Abstract
Purpose
To provide an overview of fundamental concepts in machine learning (ML), review the literature on ML applications in imaging analysis of pituitary tumors for the last 10 years, and highlight the future directions on potential applications of ML for pituitary tumor patients.
Method
We presented an overview of the fundamental concepts in ML, its various stages used in healthcare, and highlighted the key components typically present in an imaging-based tumor analysis pipeline. A search was conducted across four databases (PubMed, Ovid, Embase, and Google Scholar) to gather research articles from the past 10 years (2009-2019) involving imaging related to pituitary tumor and ML. We grouped the studies by imaging modalities and analyzed the ML tasks in terms of the data inputs, reference standards, methodologies, and limitations.
Results
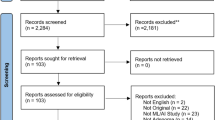
Of the 16 studies included in our analysis, 10 appeared in 2018–2019. Most of the studies utilized retrospective data and followed a semi-automatic ML pipeline. The studies included use of magnetic resonance imaging (MRI), facial photographs, surgical microscopic video, spectrometry, and spectroscopy imaging. The objectives of the studies covered 14 distinct applications and majority of the studies addressed a binary classification problem. Only five of the 11 MRI-based studies had an external validation or a holdout set to test the performance of a final trained model.
Conclusion
Through our concise evaluation and comparison of the studies using the concepts presented, we highlight future directions so that potential ML applications using different imaging modalities can be developed to benefit the clinical care of pituitary tumor patients.





Similar content being viewed by others
Abbreviations
- ML:
-
Machine learning
- AI:
-
Artificial intelligence
- NLP:
-
Natural language processing
- GBM:
-
Glioblastoma multiforme
- PA:
-
Pituitary adenoma
- ACTH:
-
Adrenocorticotrophic hormone
- TSH:
-
Thyrotropin
- OCT:
-
Optical coherence tomography
- CT:
-
Computed tomography
- MRI:
-
Magnetic resonance imaging
- CV:
-
Cross-validation
- LOOCV:
-
Leave-one-out cross validation
- DNN:
-
Deep neural network
References
Shalev-Shwartz S, Ben-David S (2014) Understanding machine learning: from theory to algorithms. Cambridge University Press, Cambridge
Young T, Hazarika D, Poria S, Cambria E (2018) Recent trends in deep learning based natural language processing. IEEE Comput Intell Mag 13:55–75
Voulodimos A, Doulamis N, Doulamis A, Protopapadakis E (2018) Deep learning for computer vision: a brief review. Comput Intell Neurosci 2018:7068349. https://doi.org/10.1155/2018/7068349
Brunetti A, Buongiorno D, Trotta GF, Bevilacqua V (2018) Computer vision and deep learning techniques for pedestrian detection and tracking: a survey. Neurocomputing 300:17–33
Garcia-Garcia A, Orts-Escolano S, Oprea S, Villena-Martinez V, Martinez-Gonzalez P, Garcia-Rodriguez J (2018) A survey on deep learning techniques for image and video semantic segmentation. Appl Soft Comput 70:41–65
Buczak AL, Guven E (2015) A survey of data mining and machine learning methods for cyber security intrusion detection. IEEE Commun Surv Tutor 18:1153–1176
Nguyen G, Dlugolinsky S, Bobák M, Tran V, García ÁL, Heredia I, Malík P, Hluchý L (2019) Machine learning and deep learning frameworks and libraries for large-scale data mining: a survey. Artif Intell Rev 52:77–124
Kim D-H, Kim TJY, Wang X, Kim M, Quan Y-J, Oh JW, Min S-H, Kim H, Bhandari B, Yang I (2018) Smart machining process using machine learning: a review and perspective on machining industry. Int J Precis Eng Manuf Technol 5:555–568
