Abstract
A statistical analysis of the variation in contents with the size of the current known smallest genomes, N. deltocephalinicola, C. ruddii, N. equitans, and M. genitalium, enabled the indication of a minimal set of codons capable of naturally building a modern-type free-living unicellular organism in an early stage of evolution. Using a linear regression model, the potential codon distribution in the minimal natural cell was predicted and compared to the composition of the smallest synthetic, JCVI-Syn3.0. The distribution of the molecular weight of potentially coded amino acids was also calculated. The main differences in the features of the minimal natural cell and H. Sapiens genome were analyzed. In this regard, the content percentage of respective amino acids and their polarization charge properties were reported and compared. The fractions of occurring nucleotides were calculated, too. Then, the estimated numbers of codons in a minimal natural cell were related to the expected numbers for random distribution. Shown increase, or decrease in the contents, relative to the calculated random filling was related to the evolutionary preferences, varying with the subsequent eras of the evolution of genetic code.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The investigation of a minimal set of genes compatible with the natural cellular life has both theoretical and practical branches (Lachance et al. 2019). The theoretical research branch brings us closer to understanding the spontaneous emergence of life (Peretó 2005). Practical approaches focus on the construction of synthetic cells (Clyde et al. 2016). The determination of the minimal set of genes necessary for survival, and so required for basic metabolism and reproduction, can be achieved by the experimental and computational analysis of the respective biochemical pathways (Ogata et al. 1999). This method is burdened with the mistake of not knowing the exact environmental conditions during the passage between abiotic and biotic eras of evolution. A parallel, more general, approach is focused on the analysis of codon usage. This important determinant of gene expression level (Zhipeng et al. 2016), indirectly reflects the level of evolutionary development, availability of nucleotides, preference of amino acids and the energetic potential of cells.
The first fully realized synthetic cell JCVI-Syn3.0 has 473 genes encoded by a 531 kbp genome (Hutchison et al. 2016). Its minimalistic genome size is less than that of its natural archetype, free-living Mycoplasma genitalium (M.g.), with 580 kbp and 525 genes (Sleator 2016), it exceeds the 491 kbp code, coding 552 genes of marine archaea symbiont Nanoarchaeum equitans (NE) (Waters et al. 2003) and the genomes of endosymbionts Carsonella ruddii (C.r.), 160 kbp, 182 genes (Tamame et al. 2007), and Nasuia deltocephalinicola (N.d.), 112 kbp and 137 genes (Bennett and Moran 2013). This list might be completed by a speculated 250 kbp genome coding 500 genes of the Last Universal Common Ancestor (LUCA) (Poole and Logan 2005) and a Forster and Church concrete theoretical design (Glass et al. 2017) of a bottom-up synthetic system, with some of the attributes of a living cell, containing 113 kbp genome coding 151 genes.
Minimal naturally emerged free-living cellular organism of a modern-type (MNC), i.e. conserving genetic code in the form we know today, and using it in the early progenotic translation, or imperfect transcription and replication processes, probably arising on the evolutionary path between the first RNA based, chemoton-like (Gánti 2003) organized system, and LUCA, or the LUCA community (Glansdorff et al. 2008). So, it was the descendant of simple RNA world replicated systems and the ancestor of all modern world organisms.
The codon distribution seems to be a more fundamental property of the early genomes than the minimum number of genes required to sustain life under ideal environmental conditions. This is because the ancient self-organizing system could have as few as two specific catalysts (Cornish-Bowden and Cárdenas 2017), instead of a big stable functional proteome. In general, the change in protein function does not require a change in the codon contents. Thus the proteome seems to be less stable than the codon contents at a given level of metabolic supply of the mass and the energy.
