Abstract
Glioblastoma is the most aggressive brain cancer with an unfavorable prognosis for patient survival. Glioma stem cells, a subpopulation of cancer cells, drive tumor initiation, self-renewal, and resistance to therapy and, together with the microenvironment, play a crucial role in glioblastoma maintenance and progression. Neurotransmitters such as noradrenaline, dopamine, and serotonin have contrasting effects on glioblastoma development, stimulating or inhibiting its progression depending on the cellular context and through their action on glioma stem cells, perhaps changing the epigenetic landscape. Recent studies have revealed that serotonin and dopamine induce chromatin modifications related to transcriptional plasticity in the mammalian brain and possibly in glioblastoma; however, this topic still needs to be explored because of its potential implications for glioblastoma treatment. Also, it is essential to consider that neurotransmitters’ effects depend on the tumor’s microenvironment since it can significantly influence the response and behavior of cancer cells. This review examines the possible role of neurotransmitters as regulators of glioblastoma development, focusing on their impact on the chromatin of glioma stem cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Glioblastoma (GB) is the most aggressive brain cancer, mainly derived from astrocytes or glioma stem cells [(GSCs), a type of cancer stem cell (CSC)] [1, 2], which are responsible for resistance to GB treatment [3,4,5]. GB patients have a poor survival prognosis, ranging from 12 to 15 months, despite the treatment, which includes surgical removal of the tumor followed by radiotherapy and chemotherapy with temozolomide [(TMZ), an alkylating agent] [4, 5]. This pathology is 1.6 times more frequent in men than in women [6,7,8,9].
GB tumors have been classified into four categories with particular gene signatures: (1) classical, overexpressing genes such as EGFR, FGFR3, PDGFA, AKT2, and NES; (2) neural, characterized by FBXO3, GABRB2, SNCG, and MBP overexpression; (3) proneural, with DLL3, NKX2-2, SOX2, ERBB3, and OLIG2 overexpression, and (4) mesenchymal, characterized by CASP1/4/5/8, ILR4, CHI3L, TRADD, TLR2/4, and RELB overexpression [10,11,12].
In malignant tumors of the central nervous system (CNS), neurotransmitters such as noradrenaline (NA), dopamine (DA), and serotonin (5-HT) have been identified [10, 13]. These neurotransmitters are possibly supplied to the tumor through innervation, given that several types of cells that constitute the GB tumors, such as glial cells (astrocytes, oligodendrocytes, and microglia), express synaptogenic proteins [14,15,16]. Supporting this hypothesis, the distribution of these neurotransmitters is higher in the peripheral regions of malignant brain tumors than in the central area [13]. Also, local synthesis in GB may exist due to the enzymes that participate in the catabolism of tyrosine and tryptophan have been detected [10].
In the GB context, neurotransmitters influence cellular metabolism through several pathways. The treatment of GB cells with NA inhibits glucose uptake and, at the same time, stimulates its release; these events are accompanied by the increase in 3′:5′-cyclic AMP and glycogen breakdown [17]. Meanwhile, DA increases the glucose reuptake in GB cell lines and induces apoptosis by stimulating the DRD2 receptor [18, 19]. Interestingly, the exposure of GB cells to fluoxetine (FLX), a selective 5-HT reuptake inhibitor (SSRI), remodels the sphingolipid distribution in the plasma membrane of GB cells, increasing the annexin V appearance, which is an indicator of intermediate stages of apoptosis [20]. Thus, these neurotransmitters play a pivotal role in the GB niche, regulating cellular proliferation, migration, apoptosis, metabolism, survival, differentiation, and angiogenesis [21].
The chromatin status plays a crucial role in the progression of GB [22]. The nucleosome is the basic structural unit of DNA packaging in eukaryotes, consisting of a core of histone proteins (H2A, H2B, H3, and H4), forming an octamer around 147 DNA base pairs. The amino acid residues at the N-terminus of the histones can be covalently modified by processes such as acylations, ubiquitin-like modifications, methylation, biotinylation, ADP ribosylation, and phosphorylation [23, 24]. Likewise, histone variants such as H2AX, H2AZ, and H3.3 regulate DNA repair, sex chromosome remodeling, nucleosome stability, and the activation of regulatory elements and gene expression [25].
In pediatric and young adult GB, mutations in H3F3A (a variant of H3 histone) are recurrent, with amino acid substitutions in histone tails K27M or G34R/G34V [26]. Moreover, several patient tumors harbor mutations of H3-K27M in oligodendroglial precursor-like cells, which are believed to be the origin of these tumors [27, 28].
Recently, it has been described that 5-HT and DA are chromatin modifiers [29, 30]. These neurotransmitters can be covalently bound to glutamine-5 or 19 on histone 3 (H3Q5/19) through the action of transglutaminase 2 (TGM2) [29, 31]. Post-translational modifications (PTMs) induce changes in gene expression, and in the case of 5-HT, they are permissive to gene expression, while in the case of DA, they alter gene transcription [29, 30]. Despite the widespread distribution of these neurotransmitters in the brain during normal and pathological conditions, their potential role in GB, especially in GSCs, remains poorly described. This review examines the role of NA, DA, and 5-HT as regulators of GB progression. Additionally, we explore their potential as chromatin modifiers in GSCs, a role minimally described in the literature.
Glioma stem cells
GBs have multiple GSC populations distributed in several tumor niches [32], including the perivascular niche (PVN), characterized by abnormal angiogenesis. The hypoxic niche, in which necrotic cells prevail, often results from vascular regression. This environment experiences an upregulation of hypoxia-inducible factor (HIF) [33], which is favorable for the self-renewal of GSCs [34]. Another niche is the invasive GB, which extends into surrounding tissues and is typically located at the border of normal brain parenchyma. It exhibits a more functional vasculature than PVN and possesses cellular heterogeneity [32, 33].
Furthermore, GSCs are characterized by the expression of several markers, including surface antigens, intermediate filaments, and transcription factors such as Prominin-1 (PROM1) or CD133, CD15, Nestin, and SOX2 [4, 35, 36]. Likewise, GSCs exhibit the capacity for self-renewal and differentiation into neurons, astrocytes, and oligodendrocytes [37, 38]. In addition, GSCs can express neural progenitor (SOX4, OLIG2, and ASCL1), astrocytic (GFAP, APOE, AQP4, CD44, CD9, and VIM), or neuronal marker genes (CD24, SOX11, and DCX) [39].
