Abstract
Background
The purpose of this study was to investigate the relationship between the expression of autophagy-related genes and prognosis in hepatocellular carcinoma (HCC).
Methods and Results
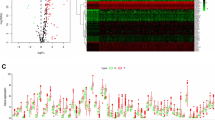
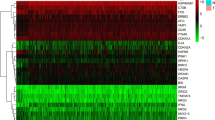
We selected three autophagy-related genes (ATG3, ATG7, and ATG9A) from gene expression data of liver cancer patients in The Cancer Genome Atlas (TCGA) database by Kaplan-Meier survival analysis, univariate and multivariate Cox regression analysis, and Gene Set Enrichment Analysis (GSEA). Human Protein Atlas (HPA) and Gene Expression Profiling Interactive Analysis (GEPIA) databases were applied to testify the credibility of our results. The expression levels of ATG3, ATG7, and ATG9A were verified by real-time quantitative PCR (RT-qPCR) in normal liver cells (L02) and three HCC cell lines (HepG2, Hep3b, and Li-7). Data analysis results from TCGA showed high ATG3, ATG7, ATG9A expression in HCC tumor tissues. Kaplan-Meier survival analysis showed that the survival rate of the high expression group of ATG3, ATG7, and ATG9A was all significantly lower than the low expression group. GSEA analysis showed that many signaling pathways (such as the regulation of autophagy, glycine serine and threonine metabolism, pathways in cancer, mitogen-activated protein kinase (MAPK) signaling pathway, mammalian target of rapamycin (mTOR) signaling pathway, as well as P53 signaling pathway) were differentially enriched in HCCs with ATG3, ATG7, and ATG9A expression. GEPIA and RT-qPCR also identified that the mRNA expression level of ATG3, ATG7, and ATG9A in normal liver cells were significantly lower than in HCC cells. High protein expression of ATG3, ATG7, and ATG9A was displayed in HCCs from the HPA database.
Conclusions
The ATG3, ATG7, ATG9A might be utilized as prognostic biomarkers for liver cancer.





Similar content being viewed by others
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A et al (2021) Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA. Cancer J Clin 71:209–249. https://doi.org/10.3322/caac.21660
Villanueva A (2019) Hepatocellular Carcinoma. N Engl J Med 380:1450–1462. https://doi.org/10.1056/NEJMra1713263.
Foerster F, Gairing SJ, Muller L, Galle PR (2022) NAFLD-driven HCC: Safety and efficacy of current and emerging treatment options. J Hepatol 76:446–457. https://doi.org/10.1016/j.jhep.2021.09.007
Chen S, Zhao E (2021) Development and validation of a robust epithelial-mesenchymal transition (EMT)-related prognostic signature for hepatocellular carcinoma. Clin Res Hepatol Gastroenterol 45:101587. https://doi.org/10.1016/j.clinre.2020.101587
Zhang Z, Teng M, Xu Z, Wang D (2020) Correlations of HACE1 expression with pathological stages, CT features and prognosis of hepatocellular carcinoma patients. J BUON 25:2570–2575
Zhang Q, Xiao Z, Sun S, Wang K, Qian J, Cui Z et al (2021) Integrated Proteomics and Bioinformatics to Identify Potential Prognostic Biomarkers in Hepatocellular Carcinoma. Cancer Manag Res 13:2307–2317. https://doi.org/10.2147/CMAR.S291811
Kroemer G, Marino G, Levine B (2010) Autophagy and the integrated stress response. Mol Cell. 40:280–293. https://doi.org/10.1016/j.molcel.2010.09.023
Kobayashi S, Liang Q (2015) Autophagy and mitophagy in diabetic cardiomyopathy. Biochim Biophys Acta. 1852:252–261. https://doi.org/10.1016/j.bbadis.2014.05.020
Nishida K, Otsu K (2016) Autophagy during cardiac remodeling. J Mol Cell Cardiol. 95:11–18. https://doi.org/10.1016/j.yjmcc.2015.12.003
Tsukada M, Ohsumi Y (1993) Isolation and characterization of autophagy-defective mutants of Saccharomyces cerevisiae. FEBS Lett 333: 169–174. https://doi.org/10.1016/0014-5793(93)80398-e
Murrow L, Malhotra R, Debnath J (2015) ATG12-ATG3 interacts with Alix to promote basal autophagic flux and late endosome function. Nat Cell Biol 17:300–310. https://doi.org/10.1038/ncb3112
Murrow L, Debnath J (2018) Atg12-Atg3 Coordinates Basal Autophagy, Endolysosomal Trafficking, and Exosome Release. Mol Cell Oncol 5:e1039191. https://doi.org/10.1080/23723556.2015.1039191
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)). Method Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Zhang H, Lu B (2020) The Roles of ceRNAs-Mediated Autophagy in Cancer Chemoresistance and Metastasis. Cancers (Basel). 12: 2926. https://doi.org/10.3390/cancers12102926
Li X, Zhou Y, Yang L, Ma Y, Peng X, Yang S et al (2020) LncRNA NEAT1 promotes autophagy via regulating miR-204/ATG3 and enhanced cell resistance to sorafenib in hepatocellular carcinoma. J Cell Physiol 235:3402–3413. https://doi.org/10.1002/jcp.29230
Guo J, Ma Y, Peng X, Jin H, Liu J (2019) LncRNA CCAT1 promotes autophagy via regulating ATG7 by sponging miR-181 in hepatocellular carcinoma. J Cell Biochem 120:17975–17983. https://doi.org/10.1002/jcb.29064
