Abstract
Background
Synovial hyperplasia caused by rheumatoid arthritis (RA), an autoimmune inflammatory disease, leads to the destruction of the articular cartilage and bone. A member of the tumor necrosis factor superfamily, Lymphotoxin-related inducible ligand that competes for glycoprotein D binding to herpes virus entry mediator on T cells (LIGHT) has been shown to correlate with the pathogenesis of RA.
Methods
We used cDNA microarray analysis to compare the expression of genes in rheumatoid fibroblast-like synoviocytes with and without LIGHT stimulation.
Results
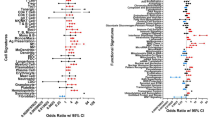
Significant changes in gene expression (P-values < 0.05 and fold change ≥ 2.0) were associated mainly with biological function categories of glycoprotein, glycosylation site as N-linked, plasma membrane part, integral to plasma membrane, intrinsic to plasma membrane, signal, plasma membrane, signal peptide, alternative splicing, and topological domain as extracellular.
Conclusions
Our results indicate that LIGHT may regulate the expression in RA-FLS of genes which are important in the differentiation of several cell types and in cellular functions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rheumatoid arthritis (RA) is causes chronic inflammation in synovial tissues, resulting in synovial hyperplasia. Pannus forms, invades articular cartilage and bone, and causes joint damage. Characteristics of the hyperplastic synovial tissues of patients with RA have been reported, including the expression of oncogenes, apoptosis resistance, and somatic mutations [1, 2].
Lymphotoxin-related inducible ligand that competes for glycoprotein D binding to herpes virus entry mediator on T cells (LIGHT) or TNFSF14 is a member of the tumor necrosis factor (TNF) superfamily and is expressed by several cell types including T cells, monocytes, granulocytes, immature dendritic cells [3], synovial fibroblasts, macrophages [4], and endothelial cells [5]. LIGHT mediates T-cell activation and contributes to inflammation and apoptosis by binding HEVN, LTβR, and decoy receptor 3 (DcR3) [6]. Serum levels of LIGHT and LIGHT-positive lymphocytes are increased in patients with RA [4, 7] and LIGHT can contribute to synovial hyperplasia and joint destruction in patients with RA [7].
DcR3/TR6/M68/TNFRSF6b is a TNF receptor that binds three ligands: Fas ligand (FasL), LIGHT, and tumor necrosis factor-like ligand 1 A (TL1A) [8]. DcR3 overexpression may contribute to tumor development by reducing available FasL, LIGHT [6], and TL1A [9], thus reducing their cytotoxic and regulatory effects. Previously, we showed that DcR3, which is overexpressed in rheumatoid arthritis fibroblast-like synoviocytes (RA-FLS) and further induced by TNFα, protects cells from Fas-induced apoptosis [10]. Further, we reported that DcR3 binds to membrane-bound TL1A and may play a role in the pathogenesis of RA [11,12,13].
In previous studies, DcR3 [14], TL1A [15], and FasL [16] regulation of gene expression in RA-FLS, as demonstrated using cDNA microarrays, suggest that DcR3 signaling, as well as that of its two ligands, TL1A and FasL, is involved in RA pathogenesis. However, the role of LIGHT in RA pathogenesis is unclear.
In this study, we used cDNA microarray analysis to investigate LIGHT-regulated gene expression in RA-FLS. Results indicate molecules with significant roles in the LIGHT-HEVN/LTβR/DcR3 signaling pathway and RA pathogenesis.
Methods
Synovial fibroblasts
RA-FLS were obtained from four patients during total knee arthroplasty who fulfilled the criteria for RA of the American College of Rheumatology (ACR) in 1987 [17] or the ACR/European Alliance of Associations for Rheumatology (EULAR) in 2010 [18]. Patients were women aged 75.5 ± 10.3 years. Their levels of C-reactive protein were 1.15 ± 2.0 mg/dl (range, 0.10–4.17 mg/dl). Drug therapy for RA included conventional synthetic disease modifying-anti-rheumatic drugs (csDMARDs): 100 mg/day of bucillamine (two patients) or 10 mg/day of leflunomide (one patient). The fourth patient was treated with prednisolone (oral; 3 mg/day). Patients were excluded from the study if they had been administered biological disease-modifying anti-rheumatic drugs (bDMARDs) or Janus kinase inhibitors (JAKi).
