Abstract
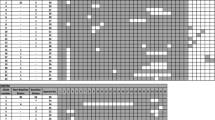
We applied multilocus microsatellite typing (MLMT) method to investigate the genetic relation between Cryptococcus neoformans var. grubii clinical and environmental isolates in São Paulo, Brazil. This MLMT method includes three functional gene sequences of C. neoformans var. grubii, which are dispersed on three chromosomes. In all, 89 strains (36 clinical and 53 environmental isolates) were analyzed. Of 36 clinical strains, 20 belonged to a major type of MLMT-13 (55.6%). They were mainly isolated from clinical specimens. About 52.8% of strains from the environment belong to a major type of MLMT-36, which are indigenous to environments and which were not isolated from clinical samples. Thus, we recognized two genotypes that distinguish majority of clinical and environmental strains. No differences were found in antifungal susceptibility and capsule size between major environmental and clinical MLMT types.

Similar content being viewed by others
References
Currie BP, Casadevall A. Estimation of the prevalence of cryptococcal infection among HIV infected individuals in New York City. Clin Infect Dis. 1994;19:1029–33.
Casadevall A, Perfect JR. Cryptococcus neoformans. Washington, DC: ASM Press; 1998. p. 41–70.
Kwon-Chung KJ, Boekhout T, Fell JW, Diaz M. Proposal to conserve the name Cryptococcus gattii against C. hondurianus, C. basillisporus (Basidiomycota, Hymenomycetes, Tremellomycetidae). Taxon. 2002;51:804–6.
Imai T, Watanabe K, Tamura M, Mikami Y, Tanaka R, Nishimura K, et al. Geographic grouping of Cryptococcus neoformans var. gattii by random amplified polymorphic DNA fingerprint patterns and ITS sequence divergence. Clin Lab. 2000;46:345–54.
Katsu M, Kidd S, Ando A, Morelli-Branchini ML, Mikami Y, Nishimura K, et al. The internal transcribed spacer and 5.8S rRNA gene show extensive diversity among isolates of the Cryptococcus neoformans species complex. FEMS Yeast Res. 2004;4:377–88.
Levitz SM. The ecology of Cryptococcus neoformans and epidemiology of cryptococcosis. Rev Infect Dis. 1991;13:1163–9.
Li A, Nishimura K, Taguchi H, Tanaka R, Wu S, Miyaji M. The isolation of Cryptococcus neoformans from pigeon droppings and serotyping of naturally and clinically sourced isolates in China. Mycopathologia. 1993;124:1–5.
Chee HY, Lee KB. Isolation of Cryptococcus neoformans var. grubii (serotype A) from pigeon dropping in Seoul, Korea. J Microbiol. 2005;43:469–72.
Boekhout T, Theelen B, Diaz M, Fwll JW, Hop WC, Abein EC, et al. Hybrid genotypes in the pathogenic Cryptococcus neoformans. Microbiology. 2001;147:891–907.
Litvintseva AP, Thakur R, Vilgalys R, Mitchell TG. Multilocus sequence typing reveals three genetic subpopulations of Cryptococcus neoformans var. grubii (serotype A), including a unique population in Botswana. Genetics. 2006;172:2223–38.
Field D, Wills C. Long polymorphic microsatellites in simple organisms. Proc Biol Sci. 1996;263:209–15.
Bretagne S, Costa JM, Besmond C, Carsique R, Calderone R. Microsatellite polymorphism in the promoter sequence of the elongation factor 3 gene of Candida albicans as basis for typing system. J Clin Microbiol. 1997;35:1777–80.
Hanafy A, Kaocharoen S, Jover-Botella A, Katsu M, Iida S, Kogure T, et al. Multilocus microsatellite typing for Cryptococcus neoformans var. grubii. Med Mycol. 2008;46:1–12.
Kwon-Chung KJ, Poacheck I, Bennett J. Improved diagnostic medium for separation of Cryptococcus neoformans var. neoformans (serotype A and D) and Cryptococcus neoformans var. gattii (serotype B and C). J Clin Microbiol. 1982;15:535–7.
Makimura K, Oguri T, Mikami Y, Kume H, Hanazawa R, Abe M, et al. Multicenter evaluation of commercial frozen plate for microdilution broth antifungal susceptibility testing of yeasts and comparison of MIC limits recommended in NCCLS M27–A2. Microbiol Immunol. 2005;49:97–106.
McClelland E, Bernhardt EP, Casadevall A. Estimating the relative contributions of virulence factors for pathogenic microbes. Infect Immun. 2006;74:1500–4.
Chang YC, Kwon-Chung KJ. Complementation of a capsule deficient mutation of Cryptococcus neoformans restores its virulence. Mol Cell Biol. 1994;14:4912–9.
