Abstract
Background
The accumulation of cadmium (Cd) in plants may compromise the growth and development of plants, thereby endangering human health through the food chain. Understanding how plants respond to Cd is important for breeding low-Cd rice cultivars.
Methods
In this study, the functions of 12-oxo-phytodienoic acid reductase 1 (OsOPR1) were predicted through bioinformatics analysis. The expression levels of OsOPR1 under Cd stress were analyzed by using qRT-PCR. Then, the role that OsOPR1 gene plays in Cd tolerance was studied in Cd-sensitive yeast strain (ycf1), and the Cd concentration of transgenic yeast was analyzed using inductively coupled plasma mass spectrometry (ICP-MS).
Results
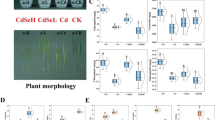
Bioinformatics analysis revealed that OsOPR1 was a protein with an Old yellow enzyme-like FMN (OYE_like_FMN) domain, and the cis-acting elements which regulate hormone synthesis or responding abiotic stress were abundant in the promoter region, which suggested that OsOPR1 may exhibit multifaceted biological functions. The expression pattern analysis showed that the expression levels of OsOPR1 were induced by Cd stress both in roots and roots of rice plants. However, the induced expression of OsOPR1 by Cd was more significant in the roots compared to that in roots. In addition, the overexpression of OsOPR1 improved the Cd tolerance of yeast cells by affecting the expression of antioxidant enzyme related genes and reducing Cd content in yeast cells.
Conclusion
Overall, these results suggested that OsOPR1 is a Cd-responsive gene and may has a potential for breeding low-Cd or Cd-tolerant rice cultivars and for phytoremediation of Cd-contaminated in farmland.








Similar content being viewed by others
Data availability
The datasets generated during and analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- ABA:
-
Abscisic acid
- ABRE:
-
Abscisic acid response element
- AOC:
-
Allene oxide cyclase
- A.thaliana :
-
Arabidopsis thaliana
- ARE:
-
Antioxidant response element
- Cd:
-
Cadmium
- E. coli :
-
Escherichia coli
- FMN:
-
Flavin mononucleotide
- GA:
-
Gibberellic acid
- GMQE:
-
Global model quality estimate
- H2O2 :
-
Hydrogen peroxide
- ICP- MS:
-
Inductively coupled plasma mass spectrometry
- JA:
-
Jasmonic acid
- MBS:
-
Drought stress response element
- MS:
-
Murashige and Skoog
- NJ:
-
Neighbor- Joining
- OPR:
-
The 12- oxo- phytodienoic acid reductas
- OYE_like_FMN:
-
Old yellow enzyme- like FMN
- OYEs:
-
Old yellow enzymes
- OPR3:
-
OPC- 8:0 by 12- oxophytodienoate reductase 3
- O3 :
-
Ozone
- PPI:
-
Protein- protein interaction
- PCR:
-
Polymerase Chain Reaction
- qRT- PCR:
-
Quantitative real time polymerase chain reaction
- SD- Ura:
-
Synthetic dropout - Uracil medium
- UBQ:
-
Ubiquitin
- UV:
-
Ultraviolet
- 4CLL4:
-
4- coumarate- CoA ligase- like4
- 4CLL8:
-
4- coumarate- CoA ligase- like 8
- 4CLL:
-
4- coumarate: CoA ligase- like
References
Gallego SM, Pena LB, Barcia RA, Azpilicueta CE, Lannone MF, Rosales EP, Zawoznik MS, Groppa MD, Benavides MP (2012) Unravelling cadmium toxicity and tolerance in plants: Insight into regulatory mechanisms. Environ Exp Bot 83:33–46. https://doi.org/10.1016/j.envexpbot.2012.04.006
Li X, Zhang X, Yang Y, Li B, Wu Y, Sun H, Yang Y (2016) Cadmium accumulation characteristics in turnip landraces from china and assessment of their phytoremediation potential for contaminated soils. Front Plant Sci 7:1862. https://doi.org/10.3389/fpls.2016.01862
