Abstract
Revealing the phylogenetic relationships of Candida krusei strains (sexual form Pichia kudriavzevii) is a prerequisite for understanding the evolution of its virulence-associated mechanisms and ecological lifestyles. Molecular phylogenetic analyses based on entire internal transcribed spacer region (ITS) and multilocus sequence typing (MLST) data were carried out with sequences available in public databases and Hungarian isolates from animals obtained for the study. The ITS haplotype network yielded a high frequency haplotype at the centre of the network (H1; n = 204) indicating that various selective pressure might resulted in population expansion from H1. MLST analysis identified three new genotypes among animal-derived isolates, therefore overall 203 sequence types were investigated to determine the population structure of C. krusei. The most commonly encountered sequence types were ST 17 and ST 67. Phylogenetic analyses showed diverse genetic construction of C. krusei population. Evidence of potential recombination events were also observed that might play some role in high intraspecies genetic variability among strains, however, the limited data of C. krusei genotypes from different countries prevented us to identify accurate evolutionary routes of commensal and pathogenic strains or species-specific lineages. Further expansion of C. krusei MLST database may promote the better understanding of the mixed evolutionary history of this species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pathogenic Candida species have widely contributed to the increasing morbidity, mortality and excess medical costs of healthcare-associated infections worldwide [1]. C. albicans remains the predominant etiological agent of candidiasis, however, the intensive use of some antifungal drugs in human healthcare has promoted a shift in the epidemiology of candidiasis towards non-albicans Candida species that are naturally less susceptible to drugs [2]. According to a global surveillance program covering twenty years (1997–2016), the four most frequently isolated non-albicans Candida species in descending order include C. glabrata (2,860; 18.7%), C. parapsilosis (2,433; 15.9%), C. tropicalis (1,418; 9.3%), and C. krusei (421; 2.8%) [3]. Among these emerging yeast pathogens C. krusei is the least-well studied despite of its high mortality rate (40–59%) especially in patients with neutropenia and hematologic malignancies [4, 5]. C. krusei is a transient commensal of the mucosal membrane of healthy individuals. In veterinary aspect, this organism has also been associated with diseases and deteriorate the health conditions in animals. C. krusei is a relevant pathogen involved in mycotic mastitis of cows resulting in reductions of milk yield and quality. Gastrointestinal diseases in birds caused by C. krusei have also been reported [6,7,8].
The emergence of antifungal resistance threatens the effective treatment of fungal infections. Azoles are one of the most common antifungal drugs used for long-term antifungal prophylactic treatment of immunocompromised individuals and applied in agricultural practices. C. krusei is a potential multidrug-resistant yeast due to its intrinsic resistance to fluconazole (with more than 96% of human clinical and veterinary isolates being fluconazole-resistant) as well as its acquired resistance developed to some other antifungal drugs (amphotericin B, echinocandins) [6, 9,10,11].
Additional specific characteristics of this species is the presence of sexual reproduction. The sexual form (teleomorph) of C. krusei is Pichia kudriavzevii that was proposed by Kurtzman and colleagues in 1980 [12]. P. kudriavzevii isolates are widely distributed in nature and due to its stress tolerance it has an increasing role in biotechnology mainly for production of bioethanol [13]. Moreover, some strains are candidate probiotics [14]. Recent whole genome sequencing supported that there is no genetic distinction between the two species [15]. Phylogenetic analysis of ribosomal DNA (rDNA) and other conserved genes along with genome sequences revealed large evolutionary distance between C. krusei and other medically relevant Candida species that further supported the re-classification of this organism in the Pichia genus [15, 16]. In accordance with the guidelines to avoid multiple denomination of any fungus species, the general usage of the name P. kudriavzevii (sexual form) has been suggested but has not yet been enacted; therefore the more conventional name, C. krusei, will be used in our article.