GSCs exhibit a proliferating or slow cycling/quiescent behavior in GB [40, 41]. In GB tumor slices, CD133 was expressed in 45% of the cells that were positive for Ki67, while Nestin was expressed in 20% [41]. Interestingly, in a forebrain organoid model generated with human induced pluripotent stem cells (hiPSCs), with tumors formed by injecting GB7 or COMI GB cell lines, Ki67 (a proliferation marker) was absent. At the same time, there was an increase in SOX2 expression, a marker for GSCs [42]. Regarding GSCs’ quiescent behavior, an F3 cell-surface receptor mRNA was recently identified as a conserved signature whose expression induction increases cellular self-renewal [43]. Furthermore, when dissociated tumoral organoid cells were injected into CD1-Nude mice brains, the resulting tumors exhibited a similar expression pattern of Ki67 and SOX2 as the inoculated organoids to the GB cell lines [42]. These findings support the notion that GSCs are a heterogeneous population with proliferating or quiescent behavior [44]; also, the GSCs share epigenetic mechanisms with neural stem cell (NSC) [45] characteristics that may play a role in maintaining their population despite GB treatments.
In addition, GSCs display changes in chromatin reorganization and remodeling, which are critical for maintenance and tumorigenicity. PTMs often drive these changes. In cultures of human GSCs, three clusters have been characterized by the presence of the trimethylation of Histone H3 Lysine 4 (H3K4me3) and Histone H3 Lysine 27 acetylation (H3K27ac) in active promoter regions of genes as P2RY1, MAPT, OLIG2, and ASCL1 (cluster 1), H3K27ac in promoter regions of CD44, ACVR1, RUNX1 and TGFBR2 (cluster 2), and H3K4me1 in the enhancer region of SOX5 (cluster 3). It is known that the expression of these genes confers GSCs’ highly proliferative and migratory characteristics (Table 1) [46,47,48].
Moreover, histone variants are incorporated into specific genomic regions throughout the cell cycle [49]. Regarding histone variants, it has been shown that H2A.Z is overexpressed in GSCs and is crucial for their maintenance [50].
The role of chromatin modification H2A.Z in neoplasias development: the prospect in glioblastoma
H2A.Z is an evolutionarily conserved H2A histone variant that shares 60% of its amino acid sequence with canonical H2A. It is related to nucleosome stability, either increasing or decreasing its mobility, which is influenced by histone variants and PTMs. Typically, this histone variant is found in promoters and enhancers. Also, the variant H2A.Z colocalizes with PTMs such as H4ac, H3K4me3, H3K4me1, and H3K27me3 [51].
H2A.Z is involved in the epithelial-mesenchymal-transition (EMT) [51,52,53] and has been related to the mechanisms involved in cancer development, such as mitosis and cellular metabolism [54, 55]. In the epithelioid pancreatic carcinoma PANC-1 cell line, the histone H2A.Z is enriched in the active gene promoters related to H3K4me3, H3K27ac marks and in gene enhancers associated with H3K27ac and H3K4me1 marks, as observed by chromatin immunoprecipitation with sequencing (ChIP-seq) analysis [56]. Pathway analysis of H2A.Z enrichment in enhancers identified genes related to focal adhesion, Wnt signaling, and HIF-1 transcriptional activity in hypoxia (among others). It is important to emphasize that HIF expression is associated with the presence of stemness markers, potentially contributing to the maintenance of the CSCs population and, consequently, cancer.
In the human LD611 cell line, derived from bladder cancer, the presence of H2A.Z is higher than in human UROtsa cells, a non-malignant uroepithelium line. When the 3–4, 5-(dimethyl-thyazol-2-yl)-2,5-diphenyltetrazolium (MTT) assay was performed, the existence of this histone was related to cellular proliferation in the LD611 cell line. In general, H2A.Z is located adjacent to the transcription start sites (TSS) of genes involved in cell cycle regulation, cellular development, cell growth, proliferation, cell death, and survival [57].
Analysis of the TCGA and Repository for Molecular Brain Neoplasia Data (REMBRANDT) datasets reveals an association between high expression of H2A.Z and poor prognosis for GB patient survival. Furthermore, a positive correlation between this histone and CD133+ or Nestin+ cells in GB has been observed. Studies conducted in a 3D model enriched with GSCs demonstrate that silencing of H2A.Z results in the formation of smaller-sized spheres. Similarly, smaller tumors are formed when cells from these spheres are injected into the brains of immunosuppressed NOD-scid IL2Rgammanull mice (NSG™), compared to those formed with cells expressing intact H2A.Z. The enrichment of H2A.Z in the epigenetic landscape of GSCs is associated with colocalization with H3K27ac, a marker related to chromatin transcriptional activity. This suggests that H2A.Z regulates the accessibility of transcriptional regulators to enhancer elements within GSC gene promoters [50].
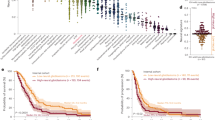
H2A.Z, as demonstrated by the DNA-barcoded nucleosome libraries technique, is susceptible to modification by tissue transglutaminase 2 (TGM2) [31], an enzyme responsible for depositing 5-HT and DA on histones [29,30,31]. This discovery prompts further investigation into whether this mechanism occurs in GSCs and its implications for their biology (Fig. 1).
Potential chromatin regulation of glioma stem cells by neurotransmitters. Glioblastoma tumors are integrated by distinct cellular types, including glioma stem cells (GSCs). It has been hypothesized that H2A.Z is susceptible to modification by 5-HT and DA, and possibly NA, which could stimulate the expression of glioma stemness markers. The figure was created with BioRender.com
This rationale encourages exploration into the role of neurotransmitters known for their chromatin remodeling function within the CNS. Such investigations may provide crucial insights into the potential regulatory mechanisms governing GSCs’ behavior, offering a foundation for targeted therapeutic interventions and advancing our comprehension of the intricate relationship between neural signaling and chromatin modulation in GB.
Neurotransmitters microenvironment in glioblastomas and their potential role in chromatin remodeling
Noradrenaline
NA enhances human GB-derived cell lines (U87 and U251) metabolism, migration, and invasion [58]. Moreover, the Human Tumor Metastasis RT2 Profiler PCR Arrays showed that exposure to NA diminishes the presence of matrix metallopeptidase-11 (MMP-11), an enzyme related to the breakdown of the extracellular matrix whose expression is higher in GB than in non-malignant-brain-tumors [59, 60].