Frutos-Pascual M, Zapirain BG (2015) Review of the use of AI techniques in serious games: decision making and machine learning. IEEE Trans Comput Intell AI Games 9:133–152
Justesen N, Bontrager P, Togelius J, Risi S (2019) Deep learning for video game playing. IEEE Trans Games. https://doi.org/10.1109/TG.2019.2896986
Zhang Q, Yang LT, Chen Z, Li P (2018) A survey on deep learning for big data. Inf Fusion 42:146–157
Esteva A, Robicquet A, Ramsundar B, Kuleshov V, DePristo M, Chou K, Cui C, Corrado G, Thrun S, Dean J (2019) A guide to deep learning in healthcare. Nat Med 25:24
Chen M, Hao Y, Hwang K, Wang L, Wang L (2017) Disease prediction by machine learning over big data from healthcare communities. IEEE Access 5:8869–8879
LeCun Y, Bengio Y, Hinton G (2015) Deep learning. Nature 521:436
Litjens G, Kooi T, Bejnordi BE, Setio AAA, Ciompi F, Ghafoorian M, van der Laak JAWM, van Ginneken B, Sánchez CI (2017) A survey on deep learning in medical image analysis. Med Image Anal 42:60–88. https://doi.org/10.1016/j.media.2017.07.005
Mazurowski MA, Buda M, Saha A, Bashir MR (2019) Deep learning in radiology: an overview of the concepts and a survey of the state of the art with focus on MRI. J Magn Reson Imaging 49:939–954. https://doi.org/10.1002/jmri.26534
Ko J, Swetter SM, Blau HM, Esteva A, Kuprel B, Novoa RA, Thrun S (2017) Dermatologist-level classification of skin cancer with deep neural networks. Nature 542:115
Buda M, Wildman-Tobriner B, Hoang JK, Thayer D, Tessler FN, Middleton WD, Mazurowski MA (2019) Management of thyroid nodules seen on US images: deep learning may match performance of radiologists. Radiology 292(3):695–701. https://doi.org/10.1148/radiol.2019181343
Bychkov D, Linder N, Turkki R, Nordling S, Kovanen PE, Verrill C, Walliander M, Lundin M, Haglund C, Lundin J (2018) Deep learning based tissue analysis predicts outcome in colorectal cancer. Sci Rep 8:3395. https://doi.org/10.1038/s41598-018-21758-3
Lenchik L, Heacock L, Weaver AA, Boutin RD, Cook TS, Itri J, Filippi CG, Gullapalli RP, Lee J, Zagurovskaya M, Retson T, Godwin K, Nicholson J, Narayana PA (2019) Automated segmentation of tissues using CT and MRI: a systematic review. Acad Radiol. https://doi.org/10.1016/j.acra.2019.07.006
Omuro A (2013) Glioblastoma and other malignant gliomas. JAMA 310(17):1842–1850. https://doi.org/10.1001/jama.2013.280319
Melmed S (2011) Pathogenesis of pituitary tumors. Nat Rev Endocrinol 7:257
Kaltsas GA, Nomikos P, Kontogeorgos G, Buchfelder M, Grossman AB (2005) Diagnosis and management of pituitary carcinomas. J Clin Endocrinol Metab 90:3089–3099. https://doi.org/10.1210/jc.2004-2231
Lake MG, Krook LS, Cruz SV (2013) Pituitary adenomas: an overview. Am Fam Physician 88:319–327
Calligaris D, Feldman DR, Norton I, Olubiyi O, Changelian AN, Machaidze R, Vestal ML, Laws ER, Dunn IF, Santagata S, Agar NYR (2015) MALDI mass spectrometry imaging analysis of pituitary adenomas for near-real-time tumor delineation. Proc Natl Acad Sci USA 112:9978–9983. https://doi.org/10.1073/pnas.1423101112
Drummond JB, Ribeiro-Oliveira A, Soares BS (2000) Non-functioning pituitary adenomas. MDText.com Inc., South Dartmouth
Lobatto DJ, Steffens ANV, Najafabadi Z, Andela AH, Pereira CD, van den Hout AM, Peul WB, Vliet WC, Vlieland TPM, Biermasz NR, van Furth WR (2018) Work disability and its determinants in patients with pituitary tumor-related disease. Pituitary 21:593–604. https://doi.org/10.1007/s11102-018-0913-3
Ezzat S, Asa SL, Couldwell WT, Barr CE, Dodge WE, Vance ML, McCutcheon IE (2004) The prevalence of pituitary adenomas: a systematic review. Cancer 101:613–619. https://doi.org/10.1002/cncr.20412
