involved in the shade-avoidance mechanism. This module depends on the red/far-red light signaling pathway, with the MpPHY–MpPIF module promoting MpSPL1 activity by repressing Mpo-MR-13 gene transcription. Downregulation of Mpo-MR-13 results in meristem dormancy (Streubel et al. 2023). This phytochrome-dependent far-red high irradiance signaling also induces the sexual reproduction program (Inoue et al. 2019), and, as shown by Streubel et al. (2023), it promotes the expression of the MpSPL2 gene (Streubel et al. 2023). However, whether it acts through MpMIR529c repression or an increase in transcription of the MpSPL2 gene was not specified. According to transcriptomic data, the expression of MpSPL1, like that of MpSPL2, is upregulated in gametangiophores (Bowman et al. 2017; Flores-Sandoval et al. 2018). Therefore, besides its role in meristem dormancy in shade conditions, MpSPL1 might be an important factor for Marchantia sexual reproduction. These data raise the intriguing possibility that the MpSPL1 gene might play a dual function in Marchantia development. Upon phytochrome signaling during shade conditions, MpSPL1 first activates the meristem dormancy program. Second, it might enable the cellular program to switch to the reproductive program, which could be cooperatively orchestrated by both MpSPL1 and MpSPL2 TFs. Collectively, based on the available data, regulatory pathways involved in the vegetative-to-reproductive phase transition in Marchantia can be proposed (Fig. 3). The MpPHY–MpPIF module acts as a master regulator to control the activation of genes related to sexual reproduction. Among those, the expression of two Marchantia SPL family members is promoted through transcriptional repression of two miRNA genes, Mpo-MR-13 and MpMIR529c. Under low far-red light conditions, Mpo-MR-13 and miR529c, along with the MpARF3 TF, inhibit the reproductive program, so they prolong vegetative growth. As miR160 downregulates MpARF3 expression, it promotes the vegetative-to-reproductive phase transition and thus acts in concordance with the phytochrome signaling pathway. An interesting question arises: do the MpPHY–MpPIF and miR160–MpARF3 modules act separately, or does one of them influence the other? There is no information concerning the influence of a high far-red light regime on MpMIR160 or MpARF3 expression. However, similar to the Mpo-MR-13 gene promoter, in the promoter region of MpMIR160, there are two PBE-box motifs. The PBE-box motif is a regulatory sequence that could potentially be recognized by the MpPIF protein (Xie et al. 2017; Streubel et al. 2023). Although it has been proposed that MpPIF mediates the repression of Mpo-MR-13 expression by recognition of the PBE-box motif in its promoter region, MpPIF could selectively promote MpMIR160 expression, as was already shown for other Marchantia genes (Inoue et al. 2019).
Another layer of germline determination within the Marchantia vegetative thallus is MpPHY–MpPIF–mediated expression activation of the basic helix-loop-helix (bHLH) TF MpBONOBO (MpBNB), an autosomal gene (Yamaoka et al. 2018). Constitutive MpBNB expression, like constitutive MpSPL2 expression, was sufficient to induce gametangium and gametangiophore production, irrespective of light conditions. However, in contrast to MpSPL2, the lack of which caused only a delay in the development of sexual structures, knockout of MpBNB completely suppressed gametangium and gametangiophore production. These observations suggest that MpBNB is a master regulator of gametangial progenitor cell programming, whereas MpSPL2 stimulates the induction of a reproductive program, probably in cooperation with other factors, for example, MpSPL1. Both these genes have similar accumulation profiles, i.e., both are upregulated in the meristematic region (Yamaoka et al. 2018; Tsuzuki et al. 2019). Interestingly, within the 5 kb promoter region of the MpBNB gene, four SBP-responsive elements can be found (SBP–SPL response element). Therefore, the miR156/529–MpSPL2 module may be one of a few coordinately working units assisting the phytochrome-mediated activation of MpBNB expression (Fig. 3). Although MpBNB expression is restricted to gametangium initial cells, this activation is a prerequisite for further gametangium development and sexual branch formation. How MpBNB influences further germ cell differentiation and gametophore development is not clear, and the mechanism is still unknown. Similar effects on sexual organ formation show MpBPCU and MpBPCV gametologs, which are essential for reproductive phase induction in females and males, respectively. However, unlike MpBNB, MpBPCU and MpBPCV are expressed already in seven-day-old female and male thalli (Iwasaki et al. 2021). Although they are present during the vegetative phase of growth, they are not sufficient to trigger a switch from the vegetative program to the reproductive program. The data showing that MpBNB is a master gene for reproductive program initiation suggest that the MpBNB protein may be important for MpBPCU and MpBPCV protein activation so that they can work together to direct gametangiophore and gamete development. It is also possible that some other undiscovered factor(s) induced by far-red light might be important for the activity of the MpBPCU and MpBPCV proteins, leading to gametogenesis (Fig. 3). Importantly, the U-chromosome–localized BPCU gene is a feminizer locus that, by repression of the SUF lncRNA, enables FGMYB gene expression, which further initiates female program continuation. BPCV has no effect on the FGMYB–SUF locus but is important for proper male program continuation (Fig. 3). Although recently it was shown that the miR319–MpRKD module plays a role in proper gemma/gemma cup development (Futagami et al. 2023), an earlier study described MpRKD as a crucial player for the final steps of egg and sperm cell development (Koi et al. 2016; Rovekamp et al. 2016). However, the effect of miR319 during reproductive organ development is unclear, as its expression profile was not examined in parallel with MpRKD expression. Additionally, though MpRKD acts downstream of the MpBPCU/V gametolog and MpFGMYB–SUF module, whether one of these TFs promotes MpRKD expression is also unknown (Fig. 3). Therefore, investigation of functional relationships among genes engaged in the successful progression of each step during the reproductive phase of the Marchantia life cycle is needed.
Although several conserved miRNA–target modules were identified in Marchantia, some of them still lack detailed functional studies, for example the miR390–TAS3 module. In angiosperms, this module controls the expression levels of clade B ARF members. Although small RNA analysis revealed the presence of tasiARFs that could target the MpARF2 gene and tasiAP2, which could regulate members of the MpAP2 family (Fig. 3), the role of the miR390–TAS3 module during the Marchantia life cycle is still unresolved. In angiosperms, the miR390–TAS3–ARF module is indispensable for the regulation of auxin signaling and for leaf morphology, among other functions (Chitwood et al. 2009; Marin et al. 2010; Skopelitis et al. 2012). As described earlier, the adaxial-abaxial cell fate program is controlled by miR165/166–HD-ZIPIII, and ARF3/ARF4 TFs are regulated by tasiARFs (McConnell et al. 2001; Otsuga et al. 2001; Emery et al. 2003; Chitwood et al. 2009; Skopelitis et al. 2012). Together, these interactions exactly define and preserve the boundaries between the adaxial and abaxial sides of the leaf. As both regulatory modules, miR166–HD-ZIPIII and miR390–TAS3–ARF, are present in the M. polymorpha genome, an interesting question arises: are there similar dependencies of the functions of these two modules in the Marchantia thallus as observed during angiosperm leaf development? The thallus clearly shows characteristic dorsiventrality, with zonation of the upper assimilatory region, forming air chambers and gemma cups, the middle storage region with parenchymal cells and the ventral epidermis with scales and rhizoids (Shimamura 2016). Therefore, detailed functional studies are needed to investigate whether these two conserved small RNA modules may act in concert to establish each region’s boundaries and whether they influence one another (Fig. 3).
Since the discovery of thousands of lncRNAs based on genome-wide analysis of plant transcriptomic data, the functional relevance of these RNAs has been demonstrated in only a few cases (Chen et al. 2020; Wierzbicki et al. 2021). Within the Marchantia transcriptome, almost 30,000 transcripts were described as undefined due to the lack of similarity to currently available plant and animal genomic data. It was suggested that some of these transcripts might be portions of long UTRs or encode lncRNAs (Lin et al. 2016). SUF is an example of such a functional lncRNA expressed exclusively in male individuals to suppress the expression of MpFGMYB (Hisanaga et al. 2019a). A similar mode of action was observed in the case of MpSUK1 lncRNA, which represses MpKNOX1 expression in antheridia (Dierschke et al. 2021). Both MpSUF and MpSUK1 are involved in sexual reproduction (Fig. 3), which is the function of many lncRNAs observed across a range of eukaryotic species (Golicz et al. 2018). Currently, lncRNAs are emerging as potentially important players in various fundamental biological processes in plants, such as organ morphogenesis, tolerance to stress and response to pathogenic infection. Therefore, further research on lncRNAs in Marchantia is needed to learn what other processes, in addition to sexual reproduction, are supervised by lncRNAs. These studies will also increase our understanding of the mechanism of lncRNA function in non-seed plants and will indicate which aspects of lncRNA biology are universal and which are locus specific.