Uraguchi S, Mori S, Kuramata M, Kawasaki A, Arao T, Ishikawa S (2009) Root-to-shoot Cd translocation via the xylem is the major process determining shoot and grain cadmium accumulation in rice. J Exp Bot 60(9):2677–2688. https://doi.org/10.1093/jxb/erp119
Shimbo S, Zhang ZW, Watanabe T, Nakatsuka H, Matsuda-Inoguchi N, Higashikawa K, Ikeda M (2001) Cadmium and lead contents in rice and other cereal products in Japan in 1998–2000. Sci Total Environ 281(1–3):165–175. https://doi.org/10.1016/S0048-9697(01)00844-0
Honda R, Swaddiwudhipong W, Nishijo M, Mahasakpan P, Teeyakasem W, Ruangyuttikarn W, Satarug S, Padungtod C, Nakagawa H (2010) Cadmium induced renal dysfunction among residents of rice farming area downstream from a zinc-mineralized belt in Thailand. Toxicol Lett 198(1):26–32. https://doi.org/10.1016/j.toxlet.2010.04.023
Wang B, Zhang M, Zhang J, Huang L, Chen X, Jiang M, Tan M (2020) Profiling of rice Cd-tolerant genes through yeast-based cDNA library survival screening. Plant Physiol Bioch 155:429–436. https://doi.org/10.1016/j.plaphy.2020.07.046
Riaz M, Kamran M, Rizwan M, Ali S, Parveen A, Malik Z, Wang X (2021) Cadmium uptake and translocation: selenium and silicon roles in Cd detoxification for the production of low Cd crops: a critical review. Chemosphere 273:129690. https://doi.org/10.1016/j.chemosphere.2021.129690
Emamverdian A, Ding Y, Mokhberdoran F, Xie Y (2015) Heavy metal stress and some mechanisms of plant defense response. Sci World J 2015:756120. https://doi.org/10.1155/2015/756120
Yu F, Liu K, Li M, Zhou Z, Deng H, Chen B (2013) Effects of cadmium on enzymatic and non-enzymatic antioxidative defences of rice (Oryza sativa L.). Int J Phytoremediat 15(6):513–521. https://doi.org/10.1080/15226514.2012.702807
Shah AA, Khan WU, Yasin NA, Akram W, Ahmad A, Abbas M, Ali A, Safdar MN (2020) Butanolide alleviated cadmium stress by improving plant growth, photosynthetic parameters and antioxidant defense system of brassica oleracea. Chemosphere 261:127728. https://doi.org/10.1016/j.chemosphere.2020.127728
Meng YT, Zhang XL, Wu Q, Shen RF, Zhu XF (2022) Transcription factor ANAC004 enhances Cd tolerance in Arabidopsis thaliana by regulating cell wall fixation, translocation and vacuolar detoxification of Cd, ABA accumulation and antioxidant capacity. J Hazard Mater 436:129121. https://doi.org/10.1016/j.jhazmat.2022.129121
Abbas T, Fan R, Hussain S, Sattar A, Khalid S, Butt M, Shahzad U, Muhammad Atif H, Batool M, Ullah S, Li Y, Al-Hashimi A, Elshikh MS, Al-Yahyai R (2022) Protective effect of jasmonic acid and potassium against cadmium stress in peas (Pisum sativum L.). Saudi J Biol Sci 29(44):2626–2633. https://doi.org/10.1016/j.sjbs.2021.12.051
Liu YK, Mu XH, Wang HX, Yan GJ (2012) A novel method for extracting green fractional vegetation cover from digital images. J Veg Sci 23(3):406–418. https://doi.org/10.1111/j.1654-1103.2011.01373.x
Mou Y, Liu Y, Tian S, Guo Q, Wen WC (2019) Genome-wide identification and characterization of the OPR gene family in wheat (Triticum aestivum L.). Int J Mol Sci 20(8):1914. https://doi.org/10.3390/ijms20081914
Raine AR, Scrutton NS, Mathews FS (1994) On the evolution of alternate core packing in eightfold beta/alpha-barrels. Protein Sci 3(10):1889–1892. https://doi.org/10.1002/pro.5560031028