There are the two main goals of the presented work. The first is the disclosure of the potential MNC codon contents free of our beliefs concerning what genes the first cell should have or not, and compare it, its proteinogenic properties, physical connotations, and nucleotide usage, with these in the real contemporary organisms. The second goal is finding the relationships between the MNC codon contents and the evolution of the genetic code. To reach this aim the potential MNC codon distribution is revealed using a statistical method of linear regression (Lane et al. 2007), explaining the dependence between the number of potential codons and the genome size, in the case of the smallest natural genomes. The smallest genomic information is represented by bacteria (Mycoplasma, Carsonella, and Nasuia) and archaea (Nanoarchaeum). In the analytical calculations of the MNC contents, for any of 64 individually analyzed codons, in each case applying the specially fitted formula of the regression model, the read value 113 kbp (Forster and Church 2006) of the investigated genome size was assumed. Repeating, the above "fit, then read" procedure, finally, the MNC codon distribution was outlined, and also the landscape of molecular weight of potential amino acids. Then the estimated frequency of each MNC triplet per thousand codons was compared to the respective usage in the synthetic JCVI-Syn3.0 indicating the specificity of the early stage of evolution. Following the universal code definition, the content percentage of potential MNC amino acids was asigned and the polarization charge were analyzed. The calculated fractions of occurring nucleotides enable the calculation of expected random filling of codons. The increase, or decrease, in the number of some codons relative to the random expectation, indicates the varying evolutionary preference, which was related to the subsequent eras in the model of the smooth evolution (Pawłowski 2019). The main characteristics of the MNC genome were presented compared to the human genome.
Materials and Methods
Potential Codon Contents for Known Smallest Natural Organisms
The numbers of codons for known smallest natural organisms used in the regression analysis also include triplets not assigned to any specific genes, which might be evolutionarily muted. Assuming the same usage of the coding and non-coding triplets, the entire potential codon pool (true codons and non-coding triplets) for a given organism was estimated according to the formula: codon usage x genome size/probe size. Probe size has been usually 3000 bp. The codon usage was taken from the well-known genomic database, CMBL at the University of Georgia (http://www.cmbl.uga.edu/software/codon_usage.html) for Ca. N.deltocephalinicola NAS ALF uid214084 (102,246 bp), Codon Usage Database (Kazusa) (http://www.kazusa.or.jp/codon/) for Ca. C. ruddii PV (150,315 bp) and N. equitans Kin4-M (456,327 bp), and STD Sequence Databases (Los Alamos NL) (http://stdgen.northwestern.edu/stdgen/ bacteria/mgen/properties.html) for M. genitalium (529,986 bp). Analyzed coding sequences on average account for 92% of all genome size, which respectively equals 112,091 bp (N.d.), 159,662 bp (C.r), 490,885 bp (N.e.), and 580,070 bp (M.g.), therefore the 8% of all estimated triplets probably do not have contemporary proteinogenic meaning.
Potential Codon Contents for MNC
The number of MNC potential codons of a given type was calculated using linear regression models of dependence between the number of codons (nc) and the genome size (gs), nc = par1 × gs + par2. The parameters par1 and par2 were attributed due to the standard statistical procedure of simple regression, applied specifically to each one-name codon in the set of smallest natural genomes. An extra point (0,0) has been added to the data, to approximately including pre-biotic self-replicating systems. The size of the wanted hypothetical MNC genome was taken as equal to 113 kbps. An example of the above procedure for the codon AUG is presented in Fig. 1. This procedure was repeated 64 times, for each nucleotide triplet. The distribution of the whole MNC genome contents was analyzed for the separate codons and the mass groups of potentially coded amino acids.
Determination of the number of AUG codons in MNC. Linear regression model of the dependence between the number of codons (nc) and the genome size (gs) was fitted to the AUG contents in the smallest natural genomes N.d., C.r., N.e., and M.g. Point (0,0) was extra included (see text). Obtained equation of the line, nc = 0.0054 × gs-138.35, analytically describing modeled dependence with a coefficient of determination, R2 = 0.9912, led to estimation the number of AUG in MNC, as nc (113 × 10+3) = 474,06,854
Statistical Comparison
For statistical comparison, the codon usage for JCVI-Syn3.0 (531,490 bp) (https://www.ncbi.nlm.nih.gov/nuccore/1009427419) was estimated with the Sequence Manipulation Suite (https://www.bioinformatics.org/ sms2/codon_usage.html) and for H. sapiens (93,487 genes, 40,662,582 codons) with the Kazusa (http://www.kazusa.or.jp/codon/). The comparison of the content percentage of the potential MNC and H. sapiens amino acids was done according to the universal genetic code rules.The comparison of amino acid polarization charge properties was done according to the following classification: hydrophobic (Ala, Ile, Leu, Phe, Val, Pro, Gly), polar (Gln, Asn, His, Ser, Thr, Cys), charged positive (Lys, Arg), charged negative (Asp, Glu), and amphipathic (Met, Tyr, Trp).
