Abstract
Key message
Comparative transcriptome analysis of wild and cultivated chrysanthemums provides valuable genomic resources and helps uncover common and divergent patterns of genome and gene evolution in these species.
Abstract
Plants are unique in that they employ polyploidy (or whole-genome duplication, WGD) as a key process for speciation and evolution. The Chrysanthemum genus is closely associated with hybridization and polyploidization, with Chrysanthemum species exhibiting diverse ploidy levels. The commercially important species, C. morifolium is an allohexaploid plant that is thought to have originated via the hybridization of several Chrysanthemum species, but the genomic and molecular evolutionary mechanisms remain poorly understood. In the present study, we sequenced and compared the transcriptomes of C. morifolium and the wild Korean diploid species, C. boreale. De novo transcriptome assembly revealed 11,318 genes in C. morifolium and 10,961 genes in C. boreale, whose functions were annotated by homology searches. An analysis of synonymous substitution rates (Ks) of paralogous and orthologous genes suggested that the two Chrysanthemum species commonly experienced the Asteraceae paleopolyploidization and recent genome duplication or triplication before the divergence of these species. Intriguingly, C. boreale probably underwent rapid diploidization, with a reduction in chromosome number, whereas C. morifolium maintained the original chromosome number. Analysis of the ratios of non-synonymous to synonymous nucleotide substitutions (Ka/Ks) between orthologous gene pairs indicated that 107 genes experienced positive selection, which may have been crucial for the adaptation, domestication, and speciation of Chrysanthemum.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Chrysanthemum shows considerable diversity in terms of ploidy level, as well as flower shape, color, and size. This genus comprises approximately 40 species that exhibit various degrees of polyploidy, from diploid to decaploid, with nine chromosomes as the basal unit (Dowrick 1952; Liu et al. 2012). The commercial species, C. morifolium, is used as an ornamental and medicinal plant worldwide and appears to be a hexaploid species (2n = 6x = 54). Cytogenetic, molecular phylogenetic, and genetic studies suggested that C. morifolium originated from the natural hybridization of species such as C. indicum, C. lavandulifolium, C. nankingense, C. vestitum, and C. zawadskii (Chen 1985; Dai et al. 1998, 2005; Ma et al. 2016; Zhou and Silan 2001). However, the extent and timing of evolutionary events such as polyploidy or whole-genome duplication (WGD) in C. morifolium remain uncertain.
WGD is one of the key factors in the evolution, speciation, and diversification of the angiosperm lineage (Soltis et al. 2009; Soltis and Soltis 2016; Van de Peer et al. 2009b; Wendel et al. 2016). Genome and transcriptome sequence analyses have revealed that angiosperms shared an ancient WGD known as paleopolyploidy and additional recent lineage-specific WGDs (Jiao et al. 2011; Tang et al. 2008; Van de Peer et al. 2009a). After WGD, gene pairs that were duplicated within species (also known as paralogs) are retained or lost in the mode of neo-, sub-, or non-functionalization followed by extensive genome rearrangement and diploidization (Panchy et al. 2016; Soltis et al. 2015; Van de Peer et al. 2009b). Such post-WGD events are variable and extensive enough to result in reproductive isolation and ultimately, speciation (Paterson et al. 2010; Vanneste et al. 2014). The cycles of WGD and the subsequent dynamic changes are recurrent events that are associated with the radiation and diversity of angiosperm species (Paterson et al. 2010; Soltis and Soltis 2016; Vanneste et al. 2014).
With the accumulation of genome-wide nucleotide sequences, evolutionary events such as WGD and speciation can be successfully inferred by evaluating the age distribution of homologous gene pairs (Blanc and Wolfe 2004; Van de Peer et al. 2009a). Synonymous nucleotide substitutions of protein-coding genes are not accompanied by amino acid changes, and are thus neutral and free from natural selection (Blanc and Wolfe 2004). Therefore, the rate of synonymous substitutions (Ks) is proportional to the time lapse since the generation of two homologous genes and is thus used to approximate the timing of the occurrence of homologs, much like a molecular clock (Blanc and Wolfe 2004). Because WGD results in the production of excessive paralogous gene pairs of a particular age, the Ks distribution of paralogs displays a peak of high density at a specific Ks value from which the timing of WGD is deduced (Blanc and Wolfe 2004). Similarly, the Ks distribution of homologous gene pairs between two different species (also known as orthologs) is used to determine the time of speciation (Blanc and Wolfe 2004). The rate of nonsynonymous substitutions (Ka) and the Ka/Ks ratio also serve as useful parameters to investigate the molecular evolution of two species that have diverged (Fay and Wu 2003). Because synonymous substitution occurs more frequently than nonsynonymous substitution, the Ka/Ks ratios of most orthologous genes are less than one, indicating that the gene pair is under purifying/negative selection (Fay and Wu 2003). By contrast, orthologs under adaptive/positive selection exhibit Ka/Ks values greater than one, providing insights into the molecular evolutionary framework underlying adaptation, divergence, and speciation (Fay and Wu 2003).