Liu S, Sun R, Zhang X, Feng Z, Wei F, Zhao L, Zhang Y, Zhu L, Feng H, Zhu H (2020) Genome-wide analysis of OPR family genes in cotton identified a role for GhOPR9 in Verticillium dahliae resistance. Genes (Basel) 11(10):1134. https://doi.org/10.3390/genes11101134
Li W, Liu B, Yu L, Feng D, Wang H, Wang J (2009) Phylogenetic analysis, structural evolution and functional divergence of the 12-oxo-phytodienoate acid reductase gene family in plants. BMC Evol Biol 9:90. https://doi.org/10.1186/1471-2148-9-90
Asfaw KG, Liu Q, Eghbalian R, Purper S, Akaberi S, Dhakarey R, Munch SW, Wehl I, Brase S, Eiche E, Hause B, Bogeski I, Schepers U, Riemann M, Nick P (2022) The jasmonate biosynthesis gene OsOPR7 can mitigate salinity induced mitochondrial oxidative stress. Plant Sci 316:111156. https://doi.org/10.1016/j.plantsci.2021.111156
Jang S, Cho K, Shibato J, Han O, Iwahashi H, Tamogami S, Zargar SM, Kubo A, Masuo Y, Agrawal GK, Rakwal R (2009) Rice OsOPRs: Transcriptional profiling responses to diverse environmental stimuli and biochemical analysis of OsOPR1. J Plant Biol 52(3):229–243. https://doi.org/10.1007/s12374-009-9022-1
Da Costa MVJ, Ramegowda V, Sreeman S, Nataraja KN (2021) Targeted phytohormone profiling identifies potential regulators of spikelet sterility in rice under combined drought and heat stress. Int J Mol Sci 22(21):11690. https://doi.org/10.3390/ijms222111690
Agrawal GK, Jwa N-S, Shibato J, Han O, Iwahashi H, Rakwal R (2003) Diverse environmental cues transiently regulate OsOPR1 of the “octadecanoid pathway” revealing its importance in rice defense/stress and development. Biochem Bioph Res Co 310(4):1073–1082. https://doi.org/10.1016/j.bbrc.2003.09.123
Li W, Zhou F, Liu B, Feng D, He Y, Qi K, Wang H, Wang J (2011) Comparative characterization, expression pattern and function analysis of the 12-oxo-phytodienoic acid reductase gene family in rice. Plant Cell Rep 30(6):981–995. https://doi.org/10.1007/s00299-011-1002-5
Zhang J, Simmons C, Yalpani N, Crane V, Wilkinson H, Kolomiets M (2005) Genomic analysis of the 12-oxo-phytodienoic acid reductase gene family of Zea mays. Plant Mol Biol 59(2):323–343. https://doi.org/10.1007/s11103-005-8883-z
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33(7):1870–1874. https://doi.org/10.1093/molbev/msw054
Zhang XD, Sun JY, You YY, Song JB, Yang ZM (2018) Identification of Cd-responsive RNA helicase genes and expression of a putative BnRH 24 mediated by miR158 in canola (Brassica napus). Ecotoxicol Environ Saf 157:159–168. https://doi.org/10.1016/j.ecoenv.2018.03.081
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, Xia R (2020) TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13(8):1194–1202. https://doi.org/10.1016/j.molp.2020.06.009
Zaynab M, Ghramh HA, Sharif Y, Fiaz S, Al-Yahyai R, Alahdal MA, Qari SH, Hessini K, Huang X, Li S (2023) Expression profiling of DUF599 genes revealed their role in regulating abiotic stress response in solanum tuberosum. J King Saud Univ Sci 35(1):102368. https://doi.org/10.1016/j.jksus.2022.102368
Shi Y, Jiang WJ, Li MY, Jiang N, Huang YY, Wang MT, Du ZY, Chen J, Li JH, Wu LY, Zhong M, Yang J, Huang J (2023) Metallochaperone protein OsHIPP17 regulates the absorption and translocation of cadmium in rice (Oryza sativa L). Int J Biol Macromol 245:125607. https://doi.org/10.1016/j.ijbiomac.2023.125607
Luo J-S, Xiao Y, Yao J, Wu Z, Yang Y, Ismail AM, Zhang Z (2020) Overexpression of a defensin-like gene CAL2 enhances cadmium accumulation in plants. Front Plant Sci 11:217. https://doi.org/10.3389/fpls.2020.00217