Nucleotides
The fractions of MNC nucleotides were calculated summing up each nucleotide in all identified codons of the genome, then dividing by the sum of all nucleotides.
MNC Estimated/Randomly Expected Codon Numbers
The randomly expected number of MNC codons was calculated according to the formula: N × f1 × f2 × f3, where N is the total number of MNC codons, and fi is the fraction of i-th nucleotide. The ratio of the estimated and the randomly expected number of codons, for the whole MNC genome, were analyzed.
Evolutionary Preferences
In the evolutionary preference analysis potential amino acids of MNC were chronologically grouped in the era classes of the emergence of contemporary code according to the model of the smooth evolution. The eras, episodes, and code emergence are summarized in Table 1.
For each set of the codons belonging to a given evolutionary class, it was calculated the mean number of the emergence episode, and the mean ratio of the MNC estimated, and the randomly expected number of codons.
Results
To reveal the potential codon distribution of the hypothetical minimal natural cell, MNC, the estimated number of each triplet was obtained due to the repetition of the linear regression-based procedure, described in Materials and Methods. The whole MNC codon distribution is presented in Fig. 2. The codons are ordered first according to the weight of potentially coded amino acids (universal code), then on subsequent positions in triplet, according to the decreasing nucleotide strength (GCAU) and the decreasing number of carbon–nitrogen rings (purines before pyrimidines). The result of the above estimation gives the total number of MNC codons = 37,660.9, and the estimated genome size = 112,982.7 bp.
The whole MNC codon contents distribution (37,660.9 codons) was obtained due to the repetition of the linear regression-based procedure described in Materials and Methods. Height nc of the vertical bar represents the number of a given codon. Colors indicate different potentially coded amino acids ordered with the increased weight. Internal order takes into account subsequently the positions of nucleotide in triplet, the decreasing nucleotide strength (GCAU), and the decreasing number of nucleotide carbon–nitrogen rings (purines before pyrimidines)
To indicate the most preferred size of potentially coded amino acids the potential MNC codons grouped in the mass groups are presented in Fig. 3, showing the possible domination of coded the average-weight (125–150 Da) molecules.
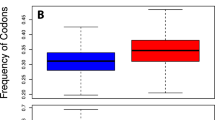
To compare the first natural and the first artificial cell, MNC, and JCVI-Syn3.0, the codon usage per 1000 codons is presented in Fig. 4. The codon order is as in Fig. 2.
MNC (blue) and JCVI-Syn3.0 (red) codon usage per 1000 codons. The codon order is as in Fig. 2
The comparison of the first cell and H. sapiens cell is presented in Figs. 5, 6, and 7, giving insight into the directions of evolution. In Fig. 5, the percentage of the content of the potential MNC and H. sapiens amino acids is presented.
In Fig. 6, the percentage content of the potential MNC and H. sapiens amino acid polarization charge properties is presented. There were hydrophobic (Ala, Ile, Leu, Phe, Val, Pro, Gly), polar (Gln, Asn, His, Ser, Thr, Cys), charged positive (Lys, Arg), charged negative (Asp, Glu), and amphipathic (Trp, Tyr, Met) amino acids, and the stop codons, grouped.
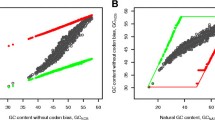
In Fig. 7, the fractions of MNC and H. sapiens nucleotides are presented to analyze the usage of the basic RNA codon bricks A, U, G, C.
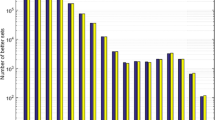
Using MNC data from Fig. 7, the ratio of the MNC estimated and the randomly expected number of codons, for the whole MNC genome is presented in Fig. 8. The codon order is as in Fig. 2.
To show evolutionary preferences, the chronological variation in the mean ratio of the MNC estimated and the randomly expected number of codons in the predefined evolutionary classes (Table 1) is presented in Fig. 9.