The Asteraceae (Compositae), one of the largest and most diverse plant families, includes 13 subfamilies, 45 tribes, 1,911 genera, and 32,913 species (Fu et al. 2016; MBG 2013). The phylogenetic relationships among the major lineages of Asteraceae have been thoroughly investigated via comparisons of genes in the chloroplast or nucleus (Linder et al. 2000; Panero et al. 2014), and information about genome evolution within Asteraceae was also recently obtained. A comparison of linkage maps between Lactuca sativa and Vitis vinifera revealed the occurrence of paleohexaploidy in Asteraceae (Truco et al. 2013), which was confirmed in four major subfamilies (Asteroideae, Cichorioideae, Carduoideae, and Mutisiodeae) by analyzing the Ks distributions for 18 species (Barker et al. 2008). Additionally, Asteroideae and Mutisiodeae experienced another round of WGD (Barker et al. 2008). These lineage-specific WGDs were also detected in several tribes or genera showing species richness (Huang et al. 2016; Panero and Crozier 2016). Similar analyses using more diverse taxa showed that there were two rounds of WGD, one prior to the divergence between Asteraceae and its sister family Calyceraceae and another specific for the core Asteraceae (Barker et al. 2016; Huang et al. 2016). Although these studies have characterized the evolutionary history in representative lineages of Asteraceae, few studies have been conducted on the Chrysanthemum genus on a genome-wide scale.
In the present study, we sequenced the transcriptome of C. morifolium and subjected it to molecular evolutionary analysis. For comparison, we included the diploid wild species, C. boreale (2n = 2x = 18), which is native to Korea. We inferred WGD and species divergence time by identifying the paralogs and orthologs within and between species and plotting their Ks distribution. Finally, we identified genes under positive selection, providing genome-wide information about genes that may have been involved in the domestication or adaption of C. morifolium.
Materials and methods
Plant materials
C. morifolium (cv. Baekma) plants were kindly provided by the National Institute of Horticultural and Herbal Science (NIHHS), Republic of Korea. C. boreale (IT121002) was collected from the Republic of Korea as previously described (Hwang et al. 2013). All plants were propagated by stem cuttings and grown in a greenhouse in NIHHS under natural light conditions.
RNA extraction, library construction, and sequencing
Total RNA was extracted from leaf tissue using TRIzol Reagent (Invitrogen, USA). RNA quantity and quality were evaluated using a NanoDrop spectrophotometer (NanoDrop Technologies, USA), electrophoresis on a 1% denaturing agarose gel, and an Agilent 2100 Bioanalyzer (Agilent Technologies, USA). GS FLX cDNA library construction, emulsion PCR, and pyrosequencing were conducted at the National Instrumentation Center for Environment Management (NICEM, Seoul National University, Republic of Korea; http://nicem.snu.ac.kr) following standard procedures. Approximately 5 µg of total RNA per sample was used for library construction, as described in the cDNA Rapid Library Preparation Method Manual provided with GS FLX Titanium Series reagents (Roche, USA). The libraries were amplified using emPCR kits (Roche, USA) and sequenced on a 454 GS FLX Titanium Sequencer (Roche, USA) according to the manufacturer’s instructions. The sequencing data were deposited in the National Center for Biotechnology Information (NCBI) Sequence Read Archive with accession numbers SRR5768981 and SRR5768982.
Assembly and functional annotation of the transcriptome
The raw data were assembled using Newbler ver. 2.6 software (Roche, USA, http://www.454.com) with the cDNA option for transcriptome assembly. Trimming and assembly of raw reads resulted in the production of isotigs or contigs (equivalent to transcripts, including splice variants), which were grouped into isogroups (equivalent to genes). Additionally, raw reads that were not assembled into isotigs or contigs were remained as singletons and preprocessed using the Lucy DNA sequence quality and vector trimming tool (Chou and Holmes 2001). After removing short singletons less than 200 bp, redundant singletons were filtered out using the CD-HIT-EST program with the option −c 0.90 (90% sequence identity) −n 10 −r 1 (comparing both strands) (Li and Godzik 2006).
The assembled transcripts and singletons were annotated by homology searches using the Basic Local Alignment Search Tool (BLASTX) against the UniProtKB/Swiss-Prot database (http://web.expasy.org/docs/swiss-prot_guideline.html) and the NCBI Viridiplantae (green plants) non-redundant (nr) protein database with an e-value threshold of 1e-5. Based on annotation using the Swiss-Prot database, Gene Ontology (GO) terms were assigned using the BLAST2GO program (Conesa et al. 2005). For each GO term, the p-value for the number of genes between the two species was calculated via Pearson’s Chi-Square test using the WEGO program (Ye et al. 2006). The GO distribution at level 2 was plotted.
Identification of putative orthologs and paralogs
Proteins and transcripts sequences for globe artichoke (Cynara cardunculus var. scolymus) were downloaded from the Globe Artichoke Genome Database (http://www.artichokegenome.unito.it). Open reading frames (ORFs) for Chrysanthemum species were determined using TransDecoder (Haas et al. 2013) with the option minimum ORF size of 300 bp and minimum protein length of 50 amino acids. For transcripts resulting in multiple ORFs, the longest ORF was used for analysis. Since redundancy via alternative splicing also biases downstream analysis, the longest transcript per isogroup or unigene was subjected to analysis. Orthologs and paralogs among the three species were identified by conducting OrthoMCL analysis (Li et al. 2003). Briefly, all-against-all BLASTP searches were performed using the translated protein sequences with an e-value threshold of 1e-10 and an identity threshold of 50%, and the Markov Clustering Algorithm was used to cluster similar sequences into orthologous and paralogous groups. Orthologous gene pairs were retrieved from orthogroups containing one gene per species. For each species, paralogous genes were determined from orthogroups containing two genes per species, regardless of the contribution of the other species.
