Abstract
One of the highly focused areas in the medical science community is segmenting tumors from brain magnetic resonance imaging (MRI). The diagnosis of malignant tumors at an early stage is necessary to provide treatment for patients. The patient’s prognosis will improve if it is detected early. Medical experts use a manual method of segmentation when making a diagnosis of brain tumors. This study proposes a new approach to simplify and automate this process. In recent research, multi-level segmentation has been widely used in medical image analysis, and the effectiveness and precision of the segmentation method are directly tied to the number of segments used. However, choosing the appropriate number of segments is often left up to the user and is challenging for many segmentation algorithms. The proposed method is a modified version of the 3D Histogram-based segmentation method, which can automatically determine an appropriate number of segments. The general algorithm contains three main steps: The first step is to use a Gaussian filter to smooth the 3D RGB histogram of an image. This eliminates unreliable and non-dominating histogram peaks that are too close together. Next, a multimodal particle swarm optimization method identifies the histogram’s peaks. In the end, pixels are placed in the cluster that best fits their characteristics based on the non-Euclidean distance. The proposed algorithm has been applied to a Cancer Imaging Archive (TCIA) and brain MRI Images for brain Tumor detection dataset. The results of the proposed method are compared with those of three clustering methods: FCM, FCM_FWCW, and FCM_FW. In the comparative analysis of the three algorithms across various MRI slices. Our algorithm consistently demonstrates superior performance. It achieves the top mean rank in all three metrics, indicating its robustness and effectiveness in clustering. The proposed method is effective in experiments, proving its capacity to find the proper clusters.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Clinicians can identify diseases earlier thanks to medical imaging, which improves patient outcomes. Appropriate medical image analysis is essential to aid specialists and promote a healthy community. The application of image processing techniques for analyzing medical images is exceedingly successful, thanks to advanced medical equipment. Image segmentation, the first step in image processing, is one of the most significant and complex challenges in image analysis, particularly in the application of medical images. Segmentation divides an image into meaningful, nonoverlapping, homogeneous, connected regions concerning color similarity. Medical image segmentation aims to isolate anatomical objects of interest for analysis and is critical in medical imaging applications [1,2,3,4]. There is a wide variety of image segmentation methods, including threshold-based, clustering-based, region-based, edge-based, etc. [5,6,7,8,9]. Other hybrid image segmentation techniques combine multiple approaches [10]. Many years of research have examined segmentation features and methods. Nevertheless, one of the restrictions is that the appropriate number of segments is a parameter that must be established a priori, and determining this value is not an easy task [11]. In addition, the problem remains tough because, as the desired number of segments increases, the problem’s computing cost increases exponentially, making it unfeasible to employ accurate methods to search for all possible solutions exhaustively.
Computerized tomography (CT), magnetic resonance imaging (MRI), electroencephalography (EEG), and positron emission tomography (PET) are the four most prevalent forms of imaging techniques for the brain. However, magnetic resonance imaging (MRI) is the most frequently utilized method. MRI does not expose patients to radiation, has minimal invasiveness, and is widely available [12]. Furthermore, the MRI can discern more clearly between fatty tissue, water, muscle, and other types of soft tissue. In addition, MRI provides greater soft tissue contrast [13]. The information contained in these images is provided to medical professionals, who can utilize it to assist in the diagnosis of a wide range of illnesses and ailments. The goal of the segmentation of tumors in the human brain is to differentiate between normal and diseased tissues of the brain, specifically cerebrospinal fluid, gray matter, and white matter of the brain. When it comes to investigating brain tumors, it is currently simple to identify abnormal tissues; nevertheless, the segmentation process is challenging, making it difficult to reproduce results, characterize abnormalities, and achieve precision [13]. The term “tumor” refers to the condition that occurs when the uncontrollable expansion of cancer cells causes it. This tumor comes in various subtypes and manifestations, each of which can be treated with a specific modality according to its unique symptoms. Brain segmentation results in an image with labels that show the different regions’ boundaries or a group of contours. Segmentation methods can be further divided into bi-level segmentation methods, which divide an image into two parts, and multi-level segmentation methods, which divide an image into more than two parts [14]. Since MRI brain images contain more than two different types of regions, each of which may correspond to a unique object, the bi-level segmentation technique cannot be effective and results in under-segmentation.