As we have shown, there are demonstrated and hypothetical interconnections among the ncRNA–target modules in different developmental processes during the Marchantia life cycle. Several are crucial for the successful progression through the vegetative phase of growth, while others are related to enabling the sexual reproduction program. However, our present understanding of these genetic networks represents only a skeletal framework, as many functional studies are still lacking. Therefore, future studies are needed to understand the biochemical functions of the yet uncharacterized module components, along with the manners in which the signaling pathways transmit the information that ultimately regulates the appropriate developmental stage.
Data availability
Enquiries about data availability should be directed to the authors.
References
Addo-Quaye C, Snyder JA, Park YB, Li YF, Sunkar R, Axtell MJ (2009) Sliced microRNA targets and precise loop-first processing of MIR319 hairpins revealed by analysis of the Physcomitrella patens degradome. RNA 15(12):2112–2121. https://doi.org/10.1261/rna.1774909
Akbergenov R, Si-Ammour A, Blevins T, Amin I, Kutter C, Vanderschuren H, Zhang P, Gruissem W, Meins F Jr., Hohn T, Pooggin MM (2006) Molecular characterization of geminivirus-derived small RNAs in different plant species. Nucleic Acids Res 34(2):462–471. https://doi.org/10.1093/nar/gkj447
Alaba S, Piszczalka P, Pietrykowska H, Pacak AM, Sierocka I, Nuc PW, Singh K, Plewka P, Sulkowska A, Jarmolowski A, Karlowski WM, Szweykowska-Kulinska Z (2015) The liverwort Pellia endiviifolia shares microtranscriptomic traits that are common to green algae and land plants. New Phytol 206(1):352–367. https://doi.org/10.1111/nph.13220
Allen RS, Li J, Stahle MI, Dubroue A, Gubler F, Millar AA (2007) Genetic analysis reveals functional redundancy and the major target genes of the Arabidopsis miR159 family. Proc Natl Acad Sci U S A 104(41):16371–16376. https://doi.org/10.1073/pnas.0707653104
Arif MA, Fattash I, Ma Z, Cho SH, Beike AK, Reski R, Axtell MJ, Frank W (2012) DICER-LIKE3 activity in Physcomitrella patens DICER-LIKE4 mutants causes severe developmental dysfunction and sterility. Mol Plant 5(6):1281–1294. https://doi.org/10.1093/mp/sss036
Arif MA, Top O, Csicsely E, Lichtenstern M, Beheshti H, Adjabi K, Frank W (2022) DICER-LIKE1a autoregulation based on intronic microRNA processing is required for stress adaptation in Physcomitrium patens. Plant J 109(1):227–240. https://doi.org/10.1111/tpj.15570
Axtell MJ (2013) Classification and comparison of small RNAs from plants. Annu Rev Plant Biol 64:137–159. https://doi.org/10.1146/annurev-arplant-050312-120043
Axtell MJ, Jan C, Rajagopalan R, Bartel DP (2006) A two-hit trigger for siRNA biogenesis in plants. Cell 127(3):565–577. https://doi.org/10.1016/j.cell.2006.09.032
Axtell MJ, Snyder JA, Bartel DP (2007) Common functions for diverse small RNAs of land plants. Plant Cell 19(6):1750–1769. https://doi.org/10.1105/tpc.107.051706
Baev V, Milev I, Naydenov M, Vachev T, Apostolova E, Mehterov N, Gozmanva M, Minkov G, Sablok G, Yahubyan G (2014) Insight into small RNA abundance and expression in high- and low-temperature stress response using deep sequencing in Arabidopsis. Plant Physiol Biochem 84:105–114. https://doi.org/10.1016/j.plaphy.2014.09.007
Bajczyk M, Jarmolowski A, Jozwiak M, Pacak A, Pietrykowska H, Sierocka I, Swida-Barteczka A, Szewc L, Szweykowska-Kulinska Z (2023) Recent Insights into Plant miRNA Biogenesis: Multiple Layers of miRNA Level Regulation. Plants https://doi.org/10.3390/plants12020342
Balzan S, Johal GS, Carraro N (2014) The role of auxin transporters in monocots development. Front Plant Sci 5:393
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116(2):281–297. https://doi.org/10.1016/s0092-8674(04)00045-5
Barton MK, Poethig RS (1993) Formation of the shoot apical meristem in Arabidopsis thaliana: an analysis of development in the wild type and in the shoot meristemless mutant. Development 119(3):823–831. https://doi.org/10.1242/dev.119.3.823
Beheshti H, Strotbek C, Arif MA, Klingl A, Top O, Frank W (2021) PpGRAS12 acts as a positive regulator of meristem formation in Physcomitrium patens. Plant Mol Biol 107(4–5):293–305. https://doi.org/10.1007/s11103-021-01125-z
Belanger S, Zhan J, Meyers BC (2023) Phylogenetic analyses of seven protein families refine the evolution of small RNA pathways in green plants. Plant Physiol 192(2):1183–1203. https://doi.org/10.1093/plphys/kiad141
Blevins T, Rajeswaran R, Shivaprasad PV, Beknazariants D, Si-Ammour A, Park HS, Vazquez F, Robertson D, Meins F Jr., Hohn T, Pooggin MM (2006) Four plant dicers mediate viral small RNA biogenesis and DNA virus induced silencing. Nucleic Acids Res 34(21):6233–6246. https://doi.org/10.1093/nar/gkl886
Bolle C (2004) The role of GRAS proteins in plant signal transduction and development. Planta 218(5):683–692. https://doi.org/10.1007/s00425-004-1203-z
Bologna NG, Voinnet O (2014) The diversity, biogenesis, and activities of endogenous silencing small RNAs in Arabidopsis. Annu Rev Plant Biol 65:473–503. https://doi.org/10.1146/annurev-arplant-050213-035728
Bologna NG, Mateos JL, Bresso EG, Palatnik JF (2009) A loop-to-base processing mechanism underlies the biogenesis of plant microRNAs miR319 and miR159. EMBO J 28(23):3646–3656. https://doi.org/10.1038/emboj.2009.292
Bowman JL (2022) The liverwort Marchantia polymorpha, a model for all ages. Curr Top Dev Biol 147:1–32. https://doi.org/10.1016/bs.ctdb.2021.12.009
Bowman JL, Sakakibara K, Furumizu C, Dierschke T (2016) Evolution in the cycles of life. Annu Rev Genet 50:133–154. https://doi.org/10.1146/annurev-genet-120215-035227
Bowman JL, Kohchi T, Yamato KT, Jenkins J, Shu SQ, Ishizaki K, Yamaoka S, Nishihama R, Nakamura Y, Berger F, Adam C, Aki SS, Althoff F, Araki T, Arteaga-Vazquez MA, Balasubrmanian S, Barry K, Bauer D, Boehm CR, Schmutz J (2017) Insights into Land Plant Evolution garnered from the Marchantia polymorpha genome. Cell 171(2):287–. https://doi.org/10.1016/j.cell.2017.09.030
Byrne ME, Barley R, Curtis M, Arroyo JM, Dunham M, Hudson A, Martienssen RA (2000) Asymmetric leaves1 mediates leaf patterning and stem cell function in Arabidopsis. Nature 408(6815):967–971. https://doi.org/10.1038/35050091
Chapman EJ, Estelle M (2009) Mechanism of auxin-regulated gene expression in plants. Annu Rev Genet 50:265–285. https://doi.org/10.1146/annurev-genet-102108-134148
Chen X (2005) MicroRNA biogenesis and function in plants. FEBS Lett 579(26):5923–5931. https://doi.org/10.1016/j.febslet.2005.07.071
Chen X, Zhang Z, Liu D, Zhang K, Li A, Mao L (2010) SQUAMOSA promoter-binding protein-like transcription factors: star players for plant growth and development. J Integr Plant Biol 52(11):946–951. https://doi.org/10.1111/j.1744-7909.2010.00987.x
