Abstract
Abstract
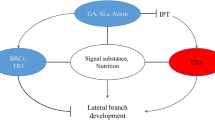
As a key integrator of shoot branching, BRANCHED 1 (BRC1) coordinates and is orchestrated by endogenous and environmental signals involved in the regulation of axillary bud outgrowth. In the present study, we characterized the regulatory roles of five BRC gene members in tobacco (Nicotiana tabacum L.) using CRISPR site-directed mutagenesis and overexpression assays. It was shown that lateral branching was negatively regulated by NtBRC1A-1, 1B-1, and 1B-2, but was unexpectedly promoted by NtBRC2A. Suppression of bud growth may be attained by direct binding of NtBRCs to the Tassels Replace Upper Ears 1 (TRU1) genes. It was speculated that NtBRC2A probably confers a dominant negative effect by interfering with the branching-inhibitory BRC1 genes. Our results suggested that highly homologous gene family members may function antagonistically in the same signaling pathway. However, the molecular mechanism underlying NtBRC2A-mediated outgrowth of axillary buds needs to be further addressed.
Key message
Axillary bud outgrowth in general is negatively regulated by the BRANCHED gene. Here we show that the BRANCHED genes play opposing regulatory roles in tobacco lateral branching.






Similar content being viewed by others
References
Aguilar-Martinez JA, Poza-Carrion C, Cubas P (2007) Arabidopsis BRANCHED1 acts as an integrator of branching signals within axillary buds. Plant Cell 19(2):458–472. https://doi.org/10.1105/tpc.106.048934
Bailey TL, Boden M, Buske FA, Frith M, Grant CE, Clementi L, Ren J, Li WW, Noble WS (2009) MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res 37:W202–208. https://doi.org/10.1093/nar/gkp335
Barbier FF, Dun EA, Kerr SC, Chabikwa TG, Beveridge CA (2019) An update on the signals controlling shoot branching. Trends Plant Sci 24(3):220–236. https://doi.org/10.1016/j.tplants.2018.12.001
Beveridge CA, Weller JL, Singer SR, Hofer JM (2003) Axillary meristem development. Budding relationships between networks controlling flowering, branching, and photoperiod responsiveness. Plant Physiol 131(3):927–934
Braun N, de Saint GA, Pillot JP, Boutet-Mercey S, Dalmais M, Antoniadi I, Li X, Maia-Grondard A, Le Signor C, Bouteiller N, Luo D, Bendahmane A, Turnbull C, Rameau C (2012) The pea TCP transcription factor PsBRC1 acts downstream of Strigolactones to control shoot branching. Plant Physiol 158(1):225–238. https://doi.org/10.1104/pp.111.182725
Chen L, Chen YQ, Ding AM, Chen H, Xia F, Wang WF, Sun YH (2016) Genome-wide analysis of TCP family in tobacco. Genet Mol Res. https://doi.org/10.4238/gmr.15027728
Chen S, Ren F, Zhang L, Liu Y, Chen X, Li Y, Zhang L, Zhu B, Zeng P, Li Z, Larkin RM, Kuang H (2018) Unstable allotetraploid tobacco genome due to frequent homeologous recombination, segmental deletion, and chromosome loss. Mol Plant 11(7):914–927. https://doi.org/10.1016/j.molp.2018.04.009
Cho SW, Kim S, Kim JM, Kim JS (2013) Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nat Biotechnol 31(3):230–232. https://doi.org/10.1038/nbt.2507
Citerne HL, Le Guilloux M, Sannier J, Nadot S, Damerval C (2013) Combining phylogenetic and syntenic analyses for understanding the evolution of TCP ECE genes in eudicots. PLoS ONE 8(9):e74803. https://doi.org/10.1371/journal.pone.0074803
Cubas P, Lauter N, Doebley J, Coen E (1999) The TCP domain: a motif found in proteins regulating plant growth and development. Plant J 18(2):215–222