In 2022, Wang and colleagues showed that NA stimulates EMT in GB cells, a mechanism for migration and metastasis. They analyzed The Glioma French, the Cancer Genome Atlas (TCGA), and the Chinese Glioma Genome Atlas datasets, reporting that the expression of Twist 1 [a gene that stimulates the stemness markers such as BMI1, CRIPTO1, DPPA2, KLF4, and SOX2 in CSC [61]] is related to a poor prognosis in GB patients. Interestingly, when the human-derived-GB cell lines (U251 and LN229) were cultured in the presence of NA, an increase in Twist 1 expression was observed (Fig. 2). In contrast, the knock-down of Twist 1 by shTwist 1 lentivirus diminishes the presence of N-cadherin, fibronectin 1, and vimentin, EMT markers [62].
Interestingly, in the context of the murine brain, it has been hypothesized that NA could be a chromatin modifier [63]. When hippocampal slices of male C57BL/6 strain mice brains were treated with NA, after dissecting the CA1 region and extracting the histones, it was observed that H3K14ac was associated with EMT [63]. Possibly, NA has a similar effect on chromatin structure in GB cells.
Dopamine
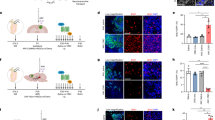
DA stimulates metabolic plasticity in GB, and it has been observed that the GB cells synthesize and secrete this catecholamine [17]. Using the intracranial injection of the HK-308-Luc or HK-374-Luc (which express luciferase) human GB cell lines for inducing tumor formation in immunosuppressed mice and followed by irradiation and subsequent subcutaneously (s.c.) administration of trifluoperazine [(TFP), an antagonist of D1 and D2 dopaminergic receptors], the tumor growth decreased and animal survival slightly improved. Interestingly, in the model mentioned above, radiation (8 Gy) stimulated the self-renewal of CSCs. Moreover, treatment with TFP reduces the expression of Sox2, Oct4, Klf4, c-Myc, and transcription factors involved in the stemness feature [64].
Glioma spheres derived from human GB tumors exposed to 7-OH-DPAT, an agonist of DA receptors, increased the number of cells. Interestingly, when the DA receptor type 2 (DRD2) was knocked down in GB cells, a decrease in the phosphorylation of STAT3, ERK, NANOG, and SOX2 expression was found [65], suggesting that one of the DA functions is to maintain a stem cell pool and therefore, participates in tumor recurrence.
In the patient-derived xenograft glioma line GBM43, the treatment with TMZ or radiation (2 Gy) increased the abundance of the H3K27ac mark in the promoter of DRD2 receptor gene. Conversely, orthotopic xenografts resulted in elevated expression of the human DRD2 receptor in the brains of nude (v/v) mice [17]. Following radiation treatment, the observed augmentation of H3K27ac presence in the DRD2 receptor promoter suggests a potential chromatin remodeling mechanism. It is well-established that transcriptionally active regions are more susceptible to impairment by ionizing radiation, and chromatin remodeling plays a crucial role in therapy [66].
The treatment of the human-derived GB cell line (T98 cell line) with an antagonist of the DA receptor D4R decreased cell viability compared to TMZ exposure [67]. Several studies suggest that DA acts as an autocrine regulator of GB development and a chromatin modifier by increasing the presence of H3K27ac, a permissive mark for gene expression (Fig. 2). The treatment with DA antagonists could become an alternative or coadjutant to conventional therapies for GB.
Serotonin
In 2019, for the first time, it was observed that 5-HT can be covalently bound to histones, specifically the H3K4me3 [29]. Furthermore, this “serotonylation” mechanism in histones is related to permissive gene expression and neuronal differentiation [29]. While in GB, the GSCs exhibit the capacity for aberrant neuronal differentiation [68], a process that could be influenced by 5-HT acting as a PTM; however, causal studies to prove this hypothesis are needed.
The function of 5-HT in GB progression has been controversial. An orthotopic model inoculated with GBM39 human cell line neurospheres in mice, treated with TMZ and FLX increased apoptosis [20, 69]. The overexpression of tryptophan hydroxylase 1 (TPH1), an enzyme limiting peripheral 5-HT synthesis [70], in two cellular lines derived from human glioblastomas (LN229 and T98G), increased proliferation, migration, and expression of genes involved in cellular adhesion, while diminishing apoptosis. Thus, TPH1 expression stimulates proliferation and motility in glioma cells [71]. It is worth mentioning that LN229 and T98G possess different karyotypes, with LN229 being XX and T98G being XY, but the effects of TPH1 overexpression are conserved between the cellular lines (Fig. 2).
Role of noradrenaline, dopamine, and serotonin in Glioblastoma progression. (A) Exposure of CSCs to NA enhances Twist 1 expression. This increment is related to the increase in SOX2, KLF4, CRIPTO1, DPPA2, and BMI1 expression, considered “stemness markers.” Also, the culture of human-derived glioblastoma cell lines LN229 and U251 with NA increases Twist 1 expression. (B) In glioblastoma orthotopic models, the use of TFP, an antagonist of dopamine receptors 1 (D1) and 2 (D2), is related to tumor size reduction and a decrease in SOX2, KLF4, and c-MYC expression, which are transcription factors characteristic of stemness. (C) The stimulus of TPH1 expression, the rate-limiting enzyme in serotonin synthesis, in the LN229 and T98G glioblastoma cell lines promotes cellular migration and proliferation. In contrast, exposure to TMZ, an alkylating drug commonly used in the treatment of glioblastoma, plus FLX, a selective inhibitor of serotonin reuptake, of tumors induced by neurospheres from the GBM39 glioblastoma cell line, resulted in the reduction of tumor size. The figure was created with BioRender.com
Metabolomic characterization of tissue samples from patients diagnosed with GB using Orbitrap secondary ion mass spectrometry (OrbiSIMS) technique shows that tumors are constituted by viable, necrotic, and non-cancerous areas where the distribution of metabolites across these regions is different. In turn, the 5-HT is present in all areas but is found in a more significant proportion in the necrotic section than in viable or non-cancerous regions [72]. Thus, 5-HT in GB could be involved in maintaining tumor cellular characteristics.