Relatively little genetic or genomic investigation has been carried out on C. krusei isolates so far. Sequence data from multiple loci facilitates determinations of population structures and epidemiological correlates of properties, such as geographical and anatomical origins of isolates and their transmission within and between hosts, therefore multilocus sequence typing (MLST) has gained widespread acceptance as a tool for exploring strain-level differences within microbial species [17,18,19,20]. Another typing scheme based on short tandem repeats for C. krusei have recently been developed that may facilitate the follow up of outbreaks in healthcare settings in the future [21]. A previous MLST study of C. krusei indicated that isolates have diploid, heterozygous genome [18], but phylogenetic analysis using the extended data deposited in MLST database to determine the genetic diversity and population structure of strains has not performed since 2007. The content of the MLST database has expanded since this original report on the genetic structure of C. krusei; therefore the authors felt that new analyses would permit a deeper understanding of the genetics and evolutionary history of this important opportunistic pathogen.
Materials and Methods
Sample Collection and Identification of Species
Eight C. krusei isolate were cultured from swab samples sent for species identification to our laboratory. All samples were obtained from the uterus of different cows. One isolate from our culture collection was also included in the study, which originated from the oesophagus of a duck diagnosed with gastrointestinal mycosis. Swab samples were inoculated on Sabouraud dextrose agar supplemented with chloramphenicol and incubated at 35 °C for 48 h. The cultures were identified by Matrix-assisted laser desorption/ionization time of flight mass spectrometry (MALDI-TOF MS) and molecular method by sequencing of internal transcribed spacer (ITS) region of fungal rDNA [22].
After culturing isolates for two days, the genomic DNA was extracted and purified directly from a single colony of each isolate by using the Fungi/yeast genomic DNA extraction kit (Favorgen, Ping-Tung, Taiwan). DNA samples were stored at − 20 °C until analysis. The complete ITS region, including ITS1 and ITS2 and the 5.8S region of rDNA were amplified with fungus-specific universal primers ITS1 (5′-TCCGTAGGTGAACCTGCGG-3′) and ITS4 (5′-TCCTCCGCTTATTGATATGC-3′). The reaction was performed in a final volume of 15 μl including 1 µl fungal DNA, 2 µl 10 × DreamTaq buffer, 0.5 µl dNTP (10 mM), 0.5 µl forward and reverse primers (10 µM each), 0.1 µl DreamTaq DNA polymerase (5 U/µl; Thermo Fisher Scientific, Waltham, MA, USA) and 10.4 µl distilled water. The reaction conditions were as follows: initial denaturation step at 95 °C for 3 min, 40 cycles of 95 °C for 30 s, annealing at 50 °C for 30 s, extension at 72 °C for 1 min, and a final extension step at 72 °C for 10 min. Both strands of purified gene fragments were sequenced using ITS1 and ITS4 primers with BigDye Terminator v3.1. cycle sequencing kit (Thermo Fisher Scientific, Waltham, MA, USA) on an ABI Prism 3130 Genetic Analyzer (Applied Biosystems). Sequences were edited and assembled using MEGA v6 software [23] and compared with sequences in database by the BLAST sequence analysis tool (https://blast.ncbi.nlm.nih.gov/Blast.cgi).
Multilocus Sequence Typing of C. krusei Isolates
The MLST scheme employed for C. krusei genotyping was based on partial amplification and sequencing of six protein-coding genes (HIS3, LEU2, NMT1, TRP1, ADE2, and LYS2D) according to the international reference MLST scheme established for this species. Six independent PCR amplifications were performed for each isolate. The primer sets and their amplicon lengths were described in detail elsewhere [18]. Experimental conditions used in PCRs and Sanger sequencing were detailed above.
Heterozygosity was identified by the presence of two coincident peaks at the same polymorphic loci on the forward and reverse sequence chromatograms and the consensus sequences of six loci of all isolates were defined [24]. Each nucleotide sequence was compared to those in the MLST database (https://pubmlst.org/organisms/candida-krusei) [25]. Sequence variation in each locus was assigned an allele number. Those sequences that did not match with any of the preexisting sequences was given a new allele number by the curator. The combination of the six allele numbers defined a unique sequence type (ST) of an isolate.
