Abstract
Genetic profiling is an increasingly useful tool for sub-classification of gliomas in adults and children. Specific gene mutations, structural rearrangements, DNA methylation patterns, and gene expression profiles are now recognized to define molecular subgroups of gliomas that arise in distinct anatomic locations and patient age groups, and also provide a better prediction of clinical outcomes for glioma patients compared to histologic assessment alone. Understanding the role of these distinctive genetic alterations in gliomagenesis is also important for the development of potential targeted therapeutic interventions. Mutations including K27M and G34R/V that affect critical amino acids within the N-terminal tail of the histone H3 variants, H3.3 and H3.1 (encoded by H3F3A and HIST1H3B genes), are prime examples of mutations in diffuse gliomas with characteristic clinical associations that can help diagnostic classification and guide effective patient management. These histone H3 mutations frequently co-occur with inactivating mutations in ATRX in association with alternative lengthening of telomeres. Telomere length can also be maintained through upregulation of telomerase reverse transcriptase (TERT) expression driven by mutation within the TERT gene promoter region, an alteration most commonly found in oligodendrogliomas and primary glioblastomas arising in adults. Interestingly, the genetic alterations perturbing histone and telomere function in pediatric gliomas tend to be different from those present in adult tumors. We present a review of these mutations affecting the histone code and telomere length, highlighting their importance in prognosis and as targets for novel therapeutics in the treatment of diffuse gliomas.



Similar content being viewed by others
References
Schwartzentruber J et al (2012) Driver mutations in histone H3.3 and chromatin remodeling genes in pediatric glioblastoma. Nature 482:226–231
Chi P, Allis CD, Wand GG (2010) Covalent histone modifications: miswritten, misinterpreted and mis-erased in human cancers. Nat Rev Cancer 10:457–469
Lee W et al (2014) PRC2 is recurrently inactivated through EED or SUZ12 loss in malignant peripheral nerve sheath tumors. Nat Genet 46:1227–1232
Buczkowicz P et al (2015) Pathology, Molecular genetics, and epigenetics of diffuse intrinsic pontine glioma. Front Oncol 5:1–9
Albig W et al (1995) The human replacement histone H3.3B gene (H3F3B). Genomics 30:264–272
Szender E, Ray-Gallet D, Almouzni G (2011) The double face of the histone variant H3.3. Cell Res 21:421–434
Kallappagoudar S et al (2015) Histone H3 mutations: a special role for H3.3 in tumorigenesis? Chromosoma 124:177–189
Lewis P et al (2010) Daxx is an H3.3 specific histone chaperone and cooperates with ATRX in replication-independent chromatin assembly at telomeres. Proc Natl Acad Sci USA 107:14075–14080
Goldberg AD et al (2010) Distinct factors control histone variant H3.3 localization at specific genomic regions. Cell 140:678–691
Drane P et al (2010) The death-associated protein DAXX is a novel histone chaperone involved in the replication-independent deposition of H3.3. Genes Dev 24:1253–1265
Blackburn EH et al (2015) Human telomere biology: a contributory and interactive factor in aging, disease risks, and protection. Science 350:1193–1198
Giardini M et al (2014) Telomere and telomerase biology. Prog Mol Biol Transl Sci 125:1–40
Conomos D et al (2013) Alternative lengthening of telomeres: remodeling the telomere architecture. Front Oncol 3:27
Kim N et al (1994) Specific association of human telomerase activity with immortal cells and cancer. Science 266:2011–2015
Killela P et al (2013) TERT promoter mutations occur frequently in gliomas and a subset of tumors derived from cells with low rates of self-renewal. Proc Natl Acad Sci USA 110:6021–6026
Huang FW et al (2013) Highly recurrent TERT promoter mutations in human melanoma. Science 339:957–959
Horn S et al (2013) TERT promoter mutations in familial and sporadic melanoma. Science 339:959–961
Bell RJ et al (2015) The transcription factor GABP selectively binds and activates the mutant TERT promoter in cancer. Science 348:1036–1039
Flynn R et al (2015) Alternative lengthening of telomeres renders cancer cells hypersensitive to ATR inhibitors. Science 347:273–277
Lovejoy C et al (2012) Loss of ATRX, genome instability, and an altered DNA damage response are hallmarks of the alternative lengthening of telomeres pathway. PLoS Genet 8:e1002772
Heaphy C et al (2011) Altered telomeres in tumors with ATRX and DAXX mutations. Science 333:425
He LJ et al (2012) Prognostic significance of overexpression of EZH2 and H3k27me3 proteins in gastric cancer. Asian Pac J Cancer Prev 13:3173–3178