Estimation of divergence time, and detection of WGD and positively selected genes
The analysis was conducted as previously described (Blanc and Wolfe 2004; Kim et al. 2014). For each ortholog or paralog, the protein sequences were aligned using ClustalW (Larkin et al. 2007) and the corresponding codons were aligned using PAL2NAL (Suyama et al. 2006) with the guidance of coding sequences (CDSs). The Ka and Ks values were calculated using the Nei–Gojobori method (Nei and Gojobori 1986) implemented in the PAML package and subjected to Fisher’s exact test to determine p-values (Yang 1997). The Ks distributions were fitted with log-Gaussian mixture models using Gaussian Mixture Models with Bayes Factors and plotted. The time of WGD and species divergence occurrence was calculated using the following equation: T = Ks/(2 × 1.5 × 10−8 substitutions/synonymous site/year) (Koch et al. 2000). To classify genes according to the selection type, the Ka/Ks score was calculated for each orthologous gene pair. Orthologous gene pairs with Ks > 0.1 were excluded from analysis to identify genes under positive selection to avoid the identification of potential paralogs (Cheng et al. 2015).
Results
Sequencing and assembly of transcriptomes
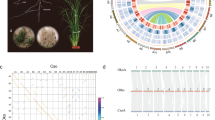
The pyrosequencing of cDNA libraries yielded 673,206 and 660,419 raw reads for C. boreale and C. morifolium, respectively, with an average length of 445 bp in both libraries (Table 1). De novo assembly of preprocessed reads resulted in the identification of 10,961 isogroups containing 13,841 isotigs for C. boreale and 11,318 isogroups containing 16,769 isotigs for C. morifolium (Table 1). Isotigs from the two species showed similar length distributions, with an average length of 1216 bp for C. boreale and 1230 bp for C. morifolium, although the number of obtained isotigs was greater in C. morifolium than in C. boreale (Fig. 1; Table 1). Additionally, 76,428 and 97,947 raw reads were retained as singletons and further preprocessed into 41,368 and 57,035 clean reads for C. boreale and C. morifolium, respectively, with an average length of ~ 480 bp (Table 1).
Functional annotation of transcripts
The transcripts were annotated based on sequence homology searches. First, BLASTX analysis against Viridiplantae proteins in the NCBI nr database revealed that 10,426 (95.12%) and 10,591 (93.58%) isogroups had significant matches in C. boreale and C. morifolium, respectively (Table 2). In both species, the species with the most BLASTX hits was Cynara cardunculus var. scolymus, followed by Vitis vinifera, Sesamum indicum, and Coffea canephora. A comparison to the UniProtKB/Swiss-Prot protein database showed that 8957 (81.72%) and 9013 (79.63%) isogroups returned positive hits in C. boreale and C. morifolium, respectively (Table 2). Overall, more than 90% of isogroups were functionally annotated by at least one public protein sequence database, which was also observed for isotigs (Table 2). However, the percentage of sequences that had significant BLAST hits was lower in singletons than in isogroups and isotigs. A total of 27,196 (65.74%) and 34,296 (60.13%) singletons were annotated in C. boreale and C. morifolium, respectively (Table 2). Similar to the isogroups, Cynara cardunculus var. scolymus, V. vinifera, and Coffea canephora were the top three species that were mostly highly represented in BLAST.
To categorize the functions of the transcripts, we assigned them to Gene Ontology (GO) terms based on Swiss-Prot annotation. In general, the GO distributions were highly similar in both species and included various categories, although several GO terms were significantly different between the two species (Fig. 2). In C. boreale, a total of 8380 (76.45%) isogroups were successfully mapped to one or more GO terms (Table 2), with 7902 (72.09%) assigned to biological process, 8259 (75.35%) to cellular component, and 7841 (71.54%) to molecular function. Likewise, in C. morifolium, 8445 (74.62%) isogroups were assigned to GO terms (Table 2), including 7919 (69.97%) to biological process, 8304 (73.37%) to cellular component, and 7888 (69.69%) to molecular function. In contrast to the isogroups, the annotation rates were lower for singletons, with 16,995 (41.08%) C. boreale and 23,918 (41.94%) C. morifolium transcripts mapped to GO terms (Table 2). The singletons included genes with relatively low expression levels and many were functionally annotated; however, since the singletons were very short compared to isogroups (Table 1), it is possible that they also included contaminants or artifacts derived from cDNA synthesis and sequencing (Meyer et al. 2009). Therefore, singletons were excluded from subsequent analyses.
Gene ontology (GO) classification of isogroups. Annotation results from Swiss-Prot were mapped to the second level of GO terms, which are represented by blue and red bars for C. boreale and C. morifolium, respectively. Statistically significant differences between the two species are indicated (**p < 0.01 and *p < 0.05)
Clustering of transcripts
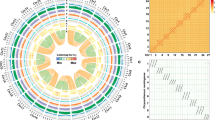
To compare the transcriptome profiles of the two species, we constructed orthologous gene clusters using OrthoMCL. Among the species whose genome sequences were available, we included the phylogenetically closest species, Cynara cardunculus var. scolymus, to investigate evolutionary events (see below). A total of 5036 orthologous clusters consisting of 21,076 genes were common to all three species (Fig. 3), 2970 of which were identified as single-copy gene clusters and used to estimate the divergence time of species. We counted 29, 51, and 2055 species-specific orthogroups in C. boreale, C. morifolium, and Cynara cardunculus var. scolymus, respectively (Fig. 3). Comparison of the two genera revealed that 773 and 2055 gene clusters were unique to Chrysanthemum and Cynara, respectively (Fig. 3).
