Abstract
Key message
A possible transcription factor TLP2 was identified to be involved in the regulation of HG biosynthesis in Arabidopsis seed mucilage. TLP2 can translocate into nucleus from plasma membrane by interacting with NF-YC3. The discovery of TLP2 gene function can further fulfill the regulatory network of pectin biosynthesis in Arabidopsis thaliana.
Abstract
Arabidopsis seed coat mucilage is an excellent model system to study the biosynthesis, function and regulation of pectin. Rhamnogalacturonan I (RG-I) and homogalacturonan (HG) are the major polysaccharides constituent of the Arabidopsis seed coat mucilage. Here, we identified a Tubby-like gene, Tubby-like protein 2 (TLP2), which was up-regulated in developing siliques when mucilage began to be produced. Ruthenium red (RR) staining of the seeds showed defective mucilage of tlp2-1 mutant after vigorous shaking compared to wild type (WT). Monosaccharide composition analysis revealed that the amount of total sugars and galacturonic acid (GalA) decreased significantly in the adherent mucilage (AM) of tlp2-1 mutant. Immunolabelling and dot immunoblotting analysis showed that unesterified HG decreased in the tlp2-1 mutant. Furthermore, TLP2 can translocate into nucleus by interacting with Nuclear Factor Y subunit C3 (NF-YC3) to function as a transcription factor. RNA-sequence and transactivation assays revealed that TLP2 could activate UDP-glucose 4-epimerase 1 (UGE1). In all, it is concluded that TLP2 could regulate the biosynthesis of HG possibly through the positive activation of UGE1.






Similar content being viewed by others
References
Akita K, Ishimizu T, Tsukamoto T, Ando T, Hase S (2002) Successive glycosyltransfer activity and enzymatic characterization of pectic polygalacturonate 4-alpha-galacturonosyltransferase solubilized from pollen tubes of Petunia axillaris using pyridylaminated oligogalacturonates as substrates. Plant Physiol 130:374–379. https://doi.org/10.1104/pp.005587
Arsovski AA, Haughn GW, Western TL (2010) Seed coat mucilage cells of Arabidopsis thaliana as a model for plant cell wall research. Plant Signal Behav 5:796–801. https://doi.org/10.4161/psb.5.7.11773
Atmodjo MA, Sakuragi Y, Zhu X, Burrell AJ, Mohanty SS, Atwood JA, Orlando R, Scheller HV, Mohnen D (2011) Galacturonosyltransferase (GAUT)1 and GAUT7 are the core of a plant cell wall pectin biosynthetic homogalacturonan:galacturonosyltransferase complex. Proc Natl Acad Sci USA 108:20225–20230. https://doi.org/10.1073/pnas.1112816108
Atmodjo MA, Hao Z, Mohnen D (2013) Evolving views of pectin biosynthesis. Annu Rev Plant Biol 64:747–779. https://doi.org/10.1146/annurev-arplant-042811-105534
Bao Y, Song W, Jin YL, Jiang C, Li B, Huang W, Liu H, Zhang HX (2014) Characterization of Arabidopsis Tubby-like proteins and redundant function of AtTLP3 and AtTLP9 in plant response to ABA and osmotic stress. Plant Mol Biol 86:471–483. https://doi.org/10.1007/s11103-014-0241-6
Batoko H, Zheng HQ, Hawes C, Moore I (2000) A Rab1 GTPase is required for transport between the endoplasmic reticulum and golgi apparatus and for normal golgi movement in plants. Plant Cell 12:2201–2217. https://doi.org/10.1105/tpc.12.11.2201
Bethke G, Thao A, Xiong G, Li B, Soltis NE, Hatsugai N, Hillmer RA, Katagiri F, Kliebenstein DJ, Pauly M, Glazebrook J (2016) Pectin biosynthesis is critical for cell wall integrity and immunity in Arabidopsis thaliana. Plant Cell 28:537–556. https://doi.org/10.1105/tpc.15.00404
