Abstract
Background
The Octopus vulgaris species complex consists of numerous morphologically similar but genetically distinct species. The current publicly available mitogenome of this species has been generated from a specimen collected from Tsukiji Fish Market, Tokyo, Japan. Octopus from the northwestern Pacific Ocean are now considered to be a separate species, Octopus sinensis. For this reason, we hypothesised that the current record of O. vulgaris was sequenced from a specimen of O. sinensis. Here, we sequenced the first complete mitogenome of a specimen of Octopus vulgaris sensu stricto that was collected from the species’ confirmed distribution areas in northeastern Atlantic.
Methods and results
The complete mitogenome was assembled de novo and annotated using 250 bp paired-end sequences. A single circular contig 15,655 bp in length with a mean read coverage of 1089 reads was reconstructed. The annotation pipeline identified 13 protein-coding genes (PCGs), 22 transfer RNAs (tRNA) and two ribosomal RNAs. A maximum likelihood phylogenetic tree recovered the assembled mitogenome as the sister taxon of a monophyletic group comprising O. sinensis and the previously published mitogenome of “O. vulgaris” from Japan. This confirms that the latter was a Japanese specimen of O. sinensis.
Conclusion
The mitogenome sequenced here is the first to be published for Octopus vulgaris sensu stricto. It represents an important first step in genetics-informed research on the evolution, conservation, and management of this commercially important species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The family Octopodidae d'Orbigny, 1840 consists of over 200 known species, many of which lack detailed taxonomic descriptions [1]. The taxonomy and phylogenetic relationships within this family have received much attention in recent years, with many cryptic species being identified across the globe [2,3,4,5]. A taxonomic group which has recently gained particular attention is the Octopus vulgaris species complex. Once thought to be a cosmopolitan species in temperate and tropical waters [6], O. vulgaris sensu stricto (s.s.) is now considered to be restricted to the Mediterranean and the northeastern Atlantic, while locations beyond this region are inhabited by morphologically similar but genetically distinct species [1, 7], some of which may still need to be formally described. As a case in point, the scientific name Octopus sinensis was recently reinstated for the East Asian common octopus previously referred to as O. vulgaris, based on a combination of molecular and morphological differences [8], while the Western Atlantic lineage was found to contain two more species, Octopus americanus [4] and Octopus insularis [3].
Octopus vulgaris is the most important commercially exploited octopus species worldwide, and off northwest Africa, it is the target of the world’s largest single-species octopus fishery. In 2010, total octopus catches for the northwest coast of Africa and Europe were 57,982 and 42,945 tonnes, respectively, consisting largely of O. vulgaris [9]. Currently, the poor state of octopus taxonomy is the single largest impediment to accurate catch statistics and comprehensive management plans for octopus species worldwide [9].
Mitochondrial DNA is a useful genetic marker for phylogenetic studies due to its maternal inheritance, conserved gene arrangement and comparatively high mutation rate [10]. Mitogenomes have been used to investigate deep phylogenetic relationships across different taxa [11], as well as delineate closely related species [12].
Within the O. vulgaris species complex, two complete mitochondrial genomes are currently available on the public databases: “O. vulgaris” (NC_006353.1 [13]) and O. sinensis (NC_052881.1 [14]). However, the original specimen used to create the first publicly available O. vulgaris mitogenome (NC_006353.1) was collected at the Tsukiji Fish Market, Tokyo, Japan in the early 2000s [13], before the scientific name O. sinensis was reinstated [8]. Since O. sinensis is distributed in the NW Pacific Ocean (from northern Japan to Taiwan), we hypothesised that this record in fact corresponds to O. sinensis. This suggests that despite its commercial and ecological importance, not a single complete mitogenome has yet been reconstructed for O. vulgaris s.s. The objective of the current study was to assemble, annotate, and describe the mitochondrial genome of O. vulgaris s.s. collected from its confirmed distribution range within northeastern Atlantic waters, and to examine its phylogenetic placement among closely related species. This study is important to improve our understanding of the taxonomic and phylogenetic relationships within the O. vulgaris species complex.
Material and methods
Sample collection, genomic library preparation and sequencing
The tissue sample of O. vulgaris s.s. was obtained from a specimen that was commercially caught near the Cies Islands in the Illas Atlanticas de Galicia National Park, Spain, in August 2022. DNA of high molecular weight was extracted from a small piece of muscle tissue using the QIAGEN DNeasy Blood & Tissue kit (Hilden, Germany). A genomic library was constructed from the extracted DNA using the NOVO kit (Novogene, Beijing, PRC). For this purpose, the DNA was first sheared into smaller fragments, and fragments of size ~ 350 bp were selected for the adaptor ligation step. The quality of the genomic library was checked using a combination of Qubit (Thermo Fisher Scientific, Waltham, USA), qPCR, and the DNA NGS 3 K assay (PerkinElmer, Waltham, USA). The quality-checked genomic library was sequenced on a NovaSeq 6000 SP platform (Illumina, San Diego, USA) using the paired-end 250 protocol.