In GB human cell lines (LN229 and U251) treated with valeric acid, a partial agonist of the 5-HT5A receptor, there is an increase in cleaved caspase 3 (CC3), an apoptosis marker, and LC3, which participates in the autophagy process [67]. Valeric acid also reduces cellular metabolism and colony formation in vitro, while in vivo, it reduces tumor growth when luciferase-LN229 and luciferase-U251 cells were subcutaneously implanted into nude mice [73].
Similarly, exposure of human GB-derived cells (U87 cell line) to combined treatment with imatinib, an anti-cancer drug that is a tyrosine kinase inhibitor, and FLX reduces the presence of a phosphorylated form of AKT (pAKT) and the percentage of proliferating cells [74]. Likewise, culturing human GB-derived cell lines (U87 or U251) with fluvoxamine, an SSRI, reduces pAKT and decreases migration and invasion by inhibiting actin polymerization. Organizing actin in the cytoskeleton is essential to cancer cell migration and invasion [75]. Thus, further research is required on the role of the serotoninergic system in GB.
In GB, the gene expression profile recapitulates the typical pattern of expression seen in human fetal neural progenitor cells (NPC) and their response under the stimulus of the 5-HT-1A receptor [76]. Activation of this receptor with its agonist, p-amino phenethyl-m-trifluoromethylphenyl piperazine (PAPP), inhibits NPC and GB expansion. Hence, it is hypothesized that GB recapitulates the typical pattern of NPC expression, supporting the parallelism between stemness and GB progression.
Concluding and future directions
GSCs are recognized as the main impediment to an effective GB treatment due to their high capacity for self-renewal and therapy resistance. GSCs utilize various strategies to maintain their population within different niches of the GB, exhibiting proliferative and quiescent behavior. Alterations in the histone variant H2A.Z within GSC can profoundly impact their cellular programs, which are essential for maintaining their unique phenotype. These changes could be induced by neurotransmitters such as NA, DA, and 5-HT, which are widely distributed throughout the CNS under normal and pathological conditions, including GBs. Recent discoveries underscore the role of 5-HT and DA as chromatin modifiers and offer insight into potential involvement in regulating GSC behavior in the context of GB. However, rigorous studies are warranted to validate the influence of neurotransmitters, particularly regarding the H2A.Z histone.
Data availability
No datasets were generated or analysed during the current study.
Abbreviations
- 5-HT:
-
5-Hydroxitriptamine
- AKT2:
-
AKT serine/threonine kinase 2
- APOE:
-
Apolipoprotein E
- AQP4:
-
Aquaporin-4
- ASCL1:
-
Achaete-scute family bHLH transcription factor 1
- CASP1/4/5/8:
-
Cysteine-aspartic acid protease 1/4/5/8
- CC3:
-
Cleaved caspase 3
- CD15:
-
Cluster of Differentiation 15 or stage-specific embryonic antigen-1 (SSEA-1)
- CD24:
-
Cluster of Differentiation 24
- CD44:
-
Cluster of Differentiation 44
- CD9:
-
Cluster of Differentiation 9
- CHI3L:
-
Chitinase-3-like protein 1
- c-Myc:
-
Cellular myelocytomatosis oncogene
- CNS:
-
Central nervous system
- CRIPTO1:
-
Teratocarcinoma-Derived Growth Factor-1 (Cripto-1)
- DA:
-
Dopamine
- DCX:
-
Doublecortin
- DLL3:
-
Delta-like ligand 3
- DPPA2:
-
Developmental pluripotency-associated 2
- DRD2:
-
Dopamine receptor type 2
- EGFR:
-
Epidermal growth factor receptor
- EMT:
-
Epithelial-mesenchymal-transition
- ERBB3:
-
Receptor tyrosine-protein kinase 3
- ERK:
-
Extracellular signal-regulated kinase
- FGFR3:
-
Fibroblast growth factor receptor 3
- FLX:
-
Fluoxetine
- GB:
-
Glioblastoma
- GFAP:
-
Glial fibrillary acidic protein
- GSCs:
-
glioma stem cells
- H3K27ac:
-
Histone H3 Lysine 27 acetylation
- H3K4me1:
-
Monomethylation of Histone H3 at Lysine 4
- H3K4me3:
-
Trimethylation of Histone H3 at Lysine 4
- HIF:
-
Hypoxia-inducible factor
- ILR4:
-
Interleukin 4 receptor
- KLF4:
-
Krüppel-like factor 4
- LC3:
-
Microtubule-associated protein 1 A/1B-light chain 3
- MAPT:
-
Microtubule-associated protein tau
- MMP-11:
-
Matrix metallopeptidase-11
- NA:
-
Noradrenaline
- NANOG:
-
NANOG homeobox
- NES:
-
Nestin
- NKX2-2:
-
NK2 Homeobox 2
- NPC:
-
Neural progenitor cells
- OLIG2:
-
Oligodendrocyte transcription factor 2
- OLIG2:
-
Oligodendrocyte transcription factor 2
- P2RY1:
-
Purinergic receptor P2Y1
- pAKT:
-
Phosphorylated form of AKT
- PAPP:
-