Danisman S (2016) TCP transcription factors at the interface between environmental challenges and the plant's growth responses. Front Plant Sci 7:1930. https://doi.org/10.3389/fpls.2016.01930
Doebley J, Stec A, Hubbard L (1997) The evolution of apical dominance in maize. Nature 386(6624):485–488. https://doi.org/10.1038/386485a0
Domagalska MA, Leyser O (2011) Signal integration in the control of shoot branching. Nat Rev Mol Cell Biol 12(4):211–221. https://doi.org/10.1038/nrm3088
Dong Z, Li W, Unger-Wallace E, Yang J, Vollbrecht E, Chuck G (2017) Ideal crop plant architecture is mediated by tassels replace upper ears1, a BTB/POZ ankyrin repeat gene directly targeted by TEOSINTE BRANCHED1. Proc Natl Acad Sci USA 114(41):E8656–E8664. https://doi.org/10.1073/pnas.1714960114
Drummond RS, Janssen BJ, Luo Z, Oplaat C, Ledger SE, Wohlers MW, Snowden KC (2015) Environmental control of branching in petunia. Plant Physiol 168(2):735–751. https://doi.org/10.1104/pp.15.00486
Dun EA, Ferguson BJ, Beveridge CA (2006) Apical dominance and shoot branching. Divergent opinions or divergent mechanisms? Plant Physiol 142(3):812–819. https://doi.org/10.1104/pp.106.086868
Finlayson SA (2007) Arabidopsis teosinte branched1-like 1 regulates axillary bud outgrowth and is homologous to monocot teosinte branched1. Plant Cell Physiol 48(5):667–677. https://doi.org/10.1093/pcp/pcm044
Friedland AE, Tzur YB, Esvelt KM, Colaiacovo MP, Church GM, Calarco JA (2013) Heritable genome editing in C. elegans via a CRISPR-Cas9 system. Nat Methods 10(8):741–743. https://doi.org/10.1038/nmeth.2532
Gallois P, Marinho P (1995) Leaf disk transformation using Agrobacterium tumefaciens-expression of heterologous genes in tobacco. Methods Mol Biol 49:39–48. https://doi.org/10.1385/0-89603-321-X:39
Gao J, Wang G, Ma S, Xie X, Wu X, Zhang X, Wu Y, Zhao P, Xia Q (2015) CRISPR/Cas9-mediated targeted mutagenesis in Nicotiana tabacum. Plant Mol Biol 87(1–2):99–110. https://doi.org/10.1007/s11103-014-0263-0
Gonzalez-Grandio E, Pajoro A, Franco-Zorrilla JM, Tarancon C, Immink RG, Cubas P (2017) Abscisic acid signaling is controlled by a BRANCHED1/HD-ZIP I cascade in Arabidopsis axillary buds. Proc Natl Acad Sci USA 114(2):E245–E254. https://doi.org/10.1073/pnas.1613199114
Ha CM, Kim GT, Kim BC, Jun JH, Soh MS, Ueno Y, Machida Y, Tsukaya H, Nam HG (2003) The BLADE-ON-PETIOLE 1 gene controls leaf pattern formation through the modulation of meristematic activity in Arabidopsis. Development 130(1):161–172. https://doi.org/10.1242/dev.00196
Herskowitz I (1987) Functional inactivation of genes by dominant negative mutations. Nature 329(6136):219–222. https://doi.org/10.1038/329219a0
Karimi M, Inze D, Depicker A (2002) GATEWAY vectors for Agrobacterium-mediated plant transformation. Trends Plant Sci 7(5):193–195
Kebrom TH, Burson BL, Finlayson SA (2006) Phytochrome B represses Teosinte Branched1 expression and induces sorghum axillary bud outgrowth in response to light signals. Plant Physiol 140(3):1109–1117. https://doi.org/10.1104/pp.105.074856
Kosugi S, Ohashi Y (2002) DNA binding and dimerization specificity and potential targets for the TCP protein family. Plant J 30(3):337–348
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and clustal X version 2.0. Bioinformatics 23(21):2947–2948. https://doi.org/10.1093/bioinformatics/btm404