Mitogenome assembly and annotation
The mitochondrial genome was assembled de novo using the GetOrganelle v.1.7 assembly pipeline [15]. The assembly parameters were set to their defaults, except for the kmer values, which were set for a combination of the following: 21, 45, 65, 85 and 105. The assembled mitogenome was then submitted to the MITOS Web Server [16] for annotation. The predicted gene boundaries were manually adjusted in MEGA11 [17] using the mitogenomes of the previously published “O. vulgaris” from Japan (NC_006353.1) and O. sinensis from China (NC_052881.1) as template references. The nucleotide composition of the complete mitogenome was calculated manually using the formulas AT-skew = (A − T)/(A + T), and GC-skew = (G − C)/(G + C) [18]. The annotated mitogenome was visualised in Chloroplot [19].
Maximum likelihood phylogenomic analysis
To reconstruct the phylogenetic relationships between the mitogenome of European O. vulgaris s.s. and those from other octopus taxa, the assembled mitochondrial genome was blast-searched against the NCBI nucleotide database. The complete mitogenomes of eight closely related species of octopus, as well as an outgroup species, Tremoctopus violaceus, were retrieved for phylogenetic analysis. The Ezsplit tool [20] was used to extract the sequences of all 13 protein coding genes (PCG) from the NCBI database, each of which was aligned separately using the codon alignment option in ClustalW with default setting [21].
A maximum likelihood phylogenetic tree was then constructed using IQ-TREE [22]. The most suitable evolutionary model was identified with ModelFinder [23]. Branch support was assessed using the ultrafast bootstrap analysis [24] based on 1000 bootstrap alignments. All other parameters were set to default. The resulting tree was visualised in Figtree v.1.4.3 [http://tree.bio.ed.ac.uk/software/figtree/].
Results and discussion
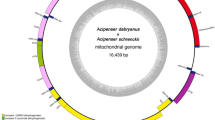
In total, the sequencing runs produced 10.4 million paired-end sequences with an average Phred quality score of 36. De novo assembly of the sequences produced a single circular contig 15,655 bp in length, with an average coverage of 1089, and a GC content of 25% (Fig. 1). The mitochondrial genome has a positive AT skew (0.096) and a positive CG skew (0.392) with base frequencies of A = 41.1%, C = 17.4%, G = 7.6% and T = 33.9%. The MITOS annotation pipeline identified 13 protein-coding genes (PCGs), 22 transfer RNAs (tRNA) and two ribosomal RNAs (rRNA) consistent with those reported from other octopus species. The plus strand contained seven PCGs (atp6, atp8, cox1, cox2, cox3, nad2, nad3) and eight tRNAs while the minus strand contained 6 PCGs (cytb, nad1, nad4, nad4l, nad5, nad6), 14 tRNAs and 2 rRNAs. The PCGs start with ATG, except for nad4, which has ATA as its start codon. The annotation pipeline also identified a single long intergenic sequence of approximately 645 bp between trnE(gaa) and cox3, which corresponds to the control region. Several (n = 21) shorter intergenic sequences ranging in length from 1 to 52 bp were also found.
The NCBI blast results showed European Octopus vulgaris s.s. to be most closely related to the only two other representatives of the O. vulgaris species complex with published mitogenomes. These are O. sinensis from China (NC_052881.1) and “O. vulgaris” from Japan (NC_006353.1), with 96.33% and 96.25% identity, respectively. Among other publicly available complete mitochondrial genomes, the next closest matches were O. bimaculoides (NC_029723.1) and O. mimus (NC_044093.1), each with roughly 85% identity.
The maximum likelihood phylogenetic tree confirmed, with 100% bootstrap support, that the assembled mitogenome is distinct from that of both O. sinensis from China and from the Japanese specimen [6] that has been deposited in the NCBI database as O. vulgaris (NC_006353.1), but it forms a monophyletic group with these specimens (Fig. 2). A blast search of the Japanese specimen showed 99.85% identity to the mitogenome of O. sinensis from China [14], but only 96.25% to that of the European O. vulgaris s.s. generated in this study. This confirms our hypothesis that the mitochondrial sequence from the Japanese specimen represents an additional mitogenome of O. sinensis. The present study thus reports the first ever mitogenome of O. vulgaris s.s.