The chronological variation in the mean ratio (y) of the MNC estimated and the randomly expected number of codons for the different classes of the evolution eras (x), as in Tab.1. Bars indicate standard deviation. Continuous line—the power-law trend. A considerable preference of the codons for the lightweight amino acids probably defined in the early eras of the evolution of the code is visible on the chart
Discussion
In the above paper, as the main goal, the distribution of minimal natural cell potential codons was revealed. Considering Mat and co. (Mat et al., 2008), the identified genome of LUCA (424 protein-encoding genes) based on the genes common to some Euryarchaea, Crenarchaea, and methanogens highly exceed the 150–340 genes estimated to be present in a minimal proteome compatible with life. Thus, LUCA was the first modern organism, but not a minimal one (Mat et al. 2008).
Determination of a minimal set of codons capable to naturally build a modern-type free-living unicellular organism in an early stage of evolution is a difficult task. The main problem is the uncertainty of natural conditions and gene requirements. One may only expect that the MNC genome is to some extent similar but smaller than the genomes of the current minimal free-living organisms. We have further assumed that the MNC genome might be not fully functional, as the system of Forster and Church (Glass et al. 2017), and to a certain extent had been similar to the contemporary endosymbionts, but where the role of the host cell had played a stable organic-rich environment. We have also presented an idea that the contents of the early genomes are covariant with the genome size, which allowed us to apply a simple regression method in the estimation of the MNC codon contents (Fig. 1).
This way of thinking lets us estimate the potential codon distribution of a whole MNC genome. Here, we used the RNA representation of the code, presumably the only one in the early forms of life (Alberts et al. 2002). Even though 92% of discussed triplets, currently code proteins of the known smallest cells, many of them might have the other meaning in the early forms of life. This is why we call them potential codons. The obtained picture (Fig. 2) shows the domination of potential codons with a high content of the nucleotides A and U, especially those potentially coding amino acids of a medium weight, from aspartate, pyruvate, and aromatic family. The highest contents exhibit triplet AAA (Lys). The codons related to the lighter amino acids (Gly,… Cys), exhibit moderate and relatively constant occurrence levels. On the other side, the codons defining heavier amino acids (Ile,… Trp) and stop codons show a more heterogeneous distribution. The distribution in the mass groups (Fig. 3) confirms domination of medium weight amino acid-related codons and reveals approximate symmetry of content in the side weight areas.
The predicted MNC genome exhibits higher usage of dominating codons and the Gly family than artificial cell JCVI-Syn3.0 (Fig. 4). The synthetic cell has more stop codons than NMC.
The MNC has fewer codons potentially coding small amino acids than H. sapiens cells (Fig. 5). It does not influence the aa electric charge properties very much (Fig. 6), preferring hydrophobicity.
The weak nucleotides represented by bases A, U dominate strong G, and C, in MNC cell. Purines prevail over pyrimidinecbs. In H. sapiens, the distribution is more homogenous (Fig. 7).
The predicted number of MNC codons differs from the randomly expected (Fig. 8). It exceeds more than twice the expected value for potentially the very light amino acid codons (Gly, Ala, Pro), and falls below expectation for the class representing very heavy amino acids (Arg, Tyr, Trp) and stop codons.
The above remarks are consistent with the vision presented in the model of the smooth evolution (Tab.1) (Pawłowski 2019), describing the emergence and development of a universal genetic code. It presents the shortest possible route obeying the simplest acceptable modifications of the evolving coding system, gradually increasing its complexity, i.e., from the first codon, and first coded amino acid, towards the current universal code with 64-codons, coding twenty proteinogenic bricks. This route approximates a certain universal bio-physicochemical attractor, common for all protoorganisms in the initial phase of the evolution of life. As a history, it can be divided into the 22 chronologically consecutive episodes, essential for the development of a code in the form we know. Logically arranged episodes obey simultaneous processes, starting approximately at the same time, and running in the same period at a comparable rate, interrelated or not, sometimes competing. They mainly include the first definition and further redefinition of amino acid for a given codon, or the code expansion for a given amino acid, which led the emergence both of new coded amino acids, and used codons, and specified the dynamic constraints for the evolutionary pathways in the following episodes. Generally, one type of amino acid is introduced in a given episode, or amino acids of the same mass. The order of the emergence of new amino acids, and newly defined codons, in the subsequent episodes, as well as the type of processes taking place, were regulated by the evolutionary dominating precedence rules (Pawłowski 2019), taking into account, sequentially, the amino acid masses, the metabolic neighbourhood, the type of mutations, and others.