Duan W, Xue B, He Y, Liao S, Li X, Li X, Liang Y-K (2023) Genome-wide identification and expression pattern analysis of dirigent members in the genus Oryza. Int J Mol Sci 24(8):7189. https://doi.org/10.3390/ijms24087189
Kim EJ, Hong WJ, Tun W, An G, Kim ST, Kim YJ, Jung KH (2021) Interaction of OsRopGEF3 protein with OsRac3 to regulate root hair elongation and reactive oxygen species formation in rice (Oryza sativa). Front Plant Sci 12:661352. https://doi.org/10.3389/fpls.2021.661352
Li Z-S, Szczypka M, Lu Y-P, Thiele D, Rea P (1996) The yeast cadmium factor protein (YCF1) is a vacuolar glutathione S-conjugate pump. J Biol Chem 271:6509–6517. https://doi.org/10.1074/jbc.271.11.6509
Hovland P, Flick J, Johnston M, Sclafani RA (1989) Galactose as a gratuitous inducer of GAL gene expression in yeasts growing on glucose. Gene 83(1):57–64. https://doi.org/10.1016/0378-1119(89)90403-4
Ruta LL, Lin YF, Kissen R, Nicolau I, Neagoe AD, Ghenea S, Bones AM, Farcasanu IC (2017) Anchoring plant metallothioneins to the inner face of the plasma membrane of cells leads to heavy metal accumulation. PLoS ONE 12(5):e0178393. https://doi.org/10.1371/journal.pone.017
Meng XX, Li WF, Shen RF, Lan P (2022) Ectopic expression of IMA small peptide genes confers tolerance to cadmium stress in Arabidopsis through activating the iron deficiency response. J Hazard Mater 422(1873–3336):126913. https://doi.org/10.1016/j.jhazmat.2021.126913
Tani T, Sobajima H, Okada K, Chujo T, Arimura S, Tsutsumi N, Nishimura M, Seto H, Nojiri H, Yamane H (2008) Identification of the OsOPR7 gene encoding 12-oxophytodienoate reductase involved in the biosynthesis of jasmonic acid in rice. Planta 227(3):517–526. https://doi.org/10.1007/s00425-007-0635-7
Marchler-Bauer A, Bo Y, Han L, He J, Lanczycki CJ, Lu S, Chitsaz F, Derbyshire MK, Geer RC, Gonzales NR, Gwadz M, Hurwitz DI, Lu F, Marchler GH, Song JS, Thanki N, Wang Z, Yamashita RA, Zhang D, Zheng C, Geer LY, Bryant SH (2016) CDD/SPARCLE: functional classification of proteins via subfamily domain architectures. Nucleic Acids Res 45(D1):D200–D203. https://doi.org/10.1093/nar/gkw1129
Wang Y, Yuan G, Yuan S, Duan W, Wang P, Bai J, Zhang F, Gao S, Zhang L, Zhao C (2016) TaOPR2 encodes a 12-oxo-phytodienoic acid reductase involved in the biosynthesis of jasmonic acid in wheat (Triticum aestivum L.). Biochem Bioph Res Co 470(1):233–238. https://doi.org/10.1016/j.bbrc.2016.01.043
Doug Chung DW, Le RKG (2013) Genome-wide analysis of gene expression. Encyclopedia of Biological Chemistry. Academic Press, pp 369–374
Chen A, Liu T, Wang Z, Chen X (2022) Plant root suberin: a layer of defence against biotic and abiotic stresses. Front Plant Sci 13:1056008. https://doi.org/10.3389/fpls.2022.1056008
Li T, Wang X, Elango D, Zhang W, Li M, Zhang F, Pan Q, Wu Y (2022) Genome-wide identification, phylogenetic and expression pattern analysis of Dof transcription factors in blueberry (Vaccinium corymbosum L.). PeerJ 10:e14087. https://doi.org/10.7717/peerj.14087
Song B, Luo X, Luo X, Liu Y, Niu Z, Zeng X (2022) Learning spatial structures of proteins improves protein-protein interaction prediction. Brief Bioinform 23(2):bbab558. https://doi.org/10.1093/bib/bbab558
Rabbani G, Baig MH, Ahmad K, Choi I (2018) Protein-protein interactions and their role in various diseases and their prediction techniques. Curr Protein Pept Sc 19(10):948–957. https://doi.org/10.2174/1389203718666170828122927