Patients provided informed written consent to participate in this study in accordance with the Declaration of Helsinki and the study was approved by the Kobe University Graduate School of Health Sciences Ethics Committee (approval no. 308). Synovial tissues were cut up, cells were dissociated in Dulbecco’s modified Eagle’s medium (#D5796, DMEM; Merck, Darmstadt, Germany) containing 0.2% collagenase (Merck) for 2 h at 37 °C in 5% CO2, and isolated cells were cultured in DMEM supplemented with 10% fetal bovine serum (#172,012, Merck) and 100 U/mL of penicillin (#45,397,387, Meiji Seika Pharma Co., Ltd., Tokyo, Japan) / streptomycin (#14,004,793, Meiji Seika Pharma Co., Ltd.) overnight at 37 °C in 5% CO2. Non-adherent cells were removed and adherent cells were provided fresh medium. Cells from passages 3 to 4 were used in this study [10].
LIGHT-regulated gene expression
Individual RA-FLS cell lines (2 × 106 cells/well of primary cultured RA-FLS) from patients 1–4 were incubated in either 1,000 ng/mL of recombinant human LIGHT protein (#664-LI/CF, R&D Systems, Minneapolis, MN) or OPTI-MEM medium (#31985-070, Thermo Fisher Scientific, Inc., Waltham, MA) for 12 h at 37 °C in 5% CO2. QIAshredder (#79,654, Qiagen, Hilden, Germany) and an RNeasy Mini kit (#74,104, Qiagen) were used to extract RNA from each cultured RA-FLS cell line, according to the manufacturer’s instructions.
Gene expression was detected using a microarray assay (Human Genome U133 Plus 2.0, GeneChip® 3’ Expression Array, Thermo Fisher Scientific), according to the manufacturer’s instructions.
Statistical analysis
Data are represented as the mean ± standard deviation unless otherwise indicated. Microarray analysis were performed using Avadis 3.3 Prophetic software (Strand Life Sciences, Bangalore, India) [19]. P-values < 0.05 in a paired t test were considered statistically significant and a fold change ≥ 2.0 in expression was used to identify differentially expressed genes. Genes were ordered into hierarchical clusters by distance (Euclidean algorithm method) and linkage (complete algorithm method).
Results
Microarray analysis (gene expression profiling of LIGHT-stimulated RA-FLS)
The expression of over 53,500 genes was assessed using microarray analysis and results demonstrated that the expression of 1843 genes in RA-FLS were regulated by LIGHT. These genes were identified using the advanced search page of the UniGene database of the National Center for Biotechnology Information (https://www.ncbi.nlm.nih.gov/gene/advanced). Of the 1843 genes, 1042 differentially upregulated genes and 801 differentially downregulated genes (P-value < 0.05 and fold change ≥ 2.0) were identified in the LIGHT-stimulated group compared with the control group. Of the 1042 differentially upregulated genes, the UniGene database was able to annotate 814 genes. The top 20 upregulated genes of the 814 annotated genes are shown in Table 1. Similarly, 593 of the 801differentially downregulated genes were annotated. The top 20 downregulated genes of the 593 annotated genes are shown in Table 2.
Functional annotation
The genes expressed differentially in response to LIGHT were classified by biological function (David Bioinformatics Database; https://david.ncifcrf.gov/). The 10 most significant categories of biological function were glycoprotein, glycosylation site as N-linked (GlcNAc…), plasma membrane part, integral to plasma membrane, intrinsic to plasma membrane, signal, plasma membrane, signal peptide, alternative splicing, and topological domain as extracellular (Table 3).