The above quoted model places the initial phase of the evolution of the genetic code in the hot era of the RNA world, dominated by self-replicating molecules of RNA, possessing some catalytic properties. They are mainly built of weak bonding light nucleotides A, and U, inducing the collective ephemeral interactions between the approaching RNA strands, or within a single RNA molecule. It can lead to the formation of the dynamic RNA sol–gel inter-phase areas, being natural reactors where the first biochemical reactions can be catalyzed. The dominant primary structure of the ancient RNA is rich in poly-A and poly-U regions. It also contains some amount of bases G, and C. They can form the first self-replicating prototRNA, and aminoacyl-tRNA synthetase protoribozyme (Pawłowski 2019).
Seen here as a result, the domination of A and U rich codons, according to the mentioned model, may be a manifestation of the heritage of the RNA-world, with the excess of weak nucleotides organized in poly-A or poly-U RNAs. Consequently, the moderate level of codons related to small amino acids may answer to the small contents of the strongly bonding nucleotides, G and C, occurring at least once at first, or the second, position of triplet codons organized in the early episodes of the evolution of genetic code. In further cooler eras, possible doublets of numerous weak nucleotides make distribution more heterogeneous. Overall domination of the medium-weight amino acids is a result of both the mentioned environmental changes in the Proto-Code Pre-Ribosomal Moderating Era and the creation of the proto-ribosome in the next era. In the first synthetic cell JCVI-Syn3.0 the Gly and mean weight aa codons seem to be relatively less important than in MNC. Likely, the bigger proteome requires other coding possibilities and more stop codons. As seen in a comparison of MNC and H. sapiens the evolution increased the role of small amino acids, not changing overall electro-physical properties in which hydrophobicity seems to be the key. Impressed in MNC, the domination of the nucleotides A and U, during the formation of a universal genetic code, later was evolutionarily damped (e.g. see the percentage of the Lys and Ile in H. sapiens, Fig. 5).
The revealed nonrandom occurrence of the strong bonding codons for the light amino acids may be a manifestation of the specific demand for the strong bonding elements determining the structure of RNA strands and ribozymes, present in the initial episodes of the evolution of life. The proposed evolutionary preference analysis, considering the chronological variation in the mean ratio of the MNC estimated and the randomly expected number of codons, in the predefined evolutionary classes, indicates clear relationships between the amino acid contents level and the date of the evolutionary era of a code emergence. The eras had been dated with the mean episode number (Fig. 9). The preference of the initially defined codons, reaching the value of about 3, chronologically decreases, likely due to deficiency of important nucleotides G and C. The exception is the slight increase in the fourth era, during the period of proto-ribosome formation and the significant usage of the nucleotides A, and U. Finally the preference falls below the value level 1 in the last era, when the codons for heavy amino acids had been defined. A power law trend, typical for scale-free phenomena (Barabási and Bonabeau 2003) was finally fitted, well reflecting observed relationships. It means that the shape of observed regularity depends less on the scale adopted on the axis describing the chronology, and indicates that the evolutionary stages followed are more important than their exact duration.
A supplementary analysis of the codon contents distribution in the mass groups (Fig. 3), was performed. It was proposed that this draft may reveal the rate of the codon incorporation into the MNC genome, for subsequent amino acids, during the evolution of the code following the weight gain. The mass groups characterized by the successive multipliers 1, 2, 3… of the hypothetical period T, consumed for incorporation of a given mass group (25 Da wide), were reported in Fig. 10. At so introduced an analog of time, as a result, there are confirmed the two characteristic exponential trends in the variation in the rate of the codon number change. The first is the accelerating increase (R2 = 0.99), then, for 3 T, in the middle of the considered range, the slowing down decrease (R2 = 0.98) starts and progressing till 6 T.
The analysis of trends of the variation in the number of MNC codons (nc) with the approximate time (t), replacing mass groups in Fig. 3. The unit T—hypothetical constant period, required for the evolutionary definition of the group of codons coding amino acids of a mass within the range of 25 DA
In light of the above, the following simple description of the kinetics of codons incorporation into the MNC genome was proposed and worked out mathematically (Appendix). With a constant assumption that codons dedicated to a given amino acid originate chronologically with time according to the increasing mass (as it is stated in the smooth evolution model), it is reasonable to formulate further approximate qualitative conclusions. First, the rate of change in the number of codons initially increases, proportionally to the actual total codon number. Secondly, above a certain total number of codons, this rate slows down proportionally to the further codon number increase. That means, the evolution may not be linear in time, and the rate of new codons incorporation may not be constant. The abrupt passage, acceleration-deceleration, in the observed kinetics, can be assigned to the early Proto-Code Ribosomal Maturation Era. The nature of it may lie in the lack of the energy efficiency of the MNC or the insufficient support of nucleotides.