A maximum likelihood phylogenetic tree reconstructed using protein coding genes under the TVM + F + I + G4 model, with a proportion of invariable sites of 0.51 and a gamma distribution shape parameter (α) of 1.54. The new mitogenome produced from the European specimen is shown in green, and that of the misidentified specimen that represents a second specimen of Octopus sinensis is shown in red. Bootstrap support values are shown to the left of the nodes. (Color fogure online)
Conclusion
Recent advances in DNA sequencing technology and high throughput computation have made it possible to assemble large numbers of mitogenomes. However, this information can only be useful in resolving taxonomic uncertainties when the specimens were correctly identified [25]; to this end, existing records need to be revisited to reflect the latest developments in how species are classified.
The phylogenetic relationships within the family Octopodidae are an ongoing topic of investigation. Mitogenomic evidence in this study shows that the current record of an “O. vulgaris” mitogenome in the NCBI database represents mitochondrial sequences from O. sinensis. Overall, our findings contribute towards improving our understanding of octopus phylogeny and taxonomy, which can ultimately inform fisheries management and improve the accuracy of catch statistics. The generation of additional mitogenomes from the broader geographical region inhabited by species within the O. vulgaris species complex will further enhance this field of knowledge.
Data availability
The annotated mitogenome of Octopus vulgaris s.s. has been submitted to the NCBI nucleotide database under accession number OR353425.
References
Norman MD, Finn JK, Hochberg FG (2013) Family Octopodidae. In: Jereb P, Roper CFE, Norman MD, Finn JK (eds) Cephalopods of the world. An annotated and illustrated catalogue of cephalopod species known to date. Volume 3. Octopods and Vampire Squids. FAO Species Catalogue for Fishery Purposes. No. 4, Vol. 3. FAO, Rome, pp 36–215
Söller R, Warnke K, Saint-Paul U, Blohm D (2000) Sequence divergence of mitochondrial DNA indicates cryptic biodiversity in Octopus vulgaris and supports the taxonomic distinctiveness of Octopus mimus (Cephalopoda: Octopodidae). Mar Biol 136:29–35. https://doi.org/10.1007/s002270050004
Leite TS, Haimovici M, Molina W, Warnke K (2008) Morphological and genetic description of Octopus insularis, a new cryptic species in the Octopus vulgaris complex (Cephalopoda: Octopodidae) from the tropical Southwestern Atlantic. J Molluscan Stud 74:63–74. https://doi.org/10.1093/mollus/eym050
Avendaño O, Roura Á, Cedillo-Robles CE, González ÁF, Rodríguez-Canul R, Velázquez-Abunader I, Guerra Á (2020) Octopus americanus: a cryptic species of the O. vulgaris species complex redescribed from the Caribbean. Aquat Ecol 4:909–925. https://doi.org/10.1007/s10452-020-09778-6
Amor MD, Hart AM (2021) Octopus djinda (Cephalopoda: Octopodidae): a new member of the Octopus vulgaris group from southwest Australia. Zootaxa 5061:145–156. https://doi.org/10.11646/zootaxa.5061.1.7
Warnke K, Söller R, Blohm D, Saint-Paul U (2004) A new look at geographic and phylogenetic relationships within the species group surrounding Octopus vulgaris (Mollusca, Cephalopoda): indications of very wide distribution from mitochondrial DNA sequences. J Zool Syst Evol Res 42:306–312. https://doi.org/10.1111/j.1439-0469.2004.00277.x
Amor MD, Norman MD, Roura A, Leite TS, Gleadall IG, Reid A, Perales-Raya C, Lu CC, Silvey CJ, Vidal EAG, Hochberg FG, Zheng X, Strugnell JM (2017) Morphological assessment of the Octopus vulgaris species complex evaluated in light of molecular-based phylogenetic inferences. Zool Scr 46:275–288. https://doi.org/10.1111/zsc.12207
Gleadall IG (2016) Octopus sinensis d’Orbigny, 1841 (Cephalopoda: Octopodidae): valid species name for the commercially valuable East Asian common octopus. Species Divers 21:31–42. https://doi.org/10.12782/sd.21.1.031
Jereb P, Roper CFE, Norman M, Finn JK (2014) Cephalopods of the world. An annotated and illustrated catalogue of cephalopod species known to date. Volume 3. Octopods and Vampire squid. FAO species catalogue for fisheries purposes. No. 4, Vol. 3. Rome, FAO. ISBN 978-92-5-107989-8
Moritz C, Dowling TE, Brown WM (1987) Evolution of animal mitochondrial DNA: relevance for population biology and systematics. Annu Rev Ecol Syst 18:269–292. https://doi.org/10.1146/annurev.es.18.110187.001413