Bouton S, Leboeuf E, Mouille G, Leydecker MT, Talbotec J, Granier F, Lahaye M, Höfte H, Truong HN (2002) QUASIMODO1 encodes a putative membrane-bound glycosyltransferase required for normal pectin synthesis and cell adhesion in Arabidopsis. Plant Cell 14:2577–2590. https://doi.org/10.1105/tpc.004259
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. https://doi.org/10.1016/0003-2697(76)90527-3
Braybrook SA, Jonsson H (2016) Shifting foundations: the mechanical cell wall and development. Curr Opin Plant Biol 29:115–120. https://doi.org/10.1016/j.pbi.2015.12.009
Bui M, Lim N, Sijacic P, Liu Z (2011) LEUNIG_HOMOLOG and LEUNIG regulate seed mucilage extrusion in Arabidopsis. J Integr Plant Biol 53:399–408. https://doi.org/10.1111/j.1744-7909.2011.01036.x
Caffall KH, Mohnen D (2009) The structure, function, and biosynthesis of plant cell wall pectic polysaccharides. Carbohydr Res 344:1879–1900. https://doi.org/10.1016/j.carres.2009.05.021
Cai M, Qiu D, Yuan T, Ding X, Li H, Duan L, Xu C, Li X, Wang S (2008) Identification of novel pathogen-responsive cis-elements and their binding proteins in the promoter of OsWRKY13, a gene regulating rice disease resistance. Plant Cell Environ 31:86–96. https://doi.org/10.1111/j.1365-3040.2007.01739.x
Carroll K, Gomez C, Shapiro L (2004) Tubby proteins: the plot thickens. Nat Rev Mol Cell Biol 5:55–63. https://doi.org/10.1038/nrm1278
Chebli Y, Geitmann A (2017) Cellular growth in plants requires regulation of cell wall biochemistry. Curr Opin Cell Biol 44:28–35. https://doi.org/10.1016/j.ceb.2017.01.002
Clausen MH, Willats WGT, Knox JP (2003) Synthetic methyl hexagalacturonate hapten inhibitors of anti-homogalacturonan monoclonal antibodies LM7, JIM5 and JIM7. Carbohydr Res 338:1797–1800. https://doi.org/10.1016/s0008-6215(03)00272-6
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743. https://doi.org/10.1046/j.1365-313x.1998.00343.x
Doblin MS, Kurek I, Jacob WD, Delmer DP (2002) Cellulose biosynthesis in plants: from genes to rosettes. Plant Cell Physiol 43:1407–1420. https://doi.org/10.1093/pcp/pcf164
Endler A, Persson S (2011) Cellulose synthases and synthesis in Arabidopsis. Mol Plant 4:199–211. https://doi.org/10.1093/mp/ssq079
Ezquer I, Mizzotti C, Nguema-Ona E, Gotte M, Beauzamy L, Viana VE, Dubrulle N, Costa de Oliveira A, Caporali E, Koroney AS, Boudaoud A, Driouich A, Colombo L (2016) The developmental regulator SEEDSTICK controls structural and mechanical properties of the Arabidopsis seed coat. Plant Cell 28:2478–2492. https://doi.org/10.1105/tpc.16.00454
Francoz E, Ranocha P, Burlat V, Dunand C (2015) Arabidopsis seed mucilage secretory cells: regulation and dynamics. Trends Plant Sci 20:515–524. https://doi.org/10.1016/j.tplants.2015.04.008
Freshour C, Clay RP, Fuller MS, Albersheim P, Darvill AG, Hahn MC (1996) Developmental and tissue-specific structural alterations of the cell-wall polysaccharides of Arabidopsis thaliana roots. Plant Physiol 110:1413–1429. https://doi.org/10.1104/pp.110.4.1413
Gnesutta N, Kumimoto RW, Swain S, Chiara M, Siriwardana C, Horner DS, Holt BF 3rd, Mantovani R (2017) CONSTANS Imparts DNA Sequence Specificity to the Histone Fold NF-YB/NF-YC Dimer. Plant Cell 29:1516–1532. https://doi.org/10.1105/tpc.16.00864
Golz JF, Allen PJ, Li SF, Parish RW, Jayawardana NU, Bacic A, Doblin MS (2018) Layers of regulation insights into the role of transcription factors controlling mucilage production in the Arabidopsis seed coat. Plant Sci 272:179–192. https://doi.org/10.1016/j.plantsci.2018.04.021
Gonzalez A, Mendenhall J, Huo Y, Lloyd A (2009) TTG1 complex MYBs, MYB5 and TT2, control outer seed coat differentiation. Dev Biol 325:412–421. https://doi.org/10.1016/j.ydbio.2008.10.005