Bae WK et al (2015) The methyltransferase EZH2 is not required for mammary cancer development, although high EZH2 and low H3K27me3 correlate with poor prognosis of ER-positive breast cancers. Mol Carcinog 54:1172–1180
He WP et al (2015) Decreased expression of H3K27me3 in human ovarian carcinomas correlates with more aggressive tumor behavior and poor patient survival. Neoplasma 62:932–937
Wei Y et al (2008) Loss of trimethylation at lysine 27 of histone H3 is a predictor of poor outcome in breast, ovarian, and pancreatic cancers. Mol Carcinog 47:701–706
Ngollo M et al (2014) The association between histone 3 lysine 27 trimethylation (H3K27me3) and prostate cancer: relationship with clinicopathological parameters. BMC Cancer 14:994
Oh EJ et al (2014) Diffuse large B-cell lymphoma with histone H3 trimethylation at lysine 27: another poor prognostic phenotype independent of c-Myc/Bcl2 coexpression. Hum Pathol 45:2043–2050
Ntziachristos P et al (2014) Contrasting roles for histone 3 lysine 27 demethylases in acute lymphoblastic leukemia. Nature 514:513–517
Khuong-Quang DA et al (2012) K27M mutation in histone H3.3 defines clinically and biologically distinct subgroups of pediatric diffuse intrinsic pontine gliomas. Acta Neuropathol 124:439–447
Wu G et al (2012) Somatic histone H3 alterations in pediatric diffuse intrinsic pontine gliomas and non-brainstem glioblastomas. Nat Genet 44:251–253
Sturm D et al (2012) Hotspot mutations in H3F3A and IDH1 define distinct epigenetic and biological subgroups of glioblastoma. Cancer Cell 4:425–437
Behjati S et al (2013) Distinct H3F3A and H3F3B driver mutations define chondroblastoma and giant cell tumor of bone. Nat Genet 45:1479–1482
Bender S et al (2013) Reduced H3K27me3 and DNA hypomethylation are major drivers of gene expression in K27M mutant pediatric high-grade gliomas. Cancer Cell 24:660–672
Solomon D et al (2016) Diffuse midline gliomas with histone H3-K27M mutation: a series of 47 cases assessing the spectrum of morphologic variation and associated genetic alterations. Brain Pathol 26:569–580
Gessi M et al (2015) High frequency of H3F3A (K27M) mutations characterizes pediatric and adult high-grade gliomas of the spinal cord. Acta Neuropathol 130:435–437
Feng J et al. (2015) The H3.3 K27M mutation results in a poorer prognosis in brainstem gliomas than thalamic glioms in adults. Human Pathol 46:1626–1632.
Buczkowicz P et al (2014) Histopathological spectrum of paediatric diffuse intrinsic pontine glioma: diagnostic and therapeutic implications. Acta Neuropathol 128:573–581
Pathak P et al (2015) Altered global histone-trimethylation code and H3F3A-ATRX mutation in pediatric GBM. J Neurooncol 121:489–497
Fontebasso AM et al (2013) Mutations in SETD2 and genes affecting histone H3K36 methylation target hemispheric high-grade gliomas. Acta Neuropathol 125:659–669
Lewis PW et al (2013) Inhibition of PRC2 activity by a gain-of function H3 mutation found in pediatric glioblastoma. Science 340:857–861
Hirabayashi Y, Suzki N, Tsuboi M et al (2009) Polycomb limits the neurogenic competence of neural precursor cells to promote astrogenic fate transition. Neuron 63:600–613
Xu W et al (2011) Oncometabolite 2-hydroxyglutarate is a competitive inhibitor of a-ketoglutarate-dependent dioxygenases. Cancer Cell 19:17–30
Chowdhury R et al (2011) The oncometabolite 2-hydroxyglutarate inhibits histone lysine demethylases. EMBO Rep 12:463–469
Lu C et al (2012) IDH mutation impairs histone demethylation and results in a block to cell differentiation. Nature 483:474–480
Arita H et al (2013) Upregulating mutations in the TERT promoter commonly occur in adult malignant gliomas and are strongly associated with total 1p19q loss. Acta Neuropathol 126:267–276
Koelsche C et al (2013) Distribution of TERT promoter mutations in pediatric and adult tumors of the nervous system. Acta Neuropathol 126:907–915
Eckel-Passow JE et al (2015) Glioma groups based on 1p/19q, IDH, and TERT promoter mutations in tumors. N Engl J Med 372:2499–2508
The Cancer Genome Atlas Research Network (2015) Comprehensive, integrative genomic analysis of diffuse lower-grade gliomas. N Engl J Med 372:2481–2497
Nonoguchi N et al (2013) TERT promoter mutations in primary and secondary glioblastomas. Acta Neuropathol 126:931–937
Labussiere M et al (2014) Combined analysis of TERT, EGFR, and IDH status defines distinct prognostic glioblastomas classes. Neurology 83:1200–1206
Castelo-Branco P et al (2013) Methylation of the TERT promoter and risk stratification of childhood brain tumours: an integrative genomic and molecular study. Lancer Oncol 14:534–542