As a result, multi-level image segmentation algorithms should be utilized to segment MRI brain images. Also, segmenting color images (RGB) is difficult due to the range of color intensities and three-color channels, unlike gray images, which have only one [15]. Therefore, various approaches can be taken to segment MRI brain images [1, 13]. Methods for segmenting MRI brain images can be divided into three broad classes: classification-based, region-based, and boundary-based methods [16, 17]. Fuzzy c-means is a typical classification-based approach widely used in medical image segmentation [18,19,20,21]. However, these clustering methods suffer from predefined values for determining the number of the proper segments.
Furthermore, the computational time is also a consideration since it depends on the number of clusters and image size. As a result, the paper’s motivation comes from automatically segmenting the image regardless of image size and selecting the appropriate number of clusters in addition to the actual peaks. To get around these problems, this study suggests using a 3D histogram and a modified multimodal particle swarm optimization method for brain MRI segmentation.
The cluster centers can be found by detecting the peaks in a three-dimensional histogram of a color image created from the RGB values of the pixels and smoothed with a Gaussian filter. The multimodal variation of particle swarm optimization (PSO) with a local search strategy has been utilized to find the global and local peaks in a histogram and the cluster centers. The contribution of the proposed clustering-based method is that a non-Euclidean distance metric has replaced the Euclidean metric for calculating the multimodal optimization algorithm’s pixel similarity and movement strategy. The number of clusters in the image can be automatically determined based on the number of PSO-detected peaks. Following the discovery of peaks, individual pixels are subsequently assigned to the cluster to which they are spatially closed using non-Euclidean distance, which results in the final segmented brain image. The proposed algorithm has been compared with the FCM [22] clustering algorithm.
The rest of this paper is organized as follows. Section 2 presents the FCM clustering. Section 3 describes the multimodal PSO algorithm. The proposed method is described in Section 4. Finally, section 5 presents the experimental and comparative results, while Section 5 concludes the paper.
2 Fuzzy c-means clustering
The k-means and c-means algorithms are two of the most well-known clustering methodologies in color image segmentation. These algorithms frequently yield good results and are widely used. Nevertheless, as mentioned before, one of the constraints is that the number of clusters is a parameter that must be determined in advance, and determining the value of this parameter is not easy. In addition, the computational time is a primary concern when solving the problem, as it depends on the required number of clusters and the image size. The c-means algorithm uses fuzzy memberships to put each pixel in the correct category.
Where xij s a representation of pixel xj’s membership in the ith cluster, vi s the center of the ith cluster, ‖∙‖ s a norm metric, and m is a constant. The fuzziness of the final partition is determined by the value of the parameter m, and usually m = 2.
The cost function is minimized when high membership values are assigned to pixels close to their clusters’ centroid. On the other hand, when low membership values are assigned to pixels with data far from the centroid, the cost function is maximized. This is because the membership function calculates the probability of a pixel being part of a particular cluster, given its location. This technique’s probability depends only on pixel distance from each cluster center. The following are the factors that contribute to the updating of the membership functions and cluster centers:
The algorithm starts by randomly selecting the clusters’ centers and each cluster’s average location. The next step that c-means conducts is to assign an entirely random membership grade to each data point for each cluster. Finally, the means attempts to place the cluster centers in the correct location within a data set and calculates the degree of membership in each cluster for each data point by updating the cluster centers and the membership grades. This iteration tries to minimize an objective function that shows how far a given data point is from the center of a cluster based on how much that data point belongs to the cluster.