Guillemin F, Guillon F, Bonnin E, Devaux MF, Chevalier T, Knox JP, Liners F, Thibault JF (2005) Distribution of pectic epitopes in cell walls of the sugar beet root. Planta 222:355–371. https://doi.org/10.1007/s00425-005-1535-3
Haughn GW, Western TL (2012) Arabidopsis seed coat mucilage is a specialized cell wall that can be used as a model for genetic analysis of plant cell wall structure and function. Front Plant Sci 3:64–68. https://doi.org/10.3389/fpls.2012.00064
Hu R, Li J, Yang X, Zhao X, Wang X, Tang Q, He G, Zhou G, Kong Y (2016) Irregular xylem 7 (IRX7) is required for anchoring seed coat mucilage in Arabidopsis. Plant Mol Biol 92:25–38. https://doi.org/10.1007/s11103-016-0493-4
Huang J, DeBowles D, Esfandiari E, Dean G, Carpita NC, Haughn GW (2011) The Arabidopsis transcription factor LUH/MUM1 is required for extrusion of seed coat mucilage. Plant Physiol 156:491–502. https://doi.org/10.1104/pp.111.172023
Ishii T (2002) A sensitive and rapid bioassay of homogalacturonan synthase using 2-aminobenzamide-labeled oligogalacturonides. Plant Cell Physiol 43:1386–1389. https://doi.org/10.1093/pcp/pcf150
Jefferson RA (1989) The GUS reporter gene system. Nature 342:837–838. https://doi.org/10.1038/342837a0
Johnson CS, Kolevski B, Smyth DR (2002) TRANSPARENT TESTA GLABRA2, a trichome and seed coat development gene of Arabidopsis, encodes a WRKY transcription factor. Plant Cell 14:1359–1375. https://doi.org/10.1105/tpc.001404
Kahle J, Baake M, Doenecke D, Albig W (2005) Subunits of the heterotrimeric transcription factor NF-Y are imported into the nucleus by distinct pathways involving importin beta and importin 13. Mol Cell Biol 25:5339–5354. https://doi.org/10.1128/MCB.25.13.5339-5354.2005
Kim JW, Kim HS, Kim SD, Park JY (2014) Insulin phosphorylates tyrosine residue 464 of tub and translocates tubby into the nucleus in HIRcB cells. Endocrinol Metab 29:163–168. https://doi.org/10.3803/EnM.2014.29.2.163
Kong Y, Zhou G, Yin Y, Xu Y, Pattathil S, Hahn MG (2011) Molecular analysis of a family of Arabidopsis genes related to galacturonosyltransferases. Plant Physiol 155:1791–1805. https://doi.org/10.1104/pp.110.163220
Kong Y, Zhou G, Abdeen AA, Schafhauser J, Richardson B, Atmodjo MA, Jung J, Wicker L, Mohnen D, Western T, Hahn MG (2013) GALACTURONOSYLTRANSFERASE-LIKE5 is involved in the production of Arabidopsis seed coat mucilage. Plant Physiol 163:1203–1217. https://doi.org/10.1104/pp.113.227041
Koornneef M (1981) The complex syndrome of the ttg mutants. Arabidopsis Inf Serv 18:45–51
Kumimoto RW, Zhang Y, Siefers N, Holt BF 3rd (2010) NF-YC3, NF-YC4 and NF-YC9 are required for CONSTANS-mediated, photoperiod-dependent flowering in Arabidopsis thaliana. Plant J 63:379–391. https://doi.org/10.1111/j.1365-313X.2010.04247.x
Kumimoto RW, Siriwardana CL, Gayler KK, Risinger JR, Siefers N, Holt BF (2013) NUCLEAR FACTOR Y transcription factors have both opposing and additive roles in ABA-mediated seed germination. PloS One 8:e59481. https://doi.org/10.1371/journal.pone.0059481
Kunieda T, Mitsuda N, Ohme TM, Takeda S, Aida M, Tasaka M, Kondo M, Nishimura M, Hara NI (2008) NAC family proteins NARS1/NAC2 and NARS2/NAM in the outer integument regulate embryogenesis in Arabidopsis. Plant Cell 20:2631–2642. https://doi.org/10.1105/tpc.108.060160
Lai CP, Lee CL, Chen PH, Wu SH, Yang CC, Shaw JF (2004) Molecular analyses of the Arabidopsis TUBBY-like protein gene family. Plant Physiol 134:1586–1597. https://doi.org/10.1104/pp.103.037820