Guilleret I et al (2002) Hypermethylation of the human telomerase catalytic subunit (hTERT) gene correlates with telomerase activity. Int J Cancer 101:335–341
Lindsey JC et al (2014) TERT promoter mutation and aberrant hypermethylation are associated with elevated expression in medulloblastoma and characterize the majority of non-infant SHH subgroup tumours. Acta Neuropathol 127:307–309
Fan Y et al (2016) Telomerase expression by aberrant methylation of the TERT promoter in melanoma arising in giant congenital nevi. J Invest Dermatol 136:339–342
Arita H et al (2013) TERT promoter mutations rather than methylation are the main mechanism for TERT upregulation in adult gliomas. Acta Neuropathol 126:939–941
Jiao Y et al (2011) DAXX/ATRX, MEN1 and mTOR pathway genes are frequently altered in pancreatic neuroendocrine tumors. Science 331:1199–1203
Mangerel J et al (2014) Alternative lengthening of telomeres is enriched in, and impacts survival of TP53 mutant pediatric malignant brain tumors. Acta Neuropathol 128:853–862
Liu X et al (2012) Frequent ATRX mutations and loss of expression in adult diffuse astrocytic tumors carrying IDH1/IDH2 and TP53 mutations. Acta Neuropathol 124:615–625
Zhang J et al (2013) Whole-genome sequencing identifies genetic alterations in pediatric low-grade gliomas. Nat Genet 45:602–612
Tabori U et al (2006) The role of telomere maintenance in the spontaneous growth arrest of pediatric low-grade gliomas. Neoplasia 8:136–142
Jiao Y et al (2012) Frequent ATRX, CIC, FUBP1 and IDH1 mutations refine the classification of malignant gliomas. Oncotarget 3:709–722
Suzuki H et al (2014) Mutational landscape and clonal architecture in grade II and III gliomas. Nat Genet 47:458–468
Abedalthagafi M et al (2013) The alternative lengthening of telomere phenotype is significantly associated with loss of ATRX expression in high-grade pediatric and adult astrocytomas: a multi-institutional study of 214 astrocytomas. Mod Pathol 26:1425–1432
Wong LH et al (2010) ATRX interacts with H3.3 in maintaining telomere structural integrity in pluripotent embryonic stem cells. Genome Res 20:351–360
Dhayalan A et al (2011) The ATRX-ADD domain binds to H3 tail peptides and reads the combined methylation state of K4 and K9. Hum Mol Genet 20:2195–2203
Law MJ et al (2010) ATR-X syndrome protein targets tandem repeats and influences allele-specific expression in a size dependent manner. Cell 143:367–378
Garcia-Cao M et al (2004) Epigenetic regulation of telomere length in mammalian cells by the Suv39h1 and Suv39h2 histone methyltransferases. Nat Genet 36:94–99
Gonzalo S et al (2006) DNA methyltransferases control telomere length and telomere recombination in mammalian cells. Nat Cell Biol 8:416–424
Gibbons RJ et al (2000) Mutations in ATRX, encoding a SWI/SNF-like protein, cause diverse changes in the pattern of DNA methylation. Nat Genet 24:368–371
Hashizume R et al (2014) Pharmacologic inhibition of histone demethylation as a therapy for pediatric brainstem glioma. Nat Med 20:1394–1396
Grasso CS et al (2015) Functionally defined therapeutic targets in diffuse intrinsic pontine glioma. Nat Med 21:555–559
Brown ZZ et al (2014) Strategy for “detoxification” of a cancer-derived histone mutant based on mapping its interaction with the methyltransferase PRC2. J Am Chem Soc 136:13498–13501
Pfister SX et al (2015) Inhibiting WEE1 selectively kills histone H3K36me3-deficient cancers by dNTP starvation. Cancer Cell 28:557–568
Bjerke L et al. (2013) Histone H3.3. mutations drive pediatric glioblastoma through upregulation of MYCN. Cancer Discov 3:512–519
Vaughan L et al. (2016) Inhibition of mTOR-kinase destabilizes MYCN and is a potential therapy for MYCN-dependent tumors. Oncotarget 7:57525–57544
Ruden M and N Puri (2013) Novel anticancer therapeutics targeting telomerase. Cancer Treatment Rev 39:444–456
Acknowledgements
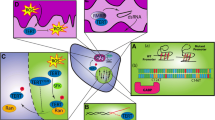
We thank Rishi Bhatnagar for creation of the illustration in Fig. 1. D.A.S. is supported by NIH Director’s Early Independence Award (DP5 OD021403) and Career Development Award from the UCSF Brain Tumor SPORE (P50 CA097257).
Author information
Authors and Affiliations
Corresponding author
Additional information
The original version of this article was revised: The second author’s name was corrected.
An erratum to this article is available at http://dx.doi.org/10.1007/s11060-017-2376-1.
Rights and permissions
About this article
Cite this article
Lee, J., Solomon, D.A. & Tihan, T. The role of histone modifications and telomere alterations in the pathogenesis of diffuse gliomas in adults and children. J Neurooncol 132, 1–11 (2017). https://doi.org/10.1007/s11060-016-2349-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-016-2349-9