Summary of OrthoMCL analysis. a Venn diagram showing OrthoMCL-based gene clusters in C. boreale, C. morifolium, and Cynara cardunculus var. scolymus. The number of gene clusters and genes (within parenthesis) is indicated for each section. b Classification of Chrysanthemum orthoclusters based on the contribution of each species
The two Chrysanthemum species shared 5729 gene clusters, whereas 1293 and 1174 gene families were specific to C. boreale and C. morifolium, respectively (Fig. 3a). We compared the number of genes per orthologous cluster in the 5729 clusters of the two species and found that 4,889 clusters contained the same number of genes in both species (Fig. 3b); in particular, 4512 were identified as single-copy orthogroups. However, 462 and 378 clusters contained more genes from C. boreale and C. morifolium, respectively, than from the other species (Fig. 3b). Specifically, clusters with a 2:1 gene ratio in C. boreale and C. morifolium were the most abundant, accounting for 325 groups, followed by the opposite (1:2) ratio for 276 groups. When we included species-specific clusters without the contribution of the other Chrysanthemum counterpart, we obtained a similar result, as 1755 and 1552 orthologous groups had surplus genes from C. boreale and C. morifolium, respectively.
In addition to orthologs, clustering analysis revealed 978, 945, and 4,326 paralogous gene clusters for C. boreale, C. morifolium, and Cynara cardunculus var. scolymus, respectively. Among these, 731, 719, and 2715 clusters included pairs of paralogous genes, which was used to examine WGD in each species.
Species divergence time and WGD
To investigate the divergence time of and WGD in two Chrysanthemum species, we examined the distribution of Ks values for orthologous gene pairs and paralogous gene pairs, respectively. At the level of the Asteraceae family, two Ks distributions for orthologs between each Chrysanthemum species and Cynara (Cb–Cc, Cm–Cc) were perfectly overlapping and showed a peak at Ks = 0.5 (Fig. 4a). Between C. boreale and C. morifolium, the Ks distribution of orthologous gene pairs (Cb–Cm) showed a prominent peak at 0.05, while a comparison of paralogous gene pairs within each species revealed a relatively sharp Ks peak at 0.1 as well as a broad peak around Ks = 0.9 for C. boreale (Cb–Cb) and Ks = 0.75 for C. morifolium (Cm–Cm) (Fig. 4a). The paralogous pairs for Cynara cardunculus var. scolymus (Cc–Cc) also showed a broad peak at Ks = 0.6 (Fig. 4a). Based on the Ks value distributions, Chrysanthemum and Cynara likely shared an ancient WGD (Ks = 0.6–0.9) and then diverged (Ks = 0.5) (Fig. 4b). Additionally, two Chrysanthemum species appear to have experienced a WGD or whole-genome triplication (WGT) event (Ks = 0.1), as well as a species divergence (Ks = 0.05) (Fig. 4b). We estimated the timing of these evolutionary events using the positions of Ks peaks and clock-like synonymous substitution rates for dicots (Koch et al. 2000). We inferred that the Asteraceae species experienced an ancient WGD event 20–30 million years ago (MYA) and that the Chrysanthemum genus diverged from Cynara 16.7 MYA (Fig. 4b). In addition, the two Chrysanthemum species experienced a WGD event 3.3 MYA and then diverged 1.7 MYA (Fig. 4b).
Divergence and whole-genome duplication in Chrysanthemum. a Distribution of synonymous nucleotide substitutions (Ks) between orthologs and paralogs from C. boreale (Cb), C. morifolium (Cm), and Cynara cardunculus var. scolymus (Cc). b A proposed model describing the evolutionary events in Chrysanthemum. Stars and circle indicate whole-genome duplication (or triplication) and diploidization, respectively. The recent polyploidy is not specific to C. morifolium but is instead shared with other members of the Chrysanthemum genus
Detection of sequence divergence
Considering the importance of non-synonymous nucleotide substitutions to protein function and speciation, we also calculated the Ka values and Ka/Ks ratios for orthologs between the two Chrysanthemum species. OrthoMCL clustering revealed a total of 4512 orthologous clusters that contained one gene each from C. boreale and C. morifolium, regardless of the contribution of Cynara cardunculus var. scolymus. Among these, 137 pairs were identical, with Ka and Ks values of zero, 53 pairs did not result in Ks values, and 118 or 630 pairs had only nonsynonymous or synonymous substitutions, respectively, which were excluded from further analysis. Additionally, 990 orthologous pairs with Ks > 0.1 were known to be potential paralogs and were also removed (Cheng et al. 2015). The remaining 2584 orthologous pairs had both types of substitutions (Fig. 5), although the two species separated very recently. The mean Ka, Ks, and Ka/Ks values were 0.01229, 0.04313, and 0.31642, respectively. We found that 2477 orthologous gene pairs (95.86%) were under purifying selection, with Ka/Ks < 1, whereas 107 pairs (4.14%) were under positive selection, with Ka/Ks > 1 (Fig. 5). Among the genes under positive selection, all except three share sequence similarity with known proteins; these genes are listed in Table S1. These genes encode proteins involved in processes at various levels, such as transcriptional regulation (NAC domain-containing protein, trihelix protein, and CBF1-interacting co-repressor), post-translational modification (heat-shock protein), signal transduction (protein phosphatase 2C-like protein, G protein alpha subunit, GYF-like protein, and TIP41), chromatin condensation (regulator of chromosome condensation repeat), disease resistance (mannose-binding lectin and Kunitz-like protease inhibitor), and so on (Table S1).