Li YQ, Moscatelli A, Cai G, Cresti M (1997) Functional interactions among cytoskeleton, membranes, and cell wall in the pollen tube of flowering plants. Int Rev Cytol 176:133–199. https://doi.org/10.1016/s0074-7696(08)61610-1
Li SF, Milliken ON, Pham H, Seyit R, Napoli R, Preston J, Koltunow AM, Parish RW (2009) The Arabidopsis MYB5 transcription factor regulates mucilage synthesis, seed coat development, and trichome morphogenesis. Plant Cell 21:72–89. https://doi.org/10.1105/tpc.108.063503
Li L, Zheng W, Zhu Y, Ye H, Tang B, Arendsee ZW, Jones D, Li R, Ortiz D, Zhao X, Du C, Nettleton D, Scott MP, Salas-Fernandez MG, Yin Y, Wurtele ES (2015) QQS orphan gene regulates carbon and nitrogen partitioning across species via NF-YC interactions. Proc Natl Acad Sci USA 112:14734–14739. https://doi.org/10.1073/pnas.1514670112
Liepman AH, Wightman R, Geshi N, Turner SR, Scheller HV (2010) Arabidopsis-a powerful model system for plant cell wall research. Plant J 61:1107–1121. https://doi.org/10.1111/j.1365-313X.2010.04161.x
Liners F, Letesson J-J, Didembourg C, Cutsem PV (1989) Monoclonal antibodies against pectin. Plant Physiol 91:1419–1424
Lionetti V (2015) PECTOPLATE: the simultaneous phenotyping of pectin methylesterases, pectinases, and oligogalacturonides in plants during biotic stresses. Front Plant Sci 6:331. https://doi.org/10.3389/fpls.2015.00331
Lionetti V, Fabri E, De Caroli M, Hansen AR, Willats WG, Piro G, Bellincampi D (2017) Three pectin methylesterase inhibitors protect cell wall integrity for Arabidopsis immunity to botrytis. Plant Physiol 173:1844–1863. https://doi.org/10.1104/pp.16.01185
Liu X, Hu P, Huang M, Tang Y, Li Y, Li L, Hou X (2016) The NF-YC-RGL2 module integrates GA and ABA signalling to regulate seed germination in Arabidopsis. Nat Commun 7:12768. https://doi.org/10.1038/ncomms12768
Macquet A, Ralet MC, Kronenberger J, Marion PA, North HM (2007) In situ, chemical and macromolecular study of the composition of Arabidopsis thaliana seed coat mucilage. Plant Cell Physiol 48:984–999. https://doi.org/10.1093/pcp/pcm068
Mitsuda N, Ohme TM (2009) Functional analysis of transcription factors in Arabidopsis. Plant Cell Physiol 50:1232–1248. https://doi.org/10.1093/pcp/pcp075
Mohnen D (2008) Pectin structure and biosynthesis. Curr Opin Plant Biol 11:266–277. https://doi.org/10.1016/j.pbi.2008.03.006
Mukhopadhyay S, Jackson dPK (2011) The tubby family proteins. Genome Biol 12:225–233. https://doi.org/10.1186/gb-2011-12-6-225
Myers ZA, Kumimoto RW, Siriwardana CL, Gayler KK, Risinger JR, Pezzetta D, Holt Iii BF (2016) NUCLEAR FACTOR Y, subunit C (NF-YC) transcription factors are positive regulators of photomorphogenesis in Arabidopsis thaliana. PLoS Genet 12:e1006333. https://doi.org/10.1371/journal.pgen.1006333
North HM, Berger A, Saez AS, Ralet MC (2014) Understanding polysaccharide production and properties using seed coat mutants: future perspectives for the exploitation of natural variants. Ann Bot 114:1251–1263. https://doi.org/10.1093/aob/mcu011
Orfila C, Sorensen SO, Harholt J, Geshi N, Crombie H, Truong HN, Reid JS, Knox JP, Scheller HV (2005) QUASIMODO1 is expressed in vascular tissue of Arabidopsis thaliana inflorescence stems, and affects homogalacturonan and xylan biosynthesis. Planta 222:613–622. https://doi.org/10.1007/s00425-005-0008-z
Pattathil S, Avci U, Baldwin D, Swennes AG, McGill JA, Popper Z, Bootten T, Albert A, Davis RH, Chennareddy C, Dong R, O’Shea B, Rossi R, Leoff C, Freshour G, Narra R, O’Neil M, York WS, Hahn MG (2010) A comprehensive toolkit of plant cell wall glycan-directed monoclonal antibodies. Plant Physiol 153:514–525. https://doi.org/10.1104/pp.109.151985