Lara D de, Filho LFSD, Prevedello DM, Otto BA, Carrau RL (2012) Application of image guidance in pituitary surgery. Surg Neurol Int 3:S73. https://doi.org/10.4103/2152-7806.95418
Chen CC, Carter BS, Wang R, Patel KS, Hess C, Bodach ME, Tumialan LM, Oyesiku NM, Patil CG, Litvack Z, Zada G, Aghi MK (2016) Congress of neurological surgeons systematic review and evidence-based guideline on preoperative imaging assessment of patients with suspected nonfunctioning pituitary adenomas. Neurosurgery 79:E524–E526. https://doi.org/10.1227/NEU.0000000000001391
Fathalla H, Cusimano MD, Di Ieva A, Lee J, Alsharif O, Goguen J, Zhang S, Smyth H (2015) Endoscopic versus microscopic approach for surgical treatment of acromegaly. Neurosurg Rev 38:541–549. https://doi.org/10.1007/s10143-015-0613-7
Qaddoura A, Shalung TN, Meier MP, Goguen J, Jing R, Zhang S, Kovacs K, Cusimano MD (2019) Recovery room cortisol predicts long-term glucocorticoid need after transsphenoidal surgery for pituitary tumors. Neurosurgery 84:616–623. https://doi.org/10.1093/neuros/nyy070
Wu V, Cusimano MD, Lee JM (2018) Extent of surgery in endoscopic transsphenoidal skull base approaches and the effects on sinonasal morbidity. Am J Rhinol Allergy 32:52–56. https://doi.org/10.2500/ajra.2018.32.4499
Di Ieva A, Rotondo F, Syro LV, Cusimano MD, Kovacs K (2014) Aggressive pituitary adenomas-diagnosis and emerging treatments. Nat Rev Endocrinol 10:423
Erickson BJ, Korfiatis P, Akkus Z, Kline TL (2017) Machine learning for medical imaging. Radiographics 37:505–515. https://doi.org/10.1148/rg.2017160130
Madabhushi A, Lee G (2016) Image analysis and machine learning in digital pathology: challenges and opportunities. Med Image Anal 33:170–175. https://doi.org/10.1016/J.MEDIA.2016.06.037
Pesapane F, Codari M, Sardanelli F (2018) Artificial intelligence in medical imaging: threat or opportunity? Radiologists again at the forefront of innovation in medicine. Eur Radiol Exp 2:35. https://doi.org/10.1186/s41747-018-0061-6
Rajpurkar P, Irvin J, Zhu K, Yang B, Mehta H, Duan T, Ding D, Bagul A, Langlotz C, Shpanskaya K, et al (2017) CheXNet: radiologist-level pneumonia detection on chest X-rays with deep learning. arXiv1711.05225.
Shickel B, Tighe PJ, Bihorac A, Rashidi P (2017) Deep EHR: a survey of recent advances in deep learning techniques for electronic health record (EHR) analysis. IEEE J Biomed Health Inf 22:1589–1604
Pellegrini E, Ballerini L, Hernandez M, del Chappell CV, González-Castro FM, Anblagan V, Danso D, Muñoz-Maniega S, Job S, Pernet D, Mair C, MacGillivray G, Trucco TJ, Wardlaw E (2018) Machine learning of neuroimaging for assisted diagnosis of cognitive impairment and dementia: a systematic review. Alzheimer Dement Diagn Assess Dis Monit 10:519–535. https://doi.org/10.1016/j.dadm.2018.07.004
Gillies RJ, Kinahan PE, Hricak H (2016) Radiomics: images are more than pictures, they are data. Radiology 278:563–577. https://doi.org/10.1148/radiol.2015151169
Lotan E, Jain R, Razavian N, Fatterpekar GM, Lui YW (2019) State of the art: machine learning applications in glioma imaging. Am J Roentgenol 212:26–37. https://doi.org/10.2214/AJR.18.20218
Zeynalova A, Kocak B, Durmaz ES, Comunoglu N, Ozcan K, Ozcan G, Turk O, Tanriover N, Kocer N, Kizilkilic O, Islak C (2019) Preoperative evaluation of tumour consistency in pituitary macroadenomas: a machine learning-based histogram analysis on conventional T2-weighted MRI. Neuroradiology 61(7):767–774. https://doi.org/10.1007/s00234-019-02211-2
Fan Y, Hua M, Mou A, Wu M, Liu X, Bao X, Wang R, Feng M (2019) Preoperative noninvasive radiomics approach predicts tumor consistency in patients with acromegaly: development and multicenter prospective validation. Front Endocrinol 10:403. https://doi.org/10.3389/fendo.2019.00403
