Abstract
Background
Chalcone isomerase (CHI; EC 5.5.1.6) is one of the key enzymes in the flavonoid biosynthetic pathway that is responsible for the intramolecular cyclization of chalcones into specific 2S-flavanones.
Methods and results
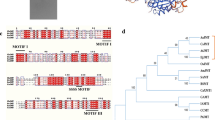
In this study, the open reading frame (ORF) of CHI was successfully isolated from the cDNA of Polygonum minus at 711-bp long, encoding for 236 amino acid residues, with a predicted molecular weight of 25.4 kDa. Multiple sequence alignment and phylogenetic analysis revealed that the conserved residues (Thr50, Tyr108, Asn115, and Ser192) in the cleft of CHI enzyme group active site are present in PmCHI protein sequence and classified as type I. PmCHI comprises more hydrophobic residues without a signal peptide and transmembrane helices. The three-dimensional (3D) structure of PmCHI predicted through homology modeling was validated by Ramachandran plot and Verify3D, with values within the acceptable range of a good model. PmCHI was cloned into pET-28b(+) plasmid, expressed in Escherichia coli BL21(DE3) at 16 °C and partially purified.
Conclusion
These findings contribute to a deeper understanding of the PmCHI protein and its potential for further characterization of its functional properties in the flavonoid biosynthetic pathway.








Similar content being viewed by others
Data availability
Data will be made available upon reasonable request.
References
Pandey RP, Parajuli P, Koffas MAG, Sohng JK (2016) Microbial production of natural and non-natural flavonoids: pathway engineering, directed evolution and systems/synthetic biology. Biotechnol Adv 34:634–662. https://doi.org/10.1016/j.biotechadv.2016.02.012
Kumar S, Pandey AK (2013) Chemistry and biological activities of flavonoids: an overview. Sci World J. https://doi.org/10.1155/2013/162750
Simkhada D, Kurumbang NP, Lee HC, Sohng JK (2010) Exploration of glycosylated flavonoids from metabolically engineered E. coli. Biotechnol Bioprocess Eng 15:754–760. https://doi.org/10.1007/s12257-010-0012-4
Hossain MK, Dayem AA, Han J et al (2016) Molecular mechanisms of the anti-obesity and anti-diabetic properties of flavonoids. Int J Mol Sci. https://doi.org/10.3390/ijms17040569
Shah FLA, Ramzi AB, Baharum SN et al (2019) Recent advancement of engineering microbial hosts for the biotechnological production of flavonoids. Mol Biol Rep 46:6647–6659. https://doi.org/10.1007/s11033-019-05066-1
Kaneko M, Hwang EI, Ohnishi Y, Horinouchi S (2003) Heterologous production of flavanones in Escherichia coli: potential for combinatorial biosynthesis of flavonoids in bacteria. J Ind Microbiol Biotechnol 30:456–461. https://doi.org/10.1007/s10295-003-0061-1
Christapher P, Parasuraman S, Christina JA et al (2015) Review on Polygonum minus. Huds, a commonly used food additive in Southeast Asia. Pharmacogn Res 7:1. https://doi.org/10.4103/0974-8490.147125
Rusdi NA, Goh HH, Baharum SN (2016) GC–MS/olfactometric characterisation and aroma extraction dilution analysis of aroma active compounds in Polygonum minus essential oil. Plant Omics 9:289–294. https://doi.org/10.21475/poj.16.09.04.p7901
Rahnamaie-Tajadod R, Goh H-H, Mohd Noor N (2019) Methyl jasmonate-induced compositional changes of volatile organic compounds in Polygonum minus leaves. J Plant Physiol 240:152994. https://doi.org/10.1016/j.jplph.2019.152994
Khairudin K, Sukiran NA, Goh H-H et al (2013) Direct discrimination of different plant populations and study on temperature effects by Fourier transform infrared spectroscopy. Metabolomics 10:11306. https://doi.org/10.1007/s11306-013-0570-5
Loke K-K, Rahnamaie-Tajadod R, Yeoh C-C et al (2017) Transcriptome analysis of Polygonum minus reveals candidate genes involved in important secondary metabolic pathways of phenylpropanoids and flavonoids. PeerJ 5:e2938. https://doi.org/10.7717/peerj.2938
Goh H-H, Khairudin K, Sukiran NA et al (2016) Metabolite profiling reveals temperature effects on the VOCs and flavonoids of different plant populations. Plant Biol 18:130–139. https://doi.org/10.1111/plb.12403