Discussion
Microarray analysis is a powerful method to investigate a wide variety of diseases, such as tumors [20], immunological diseases [21], and inflammatory diseases [22]. Previously, we used microarray analysis to characterize gene expression in DcR3 [14], TL1A [15], and FasL-stimulated [16] RA-FLS. Based on the gene expression profile of DcR3-stimulated RA-FLS, we examined in detail the expression and the function of IL-12B p40 [11], tryptophan hydroxylase 1 [13], and centrosomal protein 70 kDa [12] in DcR3-stimulated cells. In the gene expression profile of TL1A-stimulated RA-FLS, the expression of spectrin repeat-containing nuclear envelope 1, Fc receptor-like 2, PYD (pyrin domain)-containing 1, cell division cycle 45 homolog, signal transducer and activator of transcription 5B, and interferon regulatory factor 4 [15] were regulated by TL1A. In the gene expression profile of FasL-stimulated RA-FLS, FasL regulation of dual specificity phosphatase 6, epiregulin, interleukin 11, angiopoietin-like 7, protein inhibitor of activated STAT 2, and growth differentiation factor 5 [16] was identified.
In the present study, LIGHT regulation of baculoviral IAP repeat containing 3 (BIRC3), interleukin 7 receptor (IL7R), tumor necrosis factor receptor superfamily member 9 (TNFRSF9), synaptosomal-associated protein 25 kDa (SNAP25), and diaphanous-related formin 1 (DIAPH1) in RA-FLS was observed and classified into major functional clustering categories.
LIGHT upregulated the expression of BIRC3, which encodes cellular inhibitor of apoptosis protein 2 (cIAP2), to the greatest extent. cIAP2 is a component of nuclear factor kappaB and mitogen-activated protein kinase signaling pathways [23], and regulates cellular activities including differentiation, cytokine secretion, and cell death [24]. In RA, the therapeutic potential of cIAP2 is controversial. Previous reports have shown that cIAP2 deficiency is pro-inflammatory [25] and that antagonists against cIAP2 are anti-inflammatory [23]. In addition, antagonists against cIAP2 induce apoptosis in synovial fibroblasts [26].
IL7R expression was upregulated in the present study. IL7/IL7R signaling may play a role in various autoimmune or inflammatory diseases including multiple sclerosis, type 1 diabetes mellitus, RA, and ulcerative colitis. In patients with RA, IL7R expression is detected in monocytes, macrophages, synovial fibroblast cells, and endothelial cells [27, 28]. IL7/IL7R signaling may contribute to cell differentiation, such as that of T-cells, macrophages, endothelial cells, and osteoclasts, and to promote inflammation, angiogenesis, and joint destruction in RA pathogenesis [28].
Our results showed that TNFRSF9 expression was upregulated in response to LIGHT. TNFRSF9/CD137/4-1BB, an inducible costimulatory receptor, is expressed by monocytes and activated T and B cells. TNFRSF9 induces a signaling cascade in T cells that upregulates anti-apoptotic molecules, cytokine secretion, and enhanced effector function [29, 30]. TNFRSF9 signaling regulates the differentiation of Th17 cell through IL-6 expression in endothelial cells mediated by Akt and NF-kB pathways [31]. TNFRSF9 may play a role in autoimmune diseases [32] and immune response to infections [33], and may be a target molecule for cancer therapy [34]. Soluble TNFRSF9 is increased in RA patient sera [35], and the serum concentration of soluble TNFRSF9 and its ligand correlate with disease severity in RA [36].
SNAP25 was the third most downregulated gene observed in our analysis. SNAP25, a member of the soluble, N-ethylmaleimide-sensitive factor attachment protein receptor (SNARE) family, is important in neurotransmitter release and synaptic function [37]. In addition, SNAP25 is associated with the pathogenesis of cancers [38, 39] and some diseases affecting the nervous system and immune system in humans [40, 41].
DIAPH1 was downregulated in our analysis. DIAPH1 is a microtubule-binding protein that can nucleate actin filaments [42]. DIAPH1 is upregulated in laryngeal squamous cell carcinoma and may inhibit apoptosis in LSCC cells [43]. DIAPH1 knockdown promotes apoptosis in human glioblastoma cells [44].
In this study, we examined LIGHT regulation of gene expression in RA-FLS using a microarray assay. We recruited patients with a similar clinical background (never been treated with bDMARDs or JAKi and with severe knee joint destruction requiring total knee arthroplasty) to ensure sufficient sample homogeneity among the four samples.
The three ligands, TL1A, FasL, and LIGHT, bind the common decoy receptor, DcR3, and their activity is inhibited. Further study to understand the relationship between TL1A, FasL, and LIGHT regulation of gene expression should clarify the role of the TL1A/FasL/LIGHT/DcR3 signaling cascade in the pathogenesis of RA.