Discussion
In this study, we characterized the transcriptomes of wild and cultivated Chrysanthemum species. Sequencing and assembly resulted in the identification of 10,961 and 11,318 genes in C. boreale and C. morifolium, respectively, with 41,368 and 57,035 unassembled singletons, respectively (Table 1). Given that the number of genes in plant species range from 20,169 to 94,000 (Michael and Jackson 2013), the assembled transcripts represented only a portion of the genes present in each species, but the data obtained in this study were sufficient to allow us to perform evolutionary analysis between the two Chrysanthemum species. In addition, GO analysis revealed that the transcriptomes obtained in this study included functionally diverse genes in Chrysanthemum (Fig. 2).
Although the amount of raw data used for transcriptome assembly in both species was similar, the analysis of C. morifolium resulted in the discovery of more isogroups, isotigs, and singletons compared to C. boreale (Table 1), perhaps due to the differences in genome size and polyploidy. C. morifolium is a hexaploid species whose genome size is estimated to be approximately 8.83 Gbp (http://www.kew.org/cvalues), whereas C. boreale is a diploid species with a 2.94 Gbp genome (unpublished data, SY Won and J-A Jung). There is a significant positive correlation between the number of genes in an organism and genome size (Hou and Lin 2009). In polyploid species, genome doubling contributes to the expansion and diversification of gene contents, although the subsequent diploidization event involves the loss of a copy of each duplicated gene (Soltis et al. 2015). For example, 35% of genes were lost after the WGT in Brassica oleracea compared to the diploid species, Arabidopsis thaliana (Town et al. 2006). Indeed, B. oleracea was predicted to contain 45,758 genes (Liu et al. 2014), whereas the A. thaliana genome contains 25,498 genes (Initiative 2000). In particular, genes involved in the regulation of metabolic and biosynthetic processes and RNA metabolism, as well as transcription factor genes, were over-retained after WGT in B. oleracea (Liu et al. 2014).
Comparative genomic and transcriptomic analyses have revealed species- or lineage-specific genes representing major contributors to species- or lineage-specific phenotypes, adaptation, and evolution (Kaessmann 2010). Here, OrthoMCL-based clustering identified gene families that were shared among species or unique to each species (Fig. 3). Since the transcripts obtained in this study represent a subset of genes present in Chrysanthemum compared to the genome-based gene set in Cynara cardunculus var. scolymus, the 2055 Cynara-specific gene families were probably overestimated, whereas the 773 Chrysanthemum-specific gene families are indeed unique to this genus. In a comparison between Chrysanthemum species, we identified genes specific to both the wild and cultivated species, although these genes should be examined again once a more complete gene set is available. Initially, we assumed that genes in diploid species would be triplicated in hexaploid progeny if the two species diverged recently and the extra duplicated gene copies were maintained. Therefore, each orthologous cluster was considered to consist of more genes from C. morifolium than from C. boreale. Specifically, orthologous groups containing more genes from C. morifolium were likely to be more frequent than those from C. boreale. However, the clustering results showed the opposite pattern than would be expected with this assumption, perhaps due to WGD and diploidization events in C. boreale (see below).
Analysis of the Ks distribution of paralogs revealed two rounds of WGD in Chrysanthemum (Fig. 4). According to the timing of the WGD, it appears that the ancient event (Ks = 0.6–0.9) was shared with another Asteraceae member, Cynara cardunculus var. scolymus (Fig. 4a), whose Ks value overlapped with the Asteraceae-specific Ks peak at approximately 0.7–1.4 (Barker et al. 2016; Huang et al. 2016). Additionally, a peak at Ks = 0.1 was detected (Fig. 4a), which might contribute to the steep background of Ks distribution, revealing the ongoing process of gene birth and death (Blanc and Wolfe 2004). However, it cannot be ruled out that the recent peak is indicative of a hexaploidization event in C. morifolium, although it is still unclear whether this event involved autopolyploidy or allopolyploidy. A recent cytological and molecular analysis revealed segmental allohexaploidy in cultivated chrysanthemums (Klie et al. 2014). Notably, C. boreale also showed an unusual peak at the same position (Ks = 0.1) (Fig. 4a). Considering the divergence time of the two species at Ks = 0.05, one possible explanation is that a recent WGT event occurred in the common ancestor, and was followed by different evolutionary events in the two species. While C. morifolium maintained the original polyploidy status, it is likely that this particular diploid species experienced a rapid diploidization event, including a reduction in chromosome number. This explanation is supported by evidence obtained by OrthoMCL-based clustering. If the recent hexaploidization event occurred only in C. morifolium but not in C. boreale, orthologous clusters containing more genes from C. morifolium are expected to be more frequent. However, the opposite results were obtained, supporting a recent WGD or WGT in C. boreale.
Although we found that C. boreale and C. morifolium diverged from each other very recently, many characteristics of these species are quite different, such as their morphology, resistance, and growth habit, including the observed differences between wild and cultivated species and/or traits that accumulated in C. morifolium during the domestication process. One of the most effective approaches for revealing the molecular mechanisms behind these differences is to calculate the Ka/Ks ratios between orthologous gene pairs, especially due to the increasing availability of genomic and transcriptomic sequences. For orthologous gene pairs between the two Chrysanthemum species, most (over 95%) are subjected to purifying selection, with Ka/Ks less than one, resulting in the removal of deleterious mutations; this is consistent with the findings for many other plant species (Schlueter et al. 2004). However, the 107 remaining genes are under positive selection, which might influence the adaptation and evolution of each species. We found that these genes are involved in many different aspects of regulation. Among these, one representative class of genes encodes transcription factors or repressors such as NAC (NAM, ATAF1/2, and CUC2) domain-containing protein, trihelix protein, and CBF1 (C-repeat/DRE binding factor 1)-interacting co-repressor. Genes encoding regulator of chromosome condensation repeat (RCC) and Rad21/Rec8-like protein were also found to be under positive selection, conferring an additional layer of transcriptional regulation on the mode of chromosome condensation or structure. In addition, genes involved in signal transduction processes are subjected to evolutionary pressure. These include genes encoding NB-ARC (nucleotide binding adaptor shared by APAF-1, R proteins, and CED-4)-like protein, protein phosphatase 2C-like protein, G protein alpha subunit, glycine-tyrosine-phenylalanine (GYF)-like protein, sensitivity to red light reduced-like (SRR1), and a Tap42-interacting protein (TIP41) in the mTOR signaling pathway.