The exponential trends analyzed mathematically in the Appendix, are related to the initial, N0, and critical, Np, genome size, the relative rate of the genome increase, a, and the damping constant, b. At the approximate assumption that the passage time tp = 2.5, it can be estimated N0 = 779 codons, Np = 33,526 codons, a = 1.07 [1/T] and b = 1,33 [1/T]. Applied continuous formalism may surprise, but it is a natural approximation consistent with the smooth evolution model, postulating small changes during the evolution of genetic code. Mentioned 779 codons could be the non-coding kernel of the following MNC. In the first phase, showing acceleration, the environmental support in the energy and simple molecules could be enough, but not in the second phase of deceleration. Then, also the internal metabolism might not be effective enough. This could be the most important challenge of the early evolution.
In general, the distribution of MNC codons supports the prediction of the model of smooth evolution. Doubts may arise regarding the assumed size of the genome. It was indicated in the separate simulations that the absolute bottom-up limit is 96 kbp, under which the regression analysis gives the negative numbers of some codons. The increase in the assumed genome size 113 kbp of 1%, increases AU rich codons (AAA and UUU) about 2%, and GC rich codons (GGG and CCC) about 0.5%.
Statistical analysis of the MNC genome suggests that it is built fundamentally with the weak bases, A and U, encoding mainly the six amino acids and the stop codon according to the two-letter code. So, it is likely that the MNC was the last representative of the “AU world” with a deficit of G and C bases, being the final effect of the further evolution when the transitions A → G and U → C dominated other types of mutations. We think that the trace of the evolution of this type was conserved in the slowly evolving genome of Ca. C. ruddii.
The hypothetical MNC probably originated from a self-replicating AU-RNA chromosome with a high encoding density, which possessed some enzymatic properties. After the eras of incorporation of G and C into essential code positions (Pre-Code Hot Era and Proto-Code Pre-Ribosomal Warm Era), in the next eras, the ribosome, the universal genetic code, and the protein enzymatic system with the mechanism of self-splicing there were probably arisen. This leads to the origination of DNA and the other elements of modern organisms.
The above picture may raise some important questions. For example, why the nucleotides G and C were discriminated in prebiotic evolution? It could be because cytosine is harder to synthesize under plausible prebiotic conditions, and unstable (Shapiro 1999). Furthermore, guanine can fold up RNA in such a way that it cannot be an effective template for replication (Joyce and Orgel 2006).
Why early biotic evolution leading to MNC, and further, to the modern organisms, recognize the importance of discriminated bases? Why the transitions A → G and U → C became the most dominant type of mutations? Among other causes, it could be a result of changing thermal conditions (Kawamura et al. 2005). The early domination of AAA codons may be also related to the deficiency in the abiotically synthesized lysine (Orgel and Miller 1974). Together with a slight insufficiency in U, it suggests that the initial leading strand was a descendant of a polymer of A. So, one of the important goals of early biotic evolution might have been the decrease in this multi-A codon content bias. By that measure, tryptophan codon UGG should not be extremely favored by the evolution.
The premises of the analysis are that an evolutionary process links the codon composition of the smallest available genomes. This is a brave assumption, which is confirmed by the species used for the analysis existing in very different environmental conditions but falling the same linear regression. Of course, the obtained minimum set is a theoretical prediction for which no experimental validation exists. It may be treated as the next elaborated hypothesis. The presented example of the correlation between codon content and genome size happens to be AUG, the codon for methionine (Fig. 1). This one-box codon shows a good correlation with genome size. As in this minimal genome size is essentially equivalent to gene number, and, typically, genes tend to have one initiating methionine, it may indicate that the number of genes correlates with the genome size.