There are limitations to the present study. First, the study’s sample size is small. Second, microarray analysis demonstrates gene expression, but not mRNA or protein expression, which are required to confirm the expression of each gene in future studies.
Conclusions
The present study examines the regulation of gene expression in RA-FLS by LIGHT. Our results indicate that the gene expression in RA-FLS regulated by LIGHT may be important in the activation and function of lymphocytes, proliferation, apoptosis, cytokine secretion, and intracellular signaling. The actions of LIGHT may be pleiotropic in the pathogenesis of RA, by not only improving but also aggravating RA. Our results may help clarify the pathogenesis of RA and identify of new targets for RA treatment.
Data availability
The microarray assay datasets are available in the Gene Expression Omnibus (GEO) repository of the National Center for Biotechnology Information. The GEO series accession number of the datasets is GSE197057 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE197057).
Abbreviations
- RA:
-
Rheumatoid arthritis
- LIGHT:
-
Lymphotoxin-related inducible ligand that competes for glycoprotein D binding to herpes virus entry mediator on T cells
- TNF:
-
Tumor necrosis factor
- DcR3:
-
Decoy receptor 3
- FasL:
-
Fas ligand
- TL1A:
-
Tumor necrosis factor-like ligand 1 A
- RA-FLS:
-
Rheumatoid arthritis fibroblast-like synoviocytes
- ACR:
-
American College of Rheumatology
- EULAR:
-
European Alliance of Associations for Rheumatology
- csDMARDs:
-
Conventional synthetic disease modifying-anti-rheumatic drugs
- bDMARDs:
-
Biological disease-modifying anti-rheumatic drugs
- JAKi:
-
Janus kinase inhibitors
- DMEM:
-
Dulbecco’s modified Eagle’s medium
- BIRC3:
-
Baculoviral IAP repeat containing 3
- IL7R:
-
Interleukin 7 receptor
- TNFRSF9:
-
Tumor necrosis factor receptor superfamily member 9
- SNAP25:
-
Synaptosomal-associated protein 25 kDa
- DIAPH1:
-
Diaphanous-related formin 1
- cIAP2:
-
Cellular inhibitor of apoptosis protein 2
- SNARE:
-
Soluble N-ethylmaleimide-sensitive factor attachment protein receptor
References
Chou CT, Yang JS, Lee MR (2001) Apoptosis in rheumatoid arthritis–expression of Fas, Fas-L, p53, and Bcl-2 in rheumatoid synovial tissues. J Pathol 193:110–116
Yamanishi Y, Boyle DL, Rosengren S, Green DR, Zvaifler NJ, Firestein GS (2002) Regional analysis of p53 mutations in rheumatoid arthritis synovium. Proc Natl Acad Sci U S A 99:10025–10030
Mauri DN, Ebner R, Montgomery RI, Kochel KD, Cheung TC, Yu GL, Ruben S, Murphy M, Eisenberg RJ, Cohen GH, Spear PG, Ware CF (1998) LIGHT, a new member of the TNF superfamily, and lymphotoxin alpha are ligands for herpesvirus entry mediator. Immunity 8:21–30
Pierer M, Brentano F, Rethage J, Wagner U, Hantzschel H, Gay RE, Gay S, Kyburz D (2007) The TNF superfamily member LIGHT contributes to survival and activation of synovial fibroblasts in rheumatoid arthritis. Rheumatology (Oxford) 46:1063–1070
Celik S, Shankar V, Richter A, Hippe HJ, Akhavanpoor M, Bea F, Erbel C, Urban S, Blank N, Wambsganss N, Katus HA, Dengler TJ (2009) Proinflammatory and prothrombotic effects on human vascular endothelial cells of immune-cell-derived LIGHT. Eur J Med Res 14:147–156
Yu KY, Kwon B, Ni J, Zhai Y, Ebner R, Kwon BS (1999) A newly identified member of tumor necrosis factor receptor superfamily (TR6) suppresses LIGHT-mediated apoptosis. J Biol Chem 274:13733–13736
