Abstract
Background
Adipose tissue hypoxia and members of the hypoxia-inducible factor alpha (HIFA) are involved in development of obesity. However, the mechanism and functions of HIF3A, one of three HIFA paralogs, in fat deposition have not been sufficiently studied.
Methods and results
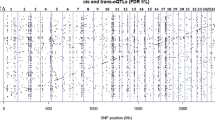
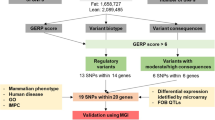
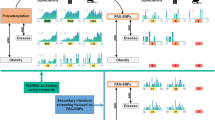
In the present study, we investigated whether Hif3a sequence variants are associated with divergent fat deposition in mouse selection lines for fatness and leanness. Sequencing and RFLP were used to analyse sequence variants within Hif3a. To identify candidate regulatory variants, we performed literature screening and used databases and bioinformatics tools like Ensembl, MethPrimer, TargetScanMouse, miRDB, PolyAsite, RISE, LncRRIsearch, RNAfold, PredictProtein, CAIcal, and switches.ELM Resource. There are 90 sequence variants in Hif3a between the two mouse lines. While most Fat line variants locate within intronic regions, Lean line variants are mainly in 3′ UTR. We constructed a map of Hif3a potential regulatory regions and identified 39 regulatory variants by integrating data on constrained and regulatory elements, CpGs, and miRNAs and lncRNAs binding sites. Moreover, 3′ UTR and two exonic variants may influence mRNA stability, translation rate and protein functionality. We propose as priority candidates for further functional studies a missense (rs37398126) and synonymous (rs37739792) variants, and intronic (rs47471302) variant that overlap conserved element in promoter region and predicted lncRNAs binding site.
Conclusion
The results indicate a potential involvement of Hif3a in fat deposition. Additionally, approach used in the present study may serve as a general guideline for constructing an integrative gene map for prioritizing candidate gene variants with phenotypic effects.




Similar content being viewed by others
References
Blüher M (2019) Obesity: global epidemiology and pathogenesis. Nat Rev Endocrinol 15:288–298. https://doi.org/10.1038/s41574-019-0176-8
Longo M, Zatterale F, Naderi J et al (2019) Adipose tissue dysfunction as determinant of obesity-associated metabolic complications. Int J Mol Sci 20:2358. https://doi.org/10.3390/ijms20092358
Gealekman O, Guseva N, Hartigan C et al (2011) Depot-specific differences and insufficient subcutaneous adipose tissue angiogenesis in human obesity. Circulation 123:186–194. https://doi.org/10.1161/CIRCULATIONAHA.110.970145
Trayhurn P (2013) Hypoxia and adipose tissue function and dysfunction in obesity. Physiol Rev 93:1–21. https://doi.org/10.1152/physrev.00017.2012
Fujisaka S, Usui I, Ikutani M et al (2013) Adipose tissue hypoxia induces inflammatory M1 polarity of macrophages in an HIF-1α-dependent and HIF-1α-independent manner in obese mice. Diabetologia 56:1403–1412. https://doi.org/10.1007/s00125-013-2885-1
Semenza GL (2012) Hypoxia-inducible factors in physiology and medicine. Cell 148:399–408. https://doi.org/10.1016/j.cell.2012.01.021
Ravenna L, Salvatori L, Russo MA (2016) HIF3α: the little we know. FEBS J 283:993–1003. https://doi.org/10.1111/febs.13572
Hatanaka M, Shimba S, Sakaue M et al (2009) Hypoxia-inducible factor-3α functions as an accelerator of 3T3-L1 adipose differentiation. Biol Pharm Bull 32:1166–1172. https://doi.org/10.1248/bpb.32.1166
Pfeiffer S, Krüger J, Maierhofer A et al (2016) Hypoxia-inducible factor 3A gene expression and methylation in adipose tissue is related to adipose tissue dysfunction. Sci Rep 6:27969. https://doi.org/10.1038/srep27969