Niu J, Zhang S, Ma S, Diao J, Zhou W, Tian J, Zang Y, Jia W (2019) Preoperative prediction of cavernous sinus invasion by pituitary adenomas using a radiomics method based on magnetic resonance images. Eur Radiol 29(3):1625–1634. https://doi.org/10.1007/s00330-018-5725-3
Zhang S, Song G, Zang Y, Jia J, Wang C, Li C, Tian J, Dong D, Zhang Y (2018) Non-invasive radiomics approach potentially predicts non-functioning pituitary adenomas subtypes before surgery. Eur Radiol 28:3692–3701. https://doi.org/10.1007/s00330-017-5180-6
Kocak B, Durmaz ES, Kadioglu P, Polat Korkmaz O, Comunoglu N, Tanriover N, Kocer N, Islak C, Kizilkilic O (2019) Predicting response to somatostatin analogues in acromegaly: machine learning-based high-dimensional quantitative texture analysis on T2-weighted MRI. Eur Radiol 29:2731–2739. https://doi.org/10.1007/s00330-018-5876-2
Liu Y, Liu X, Hong X, Liu P, Bao X, Yao Y, Xing B, Li Y, Huang Y, Zhu H, Lu L, Wang R, Feng M (2019) Prediction of recurrence after transsphenoidal surgery for Cushing’s disease: the use of machine learning algorithms. Neuroendocrinology 108:201–210. https://doi.org/10.1159/000496753
Lilja Y, Gustafsson O, Ljungberg M, Starck G, Lindblom B, Skoglund T, Bergquist H, Jakobsson K-E, Nilsson D (2017) Visual pathway impairment by pituitary adenomas: quantitative diagnostics by diffusion tensor imaging. J Neurosurg 127:569–579. https://doi.org/10.3171/2016.8.jns161290
van Griethuysen JJM, Fedorov A, Parmar C, Hosny A, Aucoin N, Narayan V, Beets-Tan RGH, Fillion-Robin J-C, Pieper S, Aerts HJWL (2017) Computational radiomics system to decode the radiographic phenotype. Cancer Res 77:e104–e107. https://doi.org/10.1158/0008-5472.CAN-17-0339
Staartjes VE, Serra C, Muscas G, Maldaner N, Akeret K, van Niftrik CHB, Fierstra J, Holzmann D, Regli L (2018) Utility of deep neural networks in predicting gross-total resection after transsphenoidal surgery for pituitary adenoma: a pilot study. Neurosurg Focus 45(5):E12. https://doi.org/10.3171/2018.8.focus18243
Egger J, Freisleben B, Nimsky C, Kapur T (2012) Template-cut: a pattern-based segmentation paradigm. Sci Rep 2:420. https://doi.org/10.1038/srep00420
Staartjes VE, Zattra CM, Akeret K, Maldaner N, Muscas G, van BasNiftrik CH, Fierstra J, Regli L, Serra C (2019) Neural network-based identification of patients at high risk for intraoperative cerebrospinal fluid leaks in endoscopic pituitary surgery. J Neurosurg. https://doi.org/10.3171/2019.4.jns19477
Ugga L, Cuocolo R, Solari D, Guadagno E, D’amico A, Somma T, Cappabianca P, De Caro LDB, Cavallo LM, Brunetti A (2019) Prediction of high proliferative index in pituitary macroadenomas using MRI-based radiomics and machine learning. Neuroradiology 61:1365–1373. https://doi.org/10.1007/s00234-019-02266-1
Schneider HJ, Kosilek RP, Günther M, Roemmler J, Stalla GK, Sievers C, Reincke M, Schopohl J, Würtz RP (2011) A novel approach to the detection of acromegaly: accuracy of diagnosis by automatic face classification. J Clin Endocrinol Metab 96(7):2074–2080. https://doi.org/10.1210/jc.2011-0237
Kong X, Gong S, Su L, Howard N, Kong Y (2018) Automatic detection of acromegaly from facial photographs using machine learning methods. EBioMedicine 27:94–102. https://doi.org/10.1016/j.ebiom.2017.12.015
Li A, Liu W, Cao P, Zheng Y, Bu Z, Zhou T (2017) Endoscopic versus microscopic transsphenoidal surgery in the treatment of pituitary adenoma: a systematic review and meta-analysis. World Neurosurg 101:236–246. https://doi.org/10.1016/j.wneu.2017.01.022