Shi Y, Yang X, Xue X, Sun D, Cai P, Song Q et al (2020) HANR Enhances Autophagy-Associated Sorafenib Resistance Through miR-29b/ATG9A Axis in Hepatocellular Carcinoma. Onco Targets Ther. 13:2127–2137. https://doi.org/10.2147/OTT.S229913
Wei H, Hu J, Pu J, Tang Q, Li W, Ma R et al (2019) Long noncoding RNA HAGLROS promotes cell proliferation, inhibits apoptosis and enhances autophagy via regulating miR-5095/ATG12 axis in hepatocellular carcinoma cells. Int Immunopharmacol 73:72–80. https://doi.org/10.1016/j.intimp.2019.04.049
Li J, Yang B, Zhou Q, Wu Y, Shang D, Guo Y et al (2013) Autophagy promotes hepatocellular carcinoma cell invasion through activation of epithelial-mesenchymal transition. Carcinogenesis 34:1343–1351. https://doi.org/10.1093/carcin/bgt063
Zhao Y, Wu H, Xing X, Ma Y, Ji S, Xu X et al (2020) CD13 Induces Autophagy to Promote Hepatocellular Carcinoma Cell Chemoresistance Through the P38/Hsp27/CREB/ATG7 Pathway. J Pharmacol Exp Ther 374:512–520. https://doi.org/10.1124/jpet.120.265637
Sundarraj K, Raghunath A, Panneerselvam L, Perumal E (2020) Fisetin Inhibits Autophagy in HepG2 Cells via PI3K/Akt/mTOR and AMPK Pathway. Nutr Cancer 73:2502–2514. https://doi.org/10.1080/01635581.2020.1836241
Cui J, Shen HM, Lim L (2020) The Role of Autophagy in Liver Cancer: Crosstalk in Signaling Pathways and Potential Therapeutic Targets. Pharmaceuticals (Basel). 13: 432. https://doi.org/10.3390/ph13120432
Wang M, Jing J, Li H, Liu J, Yuan Y, Sun L (2021) The expression characteristics and prognostic roles of autophagy-related genes in gastric cancer. PeerJ. 9:e10814. https://doi.org/10.7717/peerj.10814
Tang JY, Hsi E, Huang YC, Hsu NC, Chen YK, Chu PY et al (2013) ATG9A overexpression is associated with disease recurrence and poor survival in patients with oral squamous cell carcinoma. Virchows Arch 463:737–742. https://doi.org/10.1007/s00428-013-1482-5
Chen H, Deng Q, Wang W, Tao H, Gao Y (2020) Identification of an autophagy-related gene signature for survival prediction in patients with cervical cancer. J Ovarian Res 13:131. https://doi.org/10.1186/s13048-020-00730-8
Zhou J, Hang D, Jiang Y, Chen J, Han J, Zhou W et al (2017) Evaluation of genetic variants in autophagy pathway genes as prognostic biomarkers for breast cancer. Gene. 627:549–555. https://doi.org/10.1016/j.gene.2017.06.053
Chen M, Zhang S, Nie Z, Wen X, Gao Y (2020) Identification of an Autophagy-Related Prognostic Signature for Clear Cell Renal Cell Carcinoma. Front Oncol 10:873. https://doi.org/10.3389/fonc.2020.00873
Du H, Xie S, Guo W, Che J, Zhu L, Hang J et al (2021) Development and validation of an autophagy-related prognostic signature in esophageal cancer. Ann Transl Med 9:317. https://doi.org/10.21037/atm-20-4541
Acknowledgements
This work was supported by the WU JIEPING MEDICAL FOUNDATION of China (Grant NO. 320.6750.19089-103 and Grant NO. 320.6750.19089-75), and Beijing High-level Public Health Technical Personnel Construction Project (Grant NO. 2022-2-014).
Funding
This study was funded by the WU JIEPING MEDICAL FOUNDATION of China (Grant NO. 320.6750.19089-103 and Grant NO. 320.6750.19089-75), Beijing High-level Public Health Technical Personnel Construction Project (Grant NO. 2022-2-014).
Author information
Authors and Affiliations
Contributions
Formal analysis, Data Curation, Writing - Original Draft, Validation, Visualization, and Writing - Review & Editing were performed by Xueying Zhao. Investigation, Supervision, Data Analysis, Writing - Review & Editing were performed by Shangqi Yin. Project administration, Resources, and Data Analysis were performed by Jingren Shi, Mei Zheng, Chaonan He, Huan Meng, Ying Han, Jin Chen, and Jinyu Han. Conceptualization, Supervision, Writing - Review & Editing, Data Analysis were performed by Zhengrong Yuan and Yajie Wang. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Ethics approval and consent to participate
Patients do not need to give informed consent to this study because the analysis used anonymous clinical data from the TCGA database and retained the confidentiality of patients.
Consent for publication
The authors agree to the publication of the article and tacitly or explicitly by the responsible authorities where the work was carried out. If the article is accepted, it will not be published elsewhere in the same form, in English or any other language (including electronic versions) without the written consent of the copyright holder.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The authors Xueying Zhao and Shangqi Yin contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhao, X., Yin, S., Shi, J. et al. The association between several autophagy-related genes and their prognostic values in hepatocellular carcinoma: a study on the foundation of TCGA, GEPIA and HPA databases. Mol Biol Rep 49, 10269–10277 (2022). https://doi.org/10.1007/s11033-022-07426-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-022-07426-w