The most interesting group of genes under positive selection is related to plant defense. Among the genes for signal transduction discussed above are two genes involved in disease resistance. Specifically, ectopic expression of NB-ARC-like protein confers resistance to pathogenic fungi and bacteria in Arabidopsis (Wen et al. 2015), and a protein containing the GYF domain controls the homeostasis of nucleotide-binding leucine-rich repeat (NLR) immune receptors at the level of translational repression in Arabidopsis (Wu et al. 2017). Moreover, other genes related to disease or defense are also under positive selection, including genes encoding mannose-binding lectin (MBL), Kunitz-like protease inhibitor (also known as Kunitz-type trypsin inhibitor, KTI), avrRpt2-induced gene 1 (AIG1), disease resistance protein, cucumber mosaic virus (CMV) 1a interacting protein 1, and tobacco mosaic virus (TMV) resistance protein N-like. MBL was initially reported to be responsible for the recognition of specific carbohydrates on the surfaces of pathogens during infection (Peumans and Van Damme 1995), whereas KTI protects plants by inhibiting the digestive proteases of insects or pathogens (Kim et al. 2009). AIG1, which is induced by infection by bacteria carrying the avirulence gene avrRpt2, likely elicits differential resistance responses based on the type of avirulence gene and its recognition partner (Reuber and Ausubel 1996). In a comparison of wild versus domesticated plants, genes involved in disease resistance or stress responses were also found to be under positive selection in other Asteraceae species (Kane et al. 2011). Moreover, one outstanding trait found in the wild relatives of domesticated species is better disease and pest resistance compared to their domesticated counterparts (Warschefsky et al. 2014). Indeed, C. boreale is more resistant to white rust disease caused by Puccinia horiana than C. morifolium, (Park et al. 2014). Perhaps stress triggered by pathogens has induced the molecular evolution or positive selection of a broad range of genes for disease resistance or defense during domestication or selection by breeders, which has played an important role in the divergence of the two Chrysanthemum species.
Transcriptome sequencing has facilitated the reconstruction of the catalogs of genes in particular species, leading to the identification of genes responsible for specific conditions and helping to track evolutionary events such as WGD, adaptation, and speciation. However, the drawback of RNA sequencing is that the transcripts obtained under-represent the genes in the genome. In addition, the locations of these genes on chromosomes are not identified by this technique. Therefore, even though the present results provide a general overview of WGD, polyploidy, speciation, and adaptation in the Chrysanthemum genus, the molecular evolution of the species investigated must be confirmed at the genomic level. First, WGD or WGT could be detected based on the collinear arrangement of genes in the genome, which could exclude the recurrent birth and death of genes from analysis. Therefore, a Ks peak indicating a very recent WGD and subsequent diploidization in C. boreale could be obtained without interference from random gene duplication. Second, we expect that more genes involved in the adaptation and speciation could be identified at the genome-wide scale. Indeed, lists of genes annotated as having an unknown function include many genes that experienced rapid evolution for a long period of time and whose identities were consequently obscured; these genes might be excluded from analyses based on the transcriptome (Kane et al. 2011). Similarly, in the current study, as many as 39 positively selected genes (36.4%) were classified as encoding hypothetical, uncharacterized, or unknown proteins or were not functionally annotated. C. boreale is currently being subjected to whole-genome sequencing, which is expected to provide a fundamental resource that can be used to accurately reveal the molecular and genomic evolution of the Chrysanthemum genus.
Conclusions
Speciation in Chrysanthemum is associated with hybridization, polyploidization, and adaptation. To reconstruct these evolutionary events and to deduce their time of occurrence, we characterized the transcriptomes of a cultivated Chrysanthemum species, C. morifolium, and its wild relative, C. boreale. These analyses not only confirmed the ancient WGD event shared among members of the Asteraceae family, but they also provided evidence for a recent WGD or WGT event specific to Chrysanthemum. A comparison of the transcriptomes of these species revealed genes that likely experienced positive selection during the processes of domestication and adaptation, which would be responsible for the divergence of the two species. Among the various conditions, biotic stresses likely play important roles in selection and adaption. These transcriptome-based molecular evolutionary analyses of genomes and genes will be addressed using whole-genome sequences in the future.