3 Multimodal PSO
While unimodal optimization algorithms can only identify a single global optimum (solution) within the collection of options, multimodal optimization algorithms can find many local and global optimum solutions [23, 24]. Even though multimodal optimization approaches have not been explored nearly as extensively as unimodal optimization methods, they have recently gained much attention. However, most of them have the same problem with niching parameters. Existing approaches have trouble figuring out the right niching radius, which is their main problem [25]. In the majority of studies, basic unimodal optimization algorithms, such as the Genetic Algorithm [26,27,28] and the PSO Algorithm [29,30,31,32], have been modified to become MMO algorithms. PSO’s movement (crossover) method is suited to adapt to a multimodal form. PSO is a nature-inspired method for unimodal stochastic optimization. t has been used to solve many computational problems [33]. PSO has been extended numerous times to a multimodal form in the literature. PSO mimics the swarming behavior of foraging birds to move particles toward the best solution. Therefore, PSO depends on the movement of particles in the search space to determine the best value. Each particle keeps track of its personal best position (i.e., personal best) and the overall best position (i.e., global best) gained by the population so far. Each particle (ith) is associated with a position vector and velocity vector that is recorded as xi and vi, respectively. xi and vi are updated according to the following equation:
Where t is the iteration number, w indicates the inertia weight, \({p}_i^{best}\) and gbest correspond to the location of personal best and global best, respectively. R1 and R2 are two uniformly distributed random numbers generated inside the interval [0,1]. C1 and C2 represent the particle’s confidence and its neighbors correspondingly. Unimodal PSO cannot locate multiple solutions as all solutions move to global best (gbest). However, PSO’s mechanism for particle motion can be easily adapted to handle multimodal problems. Carrera and Coello Coello (2009) introduced a modified PSO variation for solving multimodal problems inspired by electrostatic charge interactions [32]. Each solution moves toward the solution with the greatest electrostatic interaction calculated based on the current fitness value to locate multiple optimal solutions. These interactions are determined mathematically according to the following:
Where Qij, r ≠ 0, and ε0 refer, respectively, to the electrical charges of the interacting particles, the distance between them, and the permittivity of the vacuum. To apply these ideas in an optimization context, each solution’s fitness value corresponds to the particles’ electric charge. Herein, Eq. 4 is simulated as:
In this case, the constant scalar 4πε0 s replaced by the variable α, computed based on Li [34]. gbest in Eq.4 is replaced by \({index}_i={\displaystyle \begin{array}{c} argmax\\ {}\textrm{J}=1:\textrm{M}\end{array}}{F}_{ij}\) or a constant index j. Here, M denotes the population size.
4 Proposed method
In a research paper called 3DHP, all global and local peaks within a 3D color histogram corresponding to each cluster’s center points were located using the aforementioned multimodal approaches with a local search strategy. Pixel’s color in RGB-model images is derived from a weighted average of red, green, and blue components. t can represent each pixel in an image as a three-dimensional feature vector made up of the pixel’s three component colors. 3D histograms can then be constructed from these three-color axes [35]. The presence of peaks in a histogram indicates that the image comprises multiple distinct segments, each corresponding to a particular segment. In 3DHP, a three-dimensional Gaussian filter was applied to three-dimensional histograms to reduce the effect of noise and turn them out into smoothed histograms. This procedure also eliminates insignificant smaller peaks that may have been present in the histogram. Figure 1 illustrates the three-dimensional histogram, the original color distribution, and the color distribution after the smoothing process for the image of Lena. The 3D histogram was considered an objective function, and the positions of peaks were solution space. In this case, the number of pixels in the particular position corresponds to the fitness value. Moreover, an additional local search step proposed in [36] was integrated into multimodal PSO to enhance local search ability. Finally, the fitness values are used to check out the neighbors of the ith article. So, the following changes are made to the position of the ith particle:
Graphics of the 3D histogram, RGB color distribution, and Lenna’s smoothed color distribution. a) original Lenna image, (b) three-dimensional histogram of Lenna, (c) and (d) show the normal and smoothened RGB representation of Lenna [11]
Where bestNearesti is the particle that is closest in the distance to the ith the article, D is the number of dimensions, and temp is a new position in the ith particle. A new position will then replace the particle’s position if it is determined that the new position is superior to the current position of the particle (xi). Consequently, all particles do not need to move to a single global optimum; other possible local solutions are not missed. In the next step, K dominant peaks are located. Then, K sets of peak intensity levels corresponding to cluster centers are automatically obtained for each RGB component. These peaks are represented as follows: \({p}_1^{rgb}=\left({r}_1,{g}_1,{b}_1\right),{p}_2^{rgb}=\left({r}_2,{g}_2,{b}_2\right),{p}_3^{rgb}=\left({r}_3,{g}_3,{b}_3\right)\),⋯, \({p}_k^{rgb}=\left({r}_K,{g}_K,{b}_K\right)\).
Additionally, to eliminate non-dominant clusters, it is beneficial to confine the distance that separates the two peaks as much as possible. Therefore, dominant peaks in a region eliminate all non-dominant peaks within its radius based on a distance limit parameter. It is essential to remember that this procedure is not mandatory and can be skipped if desired. For the 3DHP, this parameter was set to 80 pixels. Ultimately, every pixel will be assigned to the peak closest to it regarding the Euclidean distance. The following equation was used to calculate the Euclidean distance between the kth peak and the (i, j)th (pixel)
Also, a non-Euclidean distance criterion was proposed in [37] and then applied in [37] on color image segmentation using FCM. Therefore, this equation is calculated per:
Where A is the number of features.