Pauly M, Gille S, Liu L, Mansoori N, de Souza A, Schultink A, Xiong G (2013) Hemicellulose biosynthesis. Planta 238:627–642. https://doi.org/10.1007/s00425-013-1921-1
Peaucelle A, Braybrook S, Hofte H (2012) Cell wall mechanics and growth control in plants: the role of pectins revisited. Front Plant Sci 3:121–126. https://doi.org/10.3389/fpls.2012.00121
Penfield S, Meissner RC, Shoue DA, Carpita NC, Bevan MW (2001) MYB61 is required for mucilage deposition and extrusion in the Arabidopsis seed coat. Plant Cell 13:2777–2791. https://doi.org/10.1105/tpc.010265
Poschet G, Hannich B, Buttner M (2010) Identification and characterization of AtSTP14, a novel galactose transporter from Arabidopsis. Plant Cell Physiol 51:1571–1580. https://doi.org/10.1093/pcp/pcq100
Rautengarten C, Usadel B, Neumetzler L, Hartmann J, Bussis D, Altmann T (2008) A subtilisin-like serine protease essential for mucilage release from Arabidopsis seed coats. Plant J 54:466–480. https://doi.org/10.1111/j.1365-313X.2008.03437.x
Renak D, Dupl’akova N, Honys D (2012) Wide-scale screening of T-DNA lines for transcription factor genes affecting male gametophyte development in Arabidopsis. Sex Plant Reprod 25:39–60. https://doi.org/10.1007/s00497-011-0178-8
Rerie WG, Feldmann KA, Marks MD (1994) The GLABRA2 gene encodes a homeo domain protein required for normal tnchome development in Arabidopsis. Genes Dev 8:1388–1399. https://doi.org/10.1101/gad.8.12.1388
Rosti J, Barton CJ, Albrecht S, Dupree P, Pauly M, Findlay K, Roberts K, Seifert GJ (2007) UDP-glucose 4-epimerase isoforms UGE2 and UGE4 cooperate in providing UDP-galactose for cell wall biosynthesis and growth of Arabidopsis thaliana. Plant Cell 19:1565–1579. https://doi.org/10.1105/tpc.106.049619
Round AN, Rigby NM, MacDougall AJ, Morris VJ (2010) A new view of pectin structure revealed by acid hydrolysis and atomic force microscopy. Carbohydr Res 345:487–497. https://doi.org/10.1016/j.carres.2009.12.019
Saez AS, Ralet MC, Berger A, Botran L, Ropartz D, Marion PA, North HM (2013) PECTIN METHYLESTERASE INHIBITOR6 promotes Arabidopsis mucilage release by limiting methylesterification of homogalacturonan in seed coat epidermal cells. Plant Cell 25:308–323. https://doi.org/10.1105/tpc.112.106575
Scheller HV, Ulvskov P (2010) Hemicelluloses. Annu Rev Plant Biol 61:263–289. https://doi.org/10.1146/annurev-arplant-042809-112315
Shi DC, Wang J, Hu RB, Zhou GK, O’Neill MA, Kong YZ (2017) Boron-bridged RG-II and calcium are required to maintain the pectin network of the Arabidopsis seed mucilage ultrastructure. Plant Mol Biol 94:267–280. https://doi.org/10.1007/s11103-017-0606-8
Sterling JD, Atmodjo MA, Inwood SE, Kumar Kolli VS, Quigley HF, Hahn MG, Mohnen D (2006) Functional identification of an Arabidopsis pectin biosynthetic homogalacturonan galacturonosyltransferase. Proc Natl Acad Sci USA 103:5236–5241. https://doi.org/10.1073/pnas.0600120103
Tang Y, Liu X, Liu X, Li Y, Wu K, Hou X (2017) Arabidopsis NF-YCs Mediate the Light-Controlled Hypocotyl Elongation via Modulating Histone Acetylation. Mol Plant 10:260–273. https://doi.org/10.1016/j.molp.2016.11.007
Usadel B, Kuschinsky AM, Rosso MG, Eckermann N, Pauly M (2004) RHM2 is involved in mucilage pectin synthesis and is required for the development of the seed coat in Arabidopsis. Plant Physiol 134:286–295. https://doi.org/10.1104/pp.103.034314