Chen L, Zhu QH, Kaufmann K (2020) Long non-coding RNAs in plants: emerging modulators of gene activity in development and stress responses. Planta 252(5):92. https://doi.org/10.1007/s00425-020-03480-5
Chitwood DH, Nogueira FTS, Howell MD, Montgomery TA, Carrington JC, Timmermans MCP (2009) Pattern formation via small RNA mobility. Genes Dev 23(5):549–554. https://doi.org/10.1101/gad.1770009
Cho SH, Addo-Quaye C, Coruh C, Arif MA, Ma Z, Frank W, Axtell MJ (2008) Physcomitrella patens DCL3 is required for 22–24 nt siRNA accumulation, suppression of retrotransposon-derived transcripts, and normal development. PLoS Genet 4(12):e1000314. https://doi.org/10.1371/journal.pgen.1000314
Cho SH, Coruh C, Axtell MJ (2012) miR156 and miR390 regulate tasiRNA accumulation and developmental timing in Physcomitrella patens. Plant Cell 24(12):4837–4849. https://doi.org/10.1105/tpc.112.103176
Coruh C, Cho SH, Shahid S, Liu Q, Wierzbicki A, Axtell MJ (2015) Comprehensive Annotation of Physcomitrella patens small RNA loci reveals that the heterochromatic short interfering RNA pathway is largely conserved in land plants. Plant Cell 27(8):2148–2162. https://doi.org/10.1105/tpc.15.00228
Cove D, Bezanilla M, Harries P, Quatrano R (2006) Mosses as model systems for the study of metabolism and development. Annu Rev Plant Biol 57:497–520. https://doi.org/10.1146/annurev.arplant.57.032905.105338
Csicsely E, Oberender A, Georgiadou AS, Gutsche N, Zachgo S, Top O, Frank W (2023) Identification and characterization of DICER-LIKE genes and their roles in Marchantia polymorpha development and stress adaptation. bioRxiv. https://doi.org/10.1101/2023.02.03.526932
Deleris A, Gallego-Bartolome J, Bao J, Kasschau KD, Carrington JC, Voinnet O (2006) Hierarchical action and inhibition of plant dicer-like proteins in antiviral defense. Science 313(5783):68–71. https://doi.org/10.1126/science.1128214
Di Laurenzio L, Wysocka-Diller J, Malamy JE, Pysh L, Helariutta Y, Freshour G, Hahn MG, Feldmann KA, Benfey PN (1996) The SCARECROW gene regulates an asymmetric cell division that is essential for generating the radial organization of the Arabidopsis root. Cell 86(3):423–433. https://doi.org/10.1016/s0092-8674(00)80115-4
Dierschke T, Flores-Sandoval E, Rast-Somssich MI, Althoff F, Zachgo S, Bowman JL (2021) Gamete expression of TALE class HD genes activates the diploid sporophyte program in Marchantia polymorpha. Elife 10:e57088
Donoghue PCJ, Harrison CJ, Paps J, Schneider H (2021) The evolutionary emergence of land plants. Curr Biol 31(19):R1281–R1298. https://doi.org/10.1016/j.cub.2021.07.038
Dubos C, Stracke R, Grotewold E, Weisshaar B, Martin C, Lepiniec L (2010) MYB transcription factors in Arabidopsis. Trends Plant Sci 15(10):573–581. https://doi.org/10.1016/j.tplants.2010.06.005
Emery JF, Floyd SK, Alvarez J, Eshed Y, Hawker NP, Izhaki A, Baum SF, Bowman JL (2003) Radial patterning of Arabidopsis shoots by class IIIHD-ZIP and KANADI genes. Curr Biol 13(20):1768–1774. https://doi.org/10.1016/j.cub.2003.09.035
Endo Y, Iwakawa HO, Tomari Y (2013) Arabidopsis ARGONAUTE7 selects miR390 through multiple checkpoints during RISC assembly. EMBO Rep 14(7):652–658. https://doi.org/10.1038/embor.2013.73
Engstrom EM (2012) HAM proteins promote organ indeterminacy: but how? Plant Signal Behav 7(2):227–234. https://doi.org/10.4161/psb.18958
Fahlgren N, Montgomery TA, Howell MD, Allen E, Dvorak SK, Alexander AL, Carrington JC (2006) Regulation of AUXIN RESPONSE FACTOR3 by TAS3 ta-siRNA affects developmental timing and patterning in Arabidopsis. Curr Biol 16(9):939–944. https://doi.org/10.1016/j.cub.2006.03.065
Falz A-L, Müller-Schüssele SJ (2019) Physcomitrella as a model system for plant cell biology and organelle–organelle communication. Curr Opin Plant Biol 52:7–13. https://doi.org/10.1016/j.pbi.2019.05.007
Finet C, Jaillais Y (2012) Auxology: when auxin meets plant evo-devo. Dev Biol 369(1):19–31. https://doi.org/10.1016/j.ydbio.2012.05.039
Flores-Sandoval E, Romani F, Bowman JL (2018) Co-expression and transcriptome analysis of Marchantia polymorpha transcription factors supports class C ARFs as independent actors of an ancient auxin regulatory module. Front Plant Sci. https://doi.org/10.3389/fpls.2018.01345
Flores-Sandoval E, Eklund DM, Hong SF, Alvarez JP, Fisher TJ, Lampugnani ER, Golz JF, Vázquez‐Lobo A, Dierschke T, Lin SS (2018) Class C ARF s evolved before the origin of land plants and antagonize differentiation and developmental transitions in Marchantia polymorpha. New Phytol 218(4):1612–1630. https://doi.org/10.1111/nph.15090
Floyd SK, Bowman JL (2004) Gene regulation: ancient microRNA target sequences in plants. Nature 428(6982):485–486. https://doi.org/10.1038/428485a
Fukudome A, Kanaya A, Egami M, Nakazawa Y, Hiraguri A, Moriyama H, Fukuhara T (2011) Specific requirement of DRB4, a dsRNA-binding protein, for the in vitro dsRNA-cleaving activity of Arabidopsis Dicer-like 4. RNA 17(4):750–760. https://doi.org/10.1261/rna.2455411
Fusaro AF, Matthew L, Smith NA, Curtin SJ, Dedic-Hagan J, Ellacott GA, Watson JM, Wang MB, Brosnan C, Carroll BJ, Waterhouse PM (2006) RNA interference-inducing hairpin RNAs in plants act through the viral defence pathway. EMBO Rep 7(11):1168–1175. https://doi.org/10.1038/sj.embor.7400837
Futagami K, Tsuzuki M, Watanabe Y (2023) MpmiR319 promotes gemma/gemma cup formation in the Liverwort Marchantia polymorpha. bioRxiv. https://doi.org/10.1101/2023.09.29.560093
Garcia D, Collier SA, Byrne ME, Martienssen RA (2006) Specification of leaf polarity in Arabidopsis via the trans-acting siRNA pathway. Curr Biol 16(9):933–938. https://doi.org/10.1016/j.cub.2006.03.064
Golden TA, Schauer SE, Lang JD, Pien S, Mushegian AR, Grossniklaus U, Meinke DW, Ray A (2002) SHORT INTEGUMENTS1/SUSPENSOR1/CARPEL FACTORY, a Dicer homolog, is a maternal effect gene required for embryo development in Arabidopsis. Plant Physiol 130(2):808–822. https://doi.org/10.1104/pp.003491
Golicz AA, Bhalla PL, Singh MB (2018) lncRNAs in plant and animal sexual Reproduction. Trends Plant Sci 23(3):195–205. https://doi.org/10.1016/j.tplants.2017.12.009
Gomes GLB, Scortecci KC (2021) Auxin and its role in plant development: structure, signalling, regulation and response mechanisms. Plant Biol (Stuttg) 23(6):894–904. https://doi.org/10.1111/plb.13303
Guo MJ, Thomas J, Collins G, Timmermans MCP (2008) Direct repression of KNOX loci by the ASYMMETRIC LEAVES1 complex of Arabidopsis. Plant Cell 20(1):48–58. https://doi.org/10.1105/tpc.107.056127
Hamilton AJ, Baulcombe DC (1999) A species of small antisense RNA in posttranscriptional gene silencing in plants. Science 286(5441):950–952. https://doi.org/10.1126/science.286.5441.950
Hamilton A, Voinnet O, Chappell L, Baulcombe D (2002) Two classes of short interfering RNA in RNA silencing. EMBO J 21(17):4671–4679
Han H, Zhou Y (2022) Function and regulation of microRNA171 in plant stem cell homeostasis and developmental programing. Int J Mol Sci. https://doi.org/10.3390/ijms23052544