Amin FZ, Yamashita T, Ohneda O (2018) Deterioration of alveolar development in mice with both HIF-3α knockout and HIF-2α knockdown. BMC Res Notes 11:449. https://doi.org/10.1186/s13104-018-3563-7
Yamashita T, Ohneda O, Nagano M et al (2008) Abnormal heart development and lung remodeling in mice lacking the hypoxia-inducible factor-related basic helix-loop-helix PAS protein NEPAS. Mol Cell Biol 28:1285–1297. https://doi.org/10.1128/MCB.01332-07
Jakubauskiene E, Vilys L, Makino Y et al (2015) Increased serine-arginine (SR) protein phosphorylation changes pre-mRNA splicing in hypoxia. J Biol Chem 290:18079–18089. https://doi.org/10.1074/jbc.M115.639690
Kunej T (2021) Integrative map of HIF1A regulatory elements and variations. Genes (Basel) 12:1526. https://doi.org/10.3390/genes12101526
Kristan A, Debeljak N, Kunej T (2021) Integration and visualization of regulatory elements and variations of the EPAS1 gene in human. Genes (Basel) 12:1793. https://doi.org/10.3390/genes12111793
Ren F, Zhang N, Zhang L et al (2020) Alternative Polyadenylation: a new frontier in post transcriptional regulation. Biomark Res 8:67. https://doi.org/10.1186/s40364-020-00249-6
Wang S, Song J, Yang Y et al (2017) Interaction between obesity and the Hypoxia Inducible Factor 3 Alpha Subunit rs3826795 polymorphism in relation with plasma alanine aminotransferase. BMC Med Genet 18:80. https://doi.org/10.1186/s12881-017-0437-0
Nadeau JH, Auwerx J (2019) The virtuous cycle of human genetics and mouse models in drug discovery. Nat Rev Drug Discov 18:255–272. https://doi.org/10.1038/s41573-018-0009-9
Crowley JJ, Zhabotynsky V, Sun W et al (2015) Analyses of allele-specific gene expression in highly divergent mouse crosses identifies pervasive allelic imbalance. Nat Genet 47:353–360. https://doi.org/10.1038/ng.3222
Kelley DR (2020) Cross-species regulatory sequence activity prediction. PLoS Comput Biol 16:e1008050. https://doi.org/10.1371/journal.pcbi.1008050
Sharp GL, Hill WG, Robertson A (1984) Effects of selection on growth, body composition and food intake in mice I. Responses in selected traits. Genet Res 43:75–92. https://doi.org/10.1017/S0016672300025738
Simončič M, Režen T, Juvan P et al (2011) Obesity resistant mechanisms in the Lean polygenic mouse model as indicated by liver transcriptome and expression of selected genes in skeletal muscle. BMC Genomics 12:96. https://doi.org/10.1186/1471-2164-12-96
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–1760. https://doi.org/10.1093/bioinformatics/btp324
McKenna A, Hanna M, Banks E et al (2010) The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20:1297–1303. https://doi.org/10.1101/gr.107524.110
DePristo MA, Banks E, Poplin R et al (2011) A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet 43:491–498. https://doi.org/10.1038/ng.806
Auwera GA, Carneiro MO, Hartl C et al (2013) From FastQ data to high-confidence variant calls: the genome analysis toolkit best practices pipeline. Curr Protoc Bioinforma 43:483–492. https://doi.org/10.1002/0471250953.bi1110s43
McLaren W, Gil L, Hunt SE et al (2016) The ensembl variant effect predictor. Genome Biol 17:122. https://doi.org/10.1186/s13059-016-0974-4
Bozeman MT Golden Helix GenomeBrowse® visualization tool (version 2.X) [software]. Golden Helix, Inc. http://www.goldenhelix.com
Howe KL, Achuthan P, Allen J et al (2021) Ensembl 2021. Nucleic Acids Res 49:D884–D891. https://doi.org/10.1093/nar/gkaa942