Voiniciuc C, Dean GH, Griffiths JS, Kirchsteiger K, Hwang YT, Gillett A, Dow G, Western TL, Estelle M, Haughn GW (2013) Flying saucer1 is a transmembrane RING E3 ubiquitin ligase that regulates the degree of pectin methylesterification in Arabidopsis seed mucilage. Plant Cell 25:944–959. https://doi.org/10.1105/tpc.112.107888
Voiniciuc C, Schmidt MH, Berger A, Yang B, Ebert B, Scheller HV, North HM, Usadel B, Gunl M (2015a) MUCILAGE-RELATED10 produces galactoglucomannan that maintains pectin and cellulose architecture in Arabidopsis seed mucilage. Plant Physiol 169:403–420. https://doi.org/10.1104/pp.15.00851
Voiniciuc C, Yang B, Schmidt MH, Gunl M, Usadel B (2015b) Starting to gel: how Arabidopsis seed coat epidermal cells produce specialized secondary cell walls. Int J Mol Sci 16:3452–3473. https://doi.org/10.3390/ijms16023452
Walker M, Tehseen M, Doblin MS, Pettolino FA, Wilson SM, Bacic A, Golz JF (2011) The transcriptional regulator LEUNIG_HOMOLOG regulates mucilage release from the Arabidopsis testa. Plant Physiol 156:46–60. https://doi.org/10.1104/pp.111.172692
Wang M, Yuan D, Gao W, Li Y, Tan J, Zhang X (2013) A comparative genome analysis of PME and PMEI families reveals the evolution of pectin metabolism in plant cell walls. PloS ONE 8:e72082. https://doi.org/10.1371/journal.pone.0072082
Wang M, Xu Z, Kong Y (2018) The tubby-like proteins kingdom in animals and plants. Gene 642:16–25. https://doi.org/10.1016/j.gene.2017.10.077
Western TL, Young DS, Dean GH, Tan WL, Samuels AL, Haughn GW (2004) MUCILAGE-MODIFIED4 encodes a putative pectin biosynthetic enzyme developmentally regulated by APETALA2. TRANSPARENT TESTA GLABRA1 and GLABRA2 in the Arabidopsis seed coat. Plant Physiol 134:296–306. https://doi.org/10.1104/pp.103.035519
Willats WGT, McCartney L, Mackie W, Knox JP (2001) Pectin: cell biology and prospects for functional analysis. Plant Mol Biol 47:9–27. https://doi.org/10.1023/A:1010662911148
Yapo BM, Lerouge P, Thibault JF, Ralet MC (2007) Pectins from citrus peel cell walls contain homogalacturonans homogenous with respect to molar mass, rhamnogalacturonan I and rhamnogalacturonan II. Carbohydr Polym 69:426–435. https://doi.org/10.1016/j.carbpol.2006.12.024
Yeap WC, Lee FC, Shabari Shan DK, Musa H, Appleton DR, Kulaveerasingam H (2017) WRI1-1. ABI5, NF-YA3 and NF-YC2 increase oil biosynthesis in coordination with hormonal signaling during fruit development in oil palm. Plant J 91:97–113. https://doi.org/10.1111/tpj.13549
Yoo SD, Cho YH, Sheen J (2007) Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. Nat Protoc 2:1565–1572. https://doi.org/10.1038/nprot.2007.199
Yotsui I, Saruhashi M, Kawato T, Taji T, Hayashi T, Quatrano RS, Sakata Y (2013) ABSCISIC ACID INSENSITIVE3 regulates abscisic acid-responsive gene expression with the nuclear factor Y complex through the ACTT-core element in Physcomitrella patens. New Phytol 199:101–109. https://doi.org/10.1111/nph.12251
Yu L, Shi D, Li J, Kong Y, Yu Y, Chai G, Hu R, Wang J, Hahn MG, Zhou G (2014) CELLULOSE SYNTHASE-LIKE A2, a glucomannan synthase, is involved in maintaining adherent mucilage structure in Arabidopsis seed. Plant Physiol 164:1842–1856. https://doi.org/10.1104/pp.114.236596
Zhao X, Qiao L, Wu AM (2017) Effective extraction of Arabidopsis adherent seed mucilage by ultrasonic treatment. Sci Rep 7:40672–40679. https://doi.org/10.1038/srep40672
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31470291, 31670302), the National Key Technology R&D Program (2015BAD15B03), the Taishan Scholar Program of Shandong (to GZ) and the Elite Youth Program of Chinese Academy of Agricultural Sciences (to YK).