p-amino phenethyl-m-trifluoromethylphenyl piperazine
- PDGFA:
-
Platelet-derived growth factor subunit A
- PROM1:
-
Prominin-1 or CD133
- PTMs:
-
Post-translational modifications
- PVN:
-
Perivascular niche
- RELB:
-
RELB Proto-Oncogene, NF-KB Subunit
- SOX11:
-
SRY-box transcription factor 11
- SOX2:
-
SRY-box transcription factor 2
- SOX4:
-
SRY-box transcription factor 4
- STAT3:
-
Signal transducer and activator of transcription 3
- TFP:
-
Trifluoperazine
- TGM2:
-
Transglutaminase 2
- TLR2:
-
Toll-Like Receptor 2
- TLR4:
-
Toll-Like Receptor 4
- TMZ:
-
Temozolomide
- TPH1:
-
Tryptophan hydroxylase 1
- TRADD:
-
Tumor necrosis factor receptor type 1-associated DEATH domain protein
- TSS:
-
Transcription start site
- VIM:
-
Vimentin
References
Das S, Srikanth M, Kessler JA (2008) Nat Clin Pract Neurol 4:427–435. https://doi.org/10.1038/ncpneuro0862
Geribaldi-Doldan N, Fernandez-Ponce C, Quiroz RN, Sanchez-Gomar I, Escorcia LG, Velasquez EP, Quiroz EN (2020) Front Oncol 10:603495. https://doi.org/10.3389/fonc.2020.603495
Walker FM, Sobral LM, Danis E, Sanford B, Donthula S, Balakrishnan I, Wang D, Pierce A, Karam SD, Kargar S, Serkova NJ, Foreman NK, Venkataraman S, Dowell R, Vibhakar R, Dahl NA (2024) Nat Commun 15:4616. https://doi.org/10.1038/s41467-024-48214-3
Bao S, Wu Q, McLendon RE, Hao Y, Shi Q, Hjelmeland AB, Dewhirst MW, Bigner DD, Rich JN (2006) Nature 444:756–760. https://doi.org/10.1038/nature05236
Huang M, Zhang D, Wu JY, Xing K, Yeo E, Li C, Zhang L, Holland E, Yao L, Qin L, Binder ZA, O’Rourke DM, Brem S, Koumenis C, Gong Y, Fan Y (2020) Sci Transl Med 12. https://doi.org/10.1126/scitranslmed.aay7522
Mauvais-Jarvis F, Bairey Merz N, Barnes PJ, Brinton RD, Carrero JJ, DeMeo DL, De Vries GJ, Epperson CN, Govindan R, Klein SL, Lonardo A, Maki PM, McCullough LD, Regitz-Zagrosek V, Regensteiner JG, Rubin JB, Sandberg K, Suzuki A (2020) Lancet 396:565–582. https://doi.org/10.1016/S0140-6736(20)31561-0
Yang W, Warrington NM, Taylor SJ, Whitmire P, Carrasco E, Singleton KW, Wu N, Lathia JD, Berens ME, Kim AH, Barnholtz-Sloan JS, Swanson KR, Luo J, Rubin JB (2019) Sci Transl Med 11. https://doi.org/10.1126/scitranslmed.aao5253
Carrano A, Juarez JJ, Incontri D, Ibarra A (2021) H Guerrero Cazares Cells 10. https://doi.org/10.3390/cells10071783
Pina-Medina AG, Diaz NF, Molina-Hernandez A, Mancilla-Herrera I, Camacho-Arroyo I (2020) Life Sci 249:117536. https://doi.org/10.1016/j.lfs.2020.117536
Lam KHB, Leon AJ, Hui W, Lee SC, Batruch I, Faust K, Klekner A, Hutoczki G, Koritzinsky M, Richer M, Djuric U, Diamandis P (2022) Nat Commun 13:116. https://doi.org/10.1038/s41467-021-27667-w
Puchalski RB, Shah N, Miller J, Dalley R, Nomura SR, Yoon JG, Smith KA, Lankerovich M, Bertagnolli D, Bickley K, Boe AF, Brouner K, Butler S, Caldejon S, Chapin M, Datta S, Dee N, Desta T, Dolbeare T, Dotson N, Ebbert A, Feng D, Feng X, Fisher M, Gee G, Goldy J, Gourley L, Gregor BW, Gu G, Hejazinia N, Hohmann J, Hothi P, Howard R, Joines K, Kriedberg A, Kuan L, Lau C, Lee F, Lee H, Lemon T, Long F, Mastan N, Mott E, Murthy C, Ngo K, Olson E, Reding M, Riley Z, Rosen D, Sandman D, Shapovalova N, Slaughterbeck CR, Sodt A, Stockdale G, Szafer A, Wakeman W, Wohnoutka PE, White SJ, Marsh D, Rostomily RC, Ng L, Dang C, Jones A, Keogh B, Gittleman HR, Barnholtz-Sloan JS, Cimino PJ, Uppin MS, Keene CD, Farrokhi FR, Lathia JD, Berens ME, Iavarone A, Bernard A, Lein E, Phillips JW, Rostad SW (2018) C. Cobbs, M.J. Hawrylycz and G.D. Foltz, Science 360, 660–663 https://doi.org/10.1126/science.aaf2666
Verhaak RG, Hoadley KA, Purdom E, Wang V, Qi Y, Wilkerson MD, Miller CR, Ding L, Golub T, Mesirov JP, Alexe G, Lawrence M, O’Kelly M, Tamayo P, Weir BA, Gabriel S, Winckler W, Gupta S, Jakkula L, Feiler HS, Hodgson JG, James CD, Sarkaria JN, Brennan C, Kahn A, Spellman PT, Wilson RK, Speed TP, Gray JW, Meyerson M, Getz G, Perou CM (2010) Hayes and N. Cancer Genome Atlas Research. Cancer Cell 17:98–110. https://doi.org/10.1016/j.ccr.2009.12.020
Ikeda Y, Nakazawa S (1984) Nihon Ika Daigaku Zasshi 51:132–133. https://doi.org/10.1272/jnms1923.51.132
Anastasaki C, Gao Y, Gutmann DH (2023) Dev Cell 58:81–93. https://doi.org/10.1016/j.devcel.2022.12.011
Krishna S, Choudhury A, Keough MB, Seo K, Ni L, Kakaizada S, Lee A, Aabedi A, Popova G, Lipkin B, Cao C, Nava Gonzales C, Sudharshan R, Egladyous A, Almeida N, Zhang Y, Molinaro AM, Venkatesh HS, Daniel AGS, Shamardani K, Hyer J, Chang EF, Findlay A, Phillips JJ, Nagarajan S, Raleigh DR, Brang D, Monje M, Hervey-Jumper SL (2023) Nature 617:599–607. https://doi.org/10.1038/s41586-023-06036-1