Phylogenetic and Population Structure Analysis
To assess the intraspecific variability of the ITS region of C. krusei isolates as a universal fungal barcode, we selected sequences deposited in GenBank for phylogenetic analysis. Sequences that were not long enough (< 470 bp) were eliminated. Eight ITS sequence were excluded from the analysis as they were more closely related to Candida pseudolambica, Pichia bruneiensis and Pichia fermentans than to C. krusei (MN710485-MN710491, MT781361). Neighbor-joining (NJ) tree was constructed from 268 ITS sequences using JC + G substitution model with bootstrap of 1,000 replications. As NJ tree showed minimal differences between sequences, phylogenetic network was also generalized that better model the phylogenetic relations between isolates by creating haplotypes from diploid sequence data and considering the possibility of recombination, hybridization, gene conversion and gene transfer. Sequence polymorphism analysis was carried out with DnaSP version 6 software and haplotype network file was created [26]. Haplotypes were defined as unique combination of SNPs along a particular sequence. Haplotype diversity values range from 0 to 1, with those closer to 1 indicating higher variability. Haplotype network was generated by median-joining method [27] implemented in NETWORK v4.6.1.0 (Fluxus Technology, Suffolk, UK).
For multilocus sequence analysis, the sequenced loci were concatenated into a single sequence and single nucleotide polymorphisms (SNPs) were converted as described by Tavanti et al. (2005) [28] to label homozygous and heterozygous sites in order to allow the cluster analysis of diploid sequence data. Briefly, each base at the polymorphic sites rewritten twice for a homozygous (A, C, G, or T) datum or once each for the two component bases for a heterozygous (K, M, R, S, W, or Y) datum. All sequences available in the MLST database were involved in phylogenetic analysis. The genetic relatedness between the investigated strains was evaluated by unweighted pair group method with arithmetic averages (UPGMA) algorithm with p-distance model in MEGA v6 software. Random bootstrapping of 1,000 operations was used for the construction. Clonal complexes (CCs) of isolates that differed in sequence at only one of the six loci (single-locus variant analysis) were predicted by the goeBURST algorithm (http://www.phyloviz.net/goeburst/) to investigate the evolutionary relationships between isolates. CC is a set of STs that are all believed to be descended from the same founding genotype. STs that could not be assigned to any group were called singletons. Haplotype diversity was calculated and possible recombination events were estimated using the DnaSP v6 software [26, 29].
Results
Based on aligned ITS sequences, 76 variable sites and 32 haplotypes with haplotype diversity of 0.3869 were determined. Low intraspecific diversity was found among isolates. Most of the sequences were identical and belonged to 3 haplotypes (H1, H3 and H17). Our samples also clustered to the H1 group along with reference strain ATCC 6258. The ITS haplotype network yielded a high frequency haplotype at the centre of the network (H1; n = 204), followed by H3 (n = 26), H17 (n = 4) and other haplotypes with a single sequence indicating that various selective pressure might resulted in population expansion from H1. Several haplotypes differed in only one nucleotide from each other which is evident in network analysis but not in NJ phylogenetic reconstruction (Fig. 1).