In the proposed method, the devisor of the fraction in Eq. 6\(\left(\left\Vert {p}_i^{best}-{p}_j^{best}\right\Vert \right)\) is replaced by Eq.7. This equation could also be expressed as:
Therefore:
Also, after locating the histogram peaks, every pixel will be assigned to the peak (cluster head) closest to it in terms of the non-Euclidean distance instead of the Euclidean distance. Consequently, Eq.10 can be reformulated as:
It is worth mentioning that a preprocessing step to smooth the image by Gaussian smoothing is applied to the RGB image before calculating the 3D histogram. Meanwhile, σ and the window size for this filter are set to 0.5 and (3 × 3), (respectively)
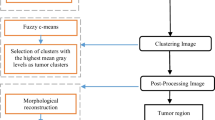
The flow diagram of the overall method is illustrated in Fig. 2.
The flow diagram of the overall method. The Gaussian filter smooths a 3D RGB histogram, eliminating non-dominating peaks. A multimodal particle swarm optimization method identifies peaks, assigning pixels to the nearest peak based on non-Euclidean distance, ensuring reliable and non-dominating histograms
5 Experimental results and performance evaluation
In this section, extensive experiments are performed on the proposed method. The results of the proposed method are compared with well-known FCM [22], FCM_FW [3], and FCM_FWCW [38]. The required parameters of the proposed method and their values are shown in Table 1. Also, the fuzziness parameter in all soft (fuzzy) clustering methods is set to 2. The maximum number of iterations for FCM, FCM_FW and FCM_FWCW is 100.
5.1 Dataset
We used the following two datasets to evaluate the proposed method:
-
The first datasets were taken from the Cancer Imaging Archive (TCIA). The National Cancer Institute supports it and contains corresponding medical imaging data for Cancer Genome Atlas (TCGA) participants.
-
The second dataset [39] is available at https://huggingface.co/datasets/miladfa7/Brain-MRI-Images-for-Brain-Tumor-Detection/tree/main.
-
The third dataset is available at https://www.kaggle.com/datasets/ammarnassanalhajali/brain-tumor.
5.2 Evaluation metrics
As the brain MRI slices are heterogeneous, qualitative (visual) evaluation of different methods is insufficient to analyze the results accurately. Therefore, quantitative metrics are needed to evaluate the results of various methods [40]. In the experiments, the following two groups of metrics are used to measure the performance of algorithms.
-
1)
Internals clustering metrics: A lower value of these metrics indicates a better segmentation result in this group.
F: this metric penalizes over-segmentation [41] (segmenting one region of the image into more than one segment):
where M and N represent the length and width of the input image, R is the number of segmented regions, Ai indicates the number of pixels in the ith segmented region ei indicates the color error in region i, and \(\sqrt{R}\) represents a penalizing term that discourages over-segmentation.
\(\overset{\acute{\mkern6mu}}{\boldsymbol{F}}\) : this metric penalizes over-segmentation and is noise-robust [42]:
Q: this metric penalizes non-homogeneous regions [42]:
-
2)
Externals clustering metrics: A higher value of these metrics indicates a better segmentation result in this group.
Accuracy: is the number of correct prediction pixels divided by the total number of pixels. This metric is calculated by Eq. (18).
Precision: It is the ratio of correct positive prediction pixels to the number of positive pixels predicted. This metric is calculated by Eq. (19).
Recall: It is the ratio of the number of correct positive prediction pixels to the number of all relevant pixels. This metric is calculated by Eq. (20).
F 1 Score: It is the harmonic mean between Precision and Recall. This metric is calculated by Eq. (21).
Specificity: The Specificity rate corresponds to the proportion of negative pixels that are correctly considered negative concerning all negative pixels. This metric is calculated by Eq. (22).
In Eq.s (18) to (22), TP, FN, TN, and FN represent True Positive, False Positive, True Negative, and False Negative, respectively.
In our experiments, these metrics are expressed as a percentage. A high percentage indicates a better performance.
5.3 Experiment 1: Visualization-based analysis using internal metrics
In this section, we evaluate the proposed method with other methods qualitatively and quantitatively. Several images have been selected for the quality assessment of each dataset. In selecting these images, we tried to select images that include different types of tumors: small and large tumor sizes, different tumor tissues, spherical and non-spherical tumor shapes, and different lighting conditions. 23 images are selected from the first, 15 from the second datasets, and 15 images from third one.