References
Barker MS, Kane NC, Matvienko M, Kozik A, Michelmore RW, Knapp SJ, Rieseberg LH (2008) Multiple paleopolyploidizations during the evolution of the compositae reveal parallel patterns of duplicate gene retention after millions of years. Mol Biol Evol 25:2445–2455
Barker MS et al (2016) Most compositae (Asteraceae) are descendants of a paleohexaploid and all share a paleotetraploid ancestor with the Calyceraceae. Am J Bot 103:1203–1211
Blanc G, Wolfe KH (2004) Widespread paleopolyploidy in model plant species inferred from age distributions of duplicate genes. Plant Cell 16:1667–1678
Chen J-y (1985) Studies on the origin of Chinese florist’s chrysanthemum. In, 1985. International Society for Horticultural Science (ISHS), Leuven, Belgium, pp 349–362. doi:10.17660/ActaHortic.1985.167.38
Cheng T, Fu B, Wu Y, Long R, Liu C, Xia Q (2015) Transcriptome sequencing and positive selected genes analysis of Bombyx mandarina. PLoS ONE 10:e0122837
Chou H-H, Holmes MH (2001) DNA sequence quality trimming and vector removal. Bioinformatics 17:1093–1104. doi:10.1093/bioinformatics/17.12.1093
Conesa A, Götz S, García-Gómez JM, Terol J, Talón M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21:3674–3676
Dai S, Chen J-Y, Li W-B (1998) Application of RAPD analysis in the study on the origin of Chinese cultivated chrysanthemum. Acta Botanica Sinica 40:1053–1059
Dai SL, Wang WK, Li MX, Xu YX (2005) Phylogenetic relationship of Dendranthema (DC.) Des Moul. revealed by fluorescent in situ hybridization. J Integr Plant Biol 47:783–791
Dowrick GJ (1952) The chromosomes of Chrysanthemum, I: the species. Heredity 6:365–375
Fay JC, Wu C-I (2003) Sequence divergence, functional constraint, and selection in protein evolution. Annu Rev Genomics Hum Genet 4:213–235
Fu ZX, Jiao BH, Nie B, Zhang GJ, Gao TG (2016) A comprehensive generic-level phylogeny of the sunflower family: implications for the systematics of Chinese Asteraceae. J Syst Evol 54:416–437
Haas BJ et al (2013) De novo transcript sequence reconstruction from RNA-Seq: reference generation and analysis with trinity. Natc Protocols. doi:10.1038/nprot.2013.1084
Hou Y, Lin S (2009) Distinct gene number-genome size relationships for eukaryotes and non-eukaryotes: gene content estimation for dinoflagellate genomes. PLoS ONE 4:e6978
Huang C-H, Zhang C, Liu M, Hu Y, Gao T, Qi J, Ma H (2016) Multiple polyploidization events across Asteraceae with two nested events in the early history revealed by nuclear phylogenomics. Mol Biol Evol 33(11):2820–2835
Hwang Y-J et al (2013) Karyomorphological analysis of wild chrysanthemum boreale collected from four natural habitats in Korea. Flower Res J 21:182–189
Initiative TAG (2000) Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408:796–815
Jiao Y et al (2011) Ancestral polyploidy in seed plants and angiosperms. Nature 473:97–100
Kaessmann H (2010) Origins, evolution, and phenotypic impact of new genes. Genome Res 20:1313–1326. doi:10.1101/gr.101386.109
Kane NC, Barker MS, Zhan SH, Rieseberg LH (2011) Molecular evolution across the Asteraceae: micro-and macroevolutionary processes. Mol Biol Evol 28:3225–3235
Kim J-Y, Park S-C, Hwang I, Cheong H, Nah J-W, Hahm K-S, Park Y (2009) Protease inhibitors from plants with antimicrobial activity. Int J Mol Sci 10:2860–2872
Kim C, Wang X, Lee T-H, Jakob K, Lee G-J, Paterson AH (2014) Comparative analysis of Miscanthus and Saccharum reveals a shared whole-genome duplication but different evolutionary fates. Plant Cell 26:2420–2429
Klie M, Schie S, Linde M, Debener T (2014) The type of ploidy of chrysanthemum is not black or white: a comparison of a molecular approach to published cytological methods. Front Plant Sci 5:479. doi:10.3389/fpls.2014.00479
Koch MA, Haubold B, Mitchell-Olds T (2000) Comparative evolutionary analysis of chalcone synthase and alcohol dehydrogenase loci in Arabidopsis, Arabis, and related genera (Brassicaceae). Mol Biol Evol 17:1483–1498
Larkin MA et al (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
Li W, Godzik A (2006) Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22:1658–1659
Li L, Stoeckert CJ, Roos DS (2003) OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res 13:2178–2189
Linder CR, Goertzen LR, Heuvel BV, Francisco-Ortega J, Jansen RK (2000) The complete external transcribed spacer of 18S-26S rDNA: amplification and phylogenetic utility at low taxonomic levels in Asteraceae and closely allied families. Mol Phylogenet Evol 14:285–303
Liu P-L, Wan Q, Guo Y-P, Yang J, Rao G-Y (2012) Phylogeny of the Genus Chrysanthemum L.: evidence from Single-Copy Nuclear Gene and Chloroplast DNA. Sequences. PLoS ONE 7:e48970. doi:10.1371/journal.pone.0048970
Liu S et al (2014) The Brassica oleracea genome reveals the asymmetrical evolution of polyploid genomes. Nat Commun 5
Ma Y-P, Chen M-M, Wei J-X, Zhao L, Liu P-L, Dai S-L, Wen J (2016) Origin of Chrysanthemum cultivars—Evidence from nuclear low-copy LFY gene sequences. Biochem Syst Ecol 65:129–136
MBG (2013) Missouri Botanical Garden. The Plant List Version 1.1 Published on the Internet http://www.theplantlist.org/. Version 1.1 Published on the Internet. http://www.theplantlist.org/. 2017