Loke KK, Rahnamaie-Tajadod R, Yeoh CC et al (2016) RNA-seq analysis for secondary metabolite pathway gene discovery in Polygonum minus. Genomics Data 7:12–13. https://doi.org/10.1016/j.gdata.2015.11.003
Roslan ND, Tan C, Ismail I, Zainal Z (2013) cDNA cloning and expression analysis of the chalcone synthase gene (CHS) from Polygonum minus. Aust J Crop Sci 7:777–783
Guo J, Zhou W, Lu Z et al (2015) Isolation and functional analysis of chalcone isomerase gene from purple-fleshed sweet potato. Plant Mol Biol Rep 33:1451–1463. https://doi.org/10.1007/s11105-014-0842-x
Jez JM, Bowman ME, Dixon RA, Noel JP (2000) Structure and mechanism of the evolutionarily unique plant enzyme chalcone isomerase. Nat Struct Biol 7:786–791. https://doi.org/10.1038/79025
Ren C, Tang X, Chen C et al (2019) Cloning and expression analysis of a new chalcone isomerase gene during flowering in safflower. Turk J Bot 43:143–150. https://doi.org/10.3906/bot-1809-25
Chang AY, Chau VWY, Landas JA, Pang Y (2017) Preparation of calcium competent Escherichia coli and heat-shock transformation. JEMI Methods 1:22–25
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Petersen TN, Brunak S, von Heijne G, Nielsen H (2011) SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods 8:785–786
Krogh A, Larsson B, Von Heijne G, Sonnhammer ELL (2001) Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol 305:567–580. https://doi.org/10.1006/jmbi.2000.4315
Armenteros JJA, Salvatore M, Emanuelsson O et al (2019) Detecting sequence signals in targeting peptides using deep learning. Life Sci Alliance 2:1–14. https://doi.org/10.26508/lsa.201900429
Yu CS, Cheng CW, Su WC et al (2014) CELLO2GO: a web server for protein subCELlular LOcalization prediction with functional gene ontology annotation. PLoS ONE 9:e99368. https://doi.org/10.1371/journal.pone.0099368
Horton P, Park KJ, Obayashi T et al (2007) WoLF PSORT: protein localization predictor. Nucleic Acids Res 35:585–587. https://doi.org/10.1093/nar/gkm259
Krieger E, Darden T, Nabuurs SB et al (2004) Making optimal use of empirical energy functions: force-field parameterization in crystal space. Proteins 57:678–683. https://doi.org/10.1002/prot.20251
Colovos C, Yeates TO (1993) Verification of protein structures: patterns of nonbonded atomic interactions. Protein Sci 2:1511–1519. https://doi.org/10.1002/pro.5560020916
Laskowski RA, MacArthur MW, Moss DS, Thornton JM (1993) PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr 26:283–291. https://doi.org/10.1107/s0021889892009944
Ramachandran GN, Ramakrishnan C, Sasisekharan V (1963) Stereochemistry of polypeptide chain configurations. J Mol Biol 7:95–99. https://doi.org/10.1016/S0022-2836(63)80023-6
Sun W, Meng X, Liang L et al (2015) Molecular and biochemical analysis of chalcone synthase from Freesia hybrida in flavonoid biosynthetic pathway. PLoS ONE 10:1–18. https://doi.org/10.1371/journal.pone.0119054
Das P, Rawal SK (2016) Cloning, expression and purification of chalcone synthase from Solanum tuberosum. IOSR J Biotechnol Biochem 2(3):7–11
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685. https://doi.org/10.1038/227680a0
Mahmood T, Yang P-C (2012) Western blot: technique, theory, and trouble shooting. N Am J Med Sci 4:429–434. https://doi.org/10.4103/1947-2714.100998
Cheng H, Li L, Cheng S et al (2011) Molecular cloning and function assay of a chalcone isomerase gene (GbCHI) from Ginkgo biloba. Plant Cell Rep 30:49–62. https://doi.org/10.1007/s00299-010-0943-4
Park SH, Lee CW, Cho SM et al (2018) Crystal structure and enzymatic properties of chalcone isomerase from the Antarctic vascular plant Deschampsia antarctica Desv. PLoS ONE 13:1–17. https://doi.org/10.1371/journal.pone.0192415
Ku Bahaudin KNA, Ramzi AB, Baharum SN et al (2018) Current progress in production of flavonoids using systems and synthetic biology platforms. Sains Malays 47:3077–3084. https://doi.org/10.17576/jsm-2018-4712-18
Yin YC, Zhang XD, Gao ZQ et al (2019) The research progress of chalcone isomerase (CHI) in plants. Mol Biotechnol 61:32–52. https://doi.org/10.1007/s12033-018-0130-3
Sun W, Shen H, Xu H et al (2019) Chalcone isomerase a key enzyme for anthocyanin biosynthesis in Ophiorrhiza japonica. Front Plant Sci 10:1–12. https://doi.org/10.3389/fpls.2019.00865