D’Souza CA, Alspaugh JA, Yue C, Cox Harashima T, GM Perfect JR, Heitman J. Cyclic AMP-dependent protein kinase controls virulence of the fungal pathogen Cryptococcus neoformans. Mol Cell Biol. 2001;21:3179–91.
Nielsen K, Cox GM, Litvintseva AP, Mylonakis E, Malliaris SD, Benjamin DK, et al. Cryptococcus neoformans strains preferentially disseminate to the central nervous system during coinfection. Infect Immunol. 2005;73:4922–33.
Maxson ME, Dadachova E, Casadevall A. Zaragoza O. Radial mass density, charge, and epitope distribution in the Cryptococcus neoformans capsule. Eukaryot Cell. 2007;6:95–109.
Mylonakis E, Ausubel FM, Perfect JR, Heitman J, Calderwood SB. Killing of Caenorhabditis elegans by Cryptococcus neoformans as a model of yeast pathogenesis. Pro Natl Acad Sci. 2002;99:15675–80.
Delgado AC, Taguchi H, Mikami Y, Miyaji M, Villares MC, Moretti ML. Human cryptococcosis: relationship of environmental and clinical strains of Cryptococcus neoformans var. neoformans from urban and rural areas. Mycopathologia. 2005;159:7–11.
Franzot SP, Hamdan JS, Currie BP, Casadevall A. Molecular epidemiology of Cryptococcus neoformans in Brazil and the United States: evidence for both local genetic differences and a global clonal population structure. J Clin Microbiol. 1997;35:2243–51.
Currie BP, Freundlich LF, Casadevall A. Restriction fragment length polymorphism analysis of Cryptococcus neoformans isolates from environmental (pigeon excreta) and clinical sources in New York City. J Clin Microbiol. 1994;32:1188–92.
Casali AK, Goulart L, Rosa e Silva LK, Ribeiro AM, Amaral AA, Amves SH, et al. Molecular subtyping of clinical and environmental Cryptococcus neoformans isolates in the Brazilian State Rio Grande Sul. J Clin Microbiol. 1995;33:3328–32.
Fries BC, Chen F, Currie BP, Casadevall A. Karyotype instability in Cryptococcus neoformans infection. J Clin Microbiol. 1996;34:1531–4.
Franzot SP, Mukherjee J, Cherniak R, Chen LC, Hamdan JS, Casadevall A. Microevolution of a standard strain of Cryptococcus neoformans resulting in differences in virulence and other phenotype. Infect Immun. 1998;66:89–97.
Fries BC, Goldman DL, Casadevall A. Phenotypic switching in Cryptococcus neoformans. Microbes Infect. 2002;4:1345–52.
Sullivan D, Haynes K, Moran G, Coleman D. Persistence, replacement, and microevolution of Cryptococcus neoformans strains in recurrent meningitis in AIDS patients. J Clin Microbiol. 1996;34:1739–44.
Fries BC, Goldman DL, Cherniak R, Ju R, Casadevall A. Phenotypic switching of Cryptococcus neoformans results in changes in cellular morphology and glucuronoxylomannan structure. Infect Immun. 1999;67:6076–83.
Loftus BJ, Fung E, Roncaglia P, Rowley D, Amedeo P, Bruno D, et al. The genome of the basidiomycetous yeast and human pathogen Cryptococcus neoformans. Science. 2005;307:1321–4.
Fraser JA, Huang JC, Pukkila-Worley R, Alspaugh JA, Mitchell TG, Heitman J. Chromosomal translocation and segmental duplication in Cryptococcus neoformans. Eukaryot Cell. 2005;4:401–6.
Walton FJ, Idnurm A, Heitman J. Novel gene functions required for melanization of the human pathogen Cryptococcus neoformans. Mol Microbiol. 2005;57:1381–96.
Fox DS, Cruz MC, Sia RA, Ke H, Cox GM, Cardenas ME, et al. Calcineurin regulatory subunit is essential for virulence and mediates interactions with FKBP12-FK506 in Cryptococcus neoformans. Mol Microbiol. 2001;39:835–49.
Acknowledgments
This work was supported by Special Coordination Funds for Promoting Science and Technology from the Ministry of Education, Culture, Sports, Science and Technology of Japan to Y. M. JZ is a recipient of grant for special program “China Nairikubu Jinzai Ikusei Jigyo”. Reference fungal strains were obtained through the National BioResource Project (NBRP) in Japan (http://www.nbrp.jp/).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhu, J., Kang, Y., Uno, J. et al. Comparison of Genotypes Between Environmental and Clinical Isolates of Cryptococcus neoformans var. grubii Based on Microsatellite Patterns. Mycopathologia 169, 47–55 (2010). https://doi.org/10.1007/s11046-009-9230-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11046-009-9230-8