Kunej T, Skok DJ, Horvat S et al (2010) The glypican 3-hosted murine Mir717 gene: sequence conservation, seed region polymorphisms and putative targets. Int J Biol Sci 6:769–772. https://doi.org/10.7150/ijbs.6.769
Beltram J, Morton NM, Kunej T, Horvat S (2016) Construction of an integrative regulatory element and variation map of the murine Tst locus. BMC Genet 17:77. https://doi.org/10.1186/s12863-016-0381-6
McCauley JL, Kenealy SJ, Margulies EH et al (2007) SNPs in Multi-Species Conserved Sequences (MCS) as useful markers in association studies: a practical approach. BMC Genomics 8:266. https://doi.org/10.1186/1471-2164-8-266
Barth DA, Prinz F, Teppan J et al (2020) Long-noncoding RNA (lncRNA) in the regulation of hypoxia-inducible factor (HIF) in cancer. Non-Coding RNA. https://doi.org/10.3390/NCRNA6030027
Gallo S, Arcidiacono MV, Tisato V et al (2018) Upregulation of the alternative splicing factor NOVA2 in colorectal cancer vasculature. Onco Targets Ther 11:6049–6056. https://doi.org/10.2147/OTT.S171678
Yi M, Li Y, Wang D et al (2020) KCNQ1OT1 exacerbates ischemia-reperfusion injury through targeted inhibition of miR-140-3P. Inflammation 435(43):1832–1845. https://doi.org/10.1007/S10753-020-01257-2
Chen C, Wang Y, Zhang Z et al (2020) Expression of lncRNA KCNQ1OT1 in human adipocyte differentiation and adipose tissue of obese people. Chin J Endocrinol Metab. https://doi.org/10.3760/cma.j.cn311282-20190806-00320
Zhang Q, Cheng T, Jin S et al (2017) Genome-wide open chromatin regions and their effects on the regulation of silk protein genes in Bombyx mori. Sci Rep 7:12919. https://doi.org/10.1038/s41598-017-13186-6
Corradin O, Scacheri PC (2014) Enhancer variants: evaluating functions in common disease. Genome Med 6:85. https://doi.org/10.1186/s13073-014-0085-3
Dick KJ, Nelson CP, Tsaprouni L et al (2014) DNA methylation and body-mass index: a genome-wide analysis. Lancet 383:1990–1998. https://doi.org/10.1016/S0140-6736(13)62674-4
Koukourakis MI, Papazoglou D, Giatromanolaki A et al (2006) C2028T polymorphism in exon 12 and dinucleotide repeat polymorphism in intron 13 of the HIF-1α gene define HIF-1α protein expression in non-small cell lung cancer. Lung Cancer 53:257–262. https://doi.org/10.1016/j.lungcan.2006.05.025
Liang S, Ren K, Li B et al (2020) LncRNA SNHG1 alleviates hypoxia-reoxygenation-induced vascular endothelial cell injury as a competing endogenous RNA through the HIF-1α/VEGF signal pathway. Mol Cell Biochem 465:1–11. https://doi.org/10.1007/s11010-019-03662-0
Shi C, Zhu L, Chen X et al (2014) IL-6 and TNF-α induced obesity-related inflammatory response through transcriptional regulation of miR-146b. J Interf Cytokine Res 34:342–348. https://doi.org/10.1089/jir.2013.0078
Sanada T, Sano T, Sotomaru Y et al (2020) Anti-inflammatory effects of miRNA-146a induced in adipose and periodontal tissues. Biochem Biophys Rep 22:100757. https://doi.org/10.1016/j.bbrep.2020.100757
Gong Q, Xie J, Li Y et al (2019) Enhanced ROBO4 is mediated by up-regulation of HIF-1α/SP1 or reduction in miR-125b-5p/miR-146a-5p in diabetic retinopathy. J Cell Mol Med 23:4723–4737. https://doi.org/10.1111/jcmm.14369
Chouvarine P, Legchenko E, Geldner J et al (2019) Hypoxia drives cardiac miRNAs and inflammation in the right and left ventricle. J Mol Med 97:1427–1438. https://doi.org/10.1007/s00109-019-01817-6
Morton NM, Nelson YB, Michailidou Z et al (2011) A stratified transcriptomics analysis of polygenic fat and lean mouse adipose tissues identifies novel candidate obesity genes. PLoS ONE 6:e23944. https://doi.org/10.1371/journal.pone.0023944