Edwards JR, Sun SG, Locklin R, Shipman CM, Adamopoulos IE, Athanasou NA, Sabokbar A (2006) LIGHT (TNFSF14), a novel mediator of bone resorption, is elevated in rheumatoid arthritis. Arthritis Rheum 54:1451–1462
Shi G, Wu Y, Zhang J, Wu J (2003) Death decoy receptor TR6/DcR3 inhibits T cell chemotaxis in vitro and in vivo. J Immunol 171:3407–3414
Migone TS, Zhang J, Luo X, Zhuang L, Chen C, Hu B, Hong JS, Perry JW, Chen SF, Zhou JX, Cho YH, Ullrich S, Kanakaraj P, Carrell J, Boyd E, Olsen HS, Hu G, Pukac L, Liu D, Ni J, Kim S, Gentz R, Feng P, Moore PA, Ruben SM, Wei P (2002) TL1A is a TNF-like ligand for DR3 and TR6/DcR3 and functions as a T cell costimulator. Immunity 16:479–492
Hayashi S, Miura Y, Nishiyama T, Mitani M, Tateishi K, Sakai Y, Hashiramoto A, Kurosaka M, Shiozawa S, Doita M (2007) Decoy receptor 3 expressed in rheumatoid synovial fibroblasts protects the cells against Fas-induced apoptosis. Arthritis Rheum 56:1067–1075
Fukuda K, Miura Y, Maeda T, Hayashi S, Kurosaka M (2016) Interleukin12B is upregulated by decoy receptor 3 in rheumatoid synovial fibroblasts. Mol Med Rep 13:3647–3652
Fukuda K, Miura Y, Maeda T, Hayashi S, Kuroda R (2018) Decoy receptor 3 down-regulates centrosomal protein 70 kDa specifically in rheumatoid synovial fibroblasts. Mod Rheumatol 28:287–292
Maeda T, Miura Y, Fukuda K, Hayashi S, Kurosaka M (2015) Decoy receptor 3 regulates the expression of tryptophan hydroxylase 1 in rheumatoid synovial fibroblasts. Mol Med Rep 12:5191–5196
Fukuda K, Miura Y, Maeda T, Takahashi M, Hayashi S, Kurosaka M (2013) Decoy receptor 3 regulates the expression of various genes in rheumatoid arthritis synovial fibroblasts. Int J Mol Med 32:910–916
Fukuda K, Miura Y, Maeda T, Hayashi S, Kuroda R (2019) Expression profiling of genes in rheumatoid fibroblast-like synoviocytes regulated by tumor necrosis factor-like ligand 1A using cDNA microarray analysis. Biomed Rep 1:1–5
Fukuda K, Miura Y, Maeda T, Hayashi S, Matsumoto T, Kuroda R (2021) Expression profiling of genes in rheumatoid fibroblast-like synoviocytes regulated by Fas ligand via cDNA microarray analysis. Exp Ther Med 22:1000
Arnett FC, Edworthy SM, Bloch DA, McShane DJ, Fries JF, Cooper NS, Healey LA, Kaplan SR, Liang MH, Luthra HS et al (1988) The American Rheumatism Association 1987 revised criteria for the classification of rheumatoid arthritis. Arthritis Rheum 31:315–324
Aletaha D, Neogi T, Silman AJ, Funovits J, Felson DT, Bingham CO 3rd, Birnbaum NS, Burmester GR, Bykerk VP, Cohen MD, Combe B, Costenbader KH, Dougados M, Emery P, Ferraccioli G, Hazes JM, Hobbs K, Huizinga TW, Kavanaugh A, Kay J, Kvien TK, Laing T, Mease P, Menard HA, Moreland LW, Naden RL, Pincus T, Smolen JS, Stanislawska-Biernat E, Symmons D, Tak PP, Upchurch KS, Vencovsky J, Wolfe F, Hawker G (2010) 2010 rheumatoid arthritis classification criteria: an American College of Rheumatology/European League against Rheumatism collaborative initiative. Ann Rheum Dis 69:1580–1588
Choi YJ, Yun HK (2016) Transcriptional profiles of Rhizobium vitis-inoculated and salicylic acid-treated ‘Tamnara’ grapevines based on microarray analysis. J Plant Biotechnol 43:37–48
Chang YC, Chen TC, Lee CT, Yang CY, Wang HW, Wang CC, Hsieh SL (2008) Epigenetic control of MHC class II expression in tumor-associated macrophages by decoy receptor 3. Blood 111:5054–5063