Lalys F, Riffaud L, Morandi X, Jannin P (2010) Automatic phases recognition in pituitary surgeries by microscope images classification. In: Lecture notes in computer science (including subseries lecture notes in artificial intelligence and lecture notes in bioinformatics), pp 34–44
Steiner G, Mackenroth L, Geiger KD, Stelling A, Pinzer T, Uckermann O, Sablinskas V, Schackert G, Koch E, Kirsch M (2012) Label-free differentiation of human pituitary adenomas by FT-IR spectroscopic imaging. Anal Bioanal Chem 403:727–735
Saha A, Yu X, Sahoo D, Mazurowski MA (2017) Effect of MRI scanner parameters on breast cancer radiomics. Expert Syst Appl 87:384–391. https://doi.org/10.1016/j.eswa.2017.06.029
Mackin D, Fave X, Zhang L, Fried D, Yang J, Taylor B, Rodriguez-Rivera E, Dodge C, Jones AK, Court L (2015) Measuring computed tomography scanner variability of radiomics features. Invest Radiol 50:757–765. https://doi.org/10.1097/rli.0000000000000180
Saha A, Harowicz MR, Mazurowski MA (2018) Breast cancer MRI radiomics: an overview of algorithmic features and impact of inter-reader variability in annotating tumors. Med Phys 45(7):3076–3085. https://doi.org/10.1002/mp.12925
Pavic M, Bogowicz M, Würms X, Glatz S, Finazzi T, Riesterer O, Roesch J, Rudofsky L, Friess M, Veit-Haibach P, Huellner M, Opitz I, Weder W, Frauenfelder T, Guckenberger M, Tanadini-Lang S (2018) Influence of inter-observer delineation variability on radiomics stability in different tumor sites. Acta Oncol 57:1070–1074. https://doi.org/10.1080/0284186X.2018.1445283
Paul JS, Plassard AJ, Landman BA, Fabbri D (2017) Deep learning for brain tumor classification. In: Proc. SPIE, vol 10137. https://doi.org/10.1117/12.2254195
Swati ZNK, Zhao Q, Kabir M, Ali F, Ali Z, Ahmed S, Lu J (2019) Brain tumor classification for MR images using transfer learning and fine-tuning. Comput Med Imaging Graph 75:34–46. https://doi.org/10.1016/j.compmedimag.2019.05.001
Deepak S, Ameer PM (2019) Brain tumor classification using deep CNN features via transfer learning. Comput Biol Med 111:103345. https://doi.org/10.1016/j.compbiomed.2019.103345
Davies BM, Carr E, Soh C, Gnanalingham KK (2018) Assessing size of pituitary adenomas: a comparison of qualitative and quantitative methods on MR. Acta Neurochir 158(4):677–683. https://doi.org/10.1007/s00701-015-2699-7
Van Essen M, Sundin A, Krenning EP, Kwekkeboom DJ (2014) Neuroendocrine tumours: the role of imaging for diagnosis and therapy. Nat Rev Endocrinol 10:102
Saha A, Harowicz MR, Grimm LJ, Kim CE, Ghate SV, Walsh R, Mazurowski MA (2018) A machine learning approach to radiogenomics of breast cancer: a study of 922 subjects and 529 DCE-MRI features. Br J Cancer 119:508–516. https://doi.org/10.1038/s41416-018-0185-8
Grossmann P, Stringfield O, El-Hachem N, Bui MM, Rios Velazquez E, Parmar C, Leijenaar RT, Haibe-Kains B, Lambin P, Gillies RJ, Aerts HJ (2017) Defining the biological basis of radiomic phenotypes in lung cancer. Elife 6:e23421. https://doi.org/10.7554/eLife.23421
Smith CP, Czarniecki M, Mehralivand S, Stoyanova R, Choyke PL, Harmon S, Turkbey B (2019) Radiomics and radiogenomics of prostate cancer. Abdom Radiol 44:2021–2029. https://doi.org/10.1007/s00261-018-1660-7
Micko ASG, Wöhrer A, Wolfsberger S, Knosp E (2015) Invasion of the cavernous sinus space in pituitary adenomas: endoscopic verification and its correlation with an MRI-based classification. J Neurosurg 122(4):803–811. https://doi.org/10.3171/2014.12.jns141083
Jacob M, Raverot G, Jouanneau E, Borson-Chazot F, Perrin G, Rabilloud M, Tilikete C, Bernard M, Vighetto A (2009) Predicting visual outcome after treatment of pituitary adenomas with optical coherence tomography. Am J Ophthalmol 147:64-70.e2. https://doi.org/10.1016/j.ajo.2008.07.016