Uribe JE, Zardoya R (2017) Revisiting the phylogeny of Cephalopoda using complete mitochondrial genomes. J Molluscan Stud 83:133–144. https://doi.org/10.1093/mollus/eyw052
Monsanto DM, van Vuuren BJ, Jagatap H, Jooste CM, Janion-Scheepers C, Teske PR, Emami-Khoyi A (2019) The complete mitogenome of the springtail Cryptopygus antarcticus travei provides evidence for speciation in the Sub-Antarctic region. Mitochondrial DNA B 4:1195–1197. https://doi.org/10.1080/23802359.2019.1591219
Yokobori SI, Fukuda N, Nakamura M, Aoyama T, Oshima T (2004) Long-term conservation of six duplicated structural genes in cephalopod mitochondrial genomes. Mol Biol Evol 21:2034–2046. https://doi.org/10.1093/molbev/msh227
Li F, Liu Y, Qin B, Bian L, Ge J, Chang Q, Liu H, Chen S (2021) Sequence and phylogenetic analysis of the mitochondrial genome for the East Asian common octopus, Octopus sinensis (Octopodidae: Octopoda). Mitochondrial DNA B 6:2120–2122. https://doi.org/10.1080/23802359.2021.1944360
Jin JJ, Yu WB, Yang JB, Song Y, Depamphilis CW, Yi TS, Li DZ (2020) GetOrganelle: a fast and versatile toolkit for accurate de novo assembly of organelle genomes. Genome Biol 21:1–31. https://doi.org/10.1186/s13059-020-02154-5
Bernt M, Donath A, Jühling F, Externbrink F, Florentz C, Fritzsch G, Pütz J, Middendorf M, Stadler PF (2013) MITOS: improved de novo metazoan mitochondrial genome annotation. Mol Phylogenet Evol 69:313–319. https://doi.org/10.1016/j.ympev.2012.08.023
Tamura K, Stecher G, Kumar S (2021) MEGA11: molecular evolutionary genetics analysis version 11. Mol Biol Evol 38:3022–3027. https://doi.org/10.1093/molbev/msab120
Perna NT, Kocher TD (1995) Patterns of nucleotide composition at fourfold degenerate sites of animal mitochondrial genomes. J Mol Evol 4:353–358. https://doi.org/10.1007/BF00186547
Zheng S, Poczai P, Hyvönen J, Tang J, Amiryousefi A (2020) Chloroplot: an online program for the versatile plotting of organelle genomes. Front Genet. https://doi.org/10.3389/fgene.2020.576124
Cucini C, Leo C, Iannotti N, Boschi S, Brunetti C, Pons J, Fanciulli PP, Frati F, Carapelli A, Nardi F (2021) EZmito: a simple and fast tool for multiple mitogenome analyses. Mitochondrial DNA B 6:1101–1109. https://doi.org/10.1080/23802359.2021.1899865
Higgins D, Thompson J, Gibson T, Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Nguyen L, Schmidt HA, von Haeseler A, Minh BQ (2015) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum likelihood phylogenies. Mol Biol Evol 32:268–274. https://doi.org/10.1093/molbev/msu300
Kalyaanamoorthy S, Minh BQ, Wong TKF, von Haeseler A, Jermiin LS (2017) ModelFinder: fast model selection for accurate phylogenetic estimates. Nat Methods 14:587–589
Hoang DT, Chernomor O, von Haeseler A, Minh BQ, Vinh LS (2017) UFBoot2: improving the ultrafast bootstrap approximation. Mol Biol Evol 35:518–522. https://doi.org/10.1093/molbev/msx281
Teske PR (2021) Mitochondrial genome announcements need to consider existing short sequences from closely related species to prevent taxonomic errors. Conserv Genet Resour 13:359–365. https://doi.org/10.1007/s12686-021-01214-7
Funding
Open access funding provided by University of Cape Town. This study was funded by a URC grant from the University of Johannesburg awarded to PRT. The postgraduate work of GNF is supported through the Sea Change Project by the Keystone Grant 542 (1001 Seaforest Species) from the Save Our Seas Foundation, as well as a scholarship from the University of Cape Town.
Author information
Authors and Affiliations
Contributions
The sample was collected by AR. Genomic library preparation and sequencing was done by AEK. Assembly, annotation and analysis were completed by AEK and GNF. GNF wrote the first draft of the manuscript and all authors contributed to editing and finalising the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Ethical approval
This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by the University of Johannesburg Faculty of Science ethics committee (2018-08-06/Zoology Department).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fee, G.N., Roura, A., Emami-Khoyi, A. et al. The complete mitochondrial genome of Octopus vulgaris. Mol Biol Rep 51, 21 (2024). https://doi.org/10.1007/s11033-023-08984-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11033-023-08984-3