Wen P, Xiao P, Xia J (2016) dbDSM: a manually curated database for deleterious synonymous mutations. Bioinformatics 32:1914–1916. https://doi.org/10.1093/bioinformatics/btw086
Jo CW, Lee JH, Song JS et al (2021) Isolated and sporadic human mesiodens is associated with a synonymous variant in the ACVR2A gene. Pediatr Dent 43:39–43
Sauna ZE, Kimchi-Sarfaty C (2011) Understanding the contribution of synonymous mutations to human disease. Nat Rev Genet 12:683–691. https://doi.org/10.1038/nrg3051
Mylonis I, Chachami G, Samiotaki M et al (2006) Identification of MAPK phosphorylation sites and their role in the localization and activity of hypoxia-inducible factor-1α. J Biol Chem 281:33095–33106. https://doi.org/10.1074/jbc.M605058200
Kasai S, Richardson MJE, Torii S et al (2017) Increase in proapoptotic activity of inhibitory PAS domain protein via phosphorylation by MK2. FEBS J 284:4115–4127. https://doi.org/10.1111/febs.14300
Corrado C, Fontana S (2020) Hypoxia and HIF signaling: one axis with divergent effects. Int J Mol Sci 21:5611. https://doi.org/10.3390/ijms21165611
Wang B, Dai T, Sun W et al (2021) Protein N-myristoylation: functions and mechanisms in control of innate immunity. Cell Mol Immunol 18:878–888. https://doi.org/10.1038/s41423-021-00663-2
Fala AM, Oliveira JF, Adamoski D et al (2015) Unsaturated fatty acids as high-affinity ligands of the C-terminal Per-ARNT-Sim domain from the Hypoxia-inducible factor 3α. Sci Rep 5:12698. https://doi.org/10.1038/srep12698
Pennington K, Chan T, Torres M, Andersen J (2018) The dynamic and stress-adaptive signaling hub of 14–3-3: emerging mechanisms of regulation and context-dependent protein–protein interactions. Oncogene 37:5587–5604. https://doi.org/10.1038/s41388-018-0348-3
Geng H, Harvey CT, Pittsenbarger J et al (2011) HDAC4 protein regulates HIF1α protein lysine acetylation and cancer cell response to hypoxia. J Biol Chem 286:38095–38102. https://doi.org/10.1074/jbc.M111.257055
Acknowledgements
The authors also want to thank Dr Tim J. Aitman (Centre for Genomic and Experimental Medicine, Institute of Genetics and Molecular Medicine, University of Edinburgh, Edinburgh, United Kingdom) for the help with WGS.
Funding
The authors acknowledge the study was financially supported by the Slovenian Research Agency under the postgraduate research program Young researchers (ŠM), P4-0220 research program and J4-2548 research project.
Author information
Authors and Affiliations
Contributions
ŠM and MŠ: Formal analysis, Writing—original draft preparation. NMM: Formal analysis. Writing – review & editing. SSA: Formal analysis. JK: Resources. PD: Resources. SH and TK: Conceptualization, Writing – review & editing, Supervision.
Corresponding authors
Ethics declarations
Competing Interests
All authors have read and approved the final version of the manuscript for submission and declare that they have no competing interests.
Ethical approval
The FLI (Fat) and FHI (Lean) selection lines have been maintained in our laboratory for more than 70 generations. All mice used in this study were maintained according to local ethical and EU regulatory guidelines under the Veterinary Administration of Republic of Slovenia permit No. U34401-23/2020/6.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mikec, Š., Šimon, M., Morton, N.M. et al. Genetic variants of the hypoxia‐inducible factor 3 alpha subunit (Hif3a) gene in the Fat and Lean mouse selection lines. Mol Biol Rep 49, 4619–4631 (2022). https://doi.org/10.1007/s11033-022-07309-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-022-07309-0