Sanders PM, Lee PY, Biesgen C, Boone JD, Beals TP, Weiler EW, Goldberg RB (2000) The arabidopsis DELAYED DEHISCENCE1 gene encodes an enzyme in the jasmonic acid synthesis pathway. Plant Cell 12(7):1041–1061. https://doi.org/10.1105/tpc.12.7.1041
Schaller F, Biesgen C, Mussig C, Altmann T, Weiler EW (2000) 12-Oxophytodienoate reductase 3 (OPR3) is the isoenzyme involved in jasmonate biosynthesis. Planta 210(6):979–984. https://doi.org/10.1007/s004250050706
Stintzi A, Browse J (2000) The Arabidopsis male-sterile mutant, opr3, lacks the 12-oxophytodienoic acid reductase required for jasmonate synthesis. P Natl Acad Sci USA 97(19):10625–10630. https://doi.org/10.1073/pnas.190264497
Koo AJK, Chung HS, Kobayashi Y, Howe GA (2006) Identification of a peroxisomal acyl-activating enzyme involved in the biosynthesis of jasmonic acid in arabidopsis. J Biol Chem 281(44):33511–33520. https://doi.org/10.1074/jbc.M607854200
Kienow L, Schneider K, Bartsch M, Stuible HP, Weng H, Miersch O, Wasternack C, Kombrink E (2008) Jasmonates meet fatty acids: functional analysis of a new acyl-coenzyme A synthetase family from Arabidopsis thaliana. J Exp Bot 59(2):403–419. https://doi.org/10.1093/jxb/erm325
Riemann M, Haga K, Shimizu T, Okada K, Ando S, Mochizuki S, Nishizawa Y, Yamanouchi U, Nick P, Yano M, Minami E, Takano M, Yamane H, Iino M (2013) Identification of rice Allene Oxide Cyclase mutants and the function of jasmonate for defence against Magnaporthe oryzae. Plant J 74(2):226–238. https://doi.org/10.1111/tpj.12115
de Nadal E, Ammerer G, Posas F (2011) Controlling gene expression in response to stress. Nat Rev Genet 12(12):833–845. https://doi.org/10.1038/nrg3055
Strucko T, Buron LD, Jarczynska ZD, Nødvig CS, Mølgaard L, Halkier BA, Mortensen UH (2017) CASCADE, a platform for controlled gene amplification for high, tunable and selection-free gene expression in yeast. Sci Rep 7:41431. https://doi.org/10.1038/srep41431
Wang FH, Qiao K, Shen YH, Wang H, Chai TY (2021) Characterization of the gene family encoding metal tolerance proteins in Triticum urartu: Phylogenetic, transcriptional, and functional analyses. Metallomics 13(7):mfab038. https://doi.org/10.1093/mtomcs/mfab038
Ali E, Hussain N, Shamsi IH, Jabeen Z, Siddiqui MH, Jiang LX (2018) Role of jasmonic acid in improving tolerance of rapeseed (Brassica napus L.) to Cd toxicity. J Zhejiang Univ Sci B 19(2):130–146. https://doi.org/10.1631/jzus.B1700191
Terra-Matos J, Teixeira MO, Santos-Pereira C, Noronha H, Domingues L, Sieiro C, Gerós H, Chaves SR, Sousa MJ, Côrte-Real M (2022) Saccharomyces cerevisiae cells lacking the zinc vacuolar transporter Zrt3 display improved ethanol productivity in lignocellulosic hydrolysates. J Fungi (Basel) 8(1):78. https://doi.org/10.3390/jof8010078
Martins D, English AM (2014) Catalase activity is stimulated by H2O2 in rich culture medium and is required for H2O2 resistance and adaptation in yeast. Redox Biol 2:308–313. https://doi.org/10.1016/j.redox.2013.12.019
Fukushima A, Iwasa M, Nakabayashi R, Kobayashi M, Nishizawa T, Okazaki Y, Saito K, Kusano M (2017) Effects of combined low glutathione with mild oxidative and low phosphorus stress on the metabolism of Arabidopsis thaliana. Front Plant Sci 8:1464. https://doi.org/10.3389/fpls.2017.01464