Phylogenetic tree of Candida krusei ITS sequences and network analysis of ITS haplotypes by median-joining method. The compressed branch of the tree contains 253 sequences. Numbers along the branches indicate bootstrap values. The size of the circumference is proportional to the haplotype frequency in the network. Mutational steps are represented by white dots. The black dots (median vectors) represent unsampled or extinct haplotypes in the population
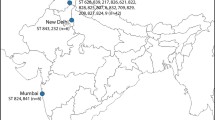
Concatenated sequence data from the six partial genes yielded a sequence of 2,847 bp for each isolate whose sequences were determined in this study. In two animal-derived isolates a shared new allele was identified in LEU2. Four isolates were assigned to previously described STs whereas another five isolates were classified in new MLST genotypes (ST numbers: 201–203) (Table 1). Together with our newly submitted data, a total of 203 STs available in the C. krusei MLST database were investigated in the study. Overall, 76 polymorphic sites were found among all examined loci. The NMT1 locus produced the highest number of polymorphic sites (n = 35), while HIS3 displayed the lowest (n = 23). The most commonly encountered strain types were ST 17 (13 isolates) and ST 67 (18 isolates). These isolates were obtained from human blood or an animal source and the geographical origins of isolation were Austria, Canada, Czech Republic, France, Hungary, Italy, Taiwan, United Kingdom, and Venezuela. Other STs were represented by less than 10 isolates. In order to reveal the phylogenetic relationship between isolates, we performed cluster analysis of STs submitted to the MLST database by the UPGMA method. C. krusei STs could be subgrouped into five clusters with an approach described by Jacobsen et al. (2007) [18], however, the bootstrap values for the nodes in the dendrogram were extremely low hampering the accurate distinction of internal structure. The largest group was subtype 1 containing 52.06% of STs, followed by subtype 2 (28.35%), subtype 4 (15.46%), subtype 3 (3.61%) and subtype 5 (0.51%) (Fig. 2). STs from human blood and animal source were the main components of group subtype 1, although, isolates from animals were submitted only from two countries (France and Hungary). Subtype 2 predominantly consisted of STs from the oropharynx (including sputum and bronchoalveolar lavage) of human patients. In contrast to other members of the group subtype 3, the anatomical origin of ST 49 and ST 87 was known (oropharynx/other superficial source and blood, respectively), while subtype 4 was prevailingly composed of blood isolates. One ST obtained from an animal formed group subtype 5 alone. In general, several STs were identified in all subtypes whose origin was unknown (Table 2, Fig. 2). Unfortunately, information on STs from different geographical origin is incomplete and thus, the correlation between subtypes and host specificity or anatomical source could not be determined (Fig. 3).
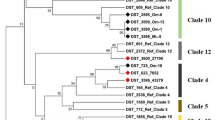
Phylogram constructed by unweighted pair group method with arithmetic mean analysis (UPGMA) of Candida krusei sequence types. The scale bar indicates P-distance. The arbitrarily selected P-distance cutoff was 0.0028. Hungarian isolates are highlighted with bold numbers. Numbers along the branches indicate bootstrap values
The goeBURST algorithm was used to identify putative ancestral STs and their closest relations. Evaluation of genotypic relationship of strains resulted 18 CCs and 94 singletons. CCs generated by goeBURST were numbered starting from 0 (for the CC with most STs). Our isolates with known STs clustered to CC-3 (ST 19 and ST 24) and CC-4 (ST 67), while newly identified STs were singletons. The ST 24 was putatively evolved from the ST 19 group founder due to loss of heterozygosity considering nucleotide positions 2,467 and 2,659 in LYS2D. In CC-4, ST 67 or ST 164 was predicted as founder of the group. Interestingly, ST 67 was the most prevalent genotype that was isolated globally (including Asia, Europe, North America and South America). Besides Hungarian isolates, C. krusei MLST data from animals were reported only from France. Of note, ST 17, ST 19, ST 24 and ST 67 were detected from animal samples in both countries. Moreover, ST 17 and ST 67 were the most prevalent genotypes identified even from humans and animals worldwide, raising the question whether strains that belong to these genotypes are better adapted to colonise or infect different hosts. Overall, ST 17, ST 19, ST 24, ST 67 and ST 68 were detected from both humans and animals and these genotypes were mostly assigned as probable ancestors of CCs (Fig. 4). To provide a more accurate representation of the relationships among genetic groups, haplotype analysis of each six coding regions and the concatenated sequences were carried out and recombination events were estimated, which revealed evidence of potential recombination among the loci assessed suggesting the mixed ancestry of strains (Table 3).