Figures 3, 4, and 5 demonstrate the results of the visual qualitative analysis of the first, second and third datasets, respectively. The segmentation results for each method are displayed by employing a distinct color set to the base image to highlight the clusters obtained. Tables 2 (first dataset), 4 (second dataset), and 6 (third dataset) contain information regarding the peak locations of the three-dimensional histogram and the cluster centroids for each cluster achieved by FCM, FCM_FW [3] and FCM_FWCW. In the same way, Tables 3, 5, and 7 show the numerical and qualitative analysis of the results from each of the three tested methods.
In our tests, the number of peaks found by M3DHP is used to figure out the number of clusters (m) for FCM, FCM_FW and FCM_FWCW. However, within the FCM_FW and FCM_FWCW algorithms, there are scenarios where some clusters end up empty without any pixels being assigned to them. The algorithm might converge to a solution where one or more clusters do not have any data points associated with them. The issue can arise from various factors, including the selection of initial cluster centers, data distribution, or the fact that the specified number of clusters does not align with the natural clustering in the data.
Based on what is shown in Figs. 3, 4, and 5 for all images on both datasets, the background is clearly distinguished when using M3DHP. On the other hand, with FCM, the backgrounds of 11 images out of 23 on the first dataset are mistakenly divided into many regions and are over-segmented. Also, in the second dataset, with FCM, the backgrounds of 5 images out of 15 on the second dataset are mistakenly divided into many regions and are over-segmented. Also, in the third dataset, with FCM, the backgrounds of 5 images out of 15 on the third dataset are mistakenly divided into many regions and are over-segmented.
For TCGA_HT_8105_19980826_26, the tumor can be distinguished more easily with the M3DHP than with the FCM. Many pixels are mistakenly assigned to the same tumor cluster by the FCM. Moreover, for TCGA_FG_6688_20020215_24, the tumor in the bottom center of the brain is very clearly distinguished when using M3DHP. However, in this case, FCM against M3DHP is not successful. For TCGA_DU_7014_19860618_30, the proposed algorithm segments the image correctly, while the segmented image by FCM represents both brain and tumor regions in a single region. In this case, the image is under-segmented with the associated image. Visual inspection reveals that the M3DHP method generally yields more homogeneous segmentation regions (Tables 4, 5, and 6).
As shown in Tables 3, 5, and 7, according to F and F′, the proposed method outperforms FCM in 12 out of 23 cases. Furthermore, according to Q, the proposed method outperforms FCM in 17 out of 23 cases. Results for the three evaluation functions F, F′, and Q suggest that the quantitative performance of both approaches is comparable when applied to the same image. It is important to note that there is not a huge disparity between these numbers; they all tend toward zero. The M3DHP approach demonstrates its efficacy by generating reliable results for the three statistical metrics F, F′, and Q. When it comes to F, F′, and Q, M3DHP typically offers superior performance to FCM in most cases. The last three rows of Tables 3, 5, and 7 indicate the average rank of algorithms according to all three performance indicators. The M3DHP ranked first according to all three performance indicators.
In the comparative analysis of the three algorithms across various MRI slices using metrics F, F ́, and Q, our algorithm consistently demonstrates superior performance. It achieves the top mean rank in all three metrics, indicating its robustness and effectiveness in clustering. FCM_FW generally ranks second, outperforming the standard FCM, which consistently ranks last. The consistent top ranking of our algorithm across all metrics underscores its potential as a preferred choice for clustering tasks in varied contexts.
5.4 Experiment 2: Numerical analysis of all images using internal metrics
In this section, we evaluate the average results obtained on both datasets’ images. As shown in Tables 8 and 9, M3DHP has the best results. Regarding F criteria, after M3DHP, the method FCM_FWCW has better results. Also, in terms of F ́ criteria, after M3DHP, FCM_FWCW has better results. Regarding Q criteria, after M3DHP, FCM_FW in the first dataset and FCM in the second dataset have better results. Methods FCM_FWCW have almost high performance in terms of F ́and F criteria and have similar F ́and F to M3DHP. However, in terms of Q criteria are much larger than M3DHP. In the third dataset, similar other datasets, our model has first rank.
These findings lead us to the conclusion that M3DHP can demonstrate competent performance during the segmentation of brain magnetic resonance images. The visual and numerical results show that the proposed M3DHP technique produces promising segmentation results. The method’s ability to automatically generate the desired number of clusters and cluster centroids proves this.