Venkatesh HS, Morishita W, Geraghty AC, Silverbush D, Gillespie SM, Arzt M, Tam LT, Espenel C, Ponnuswami A, Ni L, Woo PJ, Taylor KR, Agarwal A, Regev A, Brang D, Vogel H, Hervey-Jumper S, Bergles DE, Suva ML, Malenka RC, Monje M (2019) Nature 573:539–545. https://doi.org/10.1038/s41586-019-1563-y
Caragher SP, Shireman JM, Huang M, Miska J, Atashi F, Baisiwala S, Hong Park C, Saathoff MR, Warnke L, Xiao T, Lesniak MS, James CD, Meltzer H, Tryba AK, Ahmed AU (2019) J Neurosci 39:1982–1993. https://doi.org/10.1523/JNEUROSCI.1589-18.2018
Vo VTA, Kim S, Hua TNM, Oh J, Jeong Y (2022) Commun Biol 5:593. https://doi.org/10.1038/s42003-022-03538-y
Yang K, Xu R, Le W (2021) Oncol Rep 45. https://doi.org/10.3892/or.2021.8025
Bi J, Khan A, Tang J, Armando AM, Wu S, Zhang W, Gimple RC, Reed A, Jing H, Koga T, Wong IT, Gu Y, Miki S, Yang H, Prager B, Curtis EJ, Wainwright DA, Furnari FB, Rich JN, Cloughesy TF, Kornblum HI, Quehenberger O, Rzhetsky A, Cravatt BF, Mischel PS (2021) Cell Rep 37:109957. https://doi.org/10.1016/j.celrep.2021.109957
Bi J, Chowdhry S, Wu S, Zhang W, Masui K, Mischel PS (2020) Nat Rev Cancer 20:57–70. https://doi.org/10.1038/s41568-019-0226-5
Bagert JD, Mitchener MM, Patriotis AL, Dul BE, Wojcik F, Nacev BA, Feng L, Allis CD, Muir TW (2021) Nat Chem Biol 17:403–411. https://doi.org/10.1038/s41589-021-00738-1
Chan JC, Maze I (2020) Trends Biochem Sci 45:829–844. https://doi.org/10.1016/j.tibs.2020.05.009
Millan-Zambrano G, Burton A, Bannister AJ, Schneider R (2022) Nat Rev Genet 23:563–580. https://doi.org/10.1038/s41576-022-00468-7
Kurumizaka H, Kujirai T, Takizawa Y (2021) J Mol Biol 433:166678. https://doi.org/10.1016/j.jmb.2020.10.012
Schwartzentruber J, Korshunov A, Liu XY, Jones DT, Pfaff E, Jacob K, Sturm D, Fontebasso AM, Quang DA, Tonjes M, Hovestadt V, Albrecht S, Kool M, Nantel A, Konermann C, Lindroth A, Jager N, Rausch T, Ryzhova M, Korbel JO, Hielscher T, Hauser P, Garami M, Klekner A, Bognar L, Ebinger M, Schuhmann MU, Scheurlen W, Pekrun A, Fruhwald MC, Roggendorf W, Kramm C, Durken M, Atkinson J, Lepage P, Montpetit A, Zakrzewska M, Zakrzewski K, Liberski PP, Dong Z, Siegel P, Kulozik AE, Zapatka M, Guha A, Malkin D, Felsberg J, Reifenberger G, von Deimling A, Ichimura K, Collins VP, Witt H, Milde T, Witt O, Zhang C, Castelo-Branco P, Lichter P, Faury D, Tabori U, Plass C, Majewski J (2012) S.M. Pfister and N. Jabado, Nature 482, 226–231 https://doi.org/10.1038/nature10833
Liu I, Jiang L, Samuelsson ER, Marco Salas S, Beck A, Hack OA, Jeong D, Shaw ML, Englinger B, LaBelle J, Mire HM, Madlener S, Mayr L, Quezada MA, Trissal M, Panditharatna E, Ernst KJ, Vogelzang J, Gatesman TA, Halbert ME, Palova H, Pokorna P, Sterba J, Slaby O, Geyeregger R, Diaz A, Findlay IJ, Dun MD, Resnick A, Suva ML, Jones DTW, Agnihotri S, Svedlund J, Koschmann C, Haberler C, Czech T, Slavc I, Cotter JA, Ligon KL, Alexandrescu S, Yung WKA, Arrillaga-Romany I, Gojo J, Monje M, Nilsson M, Filbin MG (2022) Nat Genet 54:1881–1894. https://doi.org/10.1038/s41588-022-01236-3
Wu Y, Fletcher M, Gu Z, Wang Q, Costa B, Bertoni A, Man KH, Schlotter M, Felsberg J, Mangei J, Barbus M, Gaupel AC, Wang W, Weiss T, Eils R, Weller M, Liu H, Reifenberger G, Korshunov A, Angel P, Lichter P, Herrmann C, Radlwimmer B (2020) Nat Commun 11:6434. https://doi.org/10.1038/s41467-020-20225-w
Farrelly LA, Thompson RE, Zhao S, Lepack AE, Lyu Y, Bhanu NV, Zhang B, Loh YE, Ramakrishnan A, Vadodaria KC, Heard KJ, Erikson G, Nakadai T, Bastle RM, Lukasak BJ, Zebroski H 3rd, Alenina N, Bader M, Berton O, Roeder RG, Molina H, Gage FH, Shen L, Garcia BA, Li H, Muir TW, Maze I (2019) Nature 567:535–539. https://doi.org/10.1038/s41586-019-1024-7
Lepack AE, Werner CT, Stewart AF, Fulton SL, Zhong P, Farrelly LA, Smith ACW, Ramakrishnan A, Lyu Y, Bastle RM, Martin JA, Mitra S, O’Connor RM, Wang ZJ, Molina H, Turecki G, Shen L, Yan Z, Calipari ES, Dietz DM, Kenny PJ, Maze I (2020) Science 368:197–201. https://doi.org/10.1126/science.aaw8806
Lukasak BJ, Mitchener MM, Kong L, Dul BE, Lazarus CD, Ramakrishnan A, Ni J, Shen L, Maze I, Muir TW (2022) Proc Natl Acad Sci U S A 119. https://doi.org/10.1073/pnas.2208672119. e2208672119
Hambardzumyan D, Bergers G (2015) Trends Cancer 1:252–265. https://doi.org/10.1016/j.trecan.2015.10.009
Aderetti DA, Hira VVV, Molenaar RJ, van Noorden CJF (2018) Biochim Biophys Acta Rev Cancer 1869:346–354. https://doi.org/10.1016/j.bbcan.2018.04.008