Lewis R, Nicholson J (2007) Aspects of the evolution of Nicotiana tabacum L. and the status of the United States Nicotiana Germplasm collection. Genet Res Crop Evol 54:727–740. https://doi.org/10.1007/s10722-006-0024-2
Li JF, Norville JE, Aach J, McCormack M, Zhang D, Bush J, Church GM, Sheen J (2013) Multiplex and homologous recombination-mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nat Biotechnol 31(8):688–691. https://doi.org/10.1038/nbt.2654
Limpens E, Mirabella R, Fedorova E, Franken C, Franssen H, Bisseling T, Geurts R (2005) Formation of organelle-like N2-fixing symbiosomes in legume root nodules is controlled by DMI2. Proc Natl Acad Sci USA 102(29):10375–10380. https://doi.org/10.1073/pnas.0504284102
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25(4):402–408. https://doi.org/10.1006/meth.2001.1262
Mali P, Aach J, Stranges PB, Esvelt KM, Moosburner M, Kosuri S, Yang L, Church GM (2013) CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering. Nat Biotechnol 31(9):833–838. https://doi.org/10.1038/nbt.2675
Martin-Trillo M, Cubas P (2010) TCP genes: a family snapshot 10 years later. Trends Plant Sci 15(1):31–39. https://doi.org/10.1016/j.tplants.2009.11.003
Martin-Trillo M, Grandio EG, Serra F, Marcel F, Rodriguez-Buey ML, Schmitz G, Theres K, Bendahmane A, Dopazo H, Cubas P (2011) Role of tomato BRANCHED1-like genes in the control of shoot branching. Plant J 67(4):701–714. https://doi.org/10.1111/j.1365-313X.2011.04629.x
McDaniel CN, Sangrey KA, Jegla DE (1989) Cryptic floral determination: stem explants from vegetative tobacco plants have the capacity to regenerate floral shoots. Dev Biol 134(2):473–478. https://doi.org/10.1016/0012-1606(89)90120-6
Meinke DW (2013) A survey of dominant mutations in Arabidopsis thaliana. Trends Plant Sci 18(2):84–91. https://doi.org/10.1016/j.tplants.2012.08.006
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8(19):4321–4325. https://doi.org/10.1093/nar/8.19.4321
Nicolas M, Rodriguez-Buey ML, Franco-Zorrilla JM, Cubas P (2015) A recently evolved alternative splice site in the BRANCHED1a gene controls potato plant architecture. Curr Biol 25(14):1799–1809. https://doi.org/10.1016/j.cub.2015.05.053
Poza-Carrion C, Aguilar-Martinez JA, Cubas P (2007) Role of TCP gene BRANCHED1 in the control of shoot branching in Arabidopsis. Plant Signal Behav 2(6):551–552. https://doi.org/10.4161/psb.2.6.4811
Prelich G (2012) Gene overexpression: uses, mechanisms, and interpretation. Genetics 190(3):841–854. https://doi.org/10.1534/genetics.111.136911
Rameau C, Bertheloot J, Leduc N, Andrieu B, Foucher F, Sakr S (2015) Multiple pathways regulate shoot branching. Front Plant Sci 5:741
Ramsay L, Comadran J, Druka A, Marshall DF, Thomas WT, Macaulay M, MacKenzie K, Simpson C, Fuller J, Bonar N, Hayes PM, Lundqvist U, Franckowiak JD, Close TJ, Muehlbauer GJ, Waugh R (2011) INTERMEDIUM-C, a modifier of lateral spikelet fertility in barley, is an ortholog of the maize domestication gene TEOSINTE BRANCHED 1. Nat Genet 43(2):169–172. https://doi.org/10.1038/ng.745
Sarvepalli K, Nath U (2018) CIN-TCP transcription factors: transiting cell proliferation in plants. IUBMB Life 70(8):718–731. https://doi.org/10.1002/iub.1874
Shimizu-Sato S, Mori H (2001) Control of outgrowth and dormancy in axillary buds. Plant Physiol 127(4):1405–1413