Whitney LW, Becker KG, Tresser NJ, Caballero-Ramos CI, Munson PJ, Prabhu VV, Trent JM, McFarland HF, Biddison WE (1999) Analysis of gene expression in mutiple sclerosis lesions using cDNA microarrays. Ann Neurol 46:425–428
Heller RA, Schena M, Chai A, Shalon D, Bedilion T, Gilmore J, Woolley DE, Davis RW (1997) Discovery and analysis of inflammatory disease-related genes using cDNA microarrays. Proc Natl Acad Sci U S A 94:2150–2155
Mayer BA, Rehberg M, Erhardt A, Wolf A, Reichel CA, Kracht M, Krombach F, Tiegs G, Zahler S, Vollmar AM, Furst R (2011) Inhibitor of apoptosis proteins as novel targets in inflammatory processes. Arterioscler Thromb Vasc Biol 31:2240–2250
Chesi M, Mirza NN, Garbitt VM, Sharik ME, Dueck AC, Asmann YW, Akhmetzyanova I, Kosiorek HE, Calcinotto A, Riggs DL, Keane N, Ahmann GJ, Morrison KM, Fonseca R, Lacy MQ, Dingli D, Kumar SK, Ailawadhi S, Dispenzieri A, Buadi F, Gertz MA, Reeder CB, Lin Y, Chanan-Khan AA, Stewart AK, Fooksman D, Bergsagel PL (2016) IAP antagonists induce anti-tumor immunity in multiple myeloma. Nat Med 22:1411–1420
Lawlor KE, Khan N, Mildenhall A, Gerlic M, Croker BA, D’Cruz AA, Hall C, Kaur Spall S, Anderton H, Masters SL, Rashidi M, Wicks IP, Alexander WS, Mitsuuchi Y, Benetatos CA, Condon SM, Wong WW, Silke J, Vaux DL, Vince JE (2015) RIPK3 promotes cell death and NLRP3 inflammasome activation in the absence of MLKL. Nat Commun 6:6282
Lattuada D, Casnici C, Crotta K, Seneci PF, Corradini C, Truzzi M, Ingegnoli F, Marelli O (2015) Proapoptotic activity of a monomeric smac mimetic on human fibroblast-like synoviocytes from patients with rheumatoid arthritis. Inflammation 38:102–109
Haas J, Korporal M, Schwarz A, Balint B, Wildemann B (2011) The interleukin-7 receptor alpha chain contributes to altered homeostasis of regulatory T cells in multiple sclerosis. Eur J Immunol 41:845–853
Pickens SR, Chamberlain ND, Volin MV, Pope RM, Talarico NE, Mandelin AM 2nd, Shahrara S (2011) Characterization of interleukin-7 and interleukin-7 receptor in the pathogenesis of rheumatoid arthritis. Arthritis Rheum 63:2884–2893
Chester C, Sanmamed MF, Wang J, Melero I (2018) Immunotherapy targeting 4-1BB: mechanistic rationale, clinical results, and future strategies. Blood 131:49–57
Lee HW, Park SJ, Choi BK, Kim HH, Nam KO, Kwon BS (2002) 4-1BB promotes the survival of CD8 + T lymphocytes by increasing expression of Bcl-xL and Bfl-1. J Immunol 169:4882–4888
Xu L, Geng T, Zang G, Bo L, Liang Y, Zhou H, Yan J (2020) Exosome derived from CD137-modified endothelial cells regulates the Th17 responses in atherosclerosis. J Cell Mol Med 24:4659–4667
Vinay DS, Choi JH, Kim JD, Choi BK, Kwon BS (2007) Role of endogenous 4-1BB in the development of systemic lupus erythematosus. Immunology 122:394–400
Tran VG, Nguyen NNZ, Kwon B (2021) CD137 Signaling is critical in fungal clearance during systemic Candida albicans infection. J Fungi (Basel) 7:382
Chin SM, Kimberlin CR, Roe-Zurz Z, Zhang P, Xu A, Liao-Chan S, Sen D, Nager AR, Oakdale NS, Brown C, Wang F, Yang Y, Lindquist K, Yeung YA, Salek-Ardakani S, Chaparro-Riggers J (2018) Structure of the 4-1BB/4-1BBL complex and distinct binding and functional properties of utomilumab and urelumab. Nat Commun 9:4679
Michel J, Langstein J, Hofstadter F, Schwarz H (1998) A soluble form of CD137 (ILA/4-1BB), a member of the TNF receptor family, is released by activated lymphocytes and is detectable in sera of patients with rheumatoid arthritis. Eur J Immunol 28:290–295