Soeda A, Park M, Lee D, Mintz A, Androutsellis-Theotokis A, McKay RD, Engh J, Iwama T, Kunisada T, Kassam AB, Pollack IF, Park DM (2009) Oncogene 28:3949–3959. https://doi.org/10.1038/onc.2009.252
Benedetti V, Banfi F, Zaghi M, Moll-Diaz R, Massimino L, Argelich L, Bellini E, Bido S, Muggeo S, Ordazzo G, Mastrototaro G, Moneta M, Sessa A, Broccoli V (2022) Sci Adv 8:eabn3986. https://doi.org/10.1126/sciadv.abn3986
Ye C, Li H, Li Y, Zhang Y, Liu G, Mi H, Li H, Xiao Q, Niu L, Yu X (2022) iScience 25:104872. https://doi.org/10.1016/j.isci.2022.104872
Ricci-Vitiani L, Pallini R, Biffoni M, Todaro M, Invernici G, Cenci T, Maira G, Parati EA, Stassi G, Larocca LM (2010) R De Maria Nat 468:824–828. https://doi.org/10.1038/nature09557
Wang R, Chadalavada K, Wilshire J, Kowalik U, Hovinga KE, Geber A, Fligelman B, Leversha M, Brennan C, Tabar V (2010) Nature 468:829–833. https://doi.org/10.1038/nature09624
Couturier CP, Ayyadhury S, Le PU, Nadaf J, Monlong J, Riva G, Allache R, Baig S, Yan X, Bourgey M, Lee C, Wang YCD, Wee Yong V, Guiot MC, Najafabadi H, Misic B, Antel J, Bourque G, Ragoussis J, Petrecca K (2020) Nat Commun 11:3406. https://doi.org/10.1038/s41467-020-17186-5
Liau B.B., Sievers C., Donohue L.K., Gillespie S.M., Flavahan W.A., Miller T.E., Venteicher A.S., Hebert C.H., Carey C.D., Rodig S.J., Shareef S.J., Najm F.J., van Galen P., Wakimoto H., Cahill D.P., Rich J.N., Aster J.C., Suva M.L., Patel A.P., Bernstein B.E. (2017) Cell Stem Cell 20:233–246. https://doi.org/10.1016/j.stem.2016.11.003. e237
Rehfeld M, Matschke J, Hagel C, Willenborg K, Glatzel M, Bernreuther C (2021) J Cancer Res Clin Oncol 147:2969–2982. https://doi.org/10.1007/s00432-021-03704-5
Antonica F, Santomaso L, Pernici D, Petrucci L, Aiello G, Cutarelli A, Conti L, Romanel A, Miele E, Tebaldi T, Tiberi L (2022) Nat Commun 13:4767. https://doi.org/10.1038/s41467-022-32448-0
Xie XP, Laks DR, Sun D, Ganbold M, Wang Z, Pedraza AM, Bale T, Tabar V, Brennan C, Zhou X, Parada LF (2022) Dev Cell 57:32–46. https://doi.org/10.1016/j.devcel.2021.12.007. e38
Wang P, Zhao L, Gong S, Xiong S, Wang J, Zou D, Pan J, Deng Y, Yan Q, Wu N, Liao B (2021) Cell Death Dis 12:312. https://doi.org/10.1038/s41419-021-03598-8
Vinel C, Rosser G, Guglielmi L, Constantinou M, Pomella N, Zhang X, Boot JR, Jones TA, Millner TO, Dumas AA, Rakyan V, Rees J, Thompson JL, Vuononvirta J, Nadkarni S, El Assan T, Aley N, Lin YY, Liu P, Nelander S, Sheer D, Merry CLR, Marelli-Berg F, Brandner S, Marino S (2021) Nat Commun 12:6130. https://doi.org/10.1038/s41467-021-26297-6
Lu X, Maturi NP, Jarvius M, Yildirim I, Dang Y, Zhao L, Xie Y, Tan EJ, Xing P, Larsson R, Fryknas M, Uhrbom L, Chen X (2022) Nat Commun 13:2236. https://doi.org/10.1038/s41467-022-29912-2
Myers B.L., Brayer K.J., Paez-Beltran L.E., Keith M.S., Suzuki H., Newville J., Anderson R.H., Lo Y., Mertz C.M., Kollipara R., Borromeo M.D., Bachoo R.M., Johnson J.E., Vue T.Y. (2023) bioRxiv. https://doi.org/10.1101/2023.09.30.560206
Zhao K, Cui X, Wang Q, Fang C, Tan Y, Wang Y, Yi K, Yang C, You H, Shang R, Wang J, Kang C (2019) Cell Death Dis 10:877. https://doi.org/10.1038/s41419-019-2108-x
Stepniak K, Machnicka MA, Mieczkowski J, Macioszek A, Wojtas B, Gielniewski B, Poleszak K, Perycz M, Krol SK, Guzik R, Dabrowski MJ, Draminski M, Jardanowska M, Grabowicz I, Dziedzic A, Kranas H, Sienkiewicz K, Diamanti K, Kotulska K, Grajkowska W, Roszkowski M, Czernicki T, Marchel A, Komorowski J, Kaminska B, Wilczynski B (2021) Nat Commun 12:3621. https://doi.org/10.1038/s41467-021-23922-2
Yoon J, Grinchuk OV, Tirado-Magallanes R, Ngian ZK, Tay EXY, Chuah YH, Lee BWL, Feng J, Crasta KC, Ong CT, Benoukraf T, Ong DST (2022) Cell Death Differ 29:1379–1394. https://doi.org/10.1038/s41418-021-00926-5
Colino-Sanguino Y, Clark SJ, Valdes-Mora F (2022) Trends Genet 38:273–289. https://doi.org/10.1016/j.tig.2021.10.003
Domaschenz R, Kurscheid S, Nekrasov M, Han S, Tremethick DJ (2017) Cell Rep 21:943–952. https://doi.org/10.1016/j.celrep.2017.09.086
Roche J (2018) Cancers (Basel) 10. https://doi.org/10.3390/cancers10020052
Avila-Lopez PA, Guerrero G, Nunez-Martinez HN, Peralta-Alvarez CA, Hernandez-Montes G, Alvarez-Hilario LG, Herrera-Goepfert R, Albores-Saavedra J, Villegas-Sepulveda N, Cedillo-Barron L, Montes-Gomez AE, Vargas M, Schnoor M, Recillas-Targa F, Hernandez-Rivas R (2021) Oncogene 40:2065–2080. https://doi.org/10.1038/s41388-021-01664-1
Yang B, Tong R, Liu H, Wu J, Chen D, Xue Z, Ding C, Zhou L, Xie H, Wu J, Zheng S (2018) Int J Oncol 52:1235–1245. https://doi.org/10.3892/ijo.2018.4292