Hay A, Tsiantis M (2010) KNOX genes: versatile regulators of plant development and diversity. Development 137(19):3153–3165. https://doi.org/10.1242/dev.030049
Hedgethorne K, Eustermann S, Yang J-C, Ogden TE, Neuhaus D, Bloomfield G (2017) Homeodomain-like DNA binding proteins control the haploid-to-diploid transition in Dictyostelium. Sci Adv 3(9):e1602937
Hejatko J, Pernisova M, Eneva T, Palme K, Brzobohaty B (2003) The putative sensor histidine kinase CKI1 is involved in female gametophyte development in Arabidopsis. Mol Genet Genomics 269(4):443–453. https://doi.org/10.1007/s00438-003-0858-7
Henderson IR, Zhang X, Lu C, Johnson L, Meyers BC, Green PJ, Jacobsen SE (2006) Dissecting Arabidopsis thaliana DICER function in small RNA processing, gene silencing and DNA methylation patterning. Nat Genet 38(6):721–725. https://doi.org/10.1038/ng1804
Herskowitz I (1989) A regulatory hierarchy for cell specialization in yeast. Nature 342(6251):749–757. https://doi.org/10.1038/342749a0
Hisanaga T, Okahashi K, Yamaoka S, Kajiwara T, Nishihama R, Shimamura M, Yamato KT, Bowman JL, Kohchi T, Nakajima K (2019a) A cis-acting bidirectional transcription switch controls sexual dimorphism in the liverwort. Embo J. https://doi.org/10.15252/embj.2018100240
Hisanaga T, Yamaoka S, Kawashima T, Higo A, Nakajima K, Araki T, Kohchi T, Berger F (2019b) Building new insights in plant gametogenesis from an evolutionary perspective. Nat Plants. https://doi.org/10.1038/s41477-019-0466-0
Hiss M, Meyberg R, Westermann J, Haas FB, Schneider L, Schallenberg-Rudinger M, Ullrich KK, Rensing SA (2017) Sexual reproduction, sporophyte development and molecular variation in the model moss Physcomitrella patens: introducing the ecotype Reute. Plant J 90(3):606–620. https://doi.org/10.1111/tpj.13501
Honkanen S, Thamm A, Arteaga-Vazquez MA, Dolan L (2018) Negative regulation of conserved RSL class I bHLH transcription factors evolved independently among land plants. Elife. https://doi.org/10.7554/eLife.38529
Hou J, Lu D, Mason AS, Li B, Xiao M, An S, Fu D (2019) Non-coding RNAs and transposable elements in plant genomes: emergence, regulatory mechanisms and roles in plant development and stress responses. Planta 250(1):23–40. https://doi.org/10.1007/s00425-019-03166-7
Huang Y, Kendall T, Forsythe ES, Dorantes-Acosta A, Li S, Caballero-Perez J, Chen X, Arteaga-Vazquez M, Beilstein MA, Mosher RA (2015) Ancient origin and recent innovations of RNA polymerase IV and V. Mol Biol Evol 32(7):1788–1799. https://doi.org/10.1093/molbev/msv060
Husbands AY, Chitwood DH, Plavskin Y, Timmermans MC (2009) Signals and prepatterns: new insights into organ polarity in plants. Genes Dev 23(17):1986–1997. https://doi.org/10.1101/gad.1819909
Inoue K, Nishihama R, Araki T, Kohchi T (2019) Reproductive induction is a far-red high irradiance response that is mediated by phytochrome and PHYTOCHROME INTERACTING FACTOR in Marchantia polymorpha. Plant Cell Physiol 60(5):1136–1145. https://doi.org/10.1093/pcp/pcz029
Ishizaki K, Nishihama R, Yamato KT, Kohchi T (2016) Molecular Genetic Tools and techniques for Marchantia polymorpha Research. Plant Cell Physiol 57(2):262–270. https://doi.org/10.1093/pcp/pcv097
Iwasaki M, Kajiwara T, Yasui Y, Yoshitake Y, Miyazaki M, Kawamura S, Suetsugu N, Nishihama R, Yamaoka S, Wanke D, Hashimoto K, Kuchitsu K, Montgomery SA, Singh S, Tanizawa Y, Yagura M, Mochizuki T, Sakamoto M, Nakamura Y, Kohchi T (2021) Identification of the sex-determining factor in the liverwort Marchantia polymorpha reveals unique evolution of sex chromosomes in a haploid system. Curr Biol 31(24):5522–5532e5527. https://doi.org/10.1016/j.cub.2021.10.023
Jacobsen SE, Running MP, Meyerowitz EM (1999) Disruption of an RNA helicase/RNAse III gene in Arabidopsis causes unregulated cell division in floral meristems. Development 126(23):5231–5243. https://doi.org/10.1242/dev.126.23.5231
Jerome Jeyakumar JM, Ali A, Wang WM, Thiruvengadam M (2020) Characterizing the role of the miR156-SPL Network in plant development and stress response. Plants. https://doi.org/10.3390/plants9091206
Juarez M, Timmermans M (2004) MiRNAs specify dorsoventral polarity during leaf development. Dev Biol 271(2):551–552
Julca I, Ferrari C, Flores-Tornero M, Proost S, Lindner A-C, Hackenberg D, Steinbachová L, Michaelidis C, Pereira G, S., Misra CS (2021) Comparative transcriptomic analysis reveals conserved programmes underpinning organogenesis and reproduction in land plants. Nat Plants 7(8):1143–1159. https://doi.org/10.1038/s41477-021-00958-2
Katz A, Oliva M, Mosquna A, Hakim O, Ohad N (2004) FIE and CURLY LEAF polycomb proteins interact in the regulation of homeobox gene expression during sporophyte development. Plant J 37(5):707–719. https://doi.org/10.1111/j.1365-313X.2003.01996.x
Khraiwesh B, Arif MA, Seumel GI, Ossowski S, Weigel D, Reski R, Frank W (2010) Transcriptional control of Gene expression by MicroRNAs. Cell 140(1):111–122. https://doi.org/10.1016/j.cell.2009.12.023
Kidner CA, Martienssen RA (2004) Spatially restricted microRNA directs leaf polarity through ARGONAUTE1. Nature 428(6978):81–84. https://doi.org/10.1038/nature02366
Kim J, Jung JH, Reyes JL, Kim YS, Kim SY, Chung KS, Kim JA, Lee M, Lee Y, Kim N, Chua V, N. H., Park CM (2005) microRNA-directed cleavage of ATHB15 mRNA regulates vascular development in Arabidopsis inflorescence stems. Plant J 42(1):84–94. https://doi.org/10.1111/j.1365-313X.2005.02354.x
Koi S, Hisanaga T, Sato K, Shimamura M, Yamato KT, Ishizaki K, Kohchi T, Nakajima K (2016) An evolutionarily conserved plant RKD factor controls germ cell differentiation. Curr Biol 26(13):1775–1781. https://doi.org/10.1016/j.cub.2016.05.013
Koyama T, Sato F, Ohme-Takagi M (2017) Roles of miR319 and TCP Transcription Factors in Leaf Development. Plant Physiol 175(2):874–885. https://doi.org/10.1104/pp.17.00732
Kozomara A, Birgaoanu M, Griffiths-Jones S (2019) miRBase: from microRNA sequences to function. Nucleic Acids Res 47(D1):D155–D162. https://doi.org/10.1093/nar/gky1141
Krasnikova MS, Goryunov DV, Troitsky AV, Solovyev AG, Ozerova LV, Morozov SY (2013) Peculiar evolutionary history of miR390-guided TAS3-like genes in land plants. Sci World J. https://doi.org/10.1155/2013/924153
Kurihara Y, Watanabe Y (2004) Arabidopsis micro-RNA biogenesis through Dicer-like 1 protein functions. Proc Natl Acad Sci U S A 101(34):12753–12758. https://doi.org/10.1073/pnas.0403115101
Lee JH, Lin HW, Joo S, Goodenough U (2008) Early sexual origins of homeoprotein heterodimerization and evolution of the plant KNOX/BELL family. Cell 133(5):829–840. https://doi.org/10.1016/j.cell.2008.04.028
Lee MH, Kim B, Song SK, Heo JO, Yu NI, Lee SA, Kim M, Kim DG, Sohn SO, Lim CE, Chang KS, Lee MM, Lim J (2008) Large-scale analysis of the GRAS gene family in Arabidopsis thaliana. Plant Mol Biol 67(6):659–670. https://doi.org/10.1007/s11103-008-9345-1