Author information
Authors and Affiliations
Contributions
MW, ZX, YK, RH and GZ conceived the study. MW, ZX and YW performed the experiments. MW analyzed the data and ZX prepared the figures. MW, ZX, Ahmed. RI and YK drafted the manuscript. All of the authors discussed the results, edited the manuscript, and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11103_2019_827_MOESM1_ESM.pdf
Supplementary material 1 Immunolabelling and immunoblotting of CCRC-M36 in tlp2-1 mutants. A: Immnunolabeling of CCRC-M36. The first line indicates the merged images of double labeling with CCRC-M36 and calcofluor, the second line indicates the calcofluor labeling images, the third line indicates CCRC-M36 labeling. Scale bar = 100 µm. B: Immunoblotting results of RG I backbone from soluble mucilage and adherent mucilage of WT, tlp2-1, complemented tlp2-1 seeds. The extracted dried mucilage were resolved in deionized water and serially diluted, and spotted on a nitrocellulose membrane. Dot blot analyses with CCRC-M36 which recognizes RG I backbone (PDF 21598 KB)
11103_2019_827_MOESM2_ESM.pdf
Supplementary material 2 Immunolabelling of JIM7 and JIM5 in tlp2-1 mutant seed coat mucilage. A: Immunolabelling of JIM7 which recognizes esterified HG. A1-A3, merged images of double labeling with JIM7 and calcofluor. A4-A6, staining of calcofluor. A7-A9, staining of JIM7. Scale bar =50 µm. The red arrow head indicates the JIM7 signals. B: Immunolabelling of JIM5 which recognizes low esterified HG. B1-B3, merged images of double labeling with JIM5 and calcofluor. B4-B6, staining of calcofluor. B7-B9, staining of JIM5. Scale bar =50 µm (PDF 41838 KB)
11103_2019_827_MOESM3_ESM.pdf
Supplementary material 3 Immunoblotting of homogalacturonan in tlp2-1 mutant seed coat mucilage. Immunoblotting results of partially esterified and low esterified HG from soluble mucilage and adherent mucilage of WT, tlp2-1, complemented tlp2-1 seeds. The extracted dried mucilage were resolved in deionized water and serially diluted, and spotted on a nitrocellulose membrane. Dot blot analyses with JIM7, JIM5 which can bind esterified and low esterified HG, respectively (PDF 17062 KB)
11103_2019_827_MOESM4_ESM.pdf
Supplementary material 4 The expression pattern of TLP2 in different tissues. The expression patterns of TLP2 in different tissues and organs depicted by the Arabidopsis AtGenExpress database/ BAR eFP browser (PDF 790 KB)
11103_2019_827_MOESM5_ESM.pdf
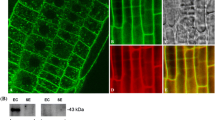
Supplementary material 5 The subcellular localization of pTLP2::TLP2-GFP. A: TLP2-GFP fusion protein subcellular localization. B: Bright field. C: Merged image (PDF 3518 KB)
Rights and permissions
About this article
Cite this article
Wang, M., Xu, Z., Ahmed, R.I. et al. Tubby-like Protein 2 regulates homogalacturonan biosynthesis in Arabidopsis seed coat mucilage. Plant Mol Biol 99, 421–436 (2019). https://doi.org/10.1007/s11103-019-00827-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-019-00827-9