Discussion
Conserved and variable rDNA regions are suitable and universally applied for comparative analysis to identify organisms at taxonomic levels and clarify phylogenetic relationships between species and populations [30]. In recent years, many studies have been denoting some limitations in using ITS for delimiting fungal species such as the lack of differentiation of closely related species and the presence of non-homologous copies of ITS in the genome [22]. Intragenomic ITS variations that was observed in Candida and Pichia species as well may derive from recombination, gene duplication and point mutations produced by DNA replication errors [22, 31, 32]. However, despite these limitations, the utilization of the rDNA ITS region for sequence analysis still appears to be the most reliable method for the rapid and accurate molecular identification of fungal pathogens. Furthermore, phylogenetic, taxonomic and population dynamics studies are substantially concerned with polymorphisms in the ITS region and are proven to be particularly useful for the delineation of ascomycetous yeasts [30, 33]. Comparative analysis of C. krusei ITS sequences including those deposited in public genomic databases showed low intraspecies variability as the majority of sequences belong to H1 haplotype, while the remaining 31 haplotypes are represented by relatively few sequences. The sampling efficiency for network analysis has been supported by the low number of median vectors which predicts the existence of unsampled or extinct haplotypes. Nevertheless, several haplotypes could be evolved from H1 with numerous mutational steps suggesting higher variability of ITS sequence of C. krusei than formerly noted [33].
The C. krusei strains were also studied by MLST analysis. MLST provides relevant information for population genetic studies and for epidemiologic investigations with higher level of reproducibility and minimal subjectivity in analysis compared with other technologies [24]. The genes chosen to be sequenced for MLST conventionally encoded metabolic functions that are subject to stabilizing selection, because diversifying selection may obscure relationships among isolates [24, 34]. C. krusei STs in MLST database (n = 203) lag far behind the number of STs of other Candida species with diploid genome that can be genotyped by the MLST approach (C. albicans, n = 3,669; C. tropicalis, n = 1,245). Jacobsen et al. [18] devised a MLST scheme ideal for strain typing of C. krusei using 129 human isolates and distinguished 94 STs. Fourteen years later, we performed an extended phylogenetic analyses with 302 isolates and 203 STs from the C. krusei MLST database. As a result of this timely update we have gained more comprehensive insight into the evolutionary history of C. krusei using multiple analyses by inclusion of strains from different source. According to our results, similar to the previous observation, STs can be subtyped. Furthermore, the majority of strains originated from animals belonged to Subtype 1. Although certain genotypes were determined from one host (e.g. ST 153–186 originated from animals) or appeared in one continent (e.g. ST 56, ST 60, ST 61 and ST 73 were noted exclusively from Australia), no unequivocal evidence of geographical associations of STs were seen due to the comparatively low numbers of isolates typed so far. This is consistent with findings that this species is the least frequently isolated among the five most common clinically relevant Candida species and typing of strains obtained from animals and environmental sources are somewhat neglected. In addition, submitting genotyped strains to MLST database requires anatomical source of isolation. Thus, strains from other sources (e.g. milk, cheese) [35] marked as unknown prevents a better resolution of phylogenetic relations. Most of the STs that originated from animals (eg., ST 19, ST 67) were putative group founders or differed only by a single allele when using cluster analysis (eg., ST 24, ST 182, ST 184, ST 186) suggesting that these strains were less exposed to changing environmental conditions. The high rate of singletons found in eBURST analysis and the evidence of recombination, which might be an explanation of lower bootstrap values in the UPGMA dendrogram, further supports the unexclusively clonal reproduction of C. krusei, a finding that coincided with previously published data [15, 18].
In conclusion, studies on population structure of C. krusei are currently incomplete. Although recombination events may play some role in the observed high intraspecies genetic variability, the limited data of C. krusei genotypes from different countries submitted to MSLT database make it difficult to reveal accurate evolutionary routes of commensal and pathogenic strains or species-specific lineages. Some genotypes were identified from humans and animals that may indicate certain genotypes are better adapted to diverse environmental niches. A significant increase of sequence data together with isolate-associated metadata in the MLST database will be needed for a more robust resolution of the population structure of C. krusei.