Leibfried A, To JP, Busch W, Stehling S, Kehle A, Demar M, Kieber JJ, Lohmann JU (2005) WUSCHEL controls meristem function by direct regulation of cytokinin-inducible response regulators. Nature 438(7071):1172–1175. https://doi.org/10.1038/nature04270
Li Y, Li C, Ding G, Jin Y (2011) Evolution of MIR159/319 microRNA genes and their post-transcriptional regulatory link to siRNA pathways. BMC Evol Biol 11:122. https://doi.org/10.1186/1471-2148-11-122
Li SB, Xie ZZ, Hu CG, Zhang JZ (2016) A review of Auxin Response factors (ARFs) in plants. Front Plant Sci 7:47. https://doi.org/10.3389/fpls.2016.00047
Li Y, Zhang S, Zhang D, Li X, Gao Z, Jiang Z (2022) The miR166-mRNA network regulates vascular tissue differentiation in Moso bamboo. Front Genet 13:893956. https://doi.org/10.3389/fgene.2022.893956
Li L, Shi F, Wang G, Guan Y, Zhang Y, Chen M, Chang J, Yang G, He G, Wang Y, Li Y (2022) Conservation and divergence of SQUAMOSA-PROMOTER BINDING PROTEIN-LIKE (SPL) gene family between wheat and rice. Int J Mol Sci. https://doi.org/10.3390/ijms23042099
Lin SS, Bowman JL (2018) MicroRNAs in Marchantia polymorpha. New Phytol 220(2):409–416. https://doi.org/10.1111/nph.15294
Lin Q, Ohashi Y, Kato M, Tsuge T, Gu H, Qu LJ, Aoyama T (2015) GLABRA2 directly suppresses Basic Helix-Loop-Helix transcription factor genes with diverse functions in Root Hair Development. Plant Cell 27(10):2894–2906. https://doi.org/10.1105/tpc.15.00607
Lin PC, Lu CW, Shen BN, Lee GZ, Bowman JL, Arteaga-Vazquez MA, Liu LYD, Hong SF, Lo CF, Su GM, Kohchi T, Ishizaki K, Zachgo S, Althoff F, Takenaka M, Yamato KT, Lin SS (2016) Identification of miRNAs and their targets in the Liverwort Marchantia polymorpha by integrating RNA-Seq and degradome analyses. Plant Cell Physiol 57(2):339–358. https://doi.org/10.1093/pcp/pcw020
Liscum E, Reed JW (2002) Genetics of Aux/IAA and ARF action in plant growth and development. Plant Mol Biol 49(3–4):387–400. https://www.ncbi.nlm.nih.gov/pubmed/12036262
Liu C, Xu Z, Chua N-h (1993) Auxin polar transport is essential for the establishment of bilateral symmetry during early plant embryogenesis. Plant Cell 5(6):621–630. https://doi.org/10.1105/tpc.5.6.621
Liu B, Li P, Li X, Liu C, Cao S, Chu C, Cao X (2005) Loss of function of OsDCL1 affects microRNA accumulation and causes developmental defects in rice. Plant Physiol 139(1):296–305
Liu F, Marquardt S, Lister C, Swiezewski S, Dean C (2010) Targeted 3’ processing of antisense transcripts triggers Arabidopsis FLC chromatin silencing. Science 327(5961):94–97. https://doi.org/10.1126/science.1180278
Liu X, Huang J, Wang Y, Khanna K, Xie Z, Owen HA, Zhao D (2010b) The role of floral organs in carpels, an Arabidopsis loss-of-function mutation in MicroRNA160a, in organogenesis and the mechanism regulating its expression. Plant J 62(3):416–428. https://doi.org/10.1111/j.1365-313X.2010.04164.x
Liu L, Guo G, Wang Z, Ji H, Mu F, Li X (2014) Auxin in plant growth and stress responses. Phytohormones. https://doi.org/10.1007/978-1-4939-0491-4_1
Long J, Barton MK (2000) Initiation of axillary and floral meristems in Arabidopsis. Dev Biol 218(2):341–353. https://doi.org/10.1006/dbio.1999.9572
Long JA, Moan EI, Medford JI, Barton MK (1996) A member of the KNOTTED class of homeodomain proteins encoded by the STM gene of Arabidopsis. Nature 379(6560):66–69. https://doi.org/10.1038/379066a0
Ludwig-Muller J (2011) Auxin conjugates: their role for plant development and in the evolution of land plants. J Exp Bot 62(6):1757–1773. https://doi.org/10.1093/jxb/erq412
Ma Z, Hu X, Cai W, Huang W, Zhou X, Luo Q, Yang H, Wang J, Huang J (2014) Arabidopsis miR171-targeted scarecrow-like proteins bind to GT cis-elements and mediate gibberellin-regulated chlorophyll biosynthesis under light conditions. PLoS Genet 10(8):e1004519. https://doi.org/10.1371/journal.pgen.1004519
Ma J, Zhao P, Liu S, Yang Q, Guo H (2020) The control of developmental phase transitions by microRNAs and Their targets in seed plants. Int J Mol Sci. https://doi.org/10.3390/ijms21061971
MacIntosh GC, Wilkerson C, Green PJ (2001) Identification and analysis of Arabidopsis expressed sequence tags characteristic of non-coding RNAs. Plant Physiol 127(3):765–776. https://www.ncbi.nlm.nih.gov/pubmed/11706161
Mallory AC, Reinhart BJ, Jones-Rhoades MW, Tang G, Zamore PD, Barton MK, Bartel DP (2004) MicroRNA control of PHABULOSA in leaf development: importance of pairing to the microRNA 5’ region. EMBO J 23(16):3356–3364. https://doi.org/10.1038/sj.emboj.7600340
Mallory AC, Bartel DP, Bartel B (2005) MicroRNA-directed regulation of Arabidopsis AUXIN RESPONSE FACTOR17 is essential for proper development and modulates expression of early auxin response genes. Plant Cell 17(5):1360–1375. https://doi.org/10.1105/tpc.105.031716
Marin E, Jouannet V, Herz A, Lokerse AS, Weijers D, Vaucheret H, Nussaume L, Crespi MD, Maizel A (2010) miR390, Arabidopsis TAS3 tasiRNAs, and their AUXIN RESPONSE FACTOR targets define an Autoregulatory Network quantitatively regulating lateral Root Growth. Plant Cell 22(4):1104–1117. https://doi.org/10.1105/tpc.109.072553
Mattick JS, Amaral PP, Dinger ME, Mercer TR, Mehler MF (2009) RNA regulation of epigenetic processes. BioEssays 31(1):51–59. https://doi.org/10.1002/bies.080099
McConnell JR, Barton MK (1998) Leaf polarity and meristem formation in Arabidopsis. Development 125(15):2935–2942. https://doi.org/10.1242/dev.125.15.2935
McConnell JR, Emery J, Eshed Y, Bao N, Bowman J, Barton MK (2001) Role of PHABULOSA and PHAVOLUTA in determining radial patterning in shoots. Nature 411(6838):709–713. https://doi.org/10.1038/35079635
Merelo P, Ram H, Pia Caggiano M, Ohno C, Ott F, Straub D, Graeff M, Cho SK, Yang SW, Wenkel S, Heisler MG (2016) Regulation of MIR165/166 by class II and class III homeodomain leucine zipper proteins establishes leaf polarity. Proc Natl Acad Sci U S A 113(42):11973–11978. https://doi.org/10.1073/pnas.1516110113
Meyberg R, Perroud PF, Haas FB, Schneider L, Heimerl T, Renzaglia KS, Rensing SA (2020) Characterisation of evolutionarily conserved key players affecting eukaryotic flagellar motility and fertility using a moss model. New Phytol 227(2):440–454. https://doi.org/10.1111/nph.16486
Montgomery TA, Howell MD, Cuperus JT, Li D, Hansen JE, Alexander AL, Chapman EJ, Fahlgren N, Allen E, Carrington JC (2008) Specificity of ARGONAUTE7-miR390 interaction and dual functionality in TAS3 trans-acting siRNA formation. Cell 133(1):128–141. https://doi.org/10.1016/j.cell.2008.02.033
Morris JL, Puttick MN, Clark JW, Edwards D, Kenrick P, Pressel S, Wellman CH, Yang Z, Schneider H, Donoghue PCJ (2018) The timescale of early land plant evolution. Proc Natl Acad Sci U S A 115(10):E2274–E2283. https://doi.org/10.1073/pnas.1719588115
Mosquna A, Katz A, Decker EL, Rensing SA, Reski R, Ohad N (2009) Regulation of stem cell maintenance by the polycomb protein FIE has been conserved during land plant evolution. Development 136(14):2433–2444. https://doi.org/10.1242/dev.035048