The thorough testing demonstrates that the proposed method performs well for image segmentation, surpassing the performance standards set by well-known methods like FCM, FCM_FWCW, and FCM_FW. The consistently superior outcomes across multiple evaluation criteria underscore the potential of the proposed method as a noteworthy contribution to the field.
5.5 Experiment 3: Analysis of all images with external metrics
In this experiment, to investigate the proposed method deeply and compare the obtained results with other state-of-the-art methods, the performance of the proposed method is evaluated with image ground truths and external evaluation metrics, such as accuracy, F1, precision, recall, and specificity. The statistical results are reported in Table 10. The visual segmentation results are also shown in Fig. 6. We test the proposed method only on the first dataset because the ground truth for the second one is not available.
Table 10 and Fig. 6 show that the proposed method has the best average performance for tested images. For image TCGA_DU_5855_19951217_23, tumor and non-tumor areas are well segmented, and due to the low light intensity, shape and texture, the other methods could not accurately detect the entire tumor area. Also, in the image TCGA_DU_7014_19860618_45, the border of the skull is segmented as a tumor area in the compared methods. However, the proposed methods are able to detect tumor areas well. This error in other methods is due mainly to the high brightness of the tissue surrounding the skull. The accuracy, F1-Score, and precision metrics rates are by an average of 95.52%, 79.68%, and 77.2257% on all testing images. This result highlights the efficacy of the feature combination employed in our method. The recall and specificity metrics in the proposed method are lower than those of the FCM_FW method. The reason is apparent: using feature weighting schemas and applying efficient extracted features can improve the results. However, the feature extraction phase is not used in our method, as are the FCM and FCM_FWCW methods.
6 Conclusion
In this paper, we suggest a modified form of the 3D Histogram-based segmentation technique that can choose the appropriate number of segments. The appropriate number of segments is determined by taking advantage of peak detection using a multimodal optimization algorithm. Using a multimodal optimization method, the optimal number of segments is calculated by exploiting peak detection. The proposed method has been applied to brain MRI to be segmented. The optimal number of clusters is unknown, making M3DHP more flexible for practice than other methods. To prove the efficiency of the proposed method, it has been compared with the well-known FCM clustering scheme. The results of the experiments demonstrate that the suggested strategy produces the desired outcomes and outperforms FCM. In our research, we developed a segmentation method for brain MRI that currently works with a single 2D MRI slice. The next step in our research will focus on extending this algorithm to handle 3D MRI segmentation.
Data availability
The brain images supporting Fig. 1 are publicly available in the Kaggle: https://www.kaggle.com/datasets/mateuszbuda/lgg-mri-segmentation
References
Pham DL, Xu C, Prince JL (2000) A survey of current methods in medical image segmentation. Annu Rev Biomed Eng 2(3):315–337
Rahkar Farshi T, Demirci R, Feizi-Derakhshi M-R (2018) Image clustering with optimization algorithms and color space. Entropy 20(4):296. https://doi.org/10.3390/e20040296
A. G. Oskouei, M. A. Balafar, and T. Akan, (2023) “A brain MRI segmentation method using feature weighting and a combination of efficient visual features,” Applied Computer Vision and Soft Computing with Interpretable AI, pp. 15–34, https://doi.org/10.1201/9781003359456-2.