Meyer E et al (2009) Sequencing and de novo analysis of a coral larval transcriptome using 454 GSFlx. BMC Genomics 10:219–219. doi:10.1186/1471-2164-10-219
Michael TP, Jackson S (2013) The first 50 plant genomes. Plant Genome. doi:10.3835/plantgenome2013.03.0001in
Nei M, Gojobori T (1986) Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol Biol Evol 3:418–426
Panchy N, Lehti-Shiu M, Shiu S-H (2016) Evolution of gene duplication in plants. Plant Physiol 171:2294–2316
Panero JL, Crozier BS (2016) Macroevolutionary dynamics in the early diversification of Asteraceae. Mol Phylogenet Evol 99:116–132
Panero JL, Freire SE, Espinar LA, Crozier BS, Barboza GE, Cantero JJ (2014) Resolution of deep nodes yields an improved backbone phylogeny and a new basal lineage to study early evolution of Asteraceae. Mol Phylogenet Evol 80:43–53
Park SK, Lim JH, Shin HK, Jung JA, Kwon YS, Kim MS, Kim KS (2014) Identification of Chrysanthemum Genetic Resources Resistant to White Rust Caused by Puccinia horiana. Plant Breeding Biotechnol 2:184–193
Paterson AH, Freeling M, Tang H, Wang X (2010) Insights from the comparison of plant genome sequences. Annu Rev Plant Biol 61:349–372
Peumans WJ, Van Damme E (1995) Lectins as plant defense proteins. Plant Physiol 109:347
Reuber TL, Ausubel FM (1996) Isolation of Arabidopsis genes that differentiate between resistance responses mediated by the RPS2 and RPM1 disease resistance genes. Plant Cell 8:241–249
Schlueter JA, Dixon P, Granger C, Grant D, Clark L, Doyle JJ, Shoemaker RC (2004) Mining EST databases to resolve evolutionary events in major crop species. Genome 47:868–876
Soltis PS, Soltis DE (2016) Ancient WGD events as drivers of key innovations in angiosperms. Curr Opin Plant Biol 30:159–165
Soltis DE et al (2009) Polyploidy and angiosperm diversification. Am J Bot 96:336–348
Soltis PS, Marchant DB, Van de Peer Y, Soltis DE (2015) Polyploidy and genome evolution in plants. Curr Opin Genet Dev 35:119–125
Suyama M, Torrents D, Bork P (2006) PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res 34:W609-W612
Tang H, Bowers JE, Wang X, Ming R, Alam M, Paterson AH (2008) Synteny and collinearity in plant genomes. Science 320:486–488
Town CD et al (2006) Comparative genomics of Brassica oleracea and Arabidopsis thaliana reveal gene loss, fragmentation, and dispersal after polyploidy. Plant Cell 18:1348–1359. doi:10.1105/tpc.106.041665
Truco MJ et al (2013) An ultra-high-density, transcript-based, genetic map of lettuce. G3: Genes Genomes Genetics 3:617–631
Van de Peer Y, Fawcett JA, Proost S, Sterck L, Vandepoele K (2009a) The flowering world: a tale of duplications. Trends Plant Sci 14:680–688
Van de Peer Y, Maere S, Meyer A (2009b) The evolutionary significance of ancient genome duplications. Nat Rev Genet 10:725–732
Vanneste K, Maere S, Van de Peer Y (2014) Tangled up in two: a burst of genome duplications at the end of the Cretaceous and the consequences for plant evolution. Philos Trans R Soc B 369:20130353. doi:10.1098/rstb.2013.0353
Warschefsky E, Penmetsa RV, Cook DR, von Wettberg EJB (2014) Back to the wilds: tapping evolutionary adaptations for resilient crops through systematic hybridization with crop wild relatives. Am J Bot 101:1791–1800. doi:10.3732/ajb.1400116
Wen Z, Yao L, Wan R, Li Z, Liu C, Wang X (2015) Ectopic expression in Arabidopsis thaliana of an NB-ARC encoding putative disease resistance gene from wild Chinese vitis pseudoreticulata enhances resistance to phytopathogenic fungi and bacteria. Front Plant Sci 6:1087. doi:10.3389/fpls.2015.01087
Wendel JF, Jackson SA, Meyers BC, Wing RA (2016) Evolution of plant genome architecture. Genome Biol 17:37
Wu Z, Huang S, Zhang X, Wu D, Xia S, Li X (2017) Regulation of plant immune receptor accumulation through translational repression by a glycine-tyrosine-phenylalanine (GYF) domain protein. eLife 6:e23684 doi:10.7554/eLife.23684
Yang Z (1997) PAML: a program package for phylogenetic analysis by maximum likelihood computer applications in the biosciences. CABIOS 13:555–556
Ye J et al (2006) WEGO: a web tool for plotting GO annotations. Nucleic Acids Res 34:W293-W297
Zhou C, Silan D (2001) AFLP analysis of some Dendranthema spp. J Beijing For Univ 24:71–75
Acknowledgements
This work was carried out with the support of the “Cooperative Research Program for Agriculture Science & Technology Development (Project No. PJ010868)” Rural Development Administration, Republic of Korea.
Author information
Authors and Affiliations
Contributions
SYW, S-JK, and T-HL designed the research. S-JK, J-AJ, JSK, S-HK, and S-HS performed the experiments. SYW and T-HL analyzed the data. SYW wrote the manuscript, which was approved by all the other authors.
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Won, S.Y., Kwon, SJ., Lee, TH. et al. Comparative transcriptome analysis reveals whole-genome duplications and gene selection patterns in cultivated and wild Chrysanthemum species. Plant Mol Biol 95, 451–461 (2017). https://doi.org/10.1007/s11103-017-0663-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-017-0663-z