Nagano H, Fukudome A, Hiraguri A, Moriyama H, Fukuhara T (2014) Distinct substrate specificities of Arabidopsis DCL3 and DCL4. Nucleic Acids Res 42(3):1845–1856. https://doi.org/10.1093/nar/gkt1077
Okano Y, Aono N, Hiwatashi Y, Murata T, Nishiyama T, Ishikawa T, Kubo M, Hasebe M (2009) A polycomb repressive complex 2 gene regulates apogamy and gives evolutionary insights into early land plant evolution. Proc Natl Acad Sci USA 106(38):16321–16326. https://doi.org/10.1073/pnas.0906997106
Ori N, Eshed Y, Chuck G, Bowman JL, Hake S (2000) Mechanisms that control knox gene expression in the Arabidopsis shoot. Development 127(24):5523–5532. https://doi.org/10.1242/dev.127.24.5523
Otsuga D, DeGuzman B, Prigge MJ, Drews GN, Clark SE (2001) REVOLUTA regulates meristem initiation at lateral positions. Plant J 25(2):223–236. https://doi.org/10.1111/j.1365-313X.2001.00959.x
Palatnik JF, Allen E, Wu X, Schommer C, Schwab R, Carrington JC, Weigel D (2003) Control of leaf morphogenesis by microRNAs. Nature 425(6955):257–263. https://doi.org/10.1038/nature01958
Palatnik JF, Wollmann H, Schommer C, Schwab R, Boisbouvier J, Rodriguez R, Warthmann N, Allen E, Dezulian T, Huson D, Carrington JC, Weigel D (2007) Sequence and expression differences underlie functional specialization of Arabidopsis microRNAs miR159 and miR319. Dev Cell 13(1):115–125. https://doi.org/10.1016/j.devcel.2007.04.012
Palos K, Yu L, Railey CE, Dittrich N, A. C., Nelson ADL (2023) Linking discoveries, mechanisms, and technologies to develop a clearer perspective on plant long noncoding RNAs. Plant Cell 35(6):1762–1786. https://doi.org/10.1093/plcell/koad027
Parent JS, Bouteiller N, Elmayan T, Vaucheret H (2015) Respective contributions of Arabidopsis DCL2 and DCL4 to RNA silencing. Plant J 81(2):223–232. https://doi.org/10.1111/tpj.12720
Park W, Li J, Song R, Messing J, Chen X (2002) CARPEL FACTORY, a Dicer homolog, and HEN1, a novel protein, act in microRNA metabolism in Arabidopsis thaliana. Curr Biol 12(17):1484–1495. https://doi.org/10.1016/s0960-9822(02)01017-5
Patel P, Mathioni SM, Hammond R, Harkess AE, Kakrana A, Arikit S, Dusia A, Meyers BC (2021) Reproductive phasiRNA loci and DICER-LIKE5, but not microRNA loci, diversified in monocotyledonous plants. Plant Physiol 185(4):1764–1782. https://doi.org/10.1093/plphys/kiab001
Peng J, Carol P, Richards DE, King KE, Cowling RJ, Murphy GP, Harberd NP (1997) The Arabidopsis GAI gene defines a signaling pathway that negatively regulates gibberellin responses. Genes Dev 11(23):3194–3205. https://doi.org/10.1101/gad.11.23.3194
Perroud PF, Haas FB, Hiss M, Ullrich KK, Alboresi A, Amirebrahimi M, Barry K, Bassi R, Bonhomme S, Chen H, Coates JC, Fujita T, Guyon-Debast A, Lang D, Lin J, Lipzen A, Nogue F, Oliver MJ, de Ponce I, Rensing SA (2018) The Physcomitrella patens gene atlas project: large-scale RNA-seq based expression data. Plant J 95(1):168–182. https://doi.org/10.1111/tpj.13940
Pietrykowska H, Sierocka I, Zielezinski A, Alisha A, Carrasco-Sanchez JC, Jarmolowski A, Karlowski WM, Szweykowska-Kulinska Z (2022) Biogenesis, conservation, and function of miRNA in liverworts. J Exp Bot 73(13):4528–4545. https://doi.org/10.1093/jxb/erac098
Pischke MS, Jones LG, Otsuga D, Fernandez DE, Drews GN, Sussman MR (2002) An Arabidopsis histidine kinase is essential for megagametogenesis. Proc Natl Acad Sci USA 99(24):15800–15805. https://doi.org/10.1073/pnas.232580499
Plavskin Y, Timmermans MC (2012) Small RNA-regulated networks and the evolution of novel structures in plants. Cold Spring Harb Symp Quant Biol 77:221–233. https://doi.org/10.1101/sqb.2013.77.014878
Plavskin Y, Nagashima A, Perroud PF, Hasebe M, Quatrano RS, Atwal GS, Timmermans MC (2016) Ancient trans-acting siRNAs Confer Robustness and Sensitivity onto the Auxin response. Dev Cell 36(3):276–289. https://doi.org/10.1016/j.devcel.2016.01.010
Preston JC, Hileman LC (2013) Functional evolution in the Plant SQUAMOSA-PROMOTER BINDING PROTEIN-LIKE (SPL) Gene Family. Front Plant Sci 4:80. https://doi.org/10.3389/fpls.2013.00080
Prigge MJ, Bezanilla M (2010) Evolutionary crossroads in developmental biology: Physcomitrella patens. Development 137(21):3535–3543. https://doi.org/10.1242/dev.049023
Proust H, Honkanen S, Jones VA, Morieri G, Prescott H, Kelly S, Ishizaki K, Kohchi T, Dolan L (2016) RSL Class I genes controlled the Development of Epidermal Structures in the common ancestor of land plants. Curr Biol 26(1):93–99. https://doi.org/10.1016/j.cub.2015.11.042
Pu X, Yang L, Liu L, Dong X, Chen S, Chen Z, Liu G, Jia Y, Yuan W, Liu L (2020) Genome-wide analysis of the MYB transcription factor superfamily in Physcomitrella patens. Int J Mol Sci. https://doi.org/10.3390/ijms21030975
Qiu YL, Palmer JD (1999) Phylogeny of early land plants: insights from genes and genomes. Trends Plant Sci 4(1):26–30. https://doi.org/10.1016/s1360-1385(98)01361-2
Rajagopalan R, Vaucheret H, Trejo J, Bartel DP (2006) A diverse and evolutionarily fluid set of microRNAs in Arabidopsis thaliana. Genes Dev 20(24):3407–3425. https://doi.org/10.1101/gad.1476406
Renner SS, Heinrichs J, Sousa A (2017) The sex chromosomes of bryophytes: recent insights, open questions, and reinvestigations of Frullania dilatata and Plagiochila asplenioides. J Syst Evol 55(4):333–339. https://doi.org/10.1111/jse.12266
Rensing SA, Lang D, Zimmer AD, Terry A, Salamov A, Shapiro H, Nishiyama T, Perroud PF, Lindquist EA, Kamisugi Y, Tanahashi T, Sakakibara K, Fujita T, Oishi K, Shin IT, Kuroki Y, Toyoda A, Suzuki Y, Hashimoto S, Boore JL (2008) The Physcomitrella genome reveals evolutionary insights into the conquest of land by plants. Science 319(5859):64–69. https://doi.org/10.1126/science.1150646
Reski R, Faust M, Wang XH, Wehe M, Abel WO (1994) Genome analysis of the moss Physcomitrella patens (Hedw.) B.S.G. Mol Gen Genet 244(4):352–359. https://doi.org/10.1007/BF00286686
Rhoades MW, Reinhart BJ, Lim LP, Burge CB, Bartel B, Bartel DP (2002) Prediction of plant microRNA targets. Cell 110(4):513–520. https://doi.org/10.1016/s0092-8674(02)00863-2
Riese M, Hohmann S, Saedler H, Munster T, Huijser P (2007) Comparative analysis of the SBP-box gene families in P. patens and seed plants. Gene 401(1–2):28–37. https://doi.org/10.1016/j.gene.2007.06.018
Riese M, Zobell O, Saedler H, Huijser P (2008) SBP-domain transcription factors as possible effectors of cryptochrome-mediated blue light signalling in the moss Physcomitrella patens. Planta 227(2):505–515. https://doi.org/10.1007/s00425-007-0661-5
Rovekamp M, Bowman JL, Grossniklaus U (2016) Marchantia MpRKD regulates the Gametophyte-Sporophyte transition by keeping Egg cells quiescent in the absence of fertilization. Curr Biol 26(13):1782–1789. https://doi.org/10.1016/j.cub.2016.05.028
Rubio-Somoza I, Weigel D (2011) MicroRNA networks and developmental plasticity in plants. Trends Plant Sci 16(5):258–264. https://doi.org/10.1016/j.tplants.2011.03.001