Jung HW, Choi SW, Choi JI, Kwon BS (2004) Serum concentrations of soluble 4-1BB and 4-1BB ligand correlated with the disease severity in rheumatoid arthritis. Exp Mol Med 36:13–22
Baker RW, Hughson FM (2016) Chaperoning SNARE assembly and disassembly. Nat Rev Mol Cell Biol 17:465–479
Di L, Gu M, Wu Y, Liu G, Zhang L, Li Y, Zhang W (2022) SNAP25 is a potential prognostic biomarker for prostate cancer. Cancer Cell Int 22:144
Yu X, Zhong P, Han Y, Huang Q, Wang J, Jia C, Lv Z (2019) Key candidate genes associated with BRAF(V600E) in papillary thyroid carcinoma on microarray analysis. J Cell Physiol 234:23369–23378
Yalin OO, Gokdogan Edgunlu T, Karakas Celik S, Emre U, Gunes T, Erdal Y, Eroglu Unal A (2019) Novel SNARE Complex Polymorphisms Associated with multiple sclerosis: signs of Synaptopathy in multiple sclerosis. Balkan Med J 36:174–178
Dagkonaki A, Avloniti M, Evangelidou M, Papazian I, Kanistras I, Tseveleki V, Lampros F, Tselios T, Jensen LT, Mobius W, Ruhwedel T, Androutsou ME, Matsoukas J, Anagnostouli M, Lassmann H, Probert L (2020) Mannan-MOG35-55 reverses experimental autoimmune encephalomyelitis, inducing a peripheral type 2 myeloid response, reducing CNS inflammation, and preserving axons in spinal cord lesions. Front Immunol 11:575451
Kovar DR, Harris ES, Mahaffy R, Higgs HN, Pollard TD (2006) Control of the assembly of ATP- and ADP-actin by formins and profilin. Cell 124:423–435
Yang J, Zhou L, Zhang Y, Zheng J, Zhou J, Wei Z, Zou J (2019) DIAPH1 Is Upregulated and Inhibits Cell Apoptosis through ATR/p53/Caspase-3 Signaling Pathway in Laryngeal Squamous Cell Carcinoma. Dis Markers 2019:6716472
Li Z, Xu Y, Zhang C, Liu X, Jiang L, Chen F (2014) Mammalian diaphanous-related formin 1 is required for motility and invadopodia formation in human U87 glioblastoma cells. Int J Mol Med 33:383–391
Acknowledgements
The authors appreciate the technical assistance of Ms. Maya Yasuda, Minako Nagata, and Kyoko Tanaka, laboratory technicians of the Department of Orthopedic Surgery, Kobe University Graduate School of Medicine.
Funding
Funding was provided by Grants-in-Aid for Scientific Research (KAKENHI) (grant nos. 15K10473 and 18K09106).
Open Access funding provided by Kobe University.
Author information
Authors and Affiliations
Contributions
KF conceived and designed the present study, was involved in data collection and analysis, confirmed the authenticity of all the raw data, and wrote the manuscript. YM conceived and designed the present study, was involved in data collection and analysis, and confirmed the authenticity of all the raw data. TosM and SH conceived and designed the present study, were involved in data collection and analysis. KK, YT, and TomM collected the data. RK conceived and designed the present study. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The Kobe University Graduate School of Health Sciences Ethics Committee approved the present study (approval no. 308). All participants provided written informed consent to participate in the present study.
Patient consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fukuda, K., Miura, Y., Maeda, T. et al. LIGHT regulated gene expression in rheumatoid synovial fibroblasts. Mol Biol Rep 51, 356 (2024). https://doi.org/10.1007/s11033-024-09311-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11033-024-09311-0