Avila-Lopez PA, Nunez-Martinez HN, Peralta-Alvarez CA, Martinez-Calvillo S, Recillas-Targa F, Hernandez-Rivas R (2022) Arch Med Res 53:840–858. https://doi.org/10.1016/j.arcmed.2022.11.010
Kim K, Punj V, Choi J, Heo K, Kim JM, Laird PW, An W (2013) Epigenetics Chromatin 6:34. https://doi.org/10.1186/1756-8935-6-34
Newburgh RW, Rosenberg RN (1972) Proc Natl Acad Sci U S A 69:1677–1680. https://doi.org/10.1073/pnas.69.7.1677
Stojic J, Hagemann C, Haas S, Herbold C, Kuhnel S, Gerngras S, Roggendorf W, Roosen K, Vince GH (2008) Neurosci Res 60:40–49. https://doi.org/10.1016/j.neures.2007.09.009
Zhong J, Shan W, Zuo Z (2021) Brain Res 1756:147280. https://doi.org/10.1016/j.brainres.2021.147280
Izadpanah MH, Forghanifard MM (2022) Genes (Basel) 13. https://doi.org/10.3390/genes13122369
Wang X, Wang Y, Xie F, Song ZT, Zhang ZQ, Zhao Y, Wang SD, Hu H, Zhang YS, Qian LJ (2022) BMC Cancer 22:213. https://doi.org/10.1186/s12885-022-09330-9
Maity S, Jarome TJ, Blair J, Lubin FD, Nguyen PV (2016) J Physiol 594:863–881. https://doi.org/10.1113/JP271432
Bhat K, Saki M, Vlashi E, Cheng F, Duhachek-Muggy S, Alli C, Yu G, Medina P, He L, Damoiseaux R, Pellegrini M, Zemke NR, Nghiemphu PL, Cloughesy TF, Liau LM, Kornblum HI, Pajonk F (2020) Proc Natl Acad Sci U S A 117:11085–11096. https://doi.org/10.1073/pnas.1920154117
Jeon HM, Oh YT, Shin YJ, Chang N, Kim D, Woo D, Yeup Y, Joo KM, Jo H, Yang H, Lee JK, Kang W, Sa J, Lee WJ, Hale J, Lathia JD, Purow B, Park MJ, Park JB, Nam DH, Lee J (2023) Neoplasia 39:100894. https://doi.org/10.1016/j.neo.2023.100894
Brambilla F, Garcia-Manteiga JM, Monteleone E, Hoelzen L, Zocchi A, Agresti A, Bianchi ME (2020) Nucleic Acids Res 48:8993–9006. https://doi.org/10.1093/nar/gkaa613
Pavletic P, Semeano A, Yano H, Bonifazi A, Giorgioni G, Piergentili A, Quaglia W, Sabbieti MG, Agas D, Santoni G, Pallini R, Ricci-Vitiani L, Sabato E, Vistoli G, Del Bello F (2022) J Med Chem 65:12124–12139. https://doi.org/10.1021/acs.jmedchem.2c00840
Beier CP, Rasmussen T, Dahlrot RH, Tenstad HB, Aaro JS, Sorensen MF, Heimisdottir SB, Sorensen MD, Svenningsen P, Riemenschneider MJ, Beier D, Kristensen BW (2018) Sci Rep 8:14965. https://doi.org/10.1038/s41598-018-33282-5
Rogakou EP, Nieves-Neira W, Boon C, Pommier Y, Bonner WM (2000) J Biol Chem 275:9390–9395. https://doi.org/10.1074/jbc.275.13.9390
McKinney J, Knappskog PM, Haavik J (2005) J Neurochem 92:311–320. https://doi.org/10.1111/j.1471-4159.2004.02850.x
Zhang J, Guo Z, Xie Q, Zhong C, Gao X, Yang Q (2022) BMC Cancer 22:457. https://doi.org/10.1186/s12885-022-09569-2
He W, Edney MK, Paine SML, Griffiths RL, Scurr DJ, Rahman R, Kim DH (2023) Anal Chem 95:5994–6001. https://doi.org/10.1021/acs.analchem.2c05807
Lu Q, Ding Y, Li Y, Lu Q (2020) Int J Biol Sci 16:2104–2115. https://doi.org/10.7150/ijbs.44906
Tzadok S, Beery E, Israeli M, Uziel O, Lahav M, Fenig E, Gil-Ad I, Weizman A, Nordenberg J (2010) Int J Oncol 37:1043–1051. https://doi.org/10.3892/ijo_00000756
Hayashi K, Michiue H, Yamada H, Takata K, Nakayama H, Wei FY, Fujimura A, Tazawa H, Asai A, Ogo N, Miyachi H, Nishiki T, Tomizawa K, Takei K, Matsui H (2016) Sci Rep 6:23372. https://doi.org/10.1038/srep23372
Diamandis P, Wildenhain J, Clarke ID, Sacher AG, Graham J, Bellows DS, Ling EK, Ward RJ, Jamieson LG, Tyers M, Dirks PB (2007). Chemical genetics reveals a complex functional ground state of neural stem cells. Nat Chem Biol 3(5) 268–273. https://doi.org/10.1038/nchembio873
Acknowledgements
This work was supported by Programa de Estancias Postdoctorales por México del Consejo Nacional de Humanidades, Ciencias y Tecnologías (CONAHCyT) to the postdoctoral fellow Jessica Romero-Reyes and by DGAPA-PAPIIT clave IN201823.
Funding
This research received no external funding.
Author information
Authors and Affiliations
Contributions
Conceptualization, JRR, NFD, and ICA.; writing-original draft preparation, JRR, NFD, and ICA.; writing-review and editing, JRR, NFD, ERVM, CCS, AMH, and ICA. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Ethical approval
Not applicable.
Consent for publication
Not applicable.
Informed consent
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Romero-Reyes, J., Vázquez-Martínez, E.R., Silva, CC. et al. Navigating glioblastoma complexity: the interplay of neurotransmitters and chromatin. Mol Biol Rep 51, 912 (2024). https://doi.org/10.1007/s11033-024-09853-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11033-024-09853-3