Moon CH, Hwang SC, Kim B-T, Ohn Y-H, Park TK (2011) Visual prognostic value of optical coherence tomography and photopic negative response in chiasmal compression. Investig Opthalmol Vis Sci 52:8527. https://doi.org/10.1167/iovs.11-8034
Buchfelder M, Schlaffer S-M (2016) Intraoperative magnetic resonance imaging for pituitary adenomas. In: Frontiers of hormone research. Karger Publishers, Berlin, pp 121–132
Sylvester PT, Evans JA, Zipfel GJ, Chole RA, Uppaluri R, Haughey BH, Getz AE, Silverstein J, Rich KM, Kim AH, Dacey RG, Chicoine MR (2015) Combined high-field intraoperative magnetic resonance imaging and endoscopy increase extent of resection and progression-free survival for pituitary adenomas. Pituitary 18(1):72–85. https://doi.org/10.1007/s11102-014-0560-2
Ijare OB, Baskin DS, Pichumani K (2019) Ex vivo 1H NMR study of pituitary adenomas to differentiate various immunohistochemical subtypes. Sci Rep 9:3007. https://doi.org/10.1038/s41598-019-38542-6
Menze BH, Jakab A, Bauer S, Kalpathy-Cramer J, Farahani K, Kirby J, Burren Y, Porz N, Slotboom J, Wiest R (2015) The multimodal brain tumor image segmentation benchmark (BRATS). IEEE Trans Med Imaging 34:1993–2024
Luo S, Li X, Li J, Luo SH (2017) Automatic Alzheimer’s disease recognition from MRI data using deep learning method. J Appl Math Phys 5:1892–1898. https://doi.org/10.4236/jamp.2017.59159
Petersen RC, Aisen PS, Beckett LA, Donohue MC, Gamst AC, Harvey DJ, Jack CR Jr, Jagust WJ, Shaw LM, Toga AW, Trojanowski JQ, Weiner MW (2010) Alzheimer’s disease neuroimaging initiative (ADNI): clinical characterization. Neurology 74:201–209. https://doi.org/10.1212/WNL.0b013e3181cb3e25
Cheng J, Huang W, Cao S, Yang R, Yang W, Yun Z, Wang Z, Feng Q (2015) Enhanced performance of brain tumor classification via tumor region augmentation and partition. PLoS ONE 10:e0140381. https://doi.org/10.1371/journal.pone.0140381
Qiao N: A systematic review on machine learning in sellar region diseases: quality and reporting items. Endocr Connect 952–960 (2019). https://doi.org/10.1530/ec-19-0156
Collins GS, Moons KGM (2019) Reporting of artificial intelligence prediction models. Lancet 393:1577–1579. https://doi.org/10.1016/S0140-6736(19)30037-6
He K, Zhang X, Ren S, Sun J (2016) Deep residual learning for image recognition. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp 770–778
Lin T-Y, Maire M, Belongie S, Bourdev L, Girshick R, Hays J, Perona P, Ramanan D, Zitnick CL, Dollár P (2014) Microsoft COCO: common objects in context. Springer, New York
Kaggle K (2019) Competitions. https://www.kaggle.com/competitions
Funding
The financial support provided through the Canadian Institute for Military and Veteran Health Research (CIMVHR) and patient donations (BrainMatters Foundation, the Dodig foundation) and Ryerson University are gratefully acknowledged. The opinions expressed in this work are authors’ and not of those the supporting agencies that provided the funding.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors do not have any other conflict of interest to disclose.
Human and animal rights
This article is a review and does not contain any studies with human participants performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Saha, A., Tso, S., Rabski, J. et al. Machine learning applications in imaging analysis for patients with pituitary tumors: a review of the current literature and future directions. Pituitary 23, 273–293 (2020). https://doi.org/10.1007/s11102-019-01026-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11102-019-01026-x