Data Availability
Generated ITS sequences were deposited in GenBank under the following accession numbers: OL470260-OL470268. Allele sequences and concatenated sequences of the six genes used for MLST are available at: https://pubmlst.org/organisms/candida-krusei. The converted concatenated dataset of loci used for cluster analysis can be found in Supplementary file 1.
References
Gómez-Gaviria M, Mora-Montes HM. Current aspects in the biology, pathogeny, and treatment of Candida krusei, a neglected fungal pathogen. Infect Drug Resist. 2020;13:1673–89. https://doi.org/10.2147/IDR.S247944.
Ksiezopolska E, Gabaldón T. Evolutionary emergence of drug resistance in Candida opportunistic pathogens. Genes (Basel). 2018;9:461. https://doi.org/10.3390/genes9090461.
Pfaller MA, Diekema DJ, Turnidge JD, Castanheira M, Jones RN. Twenty Years of the SENTRY antifungal surveillance program: results for Candida species from 1997–2016. Open Forum Infect Dis. 2019;6:S79–94. https://doi.org/10.1093/ofid/ofy358.
Wisplinghoff H, Bischoff T, Tallent SM, Seifert H, Wenzel RP, Edmond MB. Nosocomial bloodstream infections in US hospitals: analysis of 24,179 cases from a prospective nationwide surveillance study. Clin Infect Dis. 2004;39:309–17. https://doi.org/10.1086/421946.
Horn DL, Neofytos D, Anaissie EJ, Fishman JA, Steinbach WJ, Olyaei AJ, Marr KA, Pfaller MA, Chang CH, Webster KM. Epidemiology and outcomes of candidemia in 2019 patients: data from the prospective antifungal therapy alliance registry. Clin Infect Dis. 2009;48:1695–703. https://doi.org/10.1086/599039.
Du J, Wang X, Luo H, Wang Y, Liu X, Zhou X. Epidemiological investigation of non-albicans Candida species recovered from mycotic mastitis of cows in Yinchuan. Ningxia China BMC Vet Res. 2018;14:251. https://doi.org/10.1186/s12917-018-1564-3.
Donnelly KA, Wellehan JFX, Quesenberry K. Gastrointestinal disease associated with non-albicans Candida Species in six birds. J Avian Med Surg. 2019;33:413–8. https://doi.org/10.1647/2018-419.
Domán M, Makrai L, Bali K, Lengyel G, Laukó T, Bányai K. Unexpected diversity of yeast species in esophageal mycosis of waterfowls. Avian Dis. 2020;64:532–5. https://doi.org/10.1637/aviandiseases-D20-00053.
Whaley SG, Berkow EL, Rybak JM, Nishimoto AT, Barker KS, Rogers PD. Azole antifungal resistance in Candida albicans and emerging non-albicans Candida species. Front Microbiol. 2016;7:2173. https://doi.org/10.3389/fmicb.2016.02173.
Jørgensen LN, Heick TM. Azole use in agriculture, horticulture, and wood preservation - is it indispensable? Front Cell Infect Microbiol. 2021;11: 730297. https://doi.org/10.3389/fcimb.2021.730297.
Chowdhary A, Kathuria S, Xu J, Meis JF. Emergence of azole-resistant Aspergillus fumigatus strains due to agricultural azole use creates an increasing threat to human health. PLoS Pathog. 2013;9:e1003633. https://doi.org/10.1371/journal.ppat.1003633.
Kurtzman CP, Smiley MJ, Johnson CJ. Emendation of the genus Issatchenkia kudriavzev and comparison of species by deoxyribonucleic acid reassociation, mating reaction, and ascospore ultrastructure. Int J System Bacteriol. 1980. https://doi.org/10.1099/00207713-30-2-503.