Rubio-Somoza I, Zhou CM, Confraria A, Martinho C, von Born P, Baena-Gonzalez E, Wang JW, Weigel D (2014) Temporal control of leaf complexity by miRNA-regulated licensing of protein complexes. Curr Biol 24(22):2714–2719. https://doi.org/10.1016/j.cub.2014.09.058
Sakaguchi J, Watanabe Y (2012) miR165⁄166 and the development of land plants. Dev Growth Differ 54(1):93–99. https://doi.org/10.1111/j.1440-169x.2011.01318.x
Sakakibara K, Nishiyama T, Deguchi H, Hasebe M (2008) Class 1 KNOX genes are not involved in shoot development in the moss Physcomitrella patens but do function in sporophyte development. Evol Dev 10(5):555–566. https://doi.org/10.1111/j.1525-142X.2008.00271.x
Schauer SE, Jacobsen SE, Meinke DW, Ray A (2002) DICER-LIKE1: blind men and elephants in Arabidopsis development. Trends Plant Sci 7(11):487–491. https://doi.org/10.1016/s1360-1385(02)02355-5
Schommer C, Palatnik JF, Aggarwal P, Chetelat A, Cubas P, Farmer EE, Nath U, Weigel D (2008) Control of jasmonate biosynthesis and senescence by miR319 targets. PLoS Biol 6(9):e230. https://doi.org/10.1371/journal.pbio.0060230
Schommer C, Debernardi JM, Bresso EG, Rodriguez RE, Palatnik JF (2014) Repression of cell proliferation by miR319-regulated TCP4. Mol Plant 7(10):1533–1544. https://doi.org/10.1093/mp/ssu084
Scofield S, Dewitte W, Murray JAH (2007) The KNOX gene SHOOT MERISTEMLESS is required for the development of reproductive meristematic tissues in Arabidopsis. Plant J 50(5):767–781. https://doi.org/10.1111/j.1365-313X.2007.03095.x
Scofield S, Dewitte W, Murray JAH (2008) A model for Arabidopsis class-1 KNOX gene function. Plant Signal Behav 3(4):257–259. https://doi.org/10.4161/psb.3.4.5194
Seta A, Tabara M, Nishibori Y, Hiraguri A, Ohkama-Ohtsu N, Yokoyama T, Hara S, Yoshida K, Hisabori T, Fukudome A, Koiwa H, Moriyama H, Takahashi N, Fukuhara T (2017) Post-translational regulation of the Dicing activities of Arabidopsis DICER-LIKE 3 and 4 by Inorganic phosphate and the Redox State. Plant Cell Physiol 58(3):485–495. https://doi.org/10.1093/pcp/pcw226
Shaw AJ, Szovenyi P, Shaw B (2011) Bryophyte diversity and evolution: windows into the early evolution of land plants. Am J Bot 98(3):352–369. https://doi.org/10.3732/ajb.1000316
Shi ZH, Zhang C, Xu XF, Zhu J, Zhou Q, Ma LJ, Niu J, Yang ZN (2015) Overexpression of AtTTP affects ARF17 expression and leads to male sterility in Arabidopsis. PLoS ONE 10(3):e0117317. https://doi.org/10.1371/journal.pone.0117317
Shimamura M (2016) Marchantia polymorpha: taxonomy, phylogeny and morphology of a Model System. Plant Cell Physiol 57(2):230–256. https://doi.org/10.1093/pcp/pcv192
Silverstone AL, Ciampaglio CN, Sun T (1998) The Arabidopsis RGA gene encodes a transcriptional regulator repressing the gibberellin signal transduction pathway. Plant Cell 10(2):155–169. https://doi.org/10.1105/tpc.10.2.155
Singer SD, Ashton NW (2007) Revelation of ancestral roles of KNOX genes by a functional analysis of Physcomitrella homologues. Plant Cell Rep 26(12):2039–2054. https://doi.org/10.1007/s00299-007-0409-5
Singh A, Singh S, Panigrahi KC, Reski R, Sarkar AK (2014) Balanced activity of microRNA166/165 and its target transcripts from the class III homeodomain-leucine zipper family regulates root growth in Arabidopsis thaliana. Plant Cell Rep 33(6):945–953. https://doi.org/10.1007/s00299-014-1573-z
Skopelitis DS, Husbands AY, Timmermans MCP (2012) Plant small RNAs as morphogens. Curr Opin Cell Biol 24(2):217–224. https://doi.org/10.1016/j.ceb.2011.12.006
Streubel S, Deiber S, Rotzer J, Mosiolek M, Jandrasits K, Dolan L (2023) Meristem dormancy in Marchantia polymorpha is regulated by a liverwort-specific miRNA and a clade III SPL gene. Curr Biol 33(4):660–674e664. https://doi.org/10.1016/j.cub.2022.12.062
Stuurman J, Jaggi F, Kuhlemeier C (2002) Shoot meristem maintenance is controlled by a GRAS-gene mediated signal from differentiating cells. Genes Dev 16(17):2213–2218. https://doi.org/10.1101/gad.230702
Tabara M, Ohtani M, Kanekatsu M, Moriyama H, Fukuhara T (2018) Size distribution of small interfering RNAs in various organs at different developmental stages is primarily determined by the Dicing activity of Dicer-Like proteins in plants. Plant Cell Physiol 59(11):2228–2238. https://doi.org/10.1093/pcp/pcy144
Tang G, Reinhart BJ, Bartel DP, Zamore PD (2003) A biochemical framework for RNA silencing in plants. Genes Dev 17(1):49–63. https://doi.org/10.1101/gad.1048103
Thamm A, Saunders TE, Dolan L (2020) MpFEW RHIZOIDS1 miRNA-Mediated lateral inhibition controls rhizoid cell patterning in Marchantia polymorpha. Curr Biol. https://doi.org/10.1016/j.cub.2020.03.032
Tregear JW, Richaud F, Collin M, Esbelin J, Parrinello H, Cochard B, Nodichao L, Morcillo F, Adam H, Jouannic S (2022) Micro-RNA-regulated SQUAMOSA-PROMOTER BINDING PROTEIN-LIKE (SPL) gene expression and cytokinin accumulation distinguish early-developing male and female inflorescences in oil palm (Elaeis guineensis). Plants https://doi.org/10.3390/plants11050685
Tsuzuki M, Nishihama R, Ishizaki K, Kurihara Y, Matsui M, Bowman JL, Kohchi T, Hamada T, Watanabe Y (2016) Profiling and characterization of small RNAs in the Liverwort, Marchantia polymorpha, belonging to the first diverged land plants. Plant Cell Physiol 57(2):359–372. https://doi.org/10.1093/pcp/pcv182
Tsuzuki M, Futagami K, Shimamura M, Inoue C, Kunimoto K, Oogami T, Tomita Y, Inoue K, Kohchi T, Yamaoka S, Araki T, Hamada T, Watanabe Y (2019) An early arising role of the MicroRNA156/529-SPL Module in Reproductive Development revealed by the Liverwort Marchantia polymorpha. Curr Biol 29(19):3307–3314e3305. https://doi.org/10.1016/j.cub.2019.07.084
Unte US, Sorensen AM, Pesaresi P, Gandikota M, Leister D, Saedler H, Huijser P (2003) SPL8, an SBP-box gene that affects pollen sac development in Arabidopsis. Plant Cell 15(4):1009–1019. https://doi.org/10.1105/tpc.010678
Wang JW (2014) Regulation of flowering time by the miR156-mediated age pathway. J Exp Bot 65(17):4723–4730. https://doi.org/10.1093/jxb/eru246
Wang HV, Chekanova JA (2017) Long noncoding RNAs in plants. Adv Exp Med Biol 1008:133–154. https://doi.org/10.1007/978-981-10-5203-3_5
Wang H, Wang H (2015) The miR156/SPL Module, a Regulatory Hub and Versatile Toolbox, gears up crops for enhanced agronomic traits. Mol Plant 8(5):677–688. https://doi.org/10.1016/j.molp.2015.01.008
Wang JW, Czech B, Weigel D (2009) miR156-regulated SPL transcription factors define an endogenous flowering pathway in Arabidopsis thaliana. Cell 138(4):738–749. https://doi.org/10.1016/j.cell.2009.06.014
Wierzbicki AT, Haag JR, Pikaard CS (2008) Noncoding transcription by RNA polymerase Pol IVb/Pol V mediates transcriptional silencing of overlapping and adjacent genes. Cell 135(4):635–648. https://doi.org/10.1016/j.cell.2008.09.035
Wierzbicki AT, Ream TS, Haag JR, Pikaard CS (2009) RNA polymerase V transcription guides ARGONAUTE4 to chromatin. Nat Genet 41(5):630–634. https://doi.org/10.1038/ng.365