Khan ZH, Gao M, Qiu W, Islam MS, Song Z (2020) Mechanisms for cadmium adsorption by magnetic biochar composites in an aqueous solution. Chemosphere 246:125701. https://doi.org/10.1016/j.chemosphere.2019.125701
Fakhri Y, Bjørklund G, Bandpei AM, Chirumbolo S, Keramati H, Pouya RH, Asadi A, Amanidaz N, Sarafraz M, Sheikhmohammad A, Alipour M (2018) Concentrations of arsenic and lead in rice (Oryza sativa L.) in Iran: a systematic review and carcinogenic risk assessment. Food Che Toxicol 113:267–277. https://doi.org/10.1016/j.fct.2018.01.018
Huang K, Deng Y, Yuan W, Geng J, Wang G, Zou F (2020) Phospholipase D1 ameliorates apoptosis in chronic renal toxicity caused by low-dose cadmium exposure. Biomed Res Int 2020:7091053. https://doi.org/10.1155/2020/7091053
Wang J, Song L, Gong X, Xu J, Li M (2020) Functions of jasmonic acid in plant regulation and response to abiotic stress. Int J Molr Sci 21(4):1446. https://doi.org/10.3390/ijms21041446
Zhao Z, Kou M, Zhong R, Xia C, Christensen MJ, Zhang X (2021) Transcriptome analysis revealed plant hormone biosynthesis and response pathway modification by Epichloë gansuensis in Achnatherum inebrians under different soil moisture availability. J Fungi 7(8):640. https://doi.org/10.3390/jof7080640
Mosegaard S, Dipace G, Bross P, Carlsen J, Gregersen N, Olsen RKJ (2020) Riboflavin deficiency-implications for general human health and inborn errors of metabolism. Int J Mol Sci 21(11):3847. https://doi.org/10.3390/ijms21113847
Schall P, Marutschke L, Grimm B (2020) The flavoproteome of the model plant Arabidopsis thaliana. Int J Mol Sci 21(15):5371. https://doi.org/10.3390/ijms21155371
Yan Z, Chen J, Li X (2013) Methyl jasmonate as modulator of Cd toxicity in Capsicum frutescens var. fasciculatum seedlings. Ecotoi Environ Safe 98:203–209. https://doi.org/10.1016/j.ecoenv.2013.08.019
Ariño J, Ramos J, Sychrová H (2010) Alkali metal cation transport and homeostasis in yeasts. Microbiol Mol Biol Rev 74(1):95–120. https://doi.org/10.1128/MMBR.00042-09
Li H, Mu Y, Chang X, Li G, Dong Z, Sun J, Jin S, Wang X, Zhang L, Jin S (2022) Functional verification and screening of protein interacting with the slPHB3. Plant Signal Behav 17(1):2025678. https://doi.org/10.1080/15592324.2022.2025678
Cao HW, Zhao YN, Liu XS, Rono JK, Yang ZM (2022) A metal chaperone OsHIPP16 detoxifies cadmium by repressing its accumulation in rice crops. Environ Pollut 311:120058. https://doi.org/10.1016/j.envpol.2022.120058
Lage P, Sampaio-Marques B, Ludovico P, Mira NP, Mendes-Ferreira A (2019) Transcriptomic and chemogenomic analyses unveil the essential role of Com2-regulon in response and tolerance of Saccharomyces cerevisiae to stress induced by sulfur dioxide. Microbial Cell 6(11):509–523. https://doi.org/10.15698/mic2019.11.697
Harari Y, Zadok-Laviel S, Kupiec M (2017) Long telomeres do not affect cellular fitness in yeast. MBio 8:e01314–e01317. https://doi.org/10.1128/mBio.01314-17
Balaban B, Yılmaz Şahin U, Alkim C, Topaloğlu A, Kısakesen HC, Cakar ZP (2019) Evolutionary engineering of an iron-resistant saccharomyces cerevisiae mutant and its physiological and molecular characterization. Microorganisms 8:43. https://doi.org/10.3390/microorganisms8010043
Giese H, Azizan A, Kümmel A, Liao A, Peter CP, Fonseca JA, Hermann R, Duarte TM, Büchs J (2014) Liquid films on shake flask walls explain increasing maximum oxygen transfer capacities with elevating viscosity. Biotechnol Bioeng 111(2):295–308. https://doi.org/10.1002/bit.25015