H. B. Tabrizi and C. Crick, “Brain-Inspired Visual Odometry: Balancing Speed and Interpretability through a System of Systems Approach,” 2023, Accessed: Jan. 06, 2024. [Online]. Available: https://arxiv.org/abs/2312.13162v1
Mousavirad SJ, Ebrahimpour-Komleh H (2020) Human mental search-based multilevel thresholding for image segmentation. Appl Soft Comput 97:105427. https://doi.org/10.1016/J.ASOC.2019.04.002
Farag AA (2009) Edge-based image segmentation. Remote Sens Rev 6(1):95–121. https://doi.org/10.1080/02757259209532148
Slabaugh G, Unal G, Wels M, Fang T, Rao B (2009) Statistical region-based segmentation of ultrasound images. Ultrasound Med Biol 35(5):781–795. https://doi.org/10.1016/J.ULTRASMEDBIO.2008.10.014
Rahkar Farshi T, Ardabili AK (2021) A hybrid firefly and particle swarm optimization algorithm applied to multilevel image thresholding. Multimedia Systems 27(1):125–142. https://doi.org/10.1007/S00530-020-00716-Y/TABLES/7
Rahkar Farshi T, Orujpour M (2019) Multi-level image thresholding based on social spider algorithm for global optimization. Int J Inf Technol 11(4):713–718. https://doi.org/10.1007/S41870-019-00328-4/FIGURES/2
Gupta D, Anand RS (2017) A hybrid edge-based segmentation approach for ultrasound medical images. Biomed Signal Process Control 31:116–126. https://doi.org/10.1016/J.BSPC.2016.06.012
Farshi TR, Drake JH, Özcan E (2020) A multimodal particle swarm optimization-based approach for image segmentation. Expert Syst Appl 149:113233. https://doi.org/10.1016/J.ESWA.2020.113233
Xue G, Chen C, Lu Z-L, Dong Q (2010) Brain imaging techniques and their applications in decision-making research. Xin Li Xue Bao 42(1):120. https://doi.org/10.3724/SP.J.1041.2010.00120
Saritha S, Amutha Prabha N (2016) A comprehensive review: segmentation of MRI images—brain tumor. Int J Imaging Syst Technol 26(4):295–304. https://doi.org/10.1002/IMA.22201
Rahkar Farshi T, Demirci R (2021) Multilevel image thresholding with multimodal optimization. Multimed Tools Appl 80(10):15273–15289. https://doi.org/10.1007/S11042-020-10432-4/TABLES/3
Kumar S, Pant M, Kumar M, Dutt A (2018) Colour image segmentation with histogram and homogeneity histogram difference using evolutionary algorithms. Int J Mach Learn Cybern 9(1):163–183. https://doi.org/10.1007/S13042-015-0360-7/FIGURES/5
“Current Methods in the Automatic Tissue Segmentation of 3D Magnet…: Ingenta Connect.” Accessed: Jul. 09, 2022. [Online]. Available: https://www.ingentaconnect.com/content/ben/cmir/2006/00000002/00000001/art00008
Niessen WJ, Vincken KL, Weickert J, Ter Haar Romeny BM, Viergever MA (1999) Multiscale segmentation of three-dimensional MR brain images. Int J Comput Vis 31(2):185–202. https://doi.org/10.1023/A:1008070000018
Wang J, Kong J, Lu Y, Qi M, Zhang B (2008) A modified FCM algorithm for MRI brain image segmentation using both local and non-local spatial constraints. Comput Med Imaging Graph 32(8):685–698. https://doi.org/10.1016/J.COMPMEDIMAG.2008.08.004
Verma H, Verma D, Tiwari PK (2021) A population based hybrid FCM-PSO algorithm for clustering analysis and segmentation of brain image. Expert Syst Appl 167:114121. https://doi.org/10.1016/J.ESWA.2020.114121
Sikka K, Sinha N, Singh PK, Mishra AK (2009) A fully automated algorithm under modified FCM framework for improved brain MR image segmentation. Magn Reson Imaging 27(7):994–1004. https://doi.org/10.1016/J.MRI.2009.01.024
P. Wang and H. L. Wang, (2008) “A modified FCM algorithm for MRI brain image segmentation,” Proceedings - 2008 International Seminar on Future BioMedical Information Engineering, FBIE 2008, pp. 26–29, https://doi.org/10.1109/FBIE.2008.12.
Bezdek JC, Ehrlich R, Full W (1984) FCM: the fuzzy c-means clustering algorithm. Comput Geosci 10(2–3):191–203. https://doi.org/10.1016/0098-3004(84)90020-7
Rahkar Farshi T, Orujpour M (2021) A multi-modal bacterial foraging optimization algorithm. J Ambient Intell Humaniz Comput 12(11):10035–10049. https://doi.org/10.1007/S12652-020-02755-9/FIGURES/6
Orujpour M, Feizi-Derakhshi MR, Rahkar-Farshi T (2020) Multi-modal forest optimization algorithm. Neural Comput & Applic 32(10):6159–6173. https://doi.org/10.1007/S00521-019-04113-Z/FIGURES/11
Farshi TR (2022) A memetic animal migration optimizer for multimodal optimization. Evol Syst 13(1):133–144. https://doi.org/10.1007/S12530-021-09368-3/TABLES/12
Qing L, Gang W, Zaiyue Y, Qiuping W (2008) Crowding clustering genetic algorithm for multimodal function optimization. Appl Soft Comput 8(1):88–95. https://doi.org/10.1016/J.ASOC.2006.10.014
Li JP, Balazs ME, Parks GT, Clarkson PJ (2002) A species conserving genetic algorithm for multimodal function optimization. Evol Comput 10(3):207–234. https://doi.org/10.1162/106365602760234081
Liang Y, Leung KS (2011) Genetic algorithm with adaptive elitist-population strategies for multimodal function optimization. Appl Soft Comput 11(2):2017–2034. https://doi.org/10.1016/J.ASOC.2010.06.017
Wang H, Wang W, Wu Z (2013) Particle swarm optimization with adaptive mutation for multimodal optimization. Appl Math Comput 221:296–305. https://doi.org/10.1016/J.AMC.2013.06.074
Ren Z, Zhang A, Wen C, Feng Z (2014) A scatter learning particle swarm optimization algorithm for multimodal problems. IEEE Trans Cybern 44(7):1127–1140. https://doi.org/10.1109/TCYB.2013.2279802
Der Chang W (2015) A modified particle swarm optimization with multiple subpopulations for multimodal function optimization problems. Appl Soft Comput 33:170–182. https://doi.org/10.1016/J.ASOC.2015.04.002
J. Barrera and C. A. Coello Coello, (2009) “A particle swarm optimization method for multimodal optimization based on electrostatic interaction,” Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics), vol. 5845 LNAI, pp. 622–632, https://doi.org/10.1007/978-3-642-05258-3_55/COVER/.