Mukherjee V, Radecka D, Aerts G, Verstrepen KJ, Lievens B, Thevelein JM. Phenotypic landscape of non-conventional yeast species for different stress tolerance traits desirable in bioethanol fermentation. Biotechnol Biofuels. 2017;10:216. https://doi.org/10.1186/s13068-017-0899-5.
Chelliah R, Ramakrishnan SR, Prabhu PR, Antony U. Evaluation of antimicrobial activity and probiotic properties of wild-strain Pichia kudriavzevii isolated from frozen idli batter. Yeast. 2016;33:385–401. https://doi.org/10.1002/yea.3181.
Douglass AP, Offei B, Braun-Galleani S, Coughlan AY, Martos AAR, Ortiz-Merino RA, Byrne KP, Wolfe KH. Population genomics shows no distinction between pathogenic Candida krusei and environmental Pichia kudriavzevii: One species, four names. PLoS Pathog. 2018;14:e1007138. https://doi.org/10.1371/journal.ppat.1007138.
Shen XX, Zhou X, Kominek J, Kurtzman CP, Hittinger CT, Rokas A. reconstructing the backbone of the Saccharomycotina yeast phylogeny using genome-scale data. G3 (Bethesda). 2016; 6:3927–3939. doi:https://doi.org/10.1534/g3.116.034744.
Bougnoux ME, Tavanti A, Bouchier C, Gow NA, Magnier A, Davidson AD, Maiden MC, D’Enfert C, Odds FC. Collaborative consensus for optimized multilocus sequence typing of Candida albicans. J Clin Microbiol. 2003;41:5265–6. https://doi.org/10.1128/JCM.41.11.5265-5266.2003.
Jacobsen MD, Gow NA, Maiden MC, Shaw DJ, Odds FC. Strain typing and determination of population structure of Candida krusei by multilocus sequence typing. J Clin Microbiol. 2007;45:317–23. https://doi.org/10.1128/JCM.01549-06.
Tavanti A, Davidson AD, Johnson EM, Maiden MC, Shaw DJ, Gow NA, Odds FC. Multilocus sequence typing for differentiation of strains of Candida tropicalis. J Clin Microbiol. 2005;43:5593–600. https://doi.org/10.1128/JCM.43.11.5593-5600.2005.
Dodgson AR, Pujol C, Denning DW, Soll DR, Fox AJ. Multilocus sequence typing of Candida glabrata reveals geographically enriched clades. J Clin Microbiol. 2003;41:5709–17. https://doi.org/10.1128/JCM.41.12.5709-5717.2003.
van Haren MHI, de Groot T, Spruijtenburg B, Jain K, Chowdhary A, Meis JF. Development of a multiplex PCR short tandem repeat typing scheme for Candida krusei. J Clin Microbiol. 2022;60:e02032-e2121. https://doi.org/10.1128/JCM.02032-21.
Schoch CL, Seifert KA, Huhndorf S, Robert V, Spouge JL, Levesque CA, Chen W, Fungal Barcoding Consortium; Fungal Barcoding Consortium Author List. Nuclear Ribosomal Internal Transcribed Spacer (ITS) Region as a Universal DNA Barcode Marker for Fungi. Proc Natl Acad Sci U S A. 2012; 109:6241–6246. doi:https://doi.org/10.1073/pnas.1117018109.
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol Biol Evol. 2013;30:2725–9. https://doi.org/10.1093/molbev/mst197.
Odds FC, Jacobsen MD. Multilocus sequence typing of pathogenic Candida species. Eukaryot Cell. 2008;7:1075–84. https://doi.org/10.1128/EC.00062-08.
Jolley KA, Bray JE, Maiden MCJ. Open-access bacterial population genomics: BIGSdb software, the PubMLST.org website and their applications. Wellcome Open Res. 2018; 3:124. doi:https://doi.org/10.12688/wellcomeopenres.14826.1.