At first look, surprisingly, the comparison of the results with the human genome, not with the organisms “closer” to LUCA like primitive Bacteria, Archaea, and Eukaryota, shows moderate changes which indicate that evolution cannot be considered as a gradual, random train of extreme accidents. Analyzing the results, we can also assume that the early natural environment might be poor in some abiotically synthetic amino acids (Lys). Although it seems to be controversial, the attempt to link obtained MNC data to the phases of evolution described in the model of The Smooth Evolution of the Universal Genetic Code offers an impressive picture of decreasing evolutionary pressure favoring over-usage of strong nucleotides at the beginning of the codon evolution. Additionally revealed mass-dependent kinetics seems to validate this kind of thinking. According to the author, this is the best candidate to reasonably explain presented correlations, among other RNA-world hypotheses (Higgs 2009; Sengupta, and Higgs 2015; Lei and Burton 2020). It should be underlined that the GC-rich origin in the genetic code was a subject in some papers (Hartman and Smith 2019) from the metabolic first concept. If to compare Table 1 with chronological events of the genetic code based on the RNA-peptide world and GC codon-content with a more detailed mechanism (Gospodinov and Kunnev 2020) there are some similarities and differences (i.e. Arg, Val), but most agree.
In this paper, the two major goals were addressed to find potential codon contents for some of the smallest genomes and to find the relationships between these codons and the evolution of the genetic code. The minimum set used is a theoretical prediction, extrapolation to the region for which no experimental validation exists. We are sure that these and other presented questions require further more detailed investigations, especially, theoretically postulated relation between the evolution and metabolism (Wong 1975), clearly signaled in the discussion of the results in Fig. 10. We believe that they will come and allow the presented results, and also the smooth evolution model, to be further developed.
References
Alberts B et al (2002) Molecular biology of the cell. The RNA World and the Origins of Life, 4th edn. Garland Science, New York
Barabási AL, Bonabeau E (2003) Scale-free networks (PDF). Sci Am 288(5):50–59. https://doi.org/10.1038/scientificamerican0503-60
Bennett GM, Moran NA (2013) Small, smaller, smallest: the origins and evolution of ancient dual symbioses in a Phloem-feeding insect. Genome Biol Evol 5(9):1675–1688. https://doi.org/10.1093/gbe/evt118
Clyde A, Hutchison III et al (2016) Design and synthesis of a minimal bacterial genome. Science 351:6280. https://doi.org/10.1126/science.aad6253
Cornish-Bowden A, Cárdenas ML (2017) Life before LUCA. J Theor Biol 434:68–74. https://doi.org/10.1016/j.jtbi.2017.05.023
Forster AC, Church GM (2006) Towards synthesis of a minimal cell. Mol Syst Biol 2:45. https://doi.org/10.1038/msb4100090
Gánti T (2003) Chemoton theory: theory of living systems. Kluwer Academic, New York
Glansdorff N, Xu Y, Labedan B (2008) The last universal common ancestor: emergence, constitution and genetic legacy of an elusive forerunner. Biol Direct 3:29. https://doi.org/10.1186/1745-6150-3-29
Glass JI, Merryman C, Wise KS, Hutchison CA 3rd, Smith HO (2017) Minimal cells-real and imagined. Cold Spring Harb Perspect Biol 9(12):023861. https://doi.org/10.1101/cshperspect.a023861
Gospodinov A, Kunnev D (2020) Universal codons with enrichment from GC to AU nucleotide composition reveal a chronological assignment from early to late along with LUCA formation. Life (basel, Switzerland). 10:6. https://doi.org/10.3390/life10060081
Hartman H, Smith TF (2019) Origin of the genetic code is found at the transition between a Thioester World of Peptides and the Phosphoester World of Polynucleotides. Life (basel) 9(3):69. https://doi.org/10.3390/life9030069
Higgs PG (2009) A four-column theory for the origin of the genetic code: tracing the evolutionary pathways that gave rise to an optimized code. Biol Direct 4:16. https://doi.org/10.1186/1745-6150-4-16
Hutchison CA et al (2016) Design and synthesis of a minimal bacterial genome. Science 351:6253