Sierro N, Battey JN, Ouadi S, Bovet L, Goepfert S, Bakaher N, Peitsch MC, Ivanov NV (2013) Reference genomes and transcriptomes of Nicotiana sylvestris and Nicotiana tomentosiformis. Genome Biol 14(6):R60. https://doi.org/10.1186/gb-2013-14-6-r60
Sierro N, Battey JN, Ouadi S, Bakaher N, Bovet L, Willig A, Goepfert S, Peitsch MC, Ivanov NV (2014) The tobacco genome sequence and its comparison with those of tomato and potato. Nat Commun 5:3833. https://doi.org/10.1038/ncomms4833
Stafstrom JP, Sussex IM (1988) Patterns of protein synthesis in dormant and growing vegetative buds of pea. Planta 176(4):497–505. https://doi.org/10.1007/bf00397656
Takeda T, Suwa Y, Suzuki M, Kitano H, Ueguchi-Tanaka M, Ashikari M, Matsuoka M, Ueguchi C (2003) The OsTB1 gene negatively regulates lateral branching in rice. Plant J 33(3):513–520
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28(10):2731–2739. https://doi.org/10.1093/molbev/msr121
Tavakol E, Okagaki R, Verderio G, Shariati JV, Hussien A, Bilgic H, Scanlon MJ, Todt NR, Close TJ, Druka A, Waugh R, Steuernagel B, Ariyadasa R, Himmelbach A, Stein N, Muehlbauer GJ, Rossini L (2015) The barley Uniculme4 gene encodes a BLADE-ON-PETIOLE-like protein that controls tillering and leaf patterning. Plant Physiol 168(1):164–174. https://doi.org/10.1104/pp.114.252882
Teichmann T, Muhr M (2015) Shaping plant architecture Front Plant Sci 6:233. https://doi.org/10.3389/fpls.2015.00233
Veitia RA (2007) Exploring the molecular etiology of dominant-negative mutations. Plant Cell 19(12):3843–3851. https://doi.org/10.1105/tpc.107.055053
Vitha S, Benes K, Phillips JP, Gartland KM (1995) Histochemical GUS analysis. Methods Mol Biol 44:185–193. https://doi.org/10.1385/0-89603-302-3:185
Wang M, Le Moigne MA, Bertheloot J, Crespel L, Perez-Garcia MD, Oge L, Demotes-Mainard S, Hamama L, Daviere JM, Sakr S (2019) BRANCHED1: a key hub of shoot branching. Front Plant Sci 10:76. https://doi.org/10.3389/fpls.2019.00076
Xie K, Yang Y (2013) RNA-guided genome editing in plants using a CRISPR-Cas system. Mol Plant 6(6):1975–1983. https://doi.org/10.1093/mp/sst119
Xing HL, Dong L, Wang ZP, Zhang HY, Han CY, Liu B, Wang XC, Chen QJ (2014) A CRISPR/Cas9 toolkit for multiplex genome editing in plants. BMC Plant Biol 14:327. https://doi.org/10.1186/s12870-014-0327-y
Xu S, Brockmoller T, Navarro-Quezada A, Kuhl H, Gase K, Ling Z, Zhou W, Kreitzer C, Stanke M, Tang H, Lyons E, Pandey P, Pandey SP, Timmermann B, Gaquerel E, Baldwin IT (2017) Wild tobacco genomes reveal the evolution of nicotine biosynthesis. Proc Natl Acad Sci USA 114(23):6133–6138. https://doi.org/10.1073/pnas.1700073114
Yuan C, Xi L, Kou Y, Zhao Y, Zhao L (2015) Current perspectives on shoot branching regulation. Front Agric Sci Eng 2(1):38–52. https://doi.org/10.15302/J-FASE-2015053
Acknowledgments
This research was supported by a gift funding from R.J. Reynolds Tobacco Company (to SY), KTRDC Summit grant (to SY), and a gift funding from the Council for Burley Tobacco (to SY).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: ND, RD, DZ, DL and SY. Performed the experiments: ND, QQ, XW, and DL. Analyzed the data: ND, QQ, XW, RM, DZ, DL and SY. Wrote the first draft: SY.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ding, N., Qin, Q., Wu, X. et al. Antagonistic regulation of axillary bud outgrowth by the BRANCHED genes in tobacco. Plant Mol Biol 103, 185–196 (2020). https://doi.org/10.1007/s11103-020-00983-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-020-00983-3