Gupta A, Rao G (2003) A study of oxygen transfer in shake flasks using a non-invasive oxygen sensor. Biotechnol Bioeng 84(3):351–358. https://doi.org/10.1002/bit.10740
Lanclos VC, Coelho JT, Cleveland CS, Hyer AJ, McCallum MC, Savoie ER, Kosiba S, Thrash JC (2022) A cure for physiological characterization of bacterioplankton in liquid culture. J Microbiol Biol Edu 23(2):e00068–e00090. https://doi.org/10.1128/jmbe.00068-22
Gutsch A, Sergeant K, Keunen E, Prinsen E, Guerriero G, Renaut J, Hausman J-F, Cuypers A (2019) Does long-term cadmium exposure influence the composition of pectic polysaccharides in the cell wall of Medicago sativa stems? BMC Plant Biol 19(1):271. https://doi.org/10.1186/s12870-019-1859-y
Chen X, Wang H, Li X, Ma K, Zhan Y, Zeng F (2019) Molecular cloning and functional analysis of 4-Coumarate:CoA ligase 4 (4CL-like 1) from Fraxinus mandshurica and its role in abiotic stress tolerance and cell wall synthesis. BMC Plant Biol 19(1):231. https://doi.org/10.1186/s12870-019-1812-0
Eswaran N, Parameswaran S, Sathram B, Anantharaman B, Raja Krishna Kumar G, Tangirala SJ (2010) Yeast functional screen to identify genetic determinants capable of conferring abiotic stress tolerance in Jatropha curcas. BMC Biotechnol 10:23. https://doi.org/10.1186/1472-6750-10-23
Gechev T, Petrov V (2020) Reactive oxygen species and abiotic stress in plants. Int J Mol Sci 21(20):7433. https://doi.org/10.3390/ijms21207433
Zhang H-X, Zhu W-C, Feng X-H, Jin J-H, Wei A-M, Gong Z-H (2020) Transcription factor CaSBP12 negatively regulates salt stress tolerance in pepper (Capsicum annuum L.). Int J Mol Sci 21(2):444. https://doi.org/10.3390/ijms21020444
Yuan J, Sun K, Deng-Wang M-Y, Dai C-C (2016) The mechanism of ethylene signaling induced by endophytic fungus Gilmaniella sp. Al12 mediating sesquiterpenoids biosynthesis in Atractylodes lancea. Front Plant Sci 7:361. https://doi.org/10.3389/fpls.2016.00361
Adhikari B, Adhikari M, Ghimire B, Park G, Choi EH (2019) Cold atmospheric plasma-activated water irrigation induces defense hormone and gene expression in tomato seedlings. Sci Rep 9(1):16080. https://doi.org/10.1038/s41598-019-52646-z
Dong W, Wang M, Fei X, Quan T, Peng K, Xiao L, Xia G (2013) Wheat oxophytodienoate reductase gene TaOPR1 confers salinity tolerance via enhancement of abscisic acid signaling and reactive oxygen species scavenging. Plant Physiol 161(3):1217–1228. https://doi.org/10.1104/pp.112.211854
Funding
This work was supported by the National Natural Science Foundation of China (31870383) and Sichuan Provincial Financial “1 + 3” Project (2021ZYGG-002).
Author information
Authors and Affiliations
Contributions
YS: Conceptualization. RW, LW: Methodology. RW, ML: Formal analysis and investigation. LW, ZD: Writing—original draft preparation. YJ, JH, WJ: Writing—review and editing. JC, JH, BH: Funding acquisition. MZ, JY: Supervision. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Ethical approval
Not applicable.
Consent to participate
All authors have their consent to participate.
Consent for publication
All authors have their consent to publish this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wu, L., Wang, R., Li, M. et al. Functional analysis of a rice 12-oxo-phytodienoic acid reductase gene (OsOPR1) involved in Cd stress tolerance. Mol Biol Rep 51, 198 (2024). https://doi.org/10.1007/s11033-023-09159-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11033-023-09159-w