Wierzbicki AT, Blevins T, Swiezewski S (2021) Long noncoding RNAs in plants. Annu Rev Plant Biol. https://doi.org/10.1146/annurev-arplant-093020-035446
Williams L, Grigg SP, Xie MT, Christensen S, Fletcher JC (2005) Regulation of Arabidopsis shoot apical meristem and lateral organ formation by microRNA miR166g and its AtHD-ZIP target genes. Development 132(16):3657–3668. https://doi.org/10.1242/dev.01942
Wu H, Li B, Iwakawa HO, Pan Y, Tang X, Ling-Hu Q, Liu Y, Sheng S, Feng L, Zhang H, Zhang X, Tang Z, Xia X, Zhai J, Guo H (2020) Plant 22-nt siRNAs mediate translational repression and stress adaptation. Nature 581(7806):89–93. https://doi.org/10.1038/s41586-020-2231-y
Xia J, Wang X, Perroud PF, He Y, Quatrano R, Zhang W (2016) Endogenous small-noncoding RNAs and potential functions in Desiccation Tolerance in Physcomitrella Patens. Sci Rep 6:30118. https://doi.org/10.1038/srep30118
Xia R, Xu J, Meyers BC (2017) The emergence, evolution, and diversification of the miR390-TAS3-ARF pathway in land plants. Plant Cell 29(6):1232–1247. https://doi.org/10.1105/tpc.17.00185
Xie Z, Johansen LK, Gustafson AM, Kasschau KD, Lellis AD, Zilberman D, Jacobsen SE, Carrington JC (2004) Genetic and functional diversification of small RNA pathways in plants. PLoS Biol 2(5):E104. https://doi.org/10.1371/journal.pbio.0020104
Xie Z, Allen E, Wilken A, Carrington JC (2005) DICER-LIKE 4 functions in trans-acting small interfering RNA biogenesis and vegetative phase change in Arabidopsis thaliana. Proc Natl Acad Sci U S A 102(36):12984–12989. https://doi.org/10.1073/pnas.0506426102
Xie Y, Liu Y, Wang H, Ma X, Wang B, Wu G, Wang H (2017) Phytochrome-interacting factors directly suppress MIR156 expression to enhance shade-avoidance syndrome in Arabidopsis. Nat Commun 8(1):348. https://doi.org/10.1038/s41467-017-00404-y
Xie Q, Wang X, He J, Lan T, Zheng J, Li Y, Pan J, Lin L, Zhao J, Li J, Yu Y, Mo B, Chen X, Gao L, Liu L (2021) Distinct evolutionary profiles and functions of microRNA156 and microRNA529 in land plants. Int J Mol Sci. https://doi.org/10.3390/ijms222011100
Xu L, Shen WH (2008) Polycomb silencing of KNOX genes confines shoot stem cell niches in Arabidopsis. Curr Biol 18(24):1966–1971. https://doi.org/10.1016/j.cub.2008.11.019
Xu L, Xu Y, Dong AW, Sun Y, Pi LM, Xu YQ, Huang H (2003) Novel as1 and as2 defects in leaf adaxial-abaxial polarity reveal the requirement for ASYMMETRIC LEAVES1 and 2 and ERECTA functions in specifying leaf adaxial identity. Development 130(17):4097–4107. https://doi.org/10.1242/dev.00622
Xu M, Hu T, Zhao J, Park MY, Earley KW, Wu G, Yang L, Poethig RS (2016) Developmental functions of miR156-Regulated SQUAMOSA PROMOTER BINDING PROTEIN-LIKE (SPL) genes in Arabidopsis thaliana. PLoS Genet 12(8):e1006263. https://doi.org/10.1371/journal.pgen.1006263
Yamaoka S, Nishihama R, Yoshitake Y, Ishida S, Inoue K, Saito M, Okahashi K, Bao H, Nishida H, Yamaguchi K, Shigenobu S, Ishizaki K, Yamato KT, Kohchi T (2018) Generative cell specification requires transcription factors evolutionarily conserved in land plants. Curr Biol 28(3):479–. https://doi.org/10.1016/j.cub.2017.12.053
Yan J, Zhao C, Zhou J, Yang Y, Wang P, Zhu X, Tang G, Bressan RA, Zhu JK (2016) The miR165/166 mediated Regulatory Module Plays critical roles in ABA Homeostasis and response in Arabidopsis thaliana. PLoS Genet 12(11):e1006416. https://doi.org/10.1371/journal.pgen.1006416
Yelina NE, Frangedakis E, Schreier TB, Rever J, Tomaselli M, Haseloff J, Hibberd JM (2023) Streamlined regulation of chloroplast development in the liverwort Marchantia polymorpha. bioRxiv. https://doi.org/10.1101/2023.01.23.525199
Yip HK, Floyd SK, Sakakibara K, Bowman JL (2016) Class III HD-Zip activity coordinates leaf development in Physcomitrella patens. Dev Biol 419(1):184–197. https://doi.org/10.1016/j.ydbio.2016.01.012
Yoshikawa M, Peragine A, Park MY, Poethig RS (2005) A pathway for the biogenesis of trans-acting siRNAs in Arabidopsis. Genes Dev 19(18):2164–2175. https://doi.org/10.1101/gad.1352605
You C, Cui J, Wang H, Qi X, Kuo LY, Ma H, Gao L, Mo B, Chen X (2017) Conservation and divergence of small RNA pathways and microRNAs in land plants. Genome Biol 18(1):158. https://doi.org/10.1186/s13059-017-1291-2
Yu B, Wang H (2010) Translational inhibition by microRNAs in plants. Prog Mol Subcell Biol 50:41–57. https://doi.org/10.1007/978-3-642-03103-8_3
Yu Y, Zhang Y, Chen X, Chen Y (2019) Plant noncoding RNAs: hidden players in development and stress responses. Annu Rev Cell Dev Biol 35:407–431. https://doi.org/10.1146/annurev-cellbio-100818-125218
Yuan L, Liu ZN, Song XY, Johnson C, Yu XL, Sundaresan V (2016) The CKI1 histidine kinase specifies the female Gametic Precursor of the Endosperm. Dev Cell 37(1):34–46. https://doi.org/10.1016/j.devcel.2016.03.009
Zhang LL, Huang YY, Zheng YP, Liu XX, Zhou SX, Yang XM, Liu SL, Li Y, Li JL, Zhao SL, Wang H, Ji YP, Zhang JW, Pu M, Zhao ZX, Fan J, Wang WM (2022) Osa-miR535 targets SQUAMOSA promoter binding protein-like 4 to regulate blast Disease resistance in rice. Plant J 110(1):166–178. https://doi.org/10.1111/tpj.15663
Zhao H, Lu M, Singh R, Snell WJ (2001) Ectopic expression of a Chlamydomonas mt+-specific homeodomain protein in mt – gametes initiates zygote development without gamete fusion. Genes Dev 15(20):2767–2777
Zheng C, Ye M, Sang M, Wu R (2019) A regulatory network for miR156-SPL module in Arabidopsis thaliana. Int J Mol Sci. https://doi.org/10.3390/ijms20246166
Acknowledgements
We thank MSc Anna Madra-Bielewicz from Adam Mickiewicz University Poznan, Poland for helping in drawing Fig. 1.
Funding
This work was supported by National Science Centre (NCN) 2020/39/B/NZ3/00539 (to BA, HP, ZSK), 2016/21/D/NZ3/00353 (to AA, IS), by Polish Academy of Sciences—Japan Society for the Promotion of Science (PAS-JSPS) cooperation grants for 2022–2023, project no. 2 (to BA, AJ, MO, HP, ZSK, YW), MEXT KAKENHI (JP21H05652 and JP23H04191 to M.O.), JSPS KAKENHI (JP20H03271 to M.O.) and Toray Science Foundation (grant no. 19-6002 to M.O.).
The authors also received financial support from the Initiative of Excellence—Research University (05/IDUB/2019/94) at the Adam Mickiewicz University, Poznan, Poland.
Author information
Authors and Affiliations
Contributions
AA, BA, AJ, MO, HP, YW – Participated in manuscript writing and figure drawing. IS, ZSK – Designed, wrote and coordinated the manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
MO is a member of Plant Molecular Biology Editorial Board.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pietrykowska, H., Alisha, A., Aggarwal, B. et al. Conserved and non-conserved RNA–target modules in plants: lessons for a better understanding of Marchantia development. Plant Mol Biol 113, 121–142 (2023). https://doi.org/10.1007/s11103-023-01392-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-023-01392-y