R. Eberhart and J. Kennedy, (1995) “New optimizer using particle swarm theory,” Proceedings of the International Symposium on Micro Machine and Human Science, pp. 39–43, https://doi.org/10.1109/MHS.1995.494215.
X. Li, (2007) “A Multimodal Particle Swarm Optimizer Based on Fitness Euclidean-distance Ratio,” Proceedings of the 9th annual conference on Genetic and evolutionary computation - GECCO ‘07, https://doi.org/10.1145/1276958.
Navon E, Miller O, Averbuch A (2005) Color image segmentation based on adaptive local thresholds. Image Vis Comput 23(1):69–85. https://doi.org/10.1016/J.IMAVIS.2004.05.011
Qu BY, Liang JJ, Suganthan PN (2012) Niching particle swarm optimization with local search for multi-modal optimization. Inf Sci (N Y) 197:131–143. https://doi.org/10.1016/J.INS.2012.02.011
Golzari Oskouei A, Hashemzadeh M, Asheghi B, Balafar MA (2021) CGFFCM: Cluster-weight and Group-local Feature-weight learning in Fuzzy C-Means clustering algorithm for color image segmentation [Formula presented]. Appl Soft Comput 113. https://doi.org/10.1016/J.ASOC.2021.108005
Hashemzadeh M, Golzari Oskouei A, Farajzadeh N (2019) New fuzzy C-means clustering method based on feature-weight and cluster-weight learning. Appl Soft Comput 78:324–345. https://doi.org/10.1016/J.ASOC.2019.02.038
“Brain MRI Images for Brain Tumor Detection.” Accessed: 05 Jan. 2024. [Online]. Available: https://www.kaggle.com/datasets/navoneel/brain-mri-images-for-brain-tumor-detection/discussion
Chang D, Zhao Y, Liu L, Zheng C (2016) A dynamic niching clustering algorithm based on individual-connectedness and its application to color image segmentation. Pattern Recogn 60:334–347. https://doi.org/10.1016/J.PATCOG.2016.05.008
Liu J, Yang YH (1994) Multiresolution color image segmentation. IEEE Trans Pattern Anal Mach Intell 16(7):689–700. https://doi.org/10.1109/34.297949
Borsotti M, Campadelli P, Schettini R (1998) Quantitative evaluation of color image segmentation results. Pattern Recogn Lett 19(8):741–747. https://doi.org/10.1016/S0167-8655(98)00052-X
Funding
No funding for this study.
Author information
Authors and Affiliations
Contributions
All authors contributed to the research process in various forms, including original draft preparation, writing, review, and editing. TA provided core concepts, drafted the manuscript, and carried out implementations for this manuscript. AGO provided core concepts and carried out simulations for this manuscript. SA carried out implementations and drafted the manuscript. MANB drafted and proofread the manuscript and approved the final manuscript.
Corresponding author
Ethics declarations
Ethical approval and consent to participate
Not applicable.
Human ethics
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no potential competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Akan, T., Oskouei, A.G., Alp, S. et al. Brain magnetic resonance image (MRI) segmentation using multimodal optimization. Multimed Tools Appl (2024). https://doi.org/10.1007/s11042-024-19725-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11042-024-19725-4