Joyce GF, Orgel LE (2006) Progress toward understanding the origin of the RNA world. In: Gesteland RF et al (eds) The RNA world. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, pp 23–568
Kawamura K, Nagahama M, Kuranoue K (2005) Chemical evolution of RNA under hydrothermal conditions and the role of thermal copolymers of amino acids for the prebiotic degradation and formation of RNA. Adv Space Res 35(9):1626–1633
Lachance JC, Rodrigue S, Palsson BO (2019) Minimal cells, maximal knowledge. Elife 8:e45379. https://doi.org/10.7554/eLife.45379
Lane DM et al (2007) Online statistics education: a multimedia course of study. http://onlinestatbook.com/. Rice University
Lei L, Burton ZF (2020) Evolution of life on earth: tRNA, aminoacyl-tRNA synthetases and the genetic code. Life (basel) 10(3):21. https://doi.org/10.3390/life10030021
Mat WK, Xue H, Wong JT (2008) The genomics of LUCA. Front Biosci 1(13):5605–5613
Ogata H, Goto S, Sato K, Fujibuchi W, Bono H, Kanehisa M (1999) KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res 27(1):29–34. https://doi.org/10.1093/nar/27.1.29
Orgel LE, Miller SL (1974) The origins of life on the earth. p 87
Pawłowski PH (2019) The smooth evolution of the universal genetic code (main episodes). Int J Sci 09:28–51. https://doi.org/10.18483/ijSci.2180
Peretó J (2005) Controversies on the origin of life. Int Microbiol 8(1):23–31
Poole AM, Logan DT (2005) Modern mRNA proofreading and repair: clues that the last universal common ancestor possessed an RNA genome? Mol Biol Evol 22(6):1444–1455. https://doi.org/10.1093/molbev/msi132
Sengupta S, Higgs PG (2015) Pathways of genetic code evolution in ancient and modern organisms. J Mol Evol 80:229–243. https://doi.org/10.1007/s00239-015-9686-8
Shapiro R (1999) Prebiotic cytosine synthesis: a critical analysis and implications for the origin of life. Proc Natl Acad Soc USA 96(8):4396–4401. https://doi.org/10.1073/pnas.96.8.4396
Sleator RD (2016) JCVI-syn3.0-a synthetic genome stripped bare! Bioengineered 7(2):53–56. https://doi.org/10.1080/21655979.2016.1175847
Tamames J, Gil R, Latorre A, Peretó J, Silva FJ, Moya A (2007) The frontier between cell and organelle: genome analysis of Candidatus Carsonella ruddii. BMC Evol Biol 7:181. https://doi.org/10.1186/1471-2148-7-181
Waters E, Hohn MJ, Ahel I et al (2003) The genome of Nanoarchaeum equitans: insights into early archaeal evolution and derived parasitism. Proc Natl Acad Sci USA 100(22):12984–12988. https://doi.org/10.1073/pnas.1735403100
Wong JT (1975) (1975) A co-evolution theory of the genetic code. Proc Natl Acad Sci USA 72(5):1909–1912. https://doi.org/10.1073/pnas.72.5.1909
Zhipeng Z, Yunkun D, Mian Z, Lin L, Chien-hung Y, Jingjing F, She C, Yi L (2016) Codon usage is an important determinant of gene expression levels largely through its effects on transcription. Proc Natl Acad Sci USA 113(41):E6117-6125. https://doi.org/10.1073/pnas.1606724113
Acknowledgements
The authors gratefully acknowledge prof. Marcin Grynberg from IBBPAS for critical and very fruitful support during the preparation of this article.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendix
Appendix
Let’s assume that incorporation of a new codon to the MNC genome has two stages of different kinetics types, depending on the total number of codons N, where Np is the N critical value determining the passage between the stages. In the first stage, for N↕ Np, the new codons are incorporated with the rate (dN/dt) proportional to the genome size, i.e.
where t is time and a is the relative rate of the genome increase.
In the second stage, for N > Np the rate decreases linearly with the further increase in N, i.e.
where b is damping constant.
The resolution of the ordinary differential equations, Eqs. 1 and 2, are
where N0 initial MNC genome size and passage time tp = (1/a) × ln(Np/N0).
Then, the rate of genome increase can be described by
In the following of the above, the number of codons incorporated in the period T can be approximately described as
where
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pawłowski, P.H. The Codon Usage in the Minimal Natural Cell. Orig Life Evol Biosph 51, 215–230 (2021). https://doi.org/10.1007/s11084-021-09616-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11084-021-09616-1