Rozas J, Ferrer-Mata A, Sánchez-DelBarrio JC, Guirao-Rico S, Librado P, Ramos-Onsins SE, Sánchez-Gracia A. DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol Biol Evol. 2017;34:3299–302. https://doi.org/10.1093/molbev/msx248.
Bandelt HJ, Forster P, Röhl A. Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol. 1999;16:37–48. https://doi.org/10.1093/oxfordjournals.molbev.a026036.
Tavanti A, Davidson AD, Fordyce MJ, Gow NAR, Maiden MCJ, Odds FC. Population structure and properties of Candida albicans, as determined by multilocus sequence typing. J Clin Microbiol. 2005;43:5601–13. https://doi.org/10.1128/JCM.43.11.5601-5613.2005.
Hudson RR, Kaplan NL. Statistical properties of the number of recombination events in the history of a sample of DNA sequences. Genetics. 1985;111:147–64. https://doi.org/10.1093/genetics/111.1.147.
Iwen PC, Hinrichs SH, Rupp ME. Utilization of the internal transcribed spacer regions as molecular targets to detect and identify human fungal pathogens. Med Mycol. 2002;40:87–109. https://doi.org/10.1080/mmy.40.1.87.109.
Lücking R, Aime MC, Robbertse B, Miller AN, Ariyawansa HA, Aoki T, Cardinali G, Crous PW, Druzhinina IS, Geiser DM, Hawksworth DL, Hyde KD, Irinyi L, Jeewon R, Johnston PR, Kirk PM, Malosso E, May TW, Meyer W, Öpik M, Robert V, Stadler M, Thines M, Vu D, Yurkov AM, Zhang N, Schoch CL. Unambiguous identification of fungi: Where do we stand and how accurate and precise is fungal DNA barcoding? IMA Fungus. 2020;11:14. https://doi.org/10.1186/s43008-020-00033-z.
Wu ZW, Wang QM, Liu XZ, Bai FY. Intragenomic polymorphism and intergenomic recombination in the ribosomal RNA genes of strains belonging to a yeast species Pichia Membranifaciens. Mycology. 2016;7:102–11. https://doi.org/10.1080/21501203.2016.1204369.
Merseguel KB, Nishikaku AS, Rodrigues AM, Padovan AC, e Ferreira RC, de Azevedo Melo AS, Briones MR da S, Colombo AL. Genetic diversity of medically important and emerging Candida species causing invasive infection. BMC Infect Dis. 2015; 15:57. doi:https://doi.org/10.1186/s12879-015-0793-3.
Maiden MCJ. Multilocus sequence typing of bacteria. Annu Rev Microbiol. 2006;60:561–88. https://doi.org/10.1146/annurev.micro.59.030804.121325.
Lavoie K, Touchette M, St-Gelais D, Labrie S. Characterization of the fungal microflora in raw milk and specialty cheeses of the province of Quebec. Dairy Sci Technol. 2012;92:455–68. https://doi.org/10.1007/s13594-011-0051-4.
Funding
Open access funding provided by Veterinary Medical Research Institute. This work was supported by the National Research, Development and Innovation Office of Hungary-NKFIH (PD 128617).
Author information
Authors and Affiliations
Contributions
MD and KB designed the study. LM provided samples and data. MD performed experiments, data analysis and prepared the first manuscript draft. All authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors report no conflicts of interest.
Ethical Approval
The authors confirm that no ethical approval was required.
Human and Animal Rights
This work was carried out with diagnostic samples and no animal experimentation was conducted.
Consent to Participate
The authors alone are responsible for the content and writing of this paper.
Consent for Publication
Not applicable.
Additional information
Handling Editor: Ferry Hagen.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Domán, M., Makrai, L. & Bányai, K. Molecular Phylogenetic Analysis of Candida krusei. Mycopathologia 187, 333–343 (2022). https://doi.org/10.1007/s11046-022-00640-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11046-022-00640-x