Abstract
We define the Reflexive Tile Assembly Model (RTAM), which is obtained from the abstract Tile Assembly Model (aTAM) by allowing tiles to reflect across their horizontal and/or vertical axes. We show that the class of directed temperature-1 RTAM systems is not computationally universal, which is conjectured but unproven for the aTAM, and like the aTAM, the RTAM is computationally universal at temperature 2. We then show that at temperature 1, when starting from a single tile seed, the RTAM is capable of assembling \(n \times n\) squares for n odd using only n tile types, but incapable of assembling \(n \times n\) squares for n even. Moreover, we show that n is a lower bound on the number of tile types needed to assemble \(n \times n\) squares for n odd in the temperature-1 RTAM. The conjectured lower bound for temperature-1 aTAM systems is \(2n-1\). Finally, we give preliminary results toward the classification of which finite connected shapes in \(\mathbb {Z}^2\) can be assembled (strictly or weakly) by a singly seeded (i.e. seed of size 1) RTAM system, including a complete classification of which finite connected shapes can be strictly assembled by mismatch-free singly seeded RTAM systems.






















Similar content being viewed by others
Notes
It is interesting to note that there are interesting temperature-2 RTAM systems that assemble a single assembly even when individual tile orientations are taken into account. For example, rectilinear tile assembly systems in the RTAM at temperature-2 can produce a single terminal assembly (even when taking tile orientations into account) when the seed assembly of the system is L-shaped. Note that the terminal assembly cannot have width or height exceeding that of the seed. See Kari et al. (2015) for interesting examples of such rectilinear tile assembly systems with L-shaped seeds.
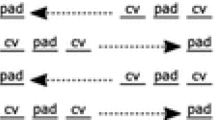
Essentially, a compact zig-zag system is one in which assembly proceeds along a single possible assembly sequence which grows one row completely from left-to-right, then immediately above that grows the next complete row right-to-left, and so on. Also, each row may grow to a length of one tile longer than the row below it.
References
Barish RD, Schulman R, Rothemund PWK, Winfree E (2009) An information-bearing seed for nucleating algorithmic self-assembly. Proc Natl Acad Sci 106(15):6054–6059
Cook M, Fu Y, Schweller RT (2011) Temperature 1 self-assembly: deterministic assembly in 3D and probabilistic assembly in 2D. In: SODA 2011: proceedings of the 22nd annual ACM-SIAM symposium on discrete algorithms. SIAM
Santini CC, Bath J, Tyrrell AM, Turberfield AJ (2013) A clocked finite state machine built from DNA. Chem Commun 49:237–239
Demaine ED, Demaine ML, Fekete SP, Patitz MJ, Schweller RT, Winslow A, Woods D (2014) One tile to rule them all: simulating any tile assembly system with a single universal tile. In: Esparza J, Fraigniaud P, Husfeldt T, Koutsoupias E (eds) Proceedings of the 41st international colloquium on automata., languages, and programming (ICALP 2014), IT University of Copenhagen, July 8–11, 2014, vol 8572 of LNCS, Springer, Berlin, pp 368–379
Doty D (2012) Theory of algorithmic self-assembly. Commun ACM 55(12):78–88
Doty D, Kari L, Masson B (2013) Negative interactions in irreversible self-assembly. Algorithmica 66(1):153–172
Doty D, Patitz MJ, Summers SM (2011) Limitations of self-assembly at temperature 1. Theor Comput Sci 412:145–158
Fekete SP, Hendricks J, Patitz MJ, Rogers TA, Schweller RT (2015) Universal computation with arbitrary polyomino tiles in non-cooperative self-assembly. In: Proceedings of the twenty-sixth annual ACM-SIAM symposium on discrete algorithms (SODA 2015), San Diego, Jan 4–6, pp 148–167
Fu B, Patitz MJ, Schweller RT, Sheline R (2012) Self-assembly with geometric tiles. In: Proceedings of the 39th international colloquium on automata, languages and programming, ICALP, pp 714–725
Ginsburg S, Spanier EH (1966) Semigroups, presburger formulas, and languages. Pac J Math 16(2):285–296
Han D, Pal S, Yang Y, Jiang S, Nangreave J, Liu Y, Yan H (2013) DNA gridiron nanostructures based on four-arm junctions. Science 339(6126):1412–1415
Kari L, Kopecki S, Meunier PÉ, Patitz MJ, Seki S (2015) Binary pattern tile set synthesis is NP-hard. In: Automata, languages, and programming—42nd international colloquium, ICALP 2015, Kyoto, July 6–10, 2015, Proceedings, Part I, pp 1022–1034
Ke Y, Ong LL, Shih WM, Yin P (2012) Three-dimensional structures self-assembled from DNA bricks. Science 338(6111):1177–1183
Kim J-W, Kim J-H, Deaton R (2011) DNA-linked nanoparticle building blocks for programmable matter. Angew Chem Int Edit 50(39):9185–9190
Lathrop JI, Lutz JH, Summers SM (2009) Strict self-assembly of discrete Sierpinski triangles. Theor Comput Sci 410:384–405
Mao C, LaBean TH, Relf JH, Seeman NC (2000) Logical computation using algorithmic self-assembly of DNA triple-crossover molecules. Nature 407(6803):493–496
Patitz MJ (2014) An introduction to tile-based self-assembly and a survey of recent results. Nat Comput 13(2):195–224
Patitz MJ, Schweller RT, Summers SM (2011) Exact shapes and turing universality at temperature 1 with a single negative glue. In: Proceedings of the 17th international conference on DNA computing and molecular programming, DNA’11. Springer, Berlin, pp 175–189
Pinheiro AV, Han D, Shih WM, Yan H (2011) Challenges and opportunities for structural DNA nanotechnology. Nat Nanotechnol 6(12):763–772
Rothemund PWK (2001) Theory and experiments in algorithmic self-assembly. PhD thesis, University of Southern California
Rothemund PWK, Papadakis N, Winfree E (2004) Algorithmic self-assembly of DNA Sierpinski triangles. PLoS Biol 2(12):e424
Rothemund PWK, Winfree E (2000) The program-size complexity of self-assembled squares (extended abstract). In: STOC ’00: Proceedings of the thirty-second annual ACM symposium on theory of computing. ACM, Portland, pp 459–468
Schulman R, Winfree E (2007) Synthesis of crystals with a programmable kinetic barrier to nucleation. Proc Nat Acad Sci 104(39):15236–15241
Winfree E (1998) Algorithmic self-assembly of DNA. PhD thesis, California Institute of Technology
Winfree E, Liu F, Wenzler LA, Seeman NC (1998) Design and self-assembly of two-dimensional DNA crystals. Nature 394(6693):539–544
Acknowledgements
We thank our anonymous reviewers for their careful reading of this work and their insightful comments that have significantly improved this paper. Supported in part by National Science Foundation Grants CCF-1117672 and CCF-1422152. T. Rogers’s research was supported by the National Science Foundation Graduate Research Fellowship Program under Grant No. DGE-1450079.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hendricks, J., Patitz, M.J. & Rogers, T.A. Reflections on tiles (in self-assembly). Nat Comput 16, 295–316 (2017). https://doi.org/10.1007/s11047-017-9617-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11047-017-9617-2