Jez JM, Noel JP (2002) Reaction mechanism of chalcone isomerase: pH dependence, diffusion control, and product binding differences. J Biol Chem 277:1361–1369. https://doi.org/10.1074/jbc.M109224200
Wan Q, Bai T, Liu M, Liu Y, Xie Y, Zhang T, Huang M, Zhang J (2022) Comparative analysis of the chalcone-flavanone isomerase genes in six citrus species and their expression analysis in sweet orange (Citrus sinensis). Front Genet 13:1–13. https://doi.org/10.3389/fgene.2022.848141
Dixon RA, Richard Blyden E, Robbins MP et al (1988) Comparative biochemistry of chalcone isomerases. Phytochemistry 27:2801–2808. https://doi.org/10.1016/0031-9422(88)80666-6
Guo D, Gao Y, Liu F et al (2019) Integrating molecular characterization and metabolites profile revealed CtCHI1’s significant role in Carthamus tinctorius L. BMC Plant Biol 19:1–13. https://doi.org/10.1186/s12870-019-1962-0
Wang L, Liu X, Meng X et al (2018) Cloning and expression analysis of a chalcone isomerase (CnCHI) gene from Chamaemelum nobile. Biotechnology 17:19–25. https://doi.org/10.3923/biotech.2018.19.25
Bednar RA, Hadcock JR (1998) Purification and characterization of chalcone isomerase from soybeans. J Biol Chem 273:30003–30011. https://doi.org/10.1074/jbc.273.45.30003
Zhou X, Li J, Fan Z (2012) Cloning and expression analysis of chalcone isomerase gene cDNA from Camellia nitidissima. Forest Res Beijing 25(1):93–99
Shahat AA, Marzouk MS (2013) 13—tannins and related compounds from medicinal plants of Africa. In: Kuete V (ed) Medicinal plant research in Africa. Elsevier, Oxford, pp 479–555
Jez JM, Bowman ME, Noel JP (2002) Role of hydrogen bonds in the reaction mechanism of chalcone isomerase. Biochemistry 41:5168–5176. https://doi.org/10.1021/bi0255266
Furumura S, Ozaki T, Sugawara A, Morishita Y, Tsukada K, Ikuta T, Inoue A, Asai T (2022) Identification and functional characterization of fungal chalcone synthase and chalcone isomerase. J Nat Prod 86:398–405. https://doi.org/10.1021/acs.jnatprod.2c01027
Khor BY, Tye GJ, Lim TS et al (2014) The structure and dynamics of BmR1 protein from Brugia malayi: in silico approaches. Int J Mol Sci 15:11082–11099. https://doi.org/10.3390/ijms150611082
Ngaki MN, Louie GV, Philippe RN et al (2012) Evolution of the chalcone-isomerase fold from fatty-acid binding to stereospecific catalysis. Nature 485:530–533. https://doi.org/10.1038/nature11009
Barh D, Barve N, Gupta K et al (2013) Exoproteome and secretome derived broad spectrum novel drug and vaccine candidates in Vibrio cholerae targeted by Piper betel derived compounds. PLoS ONE 8:e52773. https://doi.org/10.1371/journal.pone.0052773
Baneyx F (1999) Recombinant protein expression in Escherichia coli. Curr Opin Biotechnol 10:411–421. https://doi.org/10.1016/S0958-1669(99)00003-8
Acknowledgements
Fatin Lyana Azman Shah was supported by a scholarship from Jabatan Perkhidmatan Awam (JPA) Malaysia.
Funding
The work was supported through the Special Allocation for Agencies Under Academy of Science Malaysia-Ministry of Science, Technology and Innovation Grant (PKA0514F004, 6300824) and Universiti Putra Malaysia Putra Graduate Initiative (GP/IPS/2016/9482200).
Author information
Authors and Affiliations
Contributions
All authors listed have made a considerable, direct, and logical contribution to the work and approved it for publication. The experiments were designed by FLAS, SNB, HHG, TCL, ABR, SNO and SS. FLAS and SS drafted the manuscript. All authors discussed the results, edited the manuscript, and approved the final version.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest regarding the publication of this article.
Ethical approval
This article does not contain any studies involving human participants or animals performed by any of the authors.
Informed consent
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shah, F.L.A., Baharum, S.N., Goh, HH. et al. Molecular cloning and in silico analysis of chalcone isomerase from Polygonum minus. Mol Biol Rep 50, 5283–5294 (2023). https://doi.org/10.1007/s11033-023-08417-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-023-08417-1